Abstract
Atypical Bacillus cereus strains (currently classified as Bacillus tropicus) capable of causing anthrax-like disease in humans and animals have emerged in the last 2 decades. These emerging strains are characterized by the acquisition of virulence plasmids pBCXO1 and pBC210, which are homologous to the pXO1 and pXO2 virulence plasmids of Bacillus anthracis, the agent of anthrax. The aim of this study was to describe the gross, histologic, microbiologic, and molecular features of an outbreak of anthrax-like septicemia caused by B. tropicus in red kangaroos (Macropus rufus). Three red kangaroos from a wild animal preserve were found dead with no premonitory clinical signs. No changes in husbandry were reported prior to the outbreak. The peracute disease process was characterized by severe splenomegaly, associated with fibrinonecrotizing splenitis in all affected animals, in addition to segmental suppurative enteritis in 2 kangaroos and cutaneous excoriations, with underlying necrotizing cellulitis and lymphadenitis, in 1 kangaroo. Numerous intralesional, gram-positive and capsulated bacilli were identified as the formerly known B. cereus group via bacteriologic culture. Whole-genome sequencing from one of the bacterial isolates (designated 11844) revealed numerous anthrax-like virulence factors, including the pBCXO1 and pBC210 virulence plasmids. This isolate also had a close phylogenetic relationship with other B. tropicus strains carrying these virulence plasmids, including B. tropicus (formerly B. cereus) G9241. This is the first report of B. tropicus leading to anthrax-like disease in kangaroos. This disease form carries significant public health risks due to potential zoonotic transmission.
Bacillus cereus sensu lato (s.l.) is a group of gram-positive, endospore-forming bacteria that includes several Bacillus species (spp.) with closely related phylogeny such as B. cereus, Bacillus thuringiensis, B. anthracis, B. mycoides, and Bacillus tropicus, which share highly conserved chromosomal DNA but differ in their virulence encoding plasmids.15,21 The environmental reservoirs for Bacillus spp. include fresh and marine waters, decaying organic matter, vegetables and fomites, and the intestinal tract of invertebrates. 21 Transmission to a host is usually via percutaneous, inhalational, or fecal-oral routes. 4 In addition to causing food poisoning, this bacterium can also induce local and systemic infections in both immunocompetent and immunologically compromised individuals.3,8,34 In veterinary medicine, B. cereus infections have been reportedly associated with severe intestinal disease in pigs, 9 mastitis in dairy cows, 11 and gangrenous mastitis in goats. 29 The pathogenicity of B. cereus, whether intestinal or extra-intestinal, is intimately associated with tissue-destructive/reactive exoenzyme production, including 3 pore-forming enterotoxins: hemolysin BL, nonhemolytic enterotoxin, and cytotoxin K.8,27 Over the last 2 decades, several emerging strains of atypical B. cereus (currently designated as B. tropicus) have reportedly caused anthrax-like illnesses in both humans and mammals, including and apes.19,20,22,23 These closely genetically related Bacillus spp. isolates, as determined by whole-genome sequencing and/or multiple-locus sequence typing, are currently classified as B. tropicus based on updated B. cereus group taxonomic classification.10,27,38
B. anthracis, the etiological agent of anthrax, is a similar spore-forming bacillus that carries 2 virulence plasmids, pXO1 and pXO2, which encode the edema and lethal factor (composed of 3 specific protein products, namely, the edema factor, the protective antigen, and the lethal factor) and the poly-D-glutamic acid capsule, respectively. These 2 plasmids allow for the differentiation of B. anthracis from B. cereus at the molecular level.8,20 Phenotypically, B. cereus can be differentiated from B. anthracis by hemolysis, motility, γ-phage resistance, and penicillin G resistance. 24 Atypical strains of B. cereus, such as B. cereus G9241 and other Bacillus spp. G9898, LA2007, and 03BB102, carry virulence plasmids highly similar to those of B. anthracis, namely pBCXO1 and pBC210 (formerly pBC218), and may produce a unique exopolysaccharide capsule. 4 These strains cause anthrax-like disease.4,20 B. cereus biovar anthracis, which is distinct from the aforementioned atypical strains of B. cereus, is also an atypical strain that is recognized as the cause of anthrax-like disease across western Africa and carries virulence plasmids pBCXO1 and pBCXO2.4,18,23,25
Aside from these differences, all these atypical B. cereus strains belong to a single phylogenetic clade and group (Clade I Group III), which also includes B. anthracis and B. thuringiensis. The aim of this study is to describe the pathologic, bacteriologic, and molecular features of an outbreak of B. tropicus infection causing anthrax-like disease in red kangaroos in Louisiana.
Materials and Methods
Animals Included in the Study and Clinical History
An outbreak associated with sudden mortalities of unknown cause was reported from a wild animal preserve in the state of Louisiana, USA, in July 2022. Within 1 week, 3 young red kangaroos (Macropus rufus) suddenly died without premonitory clinical signs (Table 1). All kangaroos were submitted to the Louisiana Animal Disease Diagnostic Laboratory, Louisiana State University School of Veterinary Medicine for complete postmortem examinations. No changes in husbandry were reported prior to the outbreak.
