Abstract
Background
Cluster headache is a severe neurological disorder with a complex genetic background. A missense single nucleotide polymorphism (rs2653349; p.Ile308Val) in the HCRTR2 gene that encodes the hypocretin receptor 2 is the only genetic factor that is reported to be associated with cluster headache in different studies. However, as there are conflicting results between studies, we re-evaluated its role in cluster headache.
Methods
We performed a genetic association analysis for rs2653349 in our large Leiden University Cluster headache Analysis (LUCA) program study population. Systematic selection of the literature yielded three additional studies comprising five study populations, which were included in our meta-analysis. Data were extracted according to predefined criteria.
Results
A total of 575 cluster headache patients from our LUCA study and 874 controls were genotyped for HCRTR2 SNP rs2653349 but no significant association with cluster headache was found (odds ratio 0.91 (95% confidence intervals 0.75–1.10), p = 0.319). In contrast, the meta-analysis that included in total 1167 cluster headache cases and 1618 controls from the six study populations, which were part of four different studies, showed association of the single nucleotide polymorphism with cluster headache (random effect odds ratio 0.69 (95% confidence intervals 0.53–0.90), p = 0.006). The association became weaker, as the odds ratio increased to 0.80, when the meta-analysis was repeated without the initial single South European study with the largest effect size.
Conclusions
Although we did not find evidence for association of rs2653349 in our LUCA study, which is the largest investigated study population thus far, our meta-analysis provides genetic evidence for a role of HCRTR2 in cluster headache. Regardless, we feel that the association should be interpreted with caution as meta-analyses with individual populations that have limited power have diminished validity.
Introduction
Cluster headache is a primary headache disorder of largely unknown aetiology that is characterized by severe, short-lasting headache attacks accompanied by ipsilateral facial autonomic symptoms and/or restlessness occurring up to eight times per day (1,2). A role for genetic factors in the aetiology of cluster headache was overlooked for a long time, but has been considered in several studies since the early 1990s (3–8). Four family studies showed that relatives of cluster headache patients have a higher disease risk (9–12), but estimates of the relative risk varied considerably between studies, ranging from 14 to 45 (9–11,13). As cluster headache is not as rare as previously thought, it is likely that the disease risk in these studies may have been overestimated (11). In addition, one of the studies with the highest estimated relative risk (9) may have overestimated the occurrence of cluster headache by partly using heteroanamnestic information instead of direct interview or questionnaire data from all affected relatives. Recalculation of the relative risk using a cluster headache prevalence of 0.2% showed a relative risk of 5–18 for first-degree relatives and 1–3 for second-degree relatives (14), suggesting a considerably smaller but still relevant contribution of genetic factors in cluster headache (11). Case reports (3–7,15) on monozygotic twins, both affected by cluster headache, provide further support for a role of genetic factors in the disease. Complex segregation analysis suggested that a low-penetrant autosomal dominant genetic risk factor may play a role in a small subset of patients (16). Most likely, cluster headache is a complex disorder caused by both genetic and environmental factors (14).
Several genetic studies aimed to identify genes involved in cluster headache but were unsuccessful (17–20). The role of the CACNA1A gene was investigated both in a kindred with three affected family members (17) as well as in an association study (18) but results were negative. Single nucleotide polymorphism (SNP) rs1801133 in MTHFR showed no association with cluster headache either (19). A mitochondrial mutation reported to cause mitochondrial encephalopathy, lactic acidosis, and stroke-like episodes (MELAS) was identified in a single cluster headache patient without a family history of this disorder (20), but later studies (21,22) failed to replicate the involvement of mitochondrial mutations in cluster headache. Thus far, genetic research in cluster headache has only led to the identification of one replicated possible genetic susceptibility factor: SNP rs2653349 (G1246A) in HCRTR2 that encodes the hypocretin type 2 receptor (23–25). This receptor is expressed in the posterior hypothalamus, which is thought to play an important role in cluster headache (26). The role of the HCRTR2 SNP was investigated in five small cohorts in three studies (23–25), comprising in total 593 cases and 599 healthy controls. The minor A allele of SNP rs2653349 was associated with a reduced risk for cluster headache (i.e. the A allele being more frequent in controls than in cases). Rainero et al. performed a meta-analysis of all five cohorts and reported association of the HCRTR2 SNP with cluster headache (27). However, the accuracy of the estimated effect size differed largely between studies and the meta-analysis is, therefore, difficult to interpret because: (i) there were differences in the statistical models used (i.e. other models than the additive model that is thought to underlie genetic susceptibility of most complex disorders (28)); (ii) there was considerable statistical heterogeneity between the studies.
