Abstract
Background
Familial hemiplegic migraine (FHM) types 1–3 are associated with protein-altering genetic variants in
Methods
Whole-exome sequence data from 454,706 individuals from the UK Biobank were examined for self-reported and inpatient-diagnosed migraine and epilepsy. Carriers were compared with non-carriers in a burden analysis using logistic regression while accounting for age, biological sex and UK Biobank assessment center. A machine learning-based approach was employed to predict whether variants resulted in gain-of-function (GoF), loss-of-function (LoF) or neutral effects.
Results
Heterozygous carriers of GoF
Conclusions
Heterozygous genotypes in all three FHM genes were associated with migraine but not epilepsy in this genotype-focused study. Homozygous
This is a visual representation of the abstract.
Introduction
Familial hemiplegic migraine (FHM) represents a type of migraine with aura that follows an autosomal dominant inheritance pattern (1,2). Migraine attacks in FHM are characterized by distinctive aura symptoms, including unilateral muscle weakness followed by severe headache (1). Not all carriers of FHM-associated genetic variants exhibit the characteristic hemiplegic phenotype; instead, some individuals have more common aura symptoms, such as visual disturbances (3). This suggests a potential contribution of FHM-associated variants to more common types of migraine with aura (4). Probands with FHM and their first-degree relatives exhibit an increased risk of other types of migraine with aura, aside from hemiplegic aura, while showing no increased risk of migraine without aura (4). However, no conclusive evidence has yet established the role of FHM-associated genes in common forms of migraine (5). Despite its presumed rarity, with a prevalence of 0.01% (2,3), FHM has been a subject to extensive research because of its familial occurrence, offering unique insights into the genetics of migraine. This has been achieved through a large number of case reports and familial characterizations, which have resulted in the discovery of a triad of genes linked to the FHM phenotype:
FHM-associated genetic variants may have associations beyond migraine alone. Hence, several case reports have documented clusters of epilepsy cases within families affected by FHM2 (15–18). Similar co-morbidity with epilepsy has been reported in FHM1 (19–21) and FHM3 families (22,23). A recent systematic review based on data from 32 case series on FHM families suggests that 40% of family members affected by FHM also experienced seizures, with an equal division between those exhibiting focal and generalized epilepsy (24). The association between FHM-linked genetic variants and epilepsy finds further support in a study on a cohort of 318 epilepsy patients. The study found distinct
Investigations thus far have predominantly centered on characterizing the genetic profile of individuals and families exhibiting the distinctive FHM phenotype or epilepsy. By contrast, the present study adopted a genotype-first approach to examine carriers of variants in the three FHM-associated genes. This approach revealed new insights into the genetic role of these genes and their correlation with the disease. It was hypothesized that missense and protein-truncating variants within the three FHM-associated genes were associated with migraine and epilepsy. The hypothesis was tested by examining the population-based UK Biobank (UKB) (31). The UKB has enrolled approximately half a million individuals who have been extensively characterized with genetic and electronic health records being available. As exome sequence data has become available for the UKB (32), this constitutes a unique resource to investigate the potential association between sequence variants within the three FHM-associated genes and self-reported and hospital diagnoses for migraine and epilepsy.
