Abstract
In October 2020, adult male and female NSG (NOD. Cg-Prkdc scid Il2rg tm1Wjl /Sz) mice were reported for diarrhea within a mouse barrier facility. Other immunodeficient strains harboring the SCID (Prkdc scid ) or Rag (Rag null ) mutations together with the IL2rg (Il2rg null ) mutation were affected. At its peak, over 20 laboratories in 10/16 (62.5%) barrier rooms were affected. Mortality was rare except in lactating females (≥ P11). Grossly, nonlactating adult female and male mice (n = 16) had mild to moderate, small and large intestinal distension with corresponding individual cell death and sloughing of superficial enterocytes in the cecocolonic mucosa. Lactating NSG dams (n=6) had moderate to severe gastrointestinal distension and/or segmental, dark red to gray, small intestinal discoloration. In addition to the same histologic lesions seen in nonlactating female NSG mice, lactating NSG dams often had severe ulcerative inflammation affecting the jejunum, ileum, cecum, and colon. Traditional ancillary diagnostic tests including aerobic and anaerobic cultures (blood, liver, spleen, and intestines), fecal PCR, and fecal floatation failed to yield a causative organism. Further cohousing and oral gavage studies determined neither immunocompetent CD1 (Crl:CD1 [ICR]) mice nor immunodeficient NOD scid (NOD.Cg-Prkdc scid /J) and Rag2 KO (C57BL/6. Cg-Rag2 tm1.1Cgn /J) mice were susceptible to clinical disease. Extensive control barriers were implemented including a veterinary-managed NSG breeding barrier, alterations in husbandry practices, and strategic environmental disinfection, allowing for continuity of experimental studies while avoiding widespread depopulation of the barrier. Subsequent strain-resolved metagenomics and qPCR assay development identified Clostridium cuniculi and its enterotoxin exclusively within diarrheic mice.
Immunodeficient mouse strains have become a cornerstone in biomedical research. The various strategies for murine immune modulation have greatly expanded in recent years, which is partially driven by the demand for optimization of human tissue engraftment. The wide range of available immunodeficient strains has contributed immensely to the success and furtherment of cancer research, while simultaneously introducing the comparative medicine and pathology communities to novel strain-specific and engraftment-related background lesions.6,9,12,25–27
Successful housing and breeding of immunodeficient mice requires high biosecurity barrier facilities and practices, extensive rodent health surveillance, and diligent evaluation of morbidities and mortalities. Despite these efforts, opportunistic infections have become an inevitable part of managing and maintaining immunodeficient mouse strains.10,19,23,28,29
Within recent years, reports of diarrheal disease and mortality events amongst immune compromised strains have risen. In some outbreaks, etiologic agents have been identified and defined, allowing for clinical intervention, containment, and/or depopulation, when necessary.15,22,24 In other instances, despite extensive evaluation, identification of an etiopathologic agent remains unknown or, at best, correlative in nature.3,17 One key challenge in determining the causation of intestinal disease in immunodeficient mice lies in recognizing that microbial flora, which pose minimal concern in immunocompetent mice, may result in intestinal pathology due to proportional shifts in the microbiome. 3 As a result, traditional techniques for identifying etiologic agents (e.g. microbiological culture and PCR for known murine pathogens) may no longer be sufficient.
Herein we describe the epidemiology, diagnostic work-up, and clinical containment of a chronic diarrheal disease of adult male and female NSG (NOD. Cg-Prkdc scid Il2rg tm1Wjl /Szj) and NSG-related mice (high morbidity, low mortality) and lactating NSG and NSG-related dams (high morbidity, high mortality) impacting multiple rooms within our highest biosecurity facility. While conventional methods of pathogen detection initially failed to identify a causative agent, a multifaceted approach to biosecurity practices allowed for containment of disease without the need for widespread depopulation of the barrier. Subsequent strain-resolved metagenomic analysis and qPCR assays identified Clostridium cuniculi and its enterotoxin exclusively within diarrheic mice. 5 This report highlights effective strategies for disease mitigation in the context of an unknown etiologic agent and underscores the need for alternative diagnostic approaches in the context of immunocompromised mice.
Materials and Methods
Ethics Statement
All procedures were approved by the Stanford University Institutional Animal Care and Use Committee (Administrative Panel for Laboratory Animal Care) under protocols #10082 and #34205. Mice were maintained in an AAALAC-accredited institution in accordance with The Guide for the Care and Use of Laboratory Animals. 18
Mice and Barrier Husbandry Practices
NSG, Rag2 KO (C57BL/6.Cg-Rag2 tm1.1Cgn /J), and NOD scid (NOD.Cg-Prkdc scid /J) immunodeficient mice were purchased from The Jackson Laboratory (Sacramento, CA). The B6 Rag2 IL2rg double knockout (DKO; C57BL/6NTac. Cg-Rag2 tm1Fwa Il2rg tm1Wjl /Tac) mice were purchased from Taconic (Seattle, Washington and Cambridge City, Indiana) and CD-1 (Crl:CD1(ICR)) mice were purchased from Charles River Laboratories (Hollister, CA).
The mouse barrier facility was opened in 2010, and from its inception all mice were housed in Innovive irradiated, individually ventilated cages prefilled with 1/8″ corncob bedding (20 kGy–40kGy, Innocage, Innovive, San Diego, CA). Mice were fed an 18% protein, dye-free, irradiated rodent diet (20 kGy–50 kGy, 2918, Teklad, Tampa, FL) and provided with acidified (pH 2.5–3.0) bottled water (Aquavive; Innovive, San Diego, CA). All cages received enrichment consisting of irradiated, crinkled paper strips (10 kGy–40 kGy, Enviro-DriR, Shepherd Specialty Papers, Watertown, TN).
Only mice from approved vendors and directly following embryo transfer rederivation were allowed into the barrier. The excluded pathogen list includes betacoronavirus muris (mouse coronavirus), protoparvovirus rodent 1 (mouse parvoviruses), murine norovirus 1, murine rotavirus-A (murine rotavirus), cardiovirus theileri (mouse theilovirus), mammarenavirus choriomeningitidis (lymphocytic choriomeningitis virus), respirovirus muris (murine respirovirus), mastadenovirus encephalomyelitidis (mouse adenovirus 1 and 2), orthopoxvirus ectromelia (ectromelia), orthopneumovirus muris (murine orthopneumovirus), orthoreovirus mammalis (reovirus 3), K virus, gammaarterivirus lacdeh (lactate dehydrogenase-elevating virus), muromegalovirus muridbeta1 (mouse cytomegalovirus), roseolovirus muridbeta3 (mouse thymic virus), mouse polyomavirus, chaphamaparvovirus rodent1 (rodent chaphamaparvovirus 1), Mycoplasma pulmonis, Citrobacter rodentium, Clostridium piliforme, Corynebacterium bovis, Corynebacterium kutscheri, Filobacterium rodentium (CAR bacillus), Pasteurella multocida, Rodentibacter spp., Salmonella spp., Streptococcus pneumoniae, Helicobacter spp., Encephalitozoon cuniculi, Pneumocystis murina, Chlamydia muridarum, Spironucleus muris, Giardia muris, fur mites, lice, cestodes, and pinworms. Health surveillance was accomplished via quarterly testing of dirty bedding sentinels (CD-1 females; 1 cage per rack).
Within the barrier, 2 rooms were used exclusively for breeding. Both rooms operated under restricted access and were reserved for strains designated as highly valuable by individual research labs. In 1 breeding room, entry was restricted to colony managers within the Veterinary Service Center (VSC), while the other breeding room (Principal Investigator breeding room) was open to any lab, but colony management and room entry were restricted to 2 personnel per lab. Mice in both breeding rooms could enter only if delivered directly from an approved vendor or directly following rederivation. Mice from other barrier rooms were not allowed to enter the breeding barrier rooms regardless of their origin. Once mice left the breeding barrier, they were not allowed to return. See Supplemental Material S1 for detailed barrier husbandry practices.
Necropsy and Histology
Affected mice submitted for necropsy (n = 22) included diarrheic, moribund, or found deceased lactating NSG dams (n = 6) and diarrheic, nonlactating, adult female and male NSG mice (n = 16). Live mice were euthanized via carbon dioxide (CO2) inhalation in accordance with the American Veterinary Medical Association Guidelines for Euthanasia of Animals. 4 Following necropsy and ancillary diagnostic sampling, tissues were immersion-fixed in 10% neutral-buffered formalin for 72 hours. Formalin-fixed tissues were processed routinely, embedded in paraffin, sectioned at 5 µm, and stained with hematoxylin and eosin. The following organs were evaluated histologically: heart, lung, liver, kidneys, spleen, brain, adrenal glands, salivary glands, esophagus, trachea, urinary bladder, haired skin, reproductive tracts, and gastrointestinal tracts. In selected mice, elaborated microscopic evaluation of the gastrointestinal tracts included longitudinal sections of the stomach (cardia to pylorus), duodenum, jejunum, and ileum. Both halves of longitudinally bisected cecum were evaluated in addition to proximal, middle, and distal colon.
Bacteriology and Parasitology
Cultures were performed in the Stanford University VSC Animal Diagnostic Laboratory. For ease of reporting, blood agar, MacConkey agar, XLD agar, Yersinia selective agar, Campylobacter selective medium agar, chocolate agar medium, Signal blood culture bottles, and the RapID ANA II System were purchased from Remel (Lenexa, KS). Brucella blood agar, phenylethyl alcohol blood agar, and Bacteroides bile esculin and Laked kanamycin vancomycin agar bi-plates were purchased from Anaerobe Systems (Morgan Hill, CA). Enteric cultures were performed on feces from 18 mice (n = 12 diarrheic mice, n = 6 control mice). Samples were streaked to the following culture plates: blood agar, MacConkey agar, XLD agar, Yersinia selective agar, and Campylobacter selective medium agar.