Using our web-based Leiden University Cluster Headache Analysis (LUCA) study population (29), we re-evaluated the possible association of HCRTR2 SNP rs2653349 with cluster headache in the largest single study thus far and we subsequently performed new meta-analyses combining our results with those of previous studies.
Materials and methods
Patient recruitment for the LUCA study
For the LUCA study, self-reported cluster headache patients aged 18 years or older were recruited via our Dutch headache research website (www.lumc.nl/hoofdpijn), which was developed for genetic and epidemiological research on primary headache disorders. Individual diagnoses were established using an extended web-based questionnaire according to the International Classification of Headache Disorders, 2nd edition (1) and were validated in a subset of patients by a telephone interview (29). The medical ethics committee of the Leiden University Medical Centre approved our study and all participants provided written informed consent.
Genetic association study in the LUCA population
All participants who met our algorithm criteria (29) for cluster headache and had provided a DNA sample were included. Our control population consisted of anonymous blood donors, of whom no specific health information is available. Based on the low prevalence (0.2%) of cluster headache in the general population, we expected a negligible effect of possibly including one or two patients with cluster headache in our control group. Power calculations were performed using the Genetic Power Calculator (http://pngu.mgh.harvard.edu/∼purcell/gpc/)(30), under the assumption of an additive model..
Genotyping of SNP rs2653349 (i.e. DNA variant G1246A) was performed using a TaqMan assay, for which probes and primers were designed by Applied Biosystems. A standard polymerase chain reaction (PCR) was carried out using the TaqMan Universal PCR Master Mix reagent and the genotyping was performed on a Lightcycler LC-480 machine combined with LightCycler®480 1.5.0 software, version 1.5.0.39 (Roche Applied Science) to analyse the genotype clusters. Data analysis was performed using PLINK software version 1.07 (http://pngu.mgh.harvard.edu/∼purcell/plink/) by calculating odds ratios (ORs) and 95% confidence intervals (95% CI) for the association between the HCRTR2 SNP rs2653349 and cluster headache.. We assumed an additive genetic model and performed uncorrected and corrected analyses with age and sex as covariates. Association analysis was performed with the major allele (G) as reference. A significance level of 0.05 was used.
Meta-analysis
Study selection
According to the guidelines for systematic reviews of genetic association studies (31), two researchers (C.M.W. and L.A.W.) individually searched the literature for genetic studies on the role of HCRTR2 in cluster headache (see online Supplementary Material for details on search strategy and inclusion criteria). In brief, all relevant titles and abstracts were screened. Subsequently, the manuscripts of all studies meeting the predefined criteria were evaluated and studies not meeting the criteria were excluded. In case of disagreement, a third researcher (B.d.V.) was consulted.
Statistical analysis
Meta-analysis of all study populations was performed in R version 2.15 using the metagen function from the meta package. We assumed an additive genetic model and modelled the OR both as fixed effect, which assumes homogeneity across studies, and as random effect, which incorporates the between-study variability. The random effect model was defined as our primary model because of the high inter-study variability with regard to sample sizes and estimated effect sizes. The between-study heterogeneity was tested using the I2 statistic, a Cochrane’s Q statistic-based measure that describes the amount of variation due to heterogeneity rather than chance on a continuous scale from 0 to 1. I2 values of approximately 25%, 50% and 75% are indicative for low, moderate and high between-study heterogeneity, respectively (32). A funnel plot was generated using SPSS Statistics 20.0 (IBM, Armonk, NY, USA) to check for publication bias. All analyses were performed with the major allele (G) as reference and a significance level of 0.05 was used.
Results
Study populations included in the meta-analysis.