Methods
In this study, a genotype-first approach was employed, where carriers of missense and protein-truncating variants in
A single variant-level filter was applied, requiring that at least 90% of all genotypes for a given variant had a read depth of at least 10, meaning each genetic position was read at least 10 times, aiming to identify variants within the FHM-associated genes (37). The predicted consequence of the identified sequence variants within the three FHM-genes was obtained using Variant Effect Predictor (VEP) (38). Only the canonical transcript of each of the three FHM-associated genes (
The study focused on protein-altering variants (i.e. missense variants resulting in amino acid changes and protein-truncating variants, including variants resulting in premature stop (nonsense), canonical splice sites (splice-donor or splice-acceptor) or insertion/deletion variants that shifted frame (frameshift) (41). To test for association between disease outcomes and FHM sequence variants, a burden association test was applied. For each FHM gene, three different genetic scenarios were defined: (i) All missense and protein-truncating variants subdivided according to their predicted gain-of-function (GoF), LoF or neutral effects based on LoGoFunc, a machine learning approach developed to explicitly predict pathogenicity (42); (ii) missense or LoF variants that have been associated with FHM in ClinVar (43); and (iii) FHM-associated variants that were reported in ClinVar as pathogenic, likely pathogenic or pathogenic/likely pathogenic using the American College of Medical Genetics and Genomics (ACMG) classification. In Scenarios ii and iii, an assessment was made to determine whether ClinVar-reported FHM variants, regardless of pathogenicity, were associated with migraine and epilepsy, and whether the association was amplified when restricting the analysis to rare pathogenic/likely pathogenic variants (see supplemental material, Tables S1 and S2). For each of the three different genetic scenarios, we defined additional two groups: individuals having at least one heterozygous or at least one homozygous alternative sequence variants. Individuals that fulfilled both zygosity-criteria for multiple genetic variants were included in both comparative groups. Medical diagnoses were obtained from the hospital inpatient data (World Health Organization's international classification of diseases (ICD-10), UK Biobank data field: 41270). To maximize statistical power, the analyses of diagnostic subtypes were performed on all ICD-10 subcategories containing more ≥500 cases (corresponding to a cohort prevalence of 0.001; see supplemental material, Table S3). This threshold allows us to maintain statistical power without including categories where sample sizes are too small to yield meaningful insights. The following diagnoses were thus included in the subsequent genetic burden analysis: Migraine with aura (ICD-10: G43.1), migraine unspecified (G43.9), generalized idiopathic epilepsy and epileptic syndromes (G40.3) and epilepsy unspecified (G40.9). In addition to hospital diagnosis codes, self-reported cases of migraine and epilepsy (UK Biobank data field: 20002) were also included.
The burden analysis was performed by assigning each UKB participant with an indicator variable (0|1) of whether the individual was either homozygous or heterozygous for any of the identified FHM genetic variants in one of the three scenarios. For each FHM gene and scenario, a logistic regression was fitted with carrier status, defined for each gene with one of the three scenarios, as predictor and with the response variable being diagnosed or self-reported migraine or epilepsy while accounting for age at inclusion, biological sex and UKB assessment center. Note, in the analyses of being either heterozygote or homozygote carriers, individuals from the other contrast group were included among the non-carriers. The degree of association of the carrier status was determined using a likelihood ratio test,
Results
Among all UKB participants, 454,706 individuals had whole exome sequence data available. There were 38,466 more female participants than male participants with sequence data available and, on average, the female participants were slightly younger than male participants (Table 1). There were almost three times more female participants with self-reported migraine compared with male participants (Table 1). Consistently, we observed more inpatient female participants with migraine with aura or unspecified migraine. Slightly more females had reported epilepsy, whereas more males had an ICD-10 inpatient record for unspecified epilepsy (Table 1). In total, 6,065 individuals had at least one epilepsy ICD-10 code and, among them, 2,691 had reported they had epilepsy in the questionnaire. A much larger proportion of UKB participants had self-reported migraine compared with having an inpatient ICD-10 code of any migraine diagnosis (16,621 vs. 6,492, respectively). Approximately one-third of the UKB participants with a migraine diagnosis also reported migraine (i.e. 2,059 UKB participants with both diagnosed and self-reported migraine).
Descriptive statistics of the study population. Age is represented as the mean ± SD, and other numbers are represented as counts. Age comparison was tested with the Wilcoxon rank sum test, whereas the count comparisons were performed using chi-squared test. The subdiagnoses for epilepsy (G.40) and migraine (G.43) were included in further analysis based on a threshold prevalence of more than 500 cases per 500,000 individuals in the UK Biobank cohort. For details, see supplemental material, Table S3.