Blood agar, MacConkey agar, XLD agar, and Yersinia selective agar plates were incubated in non-CO2 conditions at 36°C ± 1°C or 22°C–25°C (only Yersinia selective agar plates). These plates were examined daily for up to 3 days. Campylobacter selective medium agar plates were incubated in a microaerophilic environment using the GasPak EZ CampyPouch system (BD, Franklin Lakes, NJ) in non-CO2 conditions at 42°C ± 1°C and were read at 72 hours of incubation.
Aerobic cultures of the liver (n = 7 diarrheic, n = 6 naive) and spleen (n = 7 diarrheic, n = 6 naive) were streaked onto chocolate agar, blood agar, and MacConkey agar plates along with inoculation into a thioglycollate broth media (Hardy Diagnostics, Santa Maria, CA). All plates were incubated in 5% CO2 at 36°C ± 1°C. Thioglycollate broth media were incubated in non-CO2 conditions at 36°C ± 1°C and were visually examined daily for growth. In the presence of growth, cultures were subcultured onto a chocolate agar plate and incubated in aerobic conditions (5% CO2 at 36°C ± 1°C) and onto Brucella blood agar plates, which were incubated in anaerobic conditions (non-CO2 at 36°C ± 1°C) using the GasPak EZ Anaerobe Pouch system (BD, Franklin Lakes, NJ). All aerobic culture plates were observed for growth daily for 5 days.
Anaerobic cultures of the liver (n = 7 diarrheic, n = 6 naive) and spleen (n = 7 diarrheic, n = 6 naive) were streaked onto preprepared media, including Brucella blood agar, phenylethyl alcohol blood agar, and Bacteroides bile esculin and Laked kanamycin vancomycin agar bi-plates and incubated under non-CO2 conditions at 36°C ± 1°C. Initial readings were performed after 48 hours with a final reading after an additional 3 days of incubation. All growing bacteria were subcultured under both aerobic (challenge) and anaerobic conditions. Anaerobic bacteria—those growing exclusively under anaerobic conditions—were further streaked onto blood agar and incubated anaerobically for additional testing and identification. Identification of anaerobic bacteria was performed using the RapID ANA II System or the Biolog MicroStation (Biolog, Hayward, CA).
Blood cultures (n = 6 diarrheic) were collected in broth media (Signal blood culture bottles) under aseptic conditions. Bottles were incubated at 36°C ± 1°C with gentle agitation periodically during the first 24 hours. The signal device was placed onto the blood culture bottle after 1 hour of incubation in a 5% CO2 atmosphere at 36°C ± 1°C. Blind subculturing of blood cultures was performed after 18 to 24 hours of incubation and then daily for 5 days. Each blood culture was subcultured onto chocolate agar medium, blood agar, and MacConkey agar, as well as onto anaerobic media, including Brucella blood agar, phenylethyl alcohol blood agar, and Bacteroides bile esculin and Laked kanamycin vancomycin agar bi-plates. A Gram stain was performed with each subculture.
Isolated organisms were evaluated through Gram staining, colony morphology, and biochemical tests, including the spot indole reaction, oxidase test, and catalase test. All aerobic organisms were identified using the OmniLog ID system (Biolog, Hayward, CA) within the Stanford University VSC Animal Diagnostic Laboratory. Organisms not identified via OmniLog were sent out to the University of California Davis Veterinary Medical Teaching Hospital Clinical Diagnostic Laboratory for identification using the MALDI-TOF ID system (Bruker, Billerica, MA).
Pooled small and large intestinal contents from diarrheic mice (n = 3) were submitted to the California Animal Health and Food Safety Laboratory (CAHFS, San Bernardino, CA) for repeat anaerobic cultures and Clostridium difficile anaerobic culture.
Fecal samples were collected from diarrheic (n = 6) and naive (n = 6) NSG mice, stored in sodium nitrate solution and subjected to fecal flotation tests following a standard protocol.
Molecular Screening
Fecal samples from diarrheic mice (n = 9), naive mice (n = 6), and pooled samples from pups (n = 3) cohoused with a found deceased dam were submitted to Charles River Laboratories for the Charles River Laboratories Mouse Surveillance Plus PRIA 64+ (Hanta virus, K virus, lymphocytic choriomeningitis virus, lactate dehydrogenase-elevating virus, mouse adenovirus 1 & 2, mouse cytomegalovirus, mouse coronavirus, murine norovirus, ectromelia virus, mouse parvoviruses, minute virus of mice, murine rotavirus (epizootic diarrhea of infant mice), mouse thymic virus, mouse polyomavirus, murine orthopneumovirus, reovirus (types 1–4), murine respirovirus, mouse theilovirus, ß-haemolytic Streptococcus (group A, B, C, G), Bordetella bronchiseptica, Bordetella pseudohinzii, Campylobacter spp., Citrobacter rodentium, Clostridium piliforme, Corynebacterium bovis, Corynebacterium kutscheri, Filobacterium rodentium (CAR bacillus), Helicobacter spp., Klebsiella oxytoca, Klebsiella pneumoniae, Mycoplasma pulmonis, Rodentibacter heylii, Rodentibacter pneumotropicus, Proteus mirabilis, Pseudomonas aeruginosa, Salmonella spp., Staphylococcus aureus, Streptobacillus moniliformis, Streptococcus pneumoniae, Cryptosporidium, Demodex, Encephalitozoon cuniculi, Entamoeba, Giardia, mites, pinworms, Pneumocystis, Spironucleus muris, Tritrichomonas, Chilomastix muris, Hexamastix muris, Ornithonyssus bacoti), as well as additional PCR assays for astrovirus-1, Clostridioides difficile, rodent chaphamaparvovirus 1, and Leptospira spp. (Charles River Laboratories, Wilmington MA). Molecular screening for Chlamydia muridarum was not performed on these cohorts as the PCR assay was not commercially available in 2020 and 2021. All barrier rooms were subsequently screened for Chlamydia muridarum in 2022 and were uniformly negative.
Clostridium perfringens and C. difficile Toxins Enzyme-Linked Immunosorbent Assay (ELISA)
Pooled small and large intestinal contents from diarrheic mice (n = 3) were tested for C. perfringens organisms and toxins (alpha (CPA), beta (CPB), and epsilon (ETX)) using a commercial capture ELISA kit (BIO-X, Brussels, Belgium) and for C. difficile toxins (A and B) using a commercial ELISA kit (Techlab, Blacksburg, Virginia). All procedures were conducted according to the manufacturer’s instructions.
Transmission Electron Microscopy (TEM)
For cellular and subcellular examination, samples were submitted to the Ultrastructural Pathology Service at CAHFS (University of California, Davis) for virus screening and tissue ultrastructure.
Virus screening
A sample of pooled small and large intestinal contents was collected from diarrheic mice (n = 3) and submitted for viral detection, identification, and quantification. Virus particles were extracted, cleaned, concentrated, and negatively contrasted using phosphotungstic acid as described elsewhere. 2
Tissue ultrastructure
Cecal tissue from a diarrheic female NSG mouse was collected for TEM. Briefly, following euthanasia, the cecum was isolated and immediately flushed with warm saline (37°C). Cecal tissue was immersion fixed in Karnovsky’s Fixative EM Grade (Electron Microscopy Sciences, Hatfield, PA) for 72 hours at 4°C. A section of the cecal tissue was first routinely processed and stained with hematoxylin and eosin to confirm the presence of apically adherent bacteria. A mirror image section of cecal tissue was processed in automate tissue equipment, sectioned, and contrasted as described elsewhere. 2 All samples were visualized using a JEOL 1400Plus transmission electron microscope (JEOL LTD, Peabody, MA). Images were obtained and analyzed using a OneView Camera Model 1095 with the Gatan Microscope Suite 3.0 (Gatan Inc, Pleasanton, CA).
Immunohistochemistry
Immunohistochemistry for C. perfringens was performed on formalin-fixed, paraffin-embedded 4-μm-thick sections of small and large intestines from an 8-week-old diarrheic NSG mouse, an 8-week-old control NSG mouse, and a diarrheic lactating dam. Antigen retrieval was performed using pepsin. A rabbit polyclonal anti–C. perfringens antibody (GenWay Bio, San Diego, California, 1:100) was utilized. Visualization of C. perfringens was obtained using the Dako EnVision Kit (Dako, Carpenteria, California). Positive controls included intestinal tissues of a goat in which C. perfringens had been detected by culture. Negative controls were sections of the case mice inoculated with rabbit nonimmune serum instead of the anti-C. perfringens serum.
Control Mice
A control group of naive, healthy, 7 to 8-week-old NSG mice (n = 6, 3 male and 3 female) were purchased from The Jackson Laboratory (Sacramento, CA) in August 2021. Upon arrival at our facility, they were immediately euthanized via CO2 inhalation and terminal cardiac exsanguination. Skin swabs were collected for ectoparasite evaluation (see Molecular Screening). Sterile samples of liver and spleen were collected for aerobic and anaerobic cultures (see Bacteriology and Parasitology). Fecal pellets were collected for 1) sterile aerobic and anaerobic culture (see Bacteriology and Parasitology), 2) protozoal fecal floatation (see Bacteriology and Parasitology), and 3) enteric PCR panels (see Molecular Screening). All soft tissues were collected and processed routinely for histology (see Necropsy and Histology).