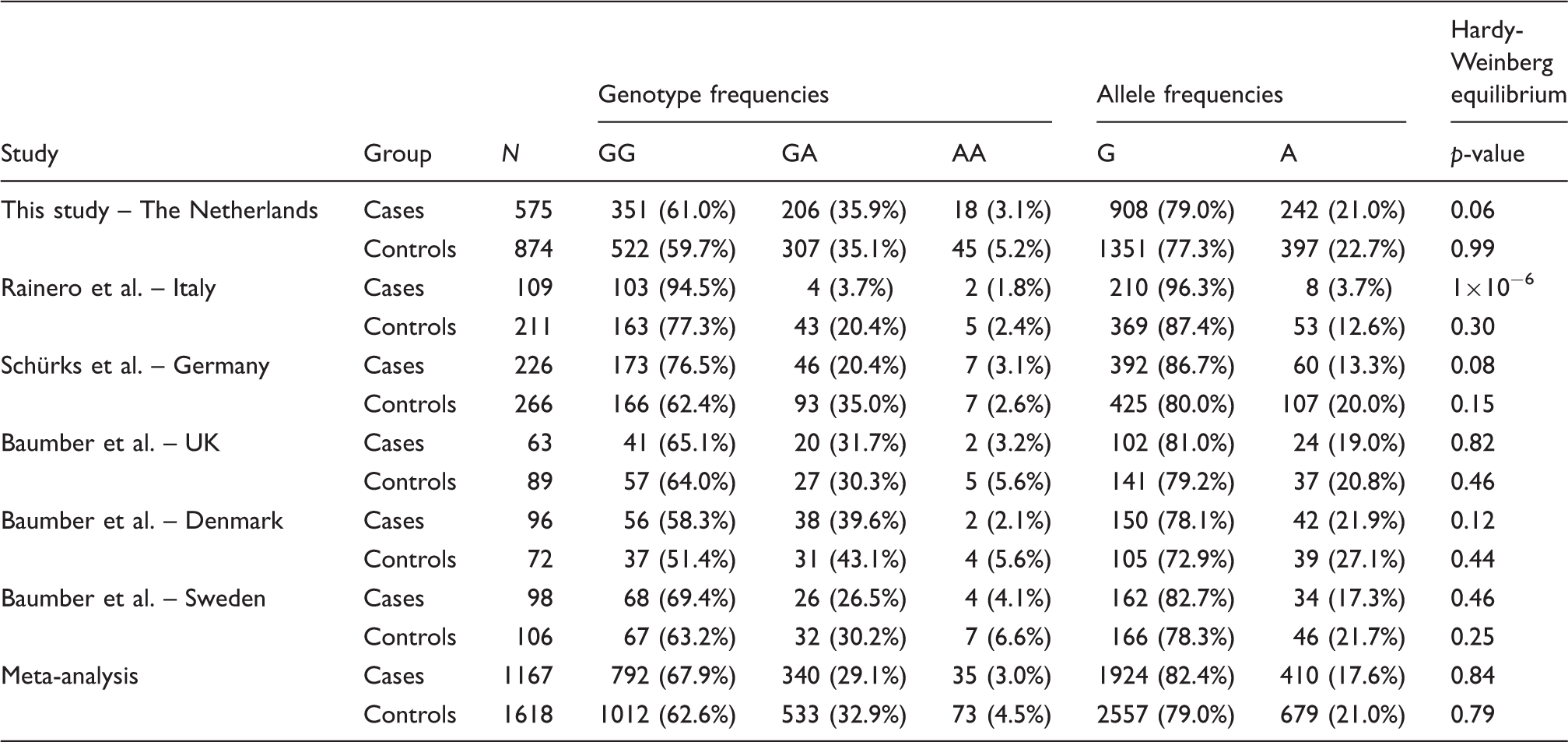
Subsequently, a meta-analysis was performed of three previously published studies (which included five study populations) and our own study (23–25) (see also online Supplementary Material). These studies comprise in total 1167 cluster headache patients and 1618 control subjects. Genotype and allele frequencies of the various study populations are shown in Table 1. Genotype frequencies in the controls were in Hardy-Weinberg equilibrium in all studies. Our meta-analysis, assuming an additive model, replicated the previously reported association between the A allele of the HCRTR2 SNP and a decreased risk of cluster headache (random effect OR 0.69 (95% CI 0.53–0.90), p = 0.006) (Figure 1)). The results of our meta-analysis indicated the presence of low to moderate between-study heterogeneity (I2 = 0.57, p = 0.042). Notably, the funnel plot graphically demonstrates asymmetry, indicating a publication bias favouring studies with an OR significantly deviating from 1, mainly caused by the initial study from Rainero et al. (23) (see Figure S1 in online Supplementary Material). That study population is the only study from Southern Europe with allele frequencies that differ greatly from the other investigated populations, which were all from Northern Europe. Remarkably, the absolute number of A alleles (8/218 alleles) among the cluster headache cases in the study population from Rainero et al. is very low compared to the other populations, which makes it difficult to get reliable estimates of the allele frequency. We, therefore, performed an additional meta-analysis that contained only the populations from Northern Europe, so combining only data of Danish, Swedish, British, German and Dutch populations. The heterogeneity across these studies was negligible (I2 = 0%, p = 0.437); hence, the random effect estimates correspond to the fixed effect estimates (OR 0.80 (95% CI 0.70–0.93), p = 0.0031). Although the effect size decreased, the association remained.