CACNA1A
Out of 4,128 identified sequence variants in
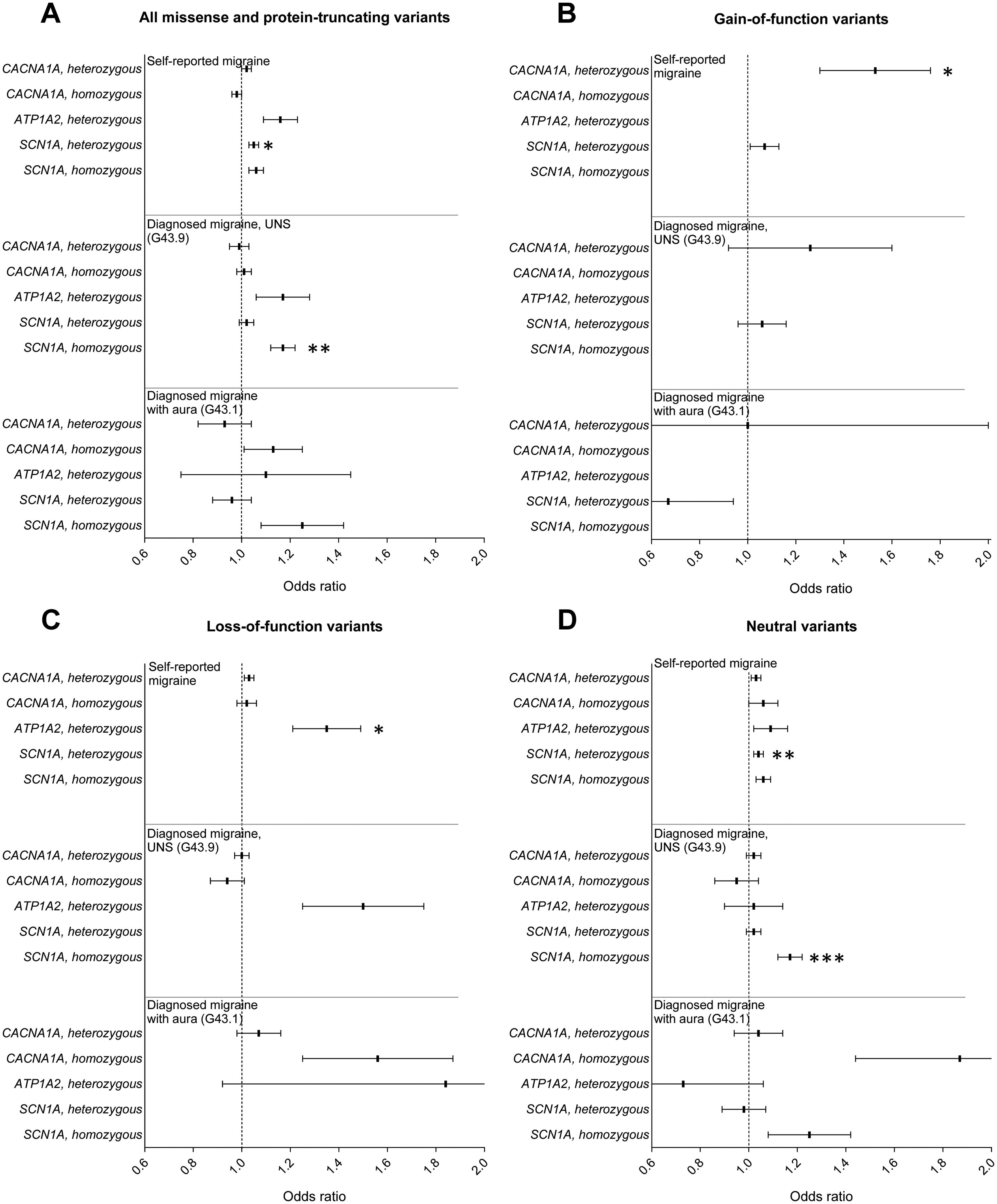
Association between migraine and variants within familial hemiplegic migraine (FHM)-associated genes;

Homozygous carriers of
Number of heterozygous and homozygous carriers of missense and protein-truncating variants (PTVs) in the three familial hemiplegic migraine-associated genes. Functional impact of variants was predicted to be gain-of-function (GoF), loss-of-function (LoF), or neutral based on a machine learning-based method (30). Variants categorized as pathogenic/likely pathogenic were based on ClinVar classifications. The number of carriers is represented as a percentage of the UK Biobank cohort within parentheses. In groups with five or less carriers, the number of carriers is not disclosed due to General Data Protection Regulation.