As part of a separate disease investigation, 3 a control group of naïve, healthy, NSG (n = 4, 2 males and 2 females) and NSG-SGM3 (NOD. Cg-Prkdc scid Il2rg tm1Wjl Tg(CMV-IL3, CSF2, KITLG)1Eav/MloySzJ; n = 4, 2 males and 2 females), were purchased from The Jackson Laboratory (Sacramento, CA) in January 2023. Upon arrival at our facility, they were immediately euthanized via CO2 inhalation and samples were collected, pooled by sex and strain, and subjected to aerobic and anaerobic cultures as well as C. perfringens and C. difficile toxin ELISA assays to screen for presence of organisms and/or toxins (C. perfringens and its toxins [CPB and ETX], C. difficile toxins A and B), which were performed as described above at CAHFS (University of California, San Bernardino).
Investigations of Diarrhea Transmissibility
Mice
Prior to development and validation of our novel qPCR assays, cohousing and gavage experiments were performed to better understand the clinical susceptibility of various mouse strains, allowing for real-time evidence-based decisions regarding husbandry and barrier practices. Immunocompetent CD-1 and immunodeficient NOD scid, Rag2 KO, and B6 Rag2 IL2rg DKO mice were either cohoused with diarrheic NSG mice (CD-1) or gavaged with intestinal homogenate prepared from nondiarrheic (control) or diarrheic NSG mice (described below).
Intestinal homogenates
Intestinal homogenates from nondiarrheic (control) or diarrheic NSG mice were prepared in sterile saline from the entirety of the small intestine, cecum, and large intestine. Homogenization was done in batches using a bead mill homogenizer (BeadBug, Benchmark Scientific, Sayreville, NJ). 5 Homogenates were pooled, centrifuged (5000 relative centrifugal force [RCF], 2 min at room temperature) to pellet large fragments, and the supernatants were added to a 15 ml tube on ice. Additional saline was added to the homogenization tube. The homogenization was repeated once and the supernatants were pooled. Gavage was performed with a polypropylene feeding tube (FTP-20-38, InsTech, Plymouth Meeting, PA). The intestinal homogenates were stored at −80°C (without freezing medium) or used immediately.
Cohousing of CD-1 mice with diarrheic NSG mice
Five immunocompetent, female CD-1 mice were cohoused with diarrheic NSG mice for several weeks as contact sentinels. Serum and fecal samples from CD-1 mice were submitted to Charles River Laboratories for pathogen detection (see Molecular Screening). The CD-1 mice were then moved to individual clean cages to monitor for evidence of diarrhea. Five naïve NSG females were then placed (1 mouse per cage) with the previously exposed CD-1 mice and cohoused for several weeks to determine if the CD-1 mice were subclinically infected.
Oral gavage of NOD scid, Rag2 KO, and B6 Rag2 IL2rg DKO with intestinal homogenate
Five B6 Rag2 IL2rg DKO females, 5 NOD scid females, and 5 Rag2 KO females were gavaged with 0.5 ml of either nondiarrheic intestinal homogenate (n = 2 mice per strain) or diarrheic intestinal homogenate (n = 3 mice per strain). Intestinal homogenates had previously been stored at −80°C. Five NSG females were treated similarly as a positive control. Mice gavaged with nondiarrheic intestinal homogenate were housed by strain separately from mice gavaged with diarrheic intestinal homogenate to avoid cross-contamination.
Another cohort of three naive male NSG, and the same mice from the three strains (B6 Rag2 IL2rg DKO, NOD scid, and Rag2 KO) were gavaged again with 0.5 ml of freshly prepared (nonfrozen) diarrheic intestinal homogenate. To test if the mice were subclinically infected, 2 naïve NSG females were added to the cages housing each strain for up to 14 days.
Identification of C. cuniculi Encoding an Enterotoxin as a Candidate Etiologic agent
Strain-resolved metagenomics using DNA from diarrheic and control fecal samples identified the genome sequence of an enterotoxin-encoding C. cuniculi as a candidate etiologic agent. A proprietary quantitative PCR (qPCR) assay was developed utilizing primers specific to a unique sequence of C. cuniculi and its enterotoxin. 5 A total of 176 NSG or NSG-related strains were screened using the novel qPCR assays. Of these, 37 were NSG or NSG-related mice with clinical signs of diarrhea within the context of this outbreak. The remaining 139 mice were either healthy, nondiarrheic NSG or NSG-related mice (n = 111), or NSG and NSG-related mice with documented alternative causes for clinical signs of diarrhea (n = 28).
Targeted microbiological culture of C. cuniculi and enterotoxin qPCR-positive samples
Pooled cecal and large intestinal samples from diarrheic mice (n = 3), confirmed positive for C. cuniculi and its enterotoxin by qPCR, were submitted to CAHFS (University of California, San Bernardino) for attempted C. cuniculi isolation. Anaerobic culture was done in cooked meat medium (Remel, Lenexa, KS) and on Brucella PRAS and egg yolk PRAS agar (Anaerobe Systems, Morgan Hill, CA) for 5 days at 37°C in an anaerobic chamber. Ten colonies from anaerobic culture agar plates were transferred to the MALDI Biotyper system (Bruker, Billerica, CA; MBT Compass Library, Version 13) for identification. A full list of clostridial species represented within the MBT Compass Library (Version 13) is presented in Supplemental Material S2.
In addition, fecal pellets from 2 C. cuniculi qPCR-positive mice were added to MEGA media 21 (pH 6.8 and pH 7) with the addition of sodium pyruvate (0.5g/L) and grown anaerobically (90% Nitrogen, 5% carbon dioxide, 5% hydrogen) in an anaerobic chamber (Sheldon Manufacturing, Cornelius, OR) at Stanford University. Every 48 hours, cultures were plated in serial dilutions. Single colonies were restreaked and then identified using the MALDI Biotyper system (Bruker, Billerica, CA) following manufacturer’s instructions. 1 A full list of clostridial species represented within the MBT Compass Library (custom library comprising BDAL 10, cfiA, and IVD 10 with additional user-added strains) is presented in Supplemental Material S3.
Results
Epidemiology
Beginning in October 2020, veterinary staff were notified of adult male and female NSG mice with diarrhea in the Principal Investigator-managed breeding room of the mouse barrier facility. The VSC-managed breeding room was not affected. Surveillance of other rooms identified additional cases of diarrhea. During the peak of the outbreak, 10/16 (62.5%) barrier rooms, representing over 20 labs, were affected (Fig. 1). Simultaneously, there was an increase in reports of lactating NSG dams dying suddenly during the mid- to late-lactation period (postnatal day 11 to postnatal day 16) resulting in orphaned pups (Fig. 2). Prior to this, cases of dams dying unexpectedly during lactation were uncommon. Dams identified prior to spontaneous mortality were moribund with perianal fecal staining.
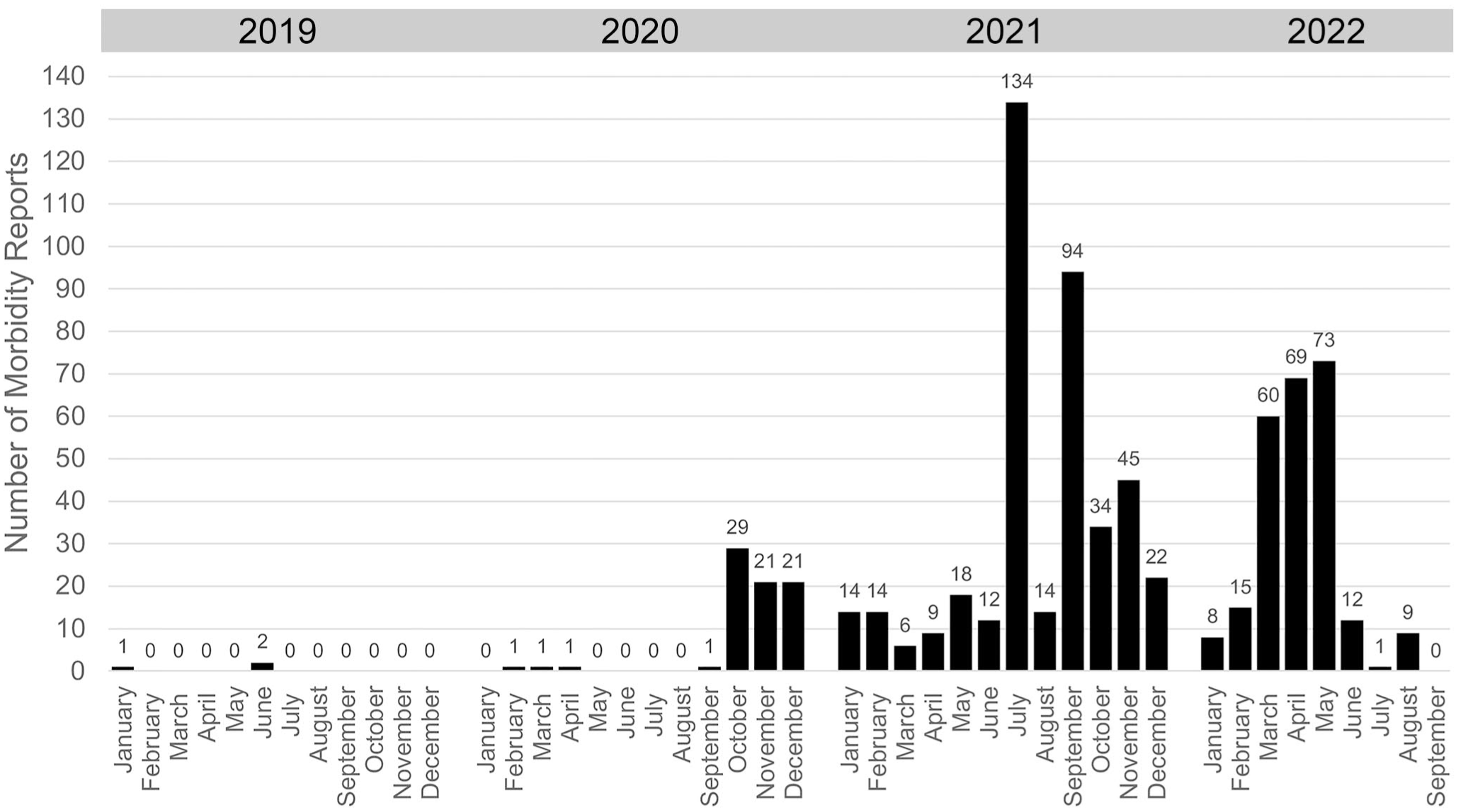
Number of morbidity reports for diarrheic mice within the mouse barrier facility from January 2019 through September 2022. Data were obtained following a retrospective analysis of the veterinary electronic medical record. Increased numbers of diarrhea-related morbidity reports began in October 2020 and persisted through mid-2022. Note: The veterinary electronic medical record did not discern between strains of mice.