Forest plot representing the results of the meta-analysis of the rs2653349 polymorphism of the HCRTR2 gene assuming an additive model.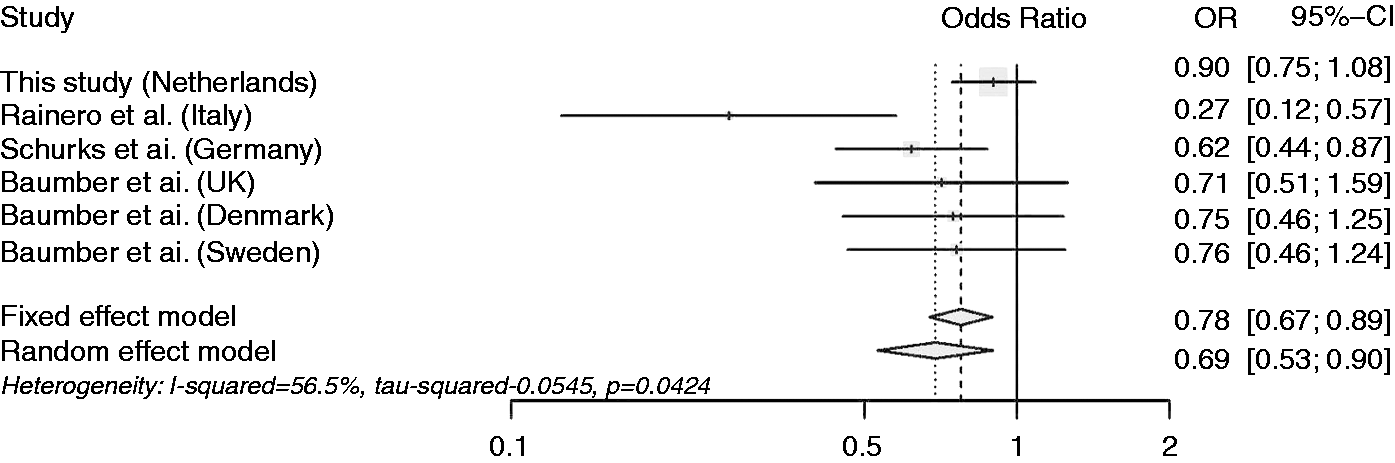
Discussion
Genetic factors have been implicated in cluster headache and several candidate gene association studies have been performed (17–21,23–25), which aimed at identifying such factors. However, most studies did not produce convincing results, except for the genetic association with SNP rs2653349, which is located in the HCRTR2 gene, that showed association for the (protective) A allele when using additive models. We decided to re-evaluate the association with this SNP by performing: (i) a genetic association study in our LUCA population: (ii) meta-analyses of data from the LUCA study with data from earlier studies.
Association with cluster headache in the study populations and the meta-analysis (additive model).
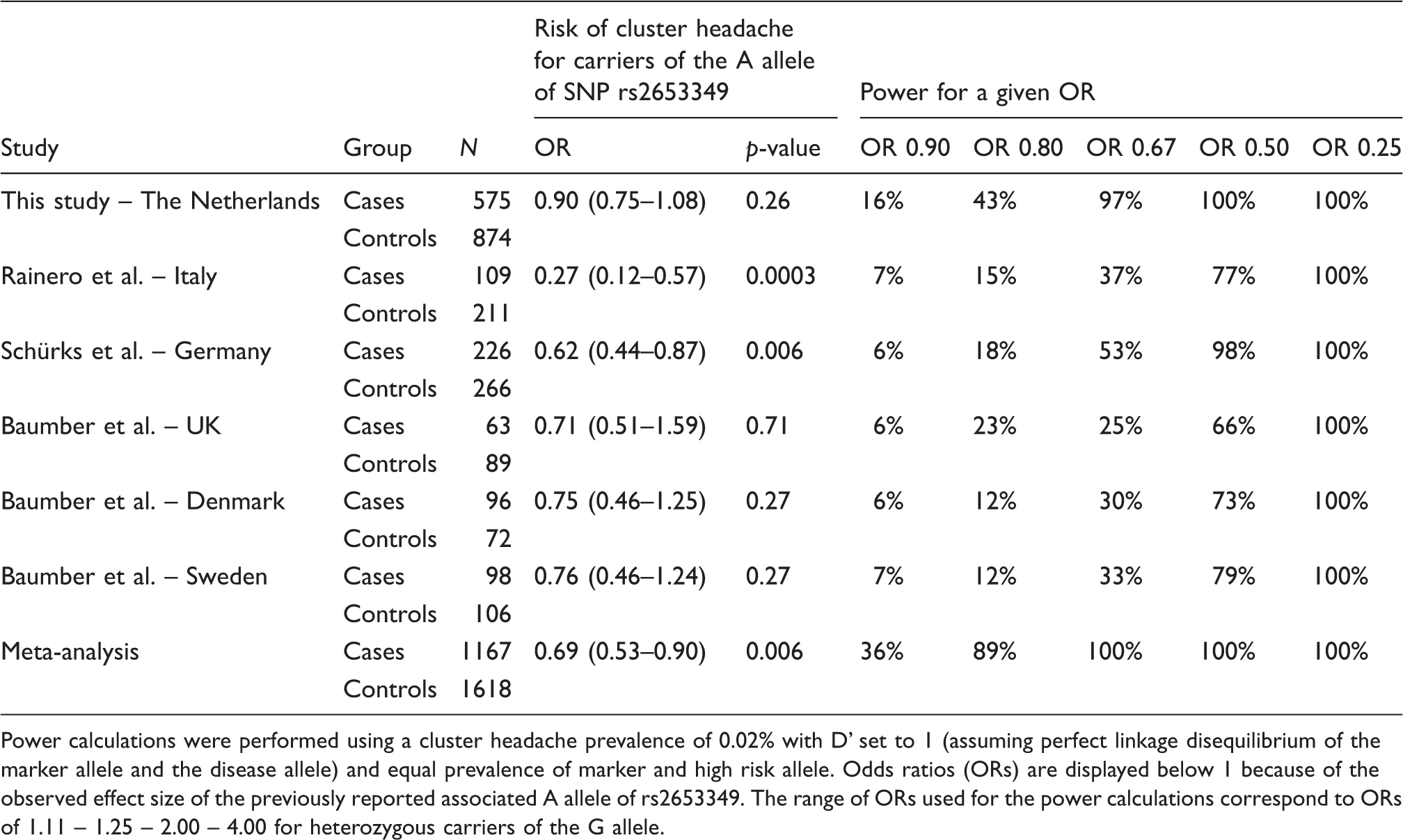
Power calculations were performed using a cluster headache prevalence of 0.02% with D’ set to 1 (assuming perfect linkage disequilibrium of the marker allele and the disease allele) and equal prevalence of marker and high risk allele. Odds ratios (ORs) are displayed below 1 because of the observed effect size of the previously reported associated A allele of rs2653349. The range of ORs used for the power calculations correspond to ORs of 1.11 – 1.25 – 2.00 – 4.00 for heterozygous carriers of the G allele.
In conclusion, our LUCA study and subsequent meta-analysis illustrate that even doubling the sample size does not lead to a definite conclusion on the role of the HCRTR2 gene in cluster headaches. We demonstrated that such a role (if any) is likely smaller than previously reported. Future genetic studies in cluster headache patients need to include much larger numbers of patients and controls and may benefit also from focusing on hypothesis-free gene variant discovery, as is the case with genome-wide association studies (GWAS).