ATP1A2
In the FHM2-associated
SCN1A
Within the
Discussion
This study adopted a genotype-first strategy to investigate the potential link between the three FHM-associated genes with migraine and epilepsy. The results unveiled a link between migraine and the genes
Although genomic research often focuses on characterizing specific gene profiles in individuals with heritable diseases, the decreasing cost of sequencing has enabled reverse phenotyping based on the gene variants. Given the complex interplay of genes in pathology, this genotype-first approach should not be viewed as definitive diagnoses but rather as indicators of potential risk for developing certain conditions. This is, however, crucial for uncovering novel genotype-disease associations, broadening the phenotypic spectrum, and facilitating further exploration of the functional significance of genetic variants (46).
This is not the first study that used a genotype-first approach to investigate the link between FHM and migraine in the UKB. Markel and Curtis (47) investigated the association with self-reported and diagnosed migraine in UKB participants carrying rare variants with minor allele frequency ≤0.01 in migraine-associated variants, including the three FHM-associated genes. It was concluded that none of the FHM-associated genes were significantly associated with migraine in the UKB (47). Notably, at the time of the study by Markel and Curtis (47), the sample size of the exome sequence data was on approximately 200,000 individuals. In contrast, the current dataset comprises 454,706 individuals (32), more than doubling the available whole-exome sequences and significantly increasing the statistical power to identify associations. The difference in sizes of the UKB cohorts may therefore account for the discrepancy of the results of these studies employing similar burden analysis. Another notable difference from previous studies is the comprehensive approach employed in the present study. This approach includes distinction between predicted GoF and LoF variants, while also considering if variants were previously associated with migraine and their pathogenicity. This multi-faceted strategy provides a more detailed and nuanced understanding of the genetic underpinnings of FHM and its relationship with epilepsy and migraine. Indeed, this approach revealed intriguing differences between the associations in the three genes. For example, an increased risk of self-reported migraine was observed for the FHM3-associated
All three migraine-associated genes addressed in the present study encode membrane ion transporters important for ion homeostasis in the brain and thus its neuronal excitability, as shown in cell culture and animal models (53). Missense GoF variants of
The identification of only a few predicted LoF variants in the
One of the limitations of the present study lies in the absence of a specific diagnosis code for FHM and the lack of FHM as a distinct self-reported disease in the UKB database. This study design carefully considered the balance between statistical robustness and the inclusiveness of disease categories to ensure the reliability of findings. Therefore, the study was restricted to exploring associations with unspecified migraine and migraine with aura. It was not possible to independently reassess whether the standardized diagnostic criteria for migraine and epilepsy were fully met in subjects assigned an ICD-10 diagnosis and in cases of self-reported disease. The study found lower prevalences of self-reported and diagnosed migraine in the UKB cohort compared to previous global estimates (62), potentially limiting the power of analysis. There may be several reasons for the lower prevalence of migraine in UKB. First, the healthy-volunteer bias, which is inherent to the UKB resource, does call for caution when interpreting the results (63). Second, migraine studies in biobanks generally have some assured delicacy (47), as not all patients with migraine have been assigned the migraine diagnosis code unless migraine was related to the contact with the hospital system. This may explain the low number of cases of diagnosed migraine without aura in the UK Biobank. Another possible explanation may be that the general trend in UK is to register migraine by the diagnostic code for Migraine, Unspecified G43.9, unless aura is present. The low prevalence of diagnosed migraine in the UKB may lead to misclassification, with undiagnosed cases in the control group, potentially underestimating the associations with migraine in this study. Due to the low number of diagnosed cases of migraine without aura, this diagnosis was excluded from our analysis. This exclusion may have biased our findings by limiting the study to certain migraine subtypes, potentially underestimating associations that could be present in broader migraine phenotypes. Another limitation of the study is the absence of stratification by biological sex due to the limited number of carriers and cases. Instead, adjustment for sex was made within the statistical analyses. Finally, in addition to clinically diagnosed migraine and epilepsy, the study included self-reported conditions, which is a limitation because self-reported conditions were not necessarily diagnosed by a healthcare professional and may lack clinical validation. The self-reported diagnoses are also potentially limited by recall bias, influencing the accuracy of participants’ recollections of past events or experiences. A previous study demonstrated a good agreement between self-reported migraine and diagnosed migraine based on the International Classification of Headache Disorders (ICHD-II) criteria (64) suggesting a value of including self-reported migraine in the analysis. Thus, migraine without aura or probable migraine without aura is confirmed in 87% of self-reported cases in females (64). By contrast, another study showed that 23.8% of migraine patients are not identified in self-reported assessment (65), potentially leading to underestimation. This may lead to bias in the analysis, but its specific direction is difficult to predict.