Red kangaroos submitted to the Louisiana Animal Disease Diagnostic Laboratory and included in this study.

Based on the preliminary postmortem results from the first 2 submitted kangaroos (cases 1 and 2), all remaining animals on the premises (n = 6) were treated by the caretakers with broad-spectrum antibiotics. Six days following the death of the first kangaroo, another young kangaroo (case 3) was found dead on the premises without any reported clinical signs and was submitted for postmortem examination at the Louisiana Animal Disease Diagnostic Laboratory.
Postmortem Examination, Sample Collection, and Histology
Postmortem examinations were performed on all kangaroos (cases 1, 2, and 3) submitted to the Louisiana Animal Disease Diagnostic Laboratory. Samples from all organs were fixed in 10% neutral-buffered formalin for 24 to 48 hours. Samples from the lungs, liver, spleen, kidneys, heart, and brain (cerebrum, cerebellum, and brainstem) were collected and stored at −80°C for ancillary testing. Impression smears from the spleen were performed at the time of postmortem examination, air-dried, and subjected to methylene blue histochemical staining following a standard procedure in the Louisiana Animal Disease Diagnostic Laboratory. Following fixation, tissues were embedded in paraffin, and sections of formalin-fixed paraffin-embedded tissues (4 μm) were stained with hematoxylin and eosin and Gram according to standard laboratory procedures prior to histologic evaluation.
Bacteriologic Culture
Samples from the spleen and liver from all kangaroos and inguinal lymph node from case 3 were inoculated onto blood agar (tryptic soy agar with 5% sheep blood), Brucella blood agar, and MacConkey agar plates (Remel, San Diego, California). Specimens for aerobic culture were inoculated on blood and MacConkey agar plates. Blood agar plates were incubated at 37°C with 5% CO2 for the first 24 hours without CO2 for the next 48 hours. MacConkey plates were incubated at 37°C for 72 hours. Specimens for anaerobic culture were inoculated on Brucella blood agar plates and incubated at 37°C with CO2 under anaerobic conditions for 7 days. Salmonella-specific cultures (tetrathionate broth with brilliant green and XLT-4 agar) from the liver and spleen of cases 1 and 2 were performed following standard procedures. Bacterial colonies were subsequently identified using a Bruker MALDI-TOF biotyper (Biotyper Compass; Bruker, Bremen, Germany) for identification.
Whole-Genome Sequencing, Assembly, and Annotation
Genomic DNA was extracted from pure bacterial colonies cultured from the spleen of case 1 using Qiagen’s DNeasy Blood and Tissue Kit (Qiagen, Germantown, Maryland) on a QIAcube automated workstation (Qiagen). DNA concentrations were determined using the Qubit dsDNA HS Assay Kit on a Qubit Flex fluorometer (ThermoFisher Scientific, Waltham, Massachusetts). DNA libraries were constructed with the Nextera XT DNA Library Preparation Kit (Illumina, San Diego, California) using 0.2 ng/µl of genomic DNA. Resultant libraries were sequenced on Illumina’s MiSeq sequencing platform using v3 sequencing chemistry with 2 × 300 bp paired-end reads per the manufacturer’s protocol. The quality assessment of the raw sequences was carried out using FastQC v0.11.9, and adapter sequences and low-quality bases were trimmed by Trimmomatic v0.39. Subsequent de novo assembly was performed using SPAdes v3.15.5, and the quality of the assembled contigs was assessed by QUAST v5.2.0. The final contigs obtained from the isolated strain, denoted as strain 11844, were subjected to BLASTn analysis to identify the reference genome with the highest identity. The draft genome of the 11844 strain was reordered utilizing Mauve v2.4.0 with B. cereus G9241 strain as the reference genome. 12 The whole-genome sequence of the 11844 strain was annotated using Bakta v1.7.0 and NCBI Prokaryotic Genome Annotation Pipeline. 35 The predicted genes were functionally annotated through BLASTp searches against the Cluster of Orthologous Group (COG) database, with an E-value threshold of <1e-5. 37 Plasmids in the 11844 genome were identified with the combination of PlasmidSPAdes and PlasmidFinder. The circular genome map of the 11844 strain, with B. cereus G9241 as the reference genome, was generated using the CGView Server (http://stothard.afns.ualberta.ca/cgview_server/ accessed on May 26, 2023). 17
Phylogenetic and Comparative Genomic Analysis
Representative species from the B. cereus group were selected for the phylogenetic analysis, with their respective nucleotide sequences retrieved from the GenBank database. Whole-genome alignments of all 16 strains were conducted using REALPHY (http://realphy.unibas.ch accessed on May 26, 2023). 5 The maximum likelihood method was employed to construct the phylogenetic tree, and evolutionary distances were computed using the general time-reversible model with gamma-distributed rates (G = 0.6) and invariant site (I) by MEGA v.11.16,36 Branch validity was assessed using the bootstrap test with 1000 replications. Average nucleotide identity values for the selected strains were calculated through the EzBioCloud server (http://www.ezbiocloud.net/tools/ani accessed on May 30, 2023), with the species boundary determined using a 95%∼96% cut-off value. 41 The genome-to-genome distance and DNA-DNA hybridization (DDH) values for the 11844 isolate and selected strains were calculated using the DSMZ service (http://ggdc.dsmz.de accessed on June 1, 2023) with a 70% cut-off for species.30,33 GenBank files of B. anthracis “Ames Ancestor” and B. cereus G9241 were used as representative reference genomes for comparative genomic analyses. The EDGAR private project was employed to the shared genes and core genomes among different strains. 6 Virulence-associated genes in the 11844 genome were identified through homology-based searches and literature-based annotation. Homologous sequences of known virulence factors were aligned against the genome of the 11844 strain using BLASTp at an E-value threshold of <1e-10.