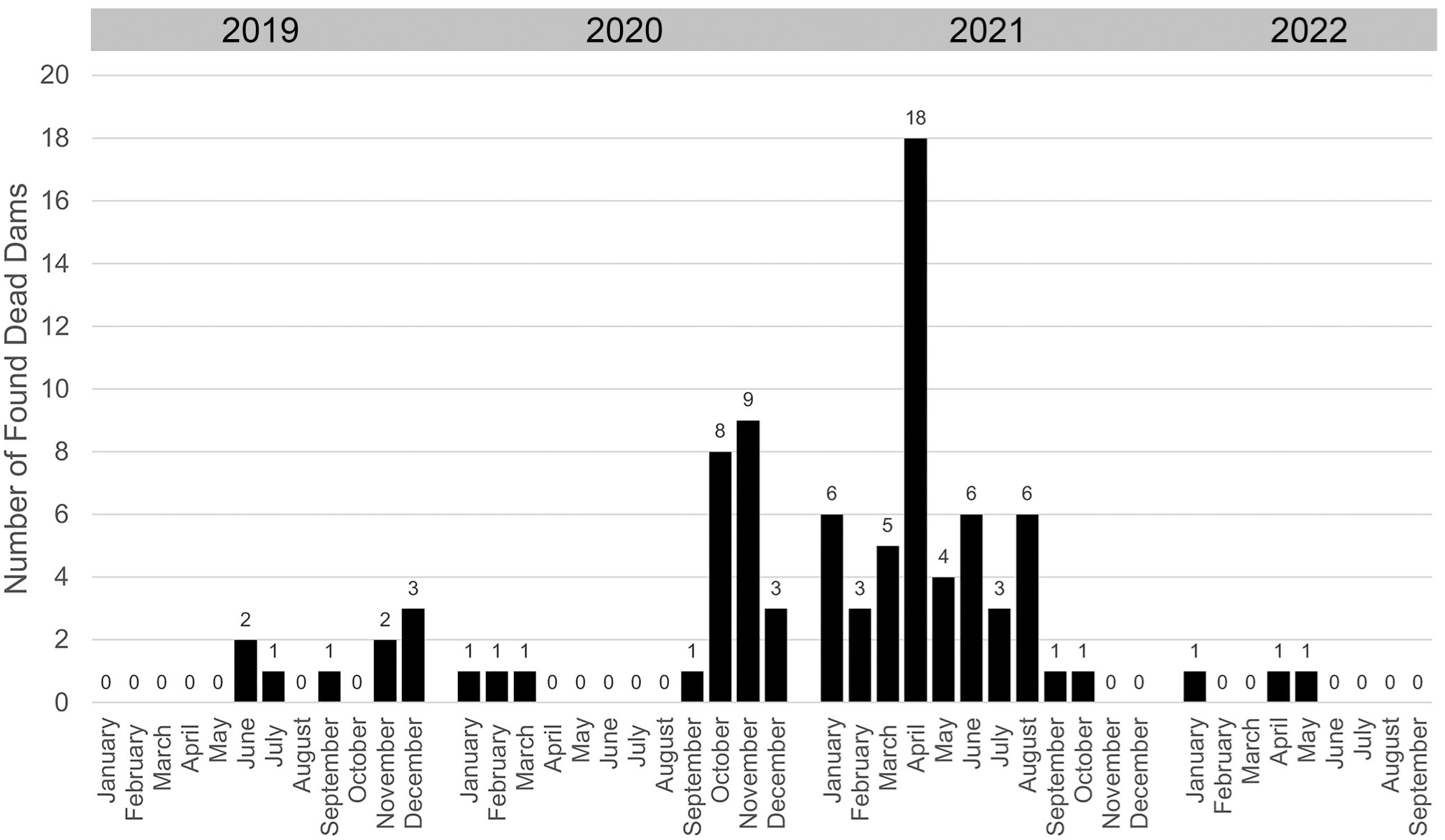
Number of found dead dams within the mouse barrier facility from January 2019 through September 2022. Data were obtained following a retrospective analysis of the veterinary electronic medical record. Increased mortalities of dams commenced in October 2020 and tapered off in mid-late 2021. Note: The veterinary electronic medical record did not discern between strains of mice.
Further investigation revealed other immunodeficient strains were affected. Affected strains, in addition to NSG, included NRG (NOD. Cg-Rag1 tm1Mom Il2rg tm1Wjl /SzJ), NBSGW (NOD. Cg-Kit W-41J Tyr+ Prkdc scid Il2rg tm1Wjl /ThomJ), NSG-SGM3, RAG2/Gc-DKO-B6, and RAG2/Gc-DKO, the latter 2 strains having Rag2 and IL2rg deletions on the C57BL/6 and BALB/c genetic backgrounds, respectively. Common to the affected strains were the presence of the SCID (Prkdc scid ) or Rag (Rag null ) mutations in combination with the IL2rg (Il2rg null ) mutation. For simplicity in this report, the affected strains will be referred to as “NSG-related” unless otherwise noted, recognizing that other genetic backgrounds other than NOD may be present and additional mutant genes may be present.
Clinical signs observed (in order of prevalence) included feces coated with corncob bedding (colloquially referred to as “popcorn poop” due to their appearance), watery or soft fecal material stuck on strip paper enrichment, fecal material smeared on cage walls, and/or overt watery diarrhea when mice were lifted by the tail (Fig. 3). For simplicity, any of the above criteria are referred to as “diarrhea.” The diarrhea occurred intermittently, occasionally resulted in unthriftiness, but did not typically result in mortality in nonlactating NSG mice.
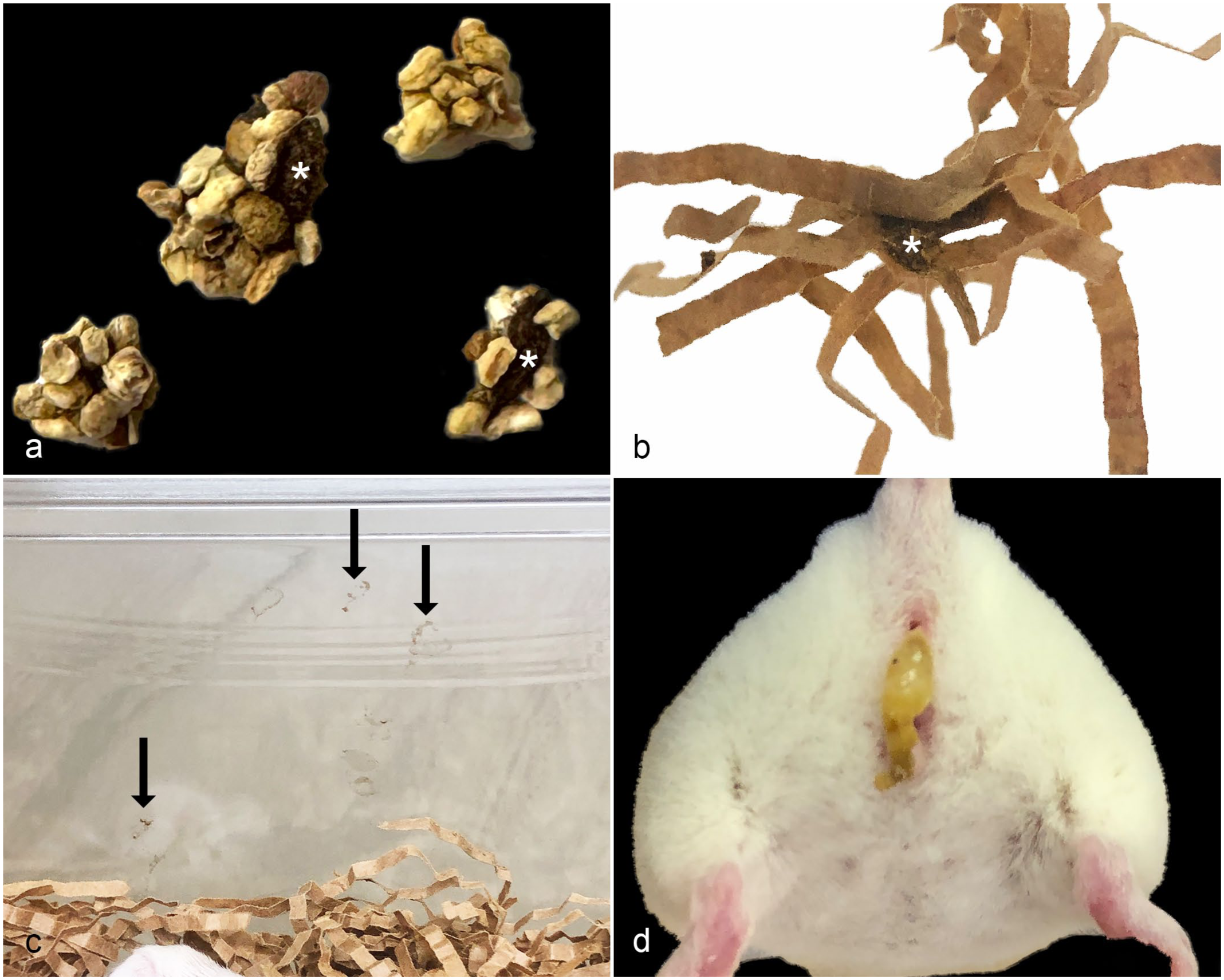
Environmental and clinical evidence of diarrhea. (a) Soft feces (white asterisks) are coated with corncob bedding (“popcorn poop”). (b) Soft feces (white asterisk) are adherent to strip paper enrichment. (c) Smears of fecal material (black arrows) on cage walls. (d) Yellow, watery diarrhea on the perineum of an NOD. Cg-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) mouse.