Article highlights
No significant association was observed in our LUCA study, which is the largest genetic study to date. Although no association was observed in our LUCA study, a meta-analysis containing also the LUCA population still showed association between SNP rs2653349 and cluster headache. Our meta-analysis results should be interpreted with caution, as we would argue that meta-analyses with individual populations that have limited power have diminished validity. The role of the HCRTR2 gene in cluster headache is likely smaller than previously reported.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
Conflict of interest
Drs De Vries, Haan, Van den Maagdenberg, Weller, as well as Ms. Koelewijn and Ms. Vijfhuizen report no disclosures. Dr. Wilbrink received industry support from Medtronic, Menarini, Allergan, Electrocore and independent support from Fonds Nuts Ohra. Dr. Ferrari reports grants and consultancy or industry support from Medtronic, Menarini, and Amgen and independent support from NWO, ZonMW, NIH, European Community, and the Dutch Heart Foundation. Dr. Terwindt reports grants and consultancy/industry support from Menarini, and independent support from NWO.
Funding
This study is funded by grants of the Netherlands Organization for Scientific Research (NWO) (903–52–291, M.D.F.; Vici 918.56.602, M.D.F.; 907–00–217, G.M.T.; Vidi 917–11–319, G.M.T.), the European Community (EC) (EUROHEAD, LSHM-CT-2004–504837, M.D.F. and A.M.J.M.v.d.M; EUROHEADPAIN nr 602633, M.D.F. and A.M.J.M.v.d.M), and the Centre for Medical Systems Biology (CMSB) and Netherlands Consortium for Systems Biology (NCSB), both within the framework of the Netherlands Genomics Initiative (NGI) (A.M.J.M.v.d.M). The funding agencies had no role in the design or conduct of the study. We are grateful to all study participants and their relatives, general practitioners and neurologists for their contributions.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