Antimicrobial Susceptibility Testing
A sample of a fresh overnight pure culture of B. tropicus isolated from the spleen from case 2 was transferred to a sterile saline tube using a sterile cotton swab until turbidity of 0.5 McFarland standard (Sensititre Nephelometer) was reached. The sample was then plated and evenly distributed onto a 150 mm Mueller Hinton agar plate (Remel, Lenexa, Kansas), with further application of antibiotic disks. The antibiotics tested included amikacin, ampicillin, trimethoprim/sulfamethoxazole, cefoxitin, cephalothin, chloramphenicol, ciprofloxacin, clarithromycin, erythromycin, ofloxacin, piperacillin/tazobactam, penicillin, amoxicillin/clavulanic acid, neomycin, tobramycin, polymyxin B, tetracycline, ceftazidime, ceftiofur, clindamycin, doxycycline, enrofloxacin, marbofloxacin, and oxacillin + 2 NaCl. The plates were then incubated overnight at 37°C. The following day, plates were read using BIOMIC software (Giles Scientific, Santa Barbara, California), which measures zone sizes for antibiotic disks and indicates susceptibility interpretations. Susceptibility results were interpreted based on the CLSI Guidelines VET01S and BD BBL Sensi-Disk Antimicrobial Susceptibility Test Disks package inserts. Penicillin susceptibility testing had no interpretation based on CLSI guidelines (VET06 1st edition), which indicates that at this time there are no known breakpoint values available for the combination of bacterial isolate, animal species, and specimen source.
Accession Numbers
The assembled whole-genome sequence of B. tropicus strain 11844 is available at GenBank under accession number JAVRFZ000000000.
Ethical Treatment of Animals
The animals included in this study were submitted to the Louisiana Animal Disease Diagnostic Laboratory for diagnostic purposes/disease investigation. These were not animals for experimental use.
Results
Postmortem and Histologic Findings
All kangaroos were in good body condition, with pale pink oral and conjunctival mucous membranes. There was severe splenomegaly in all cases, and the splenic parenchyma was replaced by multiple, white to yellow, friable to firm nodules, measuring from 0.1 to 2.0 cm in diameter (Fig. 1a).
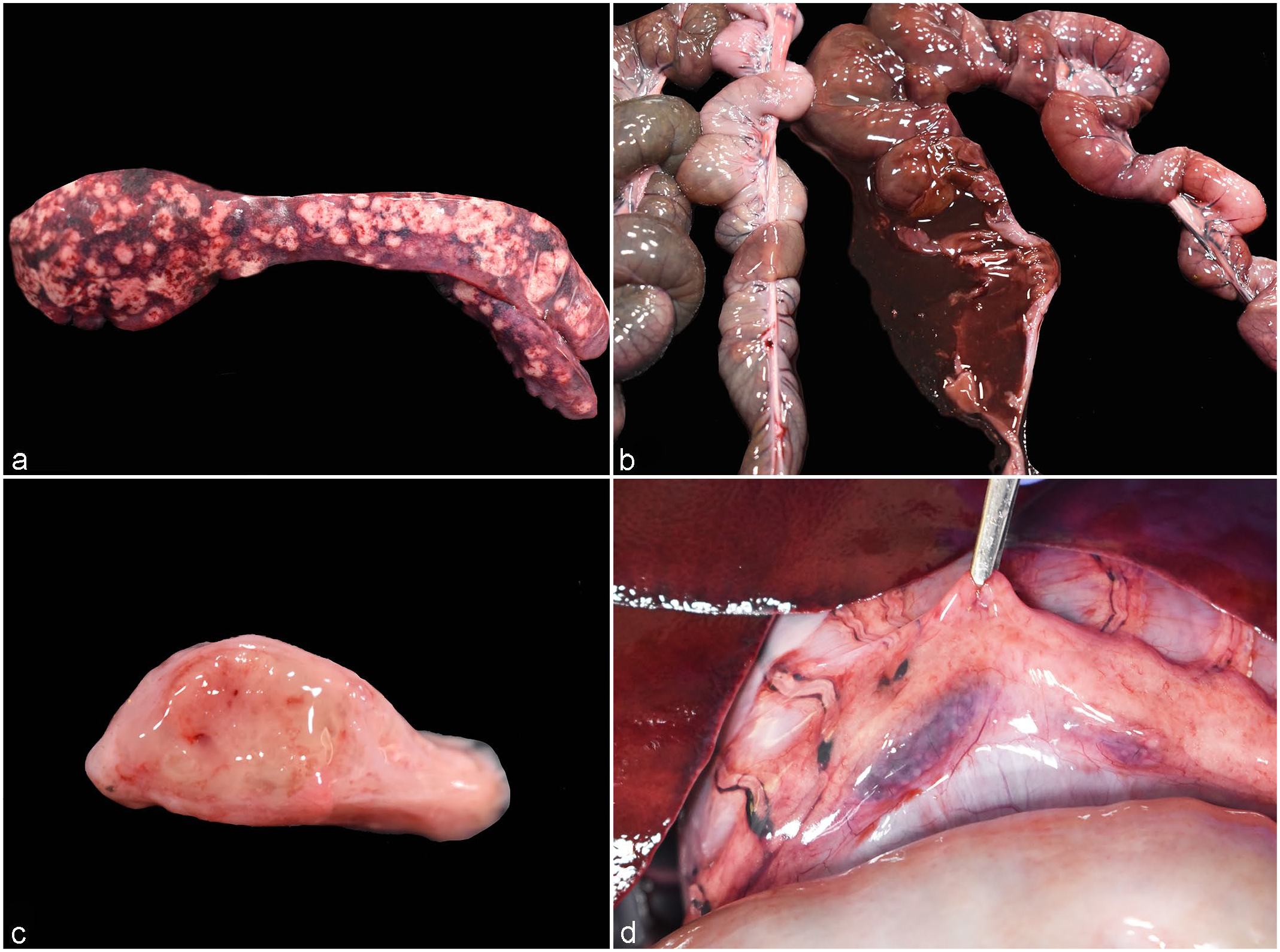
Anthrax-like disease in red kangaroos (Macropus rufus). (a) Severe splenomegaly with multifocal to coalescing, tan, raised to flat foci. Case 1. (b) The small intestinal wall is segmentally dark red, and the lumen is filled with dark red, liquid content. Case 1. (c) Markedly enlarged inguinal lymph node with petechiae and edema. Case 3. (d) Enlarged gastric lymph nodes with hemorrhage. Case 3.