Necropsy and Histology
Diarrheic nonlactating adult female and male mice (n = 16) exhibited mild to moderate gastric, small intestinal, and/or cecal distension (Fig. 4a) with a variable reduction in the number, size, and/or consistency of fecal pellets as compared to controls (Fig. 4b). The prevailing histologic lesions in this subset included cecocolonic individual cell death of superficial enterocytes with sloughing, and less frequent crypt epithelial cell death (Fig. 4c). Occasionally, a compensatory hyperplastic response was noted. A single mouse exhibited focally extensive neutrophilic and histiocytic typhlitis.
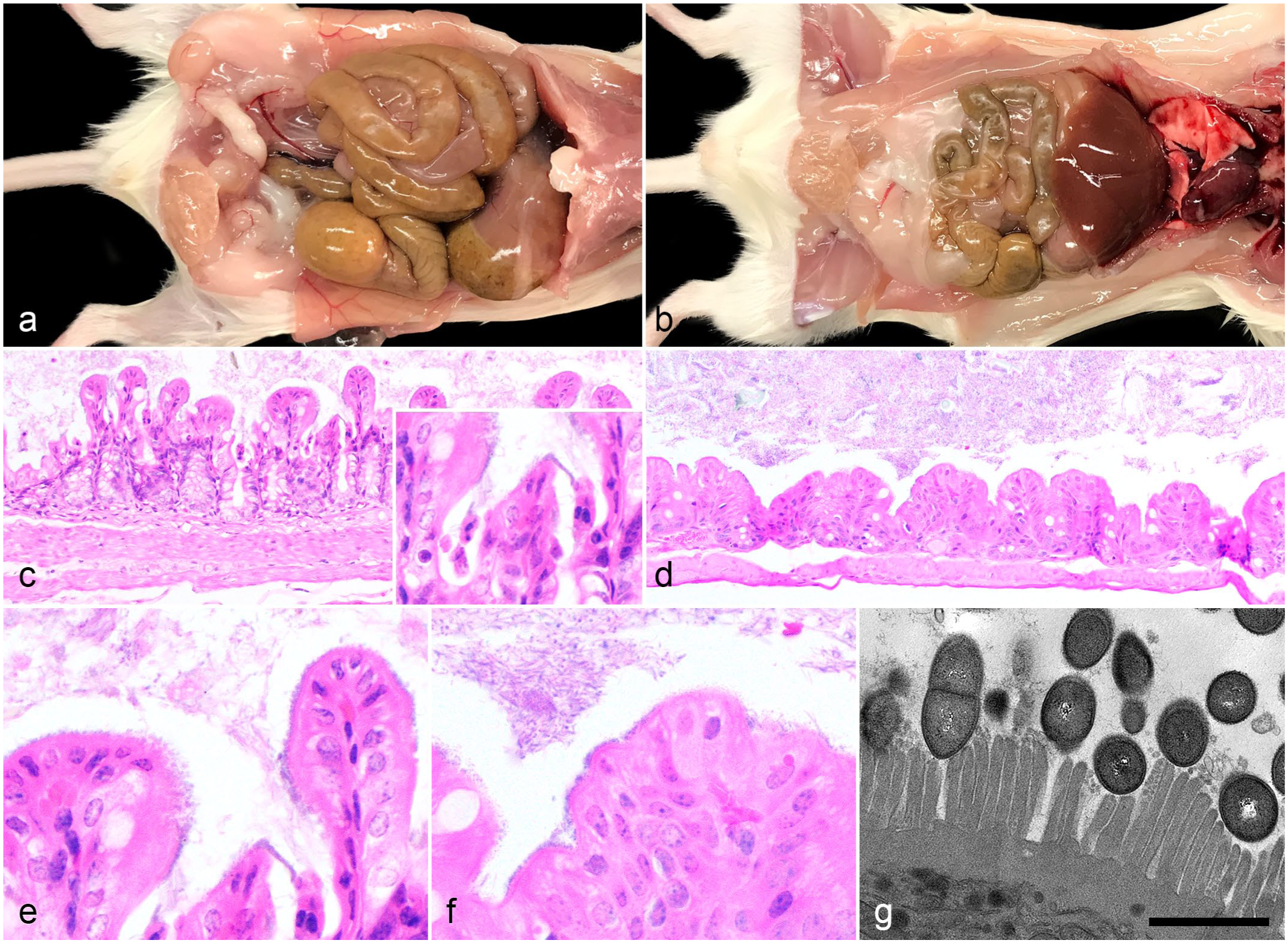
Gross and histopathologic features of NOD.Cg-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) and NSG-related mice with diarrhea. (a) Small and large intestines, diarrheic NSG mouse. Mild to moderate small and large intestinal dilation with poorly formed fecal pellets. (b) Small and large intestines, control NSG mouse. Normal small and large intestines. (c) Cecum, diarrheic NSG mouse. Individual cell death and sloughing of superficial and mid-crypt epithelial cells. Inset: High magnification of individual cell death. Hematoxylin and eosin (HE). (d) Cecum, control NSG mouse. Normal cecal mucosa. HE. (e) Cecum, diarrheic NSG mouse. Enterocytes are overlain by a carpeting of apically adherent coccobacilli. HE. (f) Cecum, control NSG mouse. Enterocytes in control NSG mice are also overlain by a carpeting of apically adherent coccobacilli. HE. (g) Cecum, diarrheic NSG mouse. Coccobacilli attached to the enterocyte apex with microvilli disarray and shortening. Transmission electron microscopy, scale bar = 1 µm.
In contrast, the gross and histologic findings in lactating dams were more severe and extensive in distribution. Grossly, lactating dams (n = 6) had moderate to severe gastrointestinal distension and/or segmental, dark red to gray, small intestinal discoloration (Fig. 5a, b). Three out of 6 dams were found deceased and were too autolyzed for histologic interpretation. Histologic lesions in the remaining 3 dams ranged from moderate, cecocolonic individual cell death of superficial enterocytes with luminal sloughing and mucosal hyperplasia (3/3) to severe ulcerative inflammation of the jejunum, ileum, cecum, and colon (2/3; Fig. 5c, d).