Common polygenic variation significantly contributes to the familial aggregation of migraine (66) as observed with
Despite these limitations, the findings of the present study hold significant promise for clinical practice. The findings provide valuable insights into the genetic component of migraine, highlighting the potential role of FHM-related genes in broader migraine phenotypes. By identifying associations between specific genetic variants and migraine, this research enhances our understanding of the genetic underpinnings of the disorder. These insights have the potential to inform future diagnostic tools and treatment strategies. Understanding mutation-related alterations in protein function can catalyze research into novel treatment paradigms, driving innovation in migraine medicine. This knowledge may pave the way for personalized treatment, allowing for targeted therapies that address the functional disturbances caused by specific genetic abnormalities. This is highly relevant in a therapeutic landscape, where, for example, choices of primary prophylactic treatment of sporadic migraine today are based on trial and error. The choice between an antihypertensive, anticonvulsant or antidepressant drug as a first and second choice is purely empirical rather than based on pathophysiological considerations.
The future development of this research will require a more detailed categorization of diagnoses that may be linked to the FHM-related mutations. It is crucial to establish clear connections between specific genetic changes and their functional impacts on the pathogenicity of migraine and related disorders. Although these initial mechanistic insights may be derived from experimental models, it is essential to consider these findings in the context of multicellular organisms and validate them in future studies within a polygenic environment similar to that of our current study.
Conclusions
This study offers a unique genotype-first analysis in the UKB, utilizing almost half a million exome-sequenced participants to investigate the link between variants in the FHM-associated genes
Clinical implications
The study highlights the role of FHM-related genes in migraine, offering insights that could inform future diagnostic tools and treatment strategies. In today's therapeutic landscape, primary prophylactic treatment of migraine is often based on trial and error. Understanding the genetic associations of This understanding of genetic abnormalities paves the way for personalized, targeted therapies that address the functional disturbances caused by these genetic variants, leading to more effective and individualized treatment for migraine patients. Homozygous carriers of
Supplemental Material
sj-docx-1-cep-10.1177_03331024241306103 - Supplemental material for Exploring the association between familial hemiplegic migraine genes (CACNA1A , ATP1A2 and SCN1A ) with migraine and epilepsy: A UK Biobank exome-wide association study
Supplemental material, sj-docx-1-cep-10.1177_03331024241306103 for Exploring the association between familial hemiplegic migraine genes (
Supplemental Material
sj-xlsx-2-cep-10.1177_03331024241306103 - Supplemental material for Exploring the association between familial hemiplegic migraine genes (CACNA1A , ATP1A2 and SCN1A ) with migraine and epilepsy: A UK Biobank exome-wide association study
Supplemental material, sj-xlsx-2-cep-10.1177_03331024241306103 for Exploring the association between familial hemiplegic migraine genes (
Supplemental Material
sj-xlsx-3-cep-10.1177_03331024241306103 - Supplemental material for Exploring the association between familial hemiplegic migraine genes (CACNA1A , ATP1A2 and SCN1A ) with migraine and epilepsy: A UK Biobank exome-wide association study
Supplemental material, sj-xlsx-3-cep-10.1177_03331024241306103 for Exploring the association between familial hemiplegic migraine genes (
Footnotes
Acknowledgements
The data used in the presented study were obtained from the UKB Resource (project ID 60032).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The research was supported by the Danish Cardiovascular Academy and Riisfort Fonden.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