In case 1, the abdominal cavity contained approximately 30 ml of red-tinged fluid, and the hepatic parenchyma had a single 1.8 × 1.7 × 1.7 cm firm area with a white to pale yellow center and a dark red rim. In case 2, the small intestinal wall was segmentally dark red, and the lumen was filled with dark red liquid content (Fig. 1b); the mesenteric lymph nodes were diffusely enlarged and dark red. The dorsal aspect of the brainstem was expanded by a focus of hemorrhage and edema. Case 3 had multiple irregularly shaped to linear excoriations on the skin overlying the sternal and flank regions, associated with multifocal subcutaneous edema and hemorrhage. The inguinal lymph nodes were bilaterally enlarged, and the nodal parenchyma was expanded by edema (Fig. 1c). The gastric and mesenteric lymph nodes were diffusely enlarged and dark red with tan foci (Fig. 1d). Throughout the glandular portion of the gastric mucosa in this kangaroo, there were multiple, variably sized ulcers (Supplemental Figure S1), and the hepatic parenchyma had multifocal, dark red pinpoint foci.
Histologically, common alterations in all animals involved the spleen, liver, and small intestine. The splenic parenchyma was affected by coalescing (cases 2 and 3) to diffuse (case 1) fibrinonecrotizing splenitis that corresponded to the yellow foci noted grossly. These areas contained numerous intralesional gram-positive bacilli with associated fibrinous exudation and were delimited by variably degenerate neutrophils (Fig. 2a, b). Within less affected areas of the splenic parenchyma, the white pulp was composed of moderately depleted lymphoid follicles, which were also multifocally disrupted by similar bacteria, and delimited by fibrinous exudation and infiltrating neutrophils (Fig. 2b). Fibrinonecrotizing lymphadenitis, with similar bacteria, was observed in case 2 (mesenteric lymph nodes) and case 3 (inguinal lymph nodes) (Fig. 2c). Necrotizing to fibrinonecrotizing enteritis (Fig. 2d) or enterocolitis with large colonies of similar bacteria were observed in cases 1 and 2, and in case 3, respectively. The area of hemorrhage and edema observed in the brainstem of case 2 corresponded to a focally extensive region of neutrophilic meningitis with large colonies of similar bacteria. The skin excoriations grossly observed in case 3 correlated histologically with epidermal necrosis, while the hemorrhagic and edematous areas in the subcutis histologically correlated with neutrophilic cellulitis with numerous intralesional similar bacteria. Several organs, including the liver, kidney, brain, thyroid gland, adrenal gland, and lungs, contained similar intravascular bacteria. Impression smears stained with methylene blue demonstrated the bacteria to be a capsulated bacillus arranged in short chains (Fig. 3). The multifocal ulcerative gastritis identified in case 3 was associated with intralesional 5 to 10 mm, round single-cell microorganisms most consistent with an amoebic species (Supplemental Figure S1).