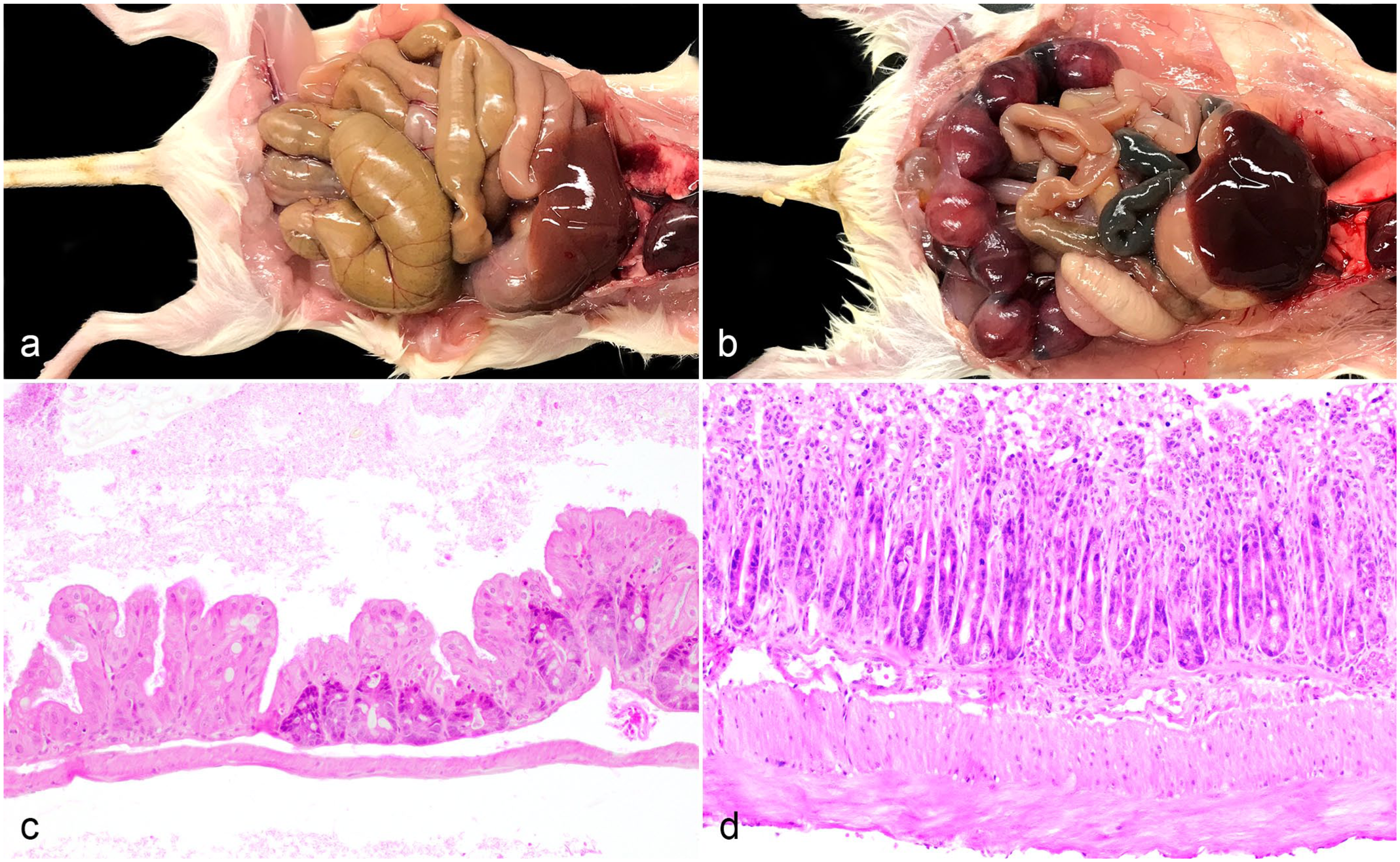
Gross and histopathologic features of diarrheic lactating NOD.Cg-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) and NSG-related dams. (a) Small and large intestines, diarrheic NSG dam. Severe small and large intestinal dilation with poorly formed fecal pellets. (b) Small and large intestines, moribund NSG dam. Segmental dark gray discoloration of the small intestine as well as thickening and opacity of the cecum. Multiple early embryos are present in the uterine horns. (c) Cecum, lactating NSG dam. Individual cell death and sloughing of mid-crypt epithelial cells. Crypts are segmentally replaced by hyperplastic epithelial cells. Hematoxylin and eosin (HE). (d) Jejunum, moribund NSG dam. Widespread ulceration with neutrophilic enteritis. HE.
In addition, in both the nonlactating adult mice (11/16, 69%) and lactating dams (2/3) cecocolonic enterocytes were segmentally overlain by a carpeting of apically adherent gram-negative coccobacilli (Fig. 4e). The presence or absence of coccobacilli was not correlated with the severity of the histologic lesions. Notably, NSG mice (6/6) received directly from the vendor and immediately processed upon receipt at Stanford University also exhibited cecocolonic carpeting of apically adherent bacteria (Fig. 4f), although to a slightly lesser extent than mice with clinical signs. No other histologic lesions were identified in NSG control mice.
Tissue Ultrastructure and Virus Screening
TEM confirmed the presence of coccobacilli attached to the apical surface of enterocyte with associated microvilli disarray and shortening (Fig. 4g). The morphologic features of the microorganisms were considered suggestive of the Enterobacteriaceae group. TEM screening of fecal material from diarrheic mice found no evidence of a viral cause.
Bacteriology, Parasitology, and Molecular Screening
Enteric aerobic and anaerobic cultures performed at the Stanford Animal Diagnostic Laboratory did not identify a causative diarrheic agent (Supplemental Table S1). Initial culture results from diarrheic adult (n = 12) and lactating NSG mice (n = 2) as well as controls (n = 6) included a mix of Enterococcus faecalis, Staphylococcus xylosus, and Lactobacillus spp. (Lactobacillus johnsonii, Lactobacillus murinus, Lactobacillus bifermentans, Lactobacillus amylovorus, Lactobacillus intestinalis, and Lactobacillus gaserri). Clostridium tertium was grown from 2 diarrheic NSG mice, as were nonspeciated flat colonies in microaerophilic media.
Pooled samples from diarrheic NSG mice (n = 3) submitted to CAHFS (San Bernardino, CA) failed to yield anaerobic growth, including C. difficile.
Aerobic and anaerobic cultures obtained from terminal cardiac blood (n = 6 diarrheic), liver (n = 7 diarrheic; n = 6 naive), and spleen (n = 7 diarrheic; n = 6 naive) were negative.
Fecal flotations performed on diarrheic (n = 6) and naive (n = 6) mice were negative.
Feces of all mice, including control mice, were PCR positive for astrovirus. Fecal PCR results were negative for all other murine pathogens included in the Charles River Laboratories Mouse Surveillance Plus PRIA (see Materials and Methods for pathogen list) as well as additional PCR assays targeting Clostridioides difficile, rodent chaphamaparvovirus 1, and Leptospira spp. (Charles River Laboratories, Wilmington MA).
C. perfringens and C. difficile ELISAs
ELISAs for C. perfringens (CPA, CPB, and ETX) and C. difficile (A and B) toxins were negative. However, pooled samples tested positive for the presence of C. perfringens organisms (not cultured anaerobically). Nevertheless, historic samples from a separate disease outbreak also demonstrated C. perfringens positivity in 2 groups of control NSG males and NSG-SGM3 females, 3 suggesting that detection of the organism is not indicative of disease.
Immunohistochemistry
The quantity and distribution of cecocolonic luminal immunoreactivity to C. perfringens was subjectively equivalent within a diarrheic NSG mouse, a diarrheic lactating NSG dam, and a control NSG mouse.
Transmissibility to CD-1 Mice
To provide real-time evidence-based guidelines for both barrier and husbandry practices during the peak of this outbreak, cohousing and gavage studies were performed prior to development and validation of our novel qPCR assays. Mice that developed diarrhea were considered susceptible to the unknown pathogen. No pathogens were detected in serum or fecal samples from CD-1 mice cohoused with diarrheic NSG mice. Despite prolonged housing with diarrheic NSG mice (several weeks), CD-1 mice never developed diarrhea and were unable to transmit diarrhea to cohoused naïve NSG females.
Transmissibility to NOD scid, Rag2 KO, and B6 Rag2 IL2rg DKO Mice
None of the strains gavaged with the nondiarrheic intestinal homogenate showed signs of diarrhea. However, NSG mice gavaged with diarrheic intestinal homogenate (positive control) developed diarrhea at 11 days post-gavage as evidenced by the presence of “popcorn poop” in their cage. B6 Rag2 IL2rg DKO, NOD scid, and Rag2 KO mice gavaged with diarrheic intestinal homogenate (previously frozen) did not show signs of diarrhea in their cage. The absence of diarrhea in the B6 Rag2 IL2rg DKO mice was unexpected given that the Principal Investigator-managed Rag2/Gc-DKO-B6 mice were susceptible to spontaneous diarrhea within the barrier.
To account for this discrepancy, and because freezing medium was not added to our −80°C frozen homogenate, this gavage study was thus repeated with freshly prepared (nonfrozen) diarrheic intestinal homogenate (Fig. 6a). The NSG mice showed evidence of diarrhea 2 days post-gavage. The B6 Rag2 IL2rg DKO, NOD scid, and Rag2 KO again did not show any evidence of diarrhea for up to 20 days post-gavage (Fig. 6b). To test if the mice were subclinically infected, 2 naïve NSG females were added to the cages housing each strain. No evidence of diarrhea was found in the NOD scid and Rag2 KO cages. In contrast, diarrhea was found in the B6 Rag2 IL2rg DKO cage 14 days after cohousing (Fig. 6c). The B6 Rag2 IL2rg DKO mice were moved to a new cage to determine whether the diarrhea originated from the B6 Rag2 IL2rg DKO or NSG mice. The B6 Rag2 IL2rg DKO cage did not show any evidence of diarrhea for up to 21 days while the NSG cage remained diarrhea positive (Fig. 6d).