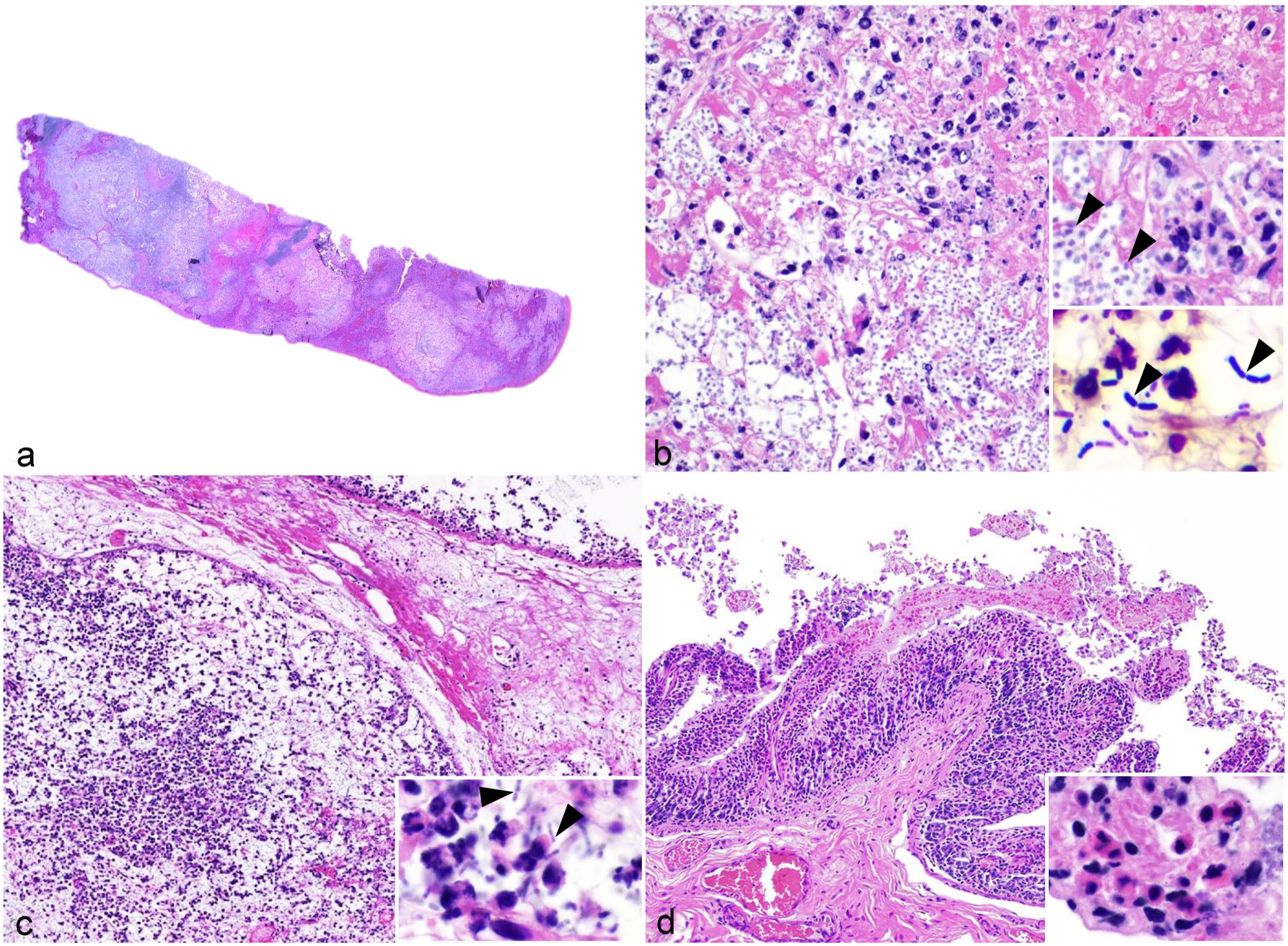
Anthrax-like disease in red kangaroos (Macropus rufus). (a) Subgross image of the spleen. The parenchyma contains extensive areas of necrosis, fibrinous exudation, and hemorrhage. Case 1. Hematoxylin and eosin (HE). (b) The areas of the splenic parenchyma affected by necrosis and fibrinous exudation contain numerous intralesional bacilli and degenerate neutrophils. Case 1. HE. Top inset: cellular debris, degenerate neutrophils, fibrinous exudation, and intralesional bacilli (arrowheads). HE. Bottom inset: intralesional bacilli are arranged in chains and are gram-positive (arrowheads). Gram stain. (c) The parenchyma of the inguinal lymph node is depleted with extensive necrosis, edema, fibrinous exudation, infiltration of degenerate neutrophils, and intralesional bacilli. Inset: degenerate neutrophils and intralesional bacilli (arrowheads). Case 3. HE. (d) The mucosa of the small intestine is infiltrated by a moderate number of neutrophils, lymphocytes, plasma cells, and histiocytes. The intestinal surface is multifocally covered by a thin fibrinous pseudomembrane. Inset: infiltrating neutrophils and fibrinous exudation within the lamina propria. Case 1. HE.
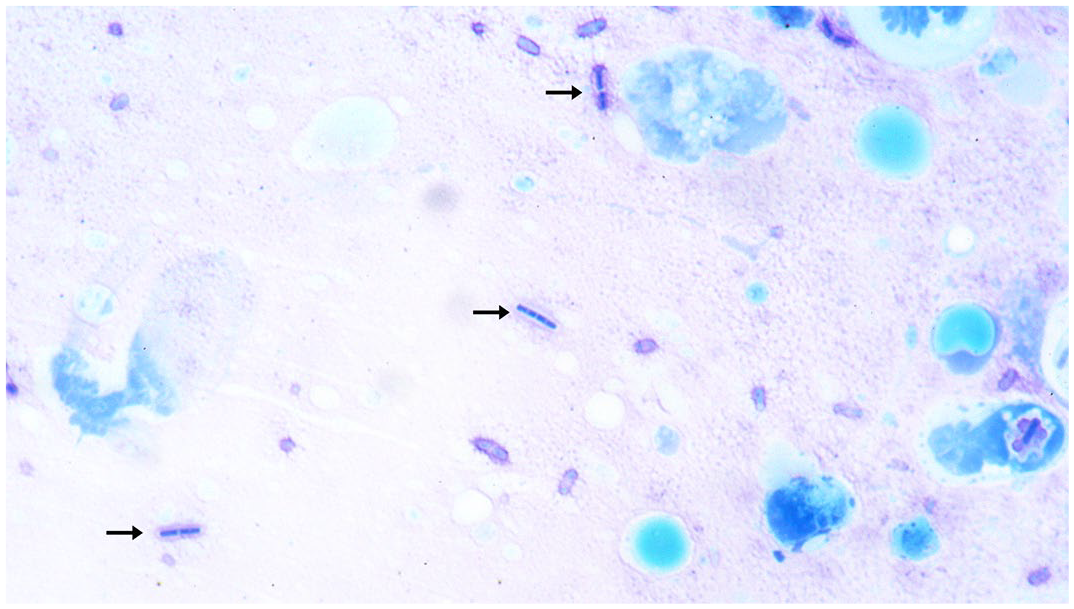
Impression smear of spleen. Methylene blue highlights intralesional capsulated bacilli (arrows) in short chains. Case 1. Methylene blue.
Bacteriology and Antibiotic Sensitivity Testing
Samples collected from the spleen and liver of all kangaroos, small intestines from cases 1 and 2, and inguinal lymph node from case 3 yielded a moderate to heavy growth of beta-hemolytic B. tropicus on blood agar plate. Cultured B. tropicus was susceptible to 2 aminoglycosides (amikacin and neomycin) and chloramphenicol. However, the cultured B. tropicus was resistant to cephalothin (cephalosporins), polymyxin B (polypeptides), and amoxicillin/clavulanic acid (Supplemental Table S1). Sensitivities were not performed in case 1. Sensitivity for penicillin was tested in cases 2 and 3; however, the lack of breakpoint values available in the literature for the species precludes the interpretation.