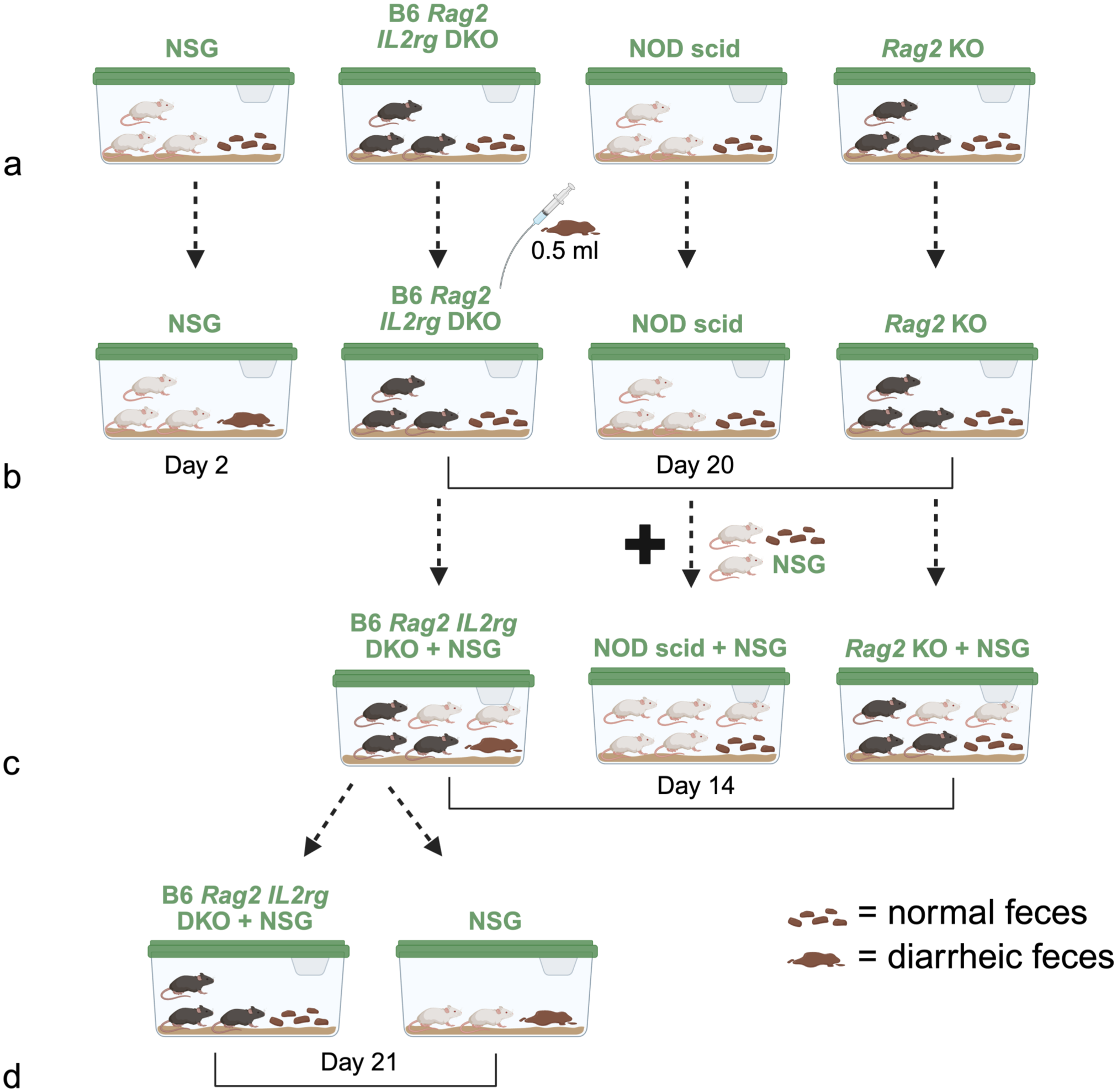
Transmissibility studies in NSG (NOD. Cg-Prkdc scid Il2rg tm1Wjl /Szj), Rag2 KO (C57BL/6. Cg-Rag2 tm1.1Cgn /J), NOD scid (NOD. Cg-Prkdc scid /J), and B6 Rag2 IL2rg DKO (C57BL/6NTac.Cg-Rag2 tm1Fwa Il2rg tm1Wjl /Tac) mice. (a) Three mice from each strain (NSG, Rag2 KO, NOD scid, and B6 Rag2 IL2rg DKO) were gavaged with 0.5 ml of freshly prepared, unfiltered, diarrheic intestinal homogenate. (b) On day 2, NSG mice exhibited diarrhea. At day 20, none of the B6 Rag2 IL2rg DKO, NOD scid, or Rag2 KO mice exhibited diarrhea. (c) Two naive NSG mice were added to each cage of B6 Rag2 IL2rg DKO, NOD scid, and Rag2 KO mice to determine if these strains could serve as subclinical carriers. On day 14, diarrhea was present in the B6 Rag2 IL2rg DKO+NSG cage. No diarrhea was observed in the NOD scid +NSG or Rag2 KO+NSG cages. (d) B6 Rag2 IL2rg DKO and NSG mice were separated to determine which strain was the source of the diarrhea. On day 21, diarrhea was present in the cage of NSG mice while B6 Rag2 IL2rg DKO mice remained asymptomatic, suggesting they can serve as subclinical carriers. Created in BioRender.
C. cuniculi and Enterotoxin qPCR
All NSG or NSG-related mice with clinical evidence of diarrhea (n = 37) were qPCR-positive, showing high copy numbers of both C. cuniculi and its enterotoxin in all fecal samples. 5 None of the clinically asymptomatic mice (n = 111) tested positive for C. cuniculi or its enterotoxin. Similarly, mice with other identified causes of diarrhea (e.g., spontaneous neoplasia, experimentally related, n = 28) tested negative. 5
Targeted Microbiological Culture of C. cuniculi and Enterotoxin qPCR-Positive Samples
Pooled cecal and large intestinal contents from diarrheic NSG mice (n = 3) were cultured with the aim of isolating C. cuniculi colonies from C. cuniculi and enterotoxin qPCR-positive samples. No colonies of Clostridium sp. were isolated. Colonies of varying morphologies were isolated from 2 fecal pellets from C. cuniculi qPCR-positive mice; however, identified colonies were neither qPCR-positive for C. cuniculi (or its enterotoxin) nor identified as Clostridium sp. via the Biotyper.
Discussion
Beginning in October 2020, a high-morbidity low-mortality diarrheal disease of NSG and NSG-related adult mice spread throughout our mouse barrier facility. Affected strains carried either the SCID or Rag null mutations in combination with the IL2rg null mutation. Affected strains included NSG, NRG, NBSGW, and NSG-SGM3. Principal Investigator-managed strains including Rag2/Gc-DKO-B6 and Rag2/Gc-DKO-c were also affected. During its peak, diarrheal disease impacted over 20 laboratories and 10/16 (62.5%) barrier rooms. During the same 1.5-year period, increased numbers of mid- to late-lactation NSG dams died suddenly. Gross findings included mild to moderate small and large intestinal distension in nonlactating mice, and severe small and large intestinal distension and/or intestinal discoloration in lactating dams. Consistent histologic lesions in all mice included cecocolonic individual cell death of superficial enterocytes with luminal sloughing. In addition, moribund dams exhibited severe ulcerative enterocolitis of the distal small and large intestine. Extensive traditional diagnostics, including aerobic and anaerobic culture (intestines, blood, liver, and spleen), fecal PCR, fecal parasitology, C. perfringens organism ELISA, C. perfringens and C. difficile toxin ELISAs, TEM virus screening, and C. perfringens immunohistochemistry, failed to identify an etiologic agent. Clinical and diagnostic efforts were implemented to (1) control the spread of a yet unidentified infectious agent, (2) understand strain susceptibility to guide barrier and husbandry practices, and (3) further explore etiopathogenesis. Utilizing strain-resolved metagenomics, an enterotoxin-encoding C. cuniculi was identified and subsequent qPCR assays were generated. 5 All mice with diarrhea of unknown cause tested positive for C. cuniculi and its enterotoxin via qPCR assays while all known healthy controls were negative. Despite extensive attempts at enteric culture, we have been unable to isolate the bacterium and thus have been unable to fulfill Koch’s postulates.
In the initial phases of this outbreak, transmissibility studies were performed to better understand if the observed diarrhea was due to an infectious agent, and if so, which strains might be susceptible to diarrhea and/or serve as carriers. Development and validation of our novel qPCR assays took several years, thus, positive outcomes in these early transmissibility studies were based on the presence or absence of diarrhea, rather than molecular identification of a pathogen. Early studies demonstrated the diarrhea was indeed contagious and transmissible to naïve NSG mice by cohousing with diarrheic mice or by the gavage of intestinal homogenates from diarrheic NSG mice, but not if the homogenate was filtered through a 0.2-µm filter. This suggested that the pathogen was filterable and therefore unlikely to be a virus. 5
The absence of diarrhea cases among other mouse strains sharing the same room as mice with infectious diarrhea suggested that not all strains were susceptible to the pathogen. However, whether the mice in a given room could be subclinically infected, and therefore serve as a reservoir, was unknown. To address this possibility, we first cohoused immunocompetent CD-1 mice with diarrheic NSG mice and showed that CD-1 mice failed to develop diarrhea and failed to transmit the diarrhea to naïve cohoused NSG mice, suggesting that they were neither susceptible nor subclinically infected. We then tested whether other immunodeficient strains (NOD scid, Rag2 KO) were susceptible to the infectious diarrhea. Following gavage with intestinal homogenates from diarrheic mice, NOD scid and Rag2 KO mice neither developed diarrhea nor transmitted it to naïve NSG mice. This suggests that, like immunocompetent CD-1 mice, immunodeficient NOD scid and Rag2 KO are neither susceptible nor capable of being subclinically infected.
Although Rag2/Gc-DKO-B6 and Rag2/Gc-DKO-c colonies in our barrier showed evidence of diarrhea during this outbreak, a commercially obtained B6 Rag2 IL2rg DKO strain did not show evidence of diarrhea following gavage with intestinal homogenates from diarrheic mice. In contrast, naïve NSG mice gavaged with the same intestinal homogenate exhibited diarrhea. Although the B6 Rag2 IL2rg DKO strain did not have evidence of diarrhea, naïve NSG became infected following cohousing suggesting that the B6 Rag2 IL2rg DKO mice were subclinically infected and therefore may serve as a reservoir. The Rag2/Gc-DKO-B6 and Rag2/Gc-DKO-c strains in our barrier were maintained as a closed colony by an investigator for over 8 years. One possibility for the discrepancy in susceptibility to infectious diarrhea between the Rag2/Gc-DKO-B6 and B6 Rag2 IL2rg DKO mice is genetic drift.