Whole-Genome Sequencing and Phylogenetic Analysis of Bacillus tropicus 11844 Isolate
Genome properties
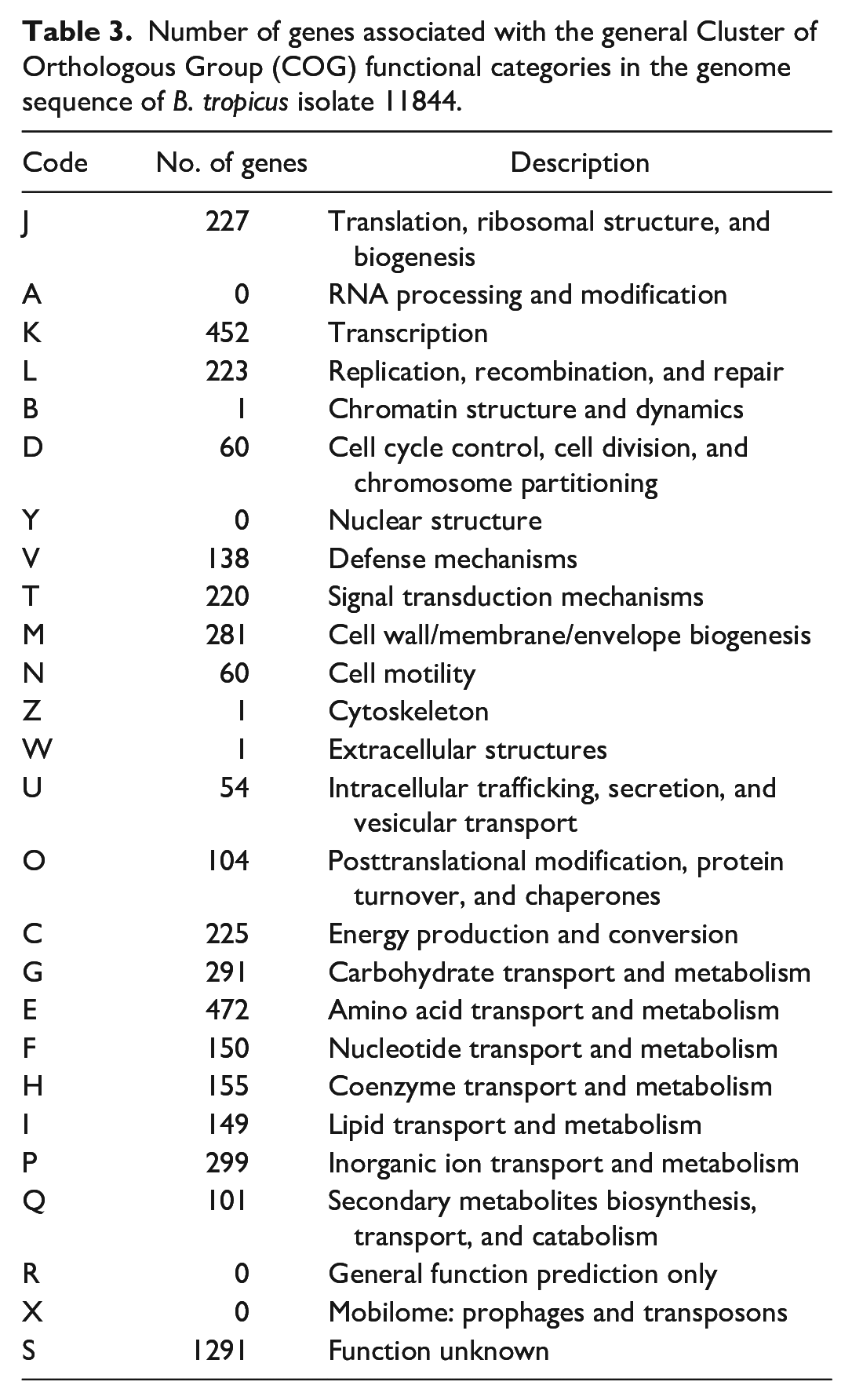
Whole-genome sequencing was performed on the Bacillus sp. 11844 isolate obtained from this outbreak of anthrax-like disease. The assembled genome contained 150 contigs, with N50 of 110,100 bp and approximately 112X sequence coverage. The full-length genome obtained had a length of 5,433,034 bp and a G+C content of 35.1%. From a total of 5642 predicted genes, 5522 were protein-coding genes, and 120 were encoded for RNA (93 tRNA, 22 rRNA, and 5 ncRNA genes) (Table 2). Approximately 87.8% of the predicted genes were assigned to 22 functional COG categories (Table 3).
Genome features of Bacillus tropicus isolate 11844 as determined by whole-genome sequencing.
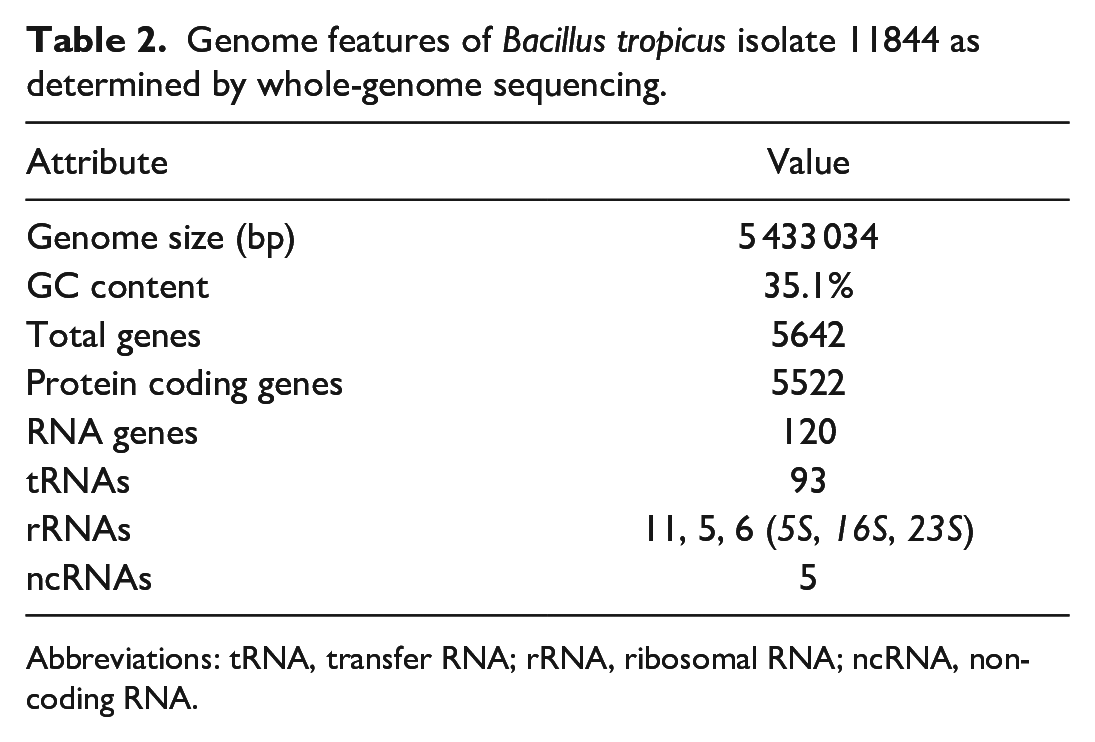
Abbreviations: tRNA, transfer RNA; rRNA, ribosomal RNA; ncRNA, non-coding RNA.