While working to understand strain-susceptibility, we simultaneously explored diagnostics to determine an underlying cause. Our initial efforts, including aerobic and anaerobic cultures, fecal parasitology, fecal PCR, ELISAs for clostridial enteric pathogens, and TEM of feces for viral pathogens, were unfruitful. Of note, apically adherent, gram-negative coccobacilli were noted along the cecal and colonic mucosa in many of the diarrheic mice and lactating dams. Our initial thought was that this coccobacillus could be contributing to the observed diarrhea. However, when reviewing histologic sections from NSG mice received directly from the vendor as a control cohort, similar coccobacilli were also observed along the cecal and colonic mucosa. Anecdotally, we now sporadically see cecocolonic apically adherent bacteria in immunodeficient mice in our colony that show no signs of diarrhea, loose stool, or cecocolonic individual cell death. Moreover, the morphology and Gram-staining characteristics of this population are not consistent with that of a Clostridium sp. Thus, we do not currently know the bacterial identity of these apically adherent bacteria, nor do we understand their contribution, if any, to the development of diarrhea. In addition, ulcerative lesions seen exclusively in lactating dams were considered part of this disease outbreak, as no other known infectious agents were identified in lactating dams. We hypothesized that the additional burden of pregnancy and lactation may have exacerbated the disease phenotype. However, because we have not done infectivity studies with pregnant dams, nor were our qPCR assays available at the onset of the outbreak, we are unable to definitively link these specific lesions to C. cuniculi.
Because of the failure of traditional diagnostics to identify an etiologic agent we resorted to a microbiome analysis using a commercial service for DNA extraction, library preparation, and sequencing (Transnetyx, Cordova, TN). Initial results focusing on changes in bacterial strains between diarrheic and control fecal samples were inconsistent, and we concluded that the observed changes were a consequence of the diarrhea and not its cause (data not shown). The processed reads were then subjected to a strain-resolved metagenomic analysis. 5 This analysis identified 11 bacterial strains that were enriched in diarrheic fecal samples. The bacterial strains were in the families Muribaculaceae (n = 3), Rikenellaceae (n = 6), and Clostridiaceae (n = 2). The 2 candidates in the Clostridiaceae were C. cuniculi and Dwaynesavagella sp000270205, a segmented filamentous bacterium. C. cuniculi is closely related to C. celatum which, along with segmented filamentous bacterium, Clostridiales bacterium VE202-01, and Bifidobacterium pseudolongum, were identified in a similar outbreak of diarrhea in NSG and NSGS mice. 17 Our qPCR analyses linked C. cuniculi, but not segmented filamentous bacterium, to the infectious diarrhea outbreak at our institution. 5
C. cuniculi is infrequently reported in the literature, but has been proposed as an inciting cause of epizootic rabbit enteropathy (ERE), a gastrointestinal disease of farmed rabbits. 8 ERE was first reported and described in France and was initially characterized by abdominal distension, small volume watery diarrhea, decreased feed intake, and high mortalities. 14 Subsequent ERE clinical reports 20 and experimental studies7,14 variably describe gross and histologic findings such as gastric, small intestinal, and/or cecal dilation; colonic mucus; reduced feces; villus atrophy and fusion; and reduced crypt depth. Similar to our findings, inoculation of enteric contents from affected to naïve rabbits results in clinical disease transmission,7,16 however, definitive identification of a causative pathogen has remained elusive in the context of ERE. Because of this, the diagnosis of ERE is based predominantly on clinical signs, resulting in somewhat nebulous reports of gross and histologic findings.
To date, we have been unable to culture C. cuniculi from C. cuniculi qPCR-positive feces, despite multiple attempts at several different institutions. It is worthwhile to note that the reference libraries used in the MALDI Biotyper system (Supplemental Materials S2 and S3) do not include C. cuniculi and thus we would not expect to confirm identification via this modality. However, none of our colonies isolated under anaerobic conditions were identified as Clostridium sp., suggesting that C. cuniculi may be noncultivable.
A source for the introduction of C. cuniculi, is unknown, though 1 possibility includes exposure via irradiated food. 13 Irradiation at the levels typically used for pelleted rodent chow (~25 kGy) does not result in sterility and spores are expected to be more resistant to irradiation. One caveat to this route of exposure is that the diarrhea outbreak did not affect the VSC-managed breeding barrier room despite the presence of NSG colonies and the provision of irradiated food. However, given the volume of food entering the barrier on a monthly basis, only a single contaminated bag would be needed to introduce a pathogen. If that bag went into a room housing NSG mice and the mice became infected, sharing of imaging modalities and procedure rooms could spread the pathogen among other colonies. The VSC-managed breeding room may have been spared due to the absence of experimental procedures and personnel restriction (VSC staff only).
The diarrhea outbreak in the barrier was managed in several steps, many of which were reliant on early strain-susceptibility data. First, all DKO mice (mice harboring the SCID or Rag mutations together with the IL2rg mutation) in an affected room were moved to a quarantine room outside the barrier. Although experiments were allowed to continue in the quarantine room, no breeding was allowed. Non-DKO mouse strains remaining in the contaminated room were moved to a spare holding room and all expendable supplies were discarded. The room was then sealed and disinfected with vaporized hydrogen peroxide. Once disinfected, the non-DKO mice were changed into clean cages, moved onto new chlorine dioxide disinfected individually ventilated cage racks, and returned to the room. This process was repeated until all rooms that housed diarrheic DKO strains were disinfected. Second, breeding of any DKO strain in the barrier was placed under the control of the VSC breeding colony managers. No lab was allowed to breed DKO strains in the barrier. The VSC-managed breeding rooms housed only DKO strains and lab personnel were not allowed in the rooms. Third, all experiments with DKO mice were restricted to 2 rooms in the barrier. No other strains were allowed in these rooms. Our justification for the separation of DKO strains and non-DKO strains was based on our demonstration that B6 Rag2 IL2rg DKO strains could be subclinically infected. Given the numerous genetically modified strains housed in the barrier, many of which are “immunovague,” we could not be certain that some were not subclinically infected during the outbreak. Fourth, DKO strains requiring in vivo bioluminescent imaging were transferred outside the barrier to a room set aside for DKO strains requiring imaging. Imaging was performed using a dedicated in vivo imaging system. This prevented DKO strains and non-DKO strains from sharing the same instrument in the barrier. Fifth, due to its low efficacy against spores, 11 Virkon-S (Lanxess, Pittsburgh, PA), the disinfectant used in the barrier during the outbreak, was changed to Vimoba (200 ppm chlorine dioxide; Quip Labs, Wilmington, DE) or Clorox Healthcare Bleach Germicidal (The Clorox Company, Oakland, CA), the latter having efficacy against C. difficile spores. Finally, because we suspected that the irradiated chow may have been the source of the C. cuniculi, we implemented autoclaved chow for all DKO strains in the barrier. Other strains in the barrier remained on irradiated chow. Having implemented these changes, we have not seen any cases of C. cuniculi diarrhea in the barrier to date, although cases of C. cuniculi diarrhea are still present in conventional facilities on campus.
In summary, the low-mortality, high-morbidity outbreak of diarrheal disease in NSG and NSG-related mice was strongly associated with the presence of C. cuniculi. Spontaneous infection suggests that strains harboring the SCID or Rag null mutations together with the IL2rg null mutation are susceptible to infection. In our studies, CD-1, NOD scid, and Rag2 KO were not susceptible to infection. Transmissibility studies further suggest that B6 Rag2 IL2rg DKO mice may be able to serve as asymptomatic carriers following exposure to C. cuniculi. Further studies using our novel qPCR assays are in progress to validate the strain-susceptibility of various immunodeficient mouse strains. The epidemiology of this outbreak highlights the unique susceptibility of immunocompromised mice to novel pathogens and underscores the utility of alternative diagnostics in determining cause.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858251372565 – Supplemental material for Clostridium cuniculi is associated with chronic high-morbidity low-mortality diarrhea in NSG and NSG-related mouse strains
Supplemental material, sj-pdf-1-vet-10.1177_03009858251372565 for Clostridium cuniculi is associated with chronic high-morbidity low-mortality diarrhea in NSG and NSG-related mouse strains by Kerriann M. Casey, Rina Barouch-Bentov, Wencke Reineking, Flavio H. Alonso, Roberta Moorhead, Minoo Fazel, Anibal G. Armien, Francisco A. Uzal, Rachael B. Chanin, Ami S. Bhatt, Sherril L. Green, Stephen A. Felt and Claude M. Nagamine in Veterinary Pathology
Footnotes
Acknowledgements
The authors acknowledge Greg Nelson for gathering epidemiology data, Elias Godoy for necropsy assistance, and Jimena Guiterrez and Tanya Watarastaporn for their technical assistance with the transmissibility studies. The authors thank Paula Yant and Jeannie Mullen for their assistance with microbiology. Erin Brooks and Yishay Pinto provided technical assistance and analysis with next-generation sequencing of tissue from formalin-fixed paraffin-embedded blocks. Maggie Madrigal-Moeller and Justin Sonnenburg graciously provided access to the MALDI Biotyper system at Stanford University.
Supplemental material for this article is available online.
Authors’ Contributions
Concept and design: KMC, FAU, SLG, SAF, CMN.
Acquisition of data or analysis: KMC, RB-B, WR, RM, MF, AGA, FAU, RBC, ASB, CMN.
Interpretation of data: KMC, RB-B, WR, FHA, RM, MF, AGA, FAU, RBC, ASB, SLG, SAF, CMN.
Manuscript preparation: KMC, CMN.
Manuscript critical revision: KMC, RB-B, WR, FHA, RM, MF, AGA, FAU, RBC, ASB, SLG, SAF, CMN.
Publication approval: KMC, RB-B, WR, FHA, RM, MF, AGA, FAU, RBC, ASB, SLG, SAF, CMN.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded, in part, by the Stanford University Discovery Innovation Fund, the A.P. Giannini Foundation, and the Stanford University Department of Comparative Medicine.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