Number of genes associated with the general Cluster of Orthologous Group (COG) functional categories in the genome sequence of B. tropicus isolate 11844.
Phylogenetic analysis
Maximum-likelihood phylogenetic analysis using the whole-genome sequences of Bacillus spp. available in the GenBank revealed that the Bacillus sp. 11844 isolate clustered together with B. cereus BC-AK, B. tropicus FDAARGOS_897, B. cereus G9241, and B. cereus 03BB87 strains (Fig. 4). The genome sequence of the Bacillus sp. 11844 isolate was 96% identical to the type strain N24 of B. tropicus and shared more than 86% similarity with B. tropicus genome sequences available in the GenBank database. Next, genome sequence-based classification was performed to determine taxonomic classification employing average nucleotide identity and digital DNA-DNA hybridization (dDDH) methods. As shown in Table 4, the 11844 genome displayed the highest average nucleotide identity value (over 99.9%) and the lowest genome-to-genome distance calculation value (0.0002) to B. cereus G9241 and B. tropicus FDAARGOS_897 genome sequences, indicating that this isolate belonged to the same species and was therefore designated as B. tropicus 11844. The DDH values DDH ≤70% are interpreted as 2 distinct bacterial species, whereas DDH values ≥70% are inferred as 2 tested organisms belonging to the same species. Notably, the highest DDH value (99.9%) was observed between the whole genomes of the B. tropicus 11844 isolate and the B. cereus G9241 and B. tropicus FDAARGOS_897 strains. The data altogether indicated that the 11844 isolate belongs to B. tropicus species, previously known as atypical B. cereus. 26
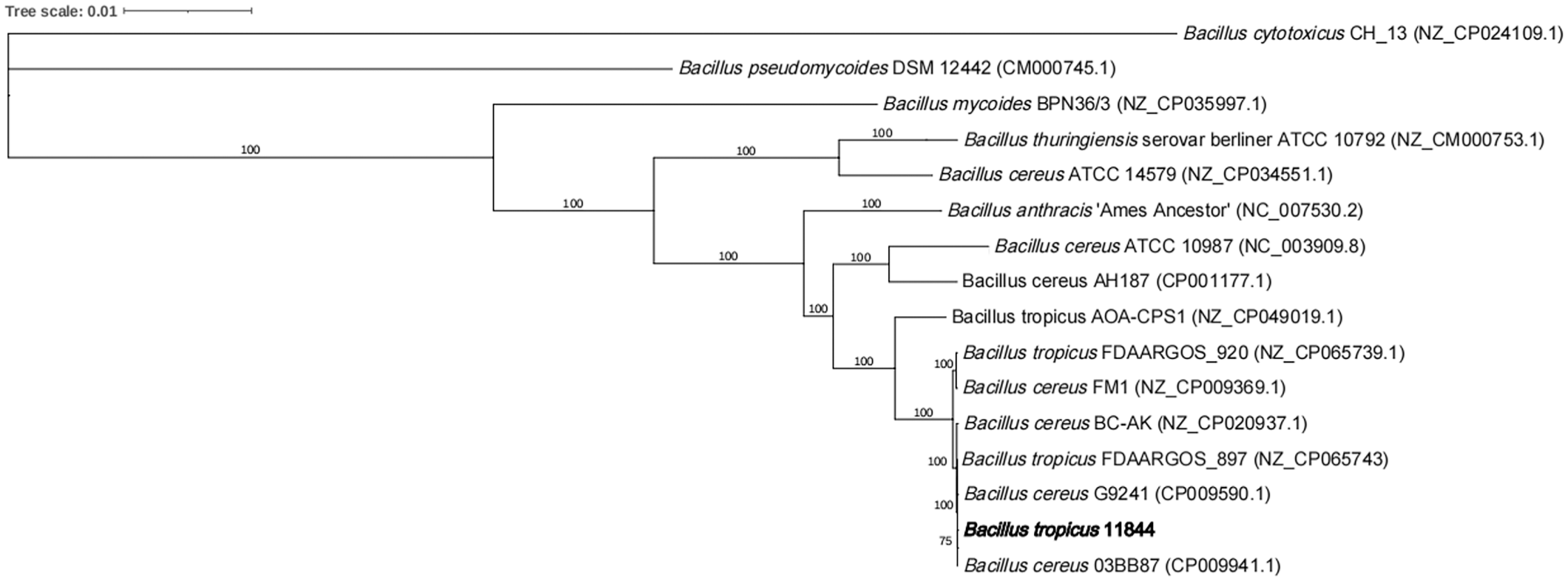
Phylogenetic analysis using whole-genome sequences of different species from the genus Bacillus identified that the isolate 11844 (in bold) clustered together with Bacillus cereus BC-AK, Bacillus tropicus FDAARGOS_897, Bacillus cereus G9241, and Bacillus cereus 03BB87 (B. cereus G9241 and B. cereus 03BB87 are classified as newly described B. tropicus due to recent taxonomic updates 28 ).
Average nucleotide identity (ANI), genome-to-genome distance calculation (GGDC), and DNA-DNA hybridization (DDH) values between B. tropicus isolate 11844 and other selected Bacillus species.
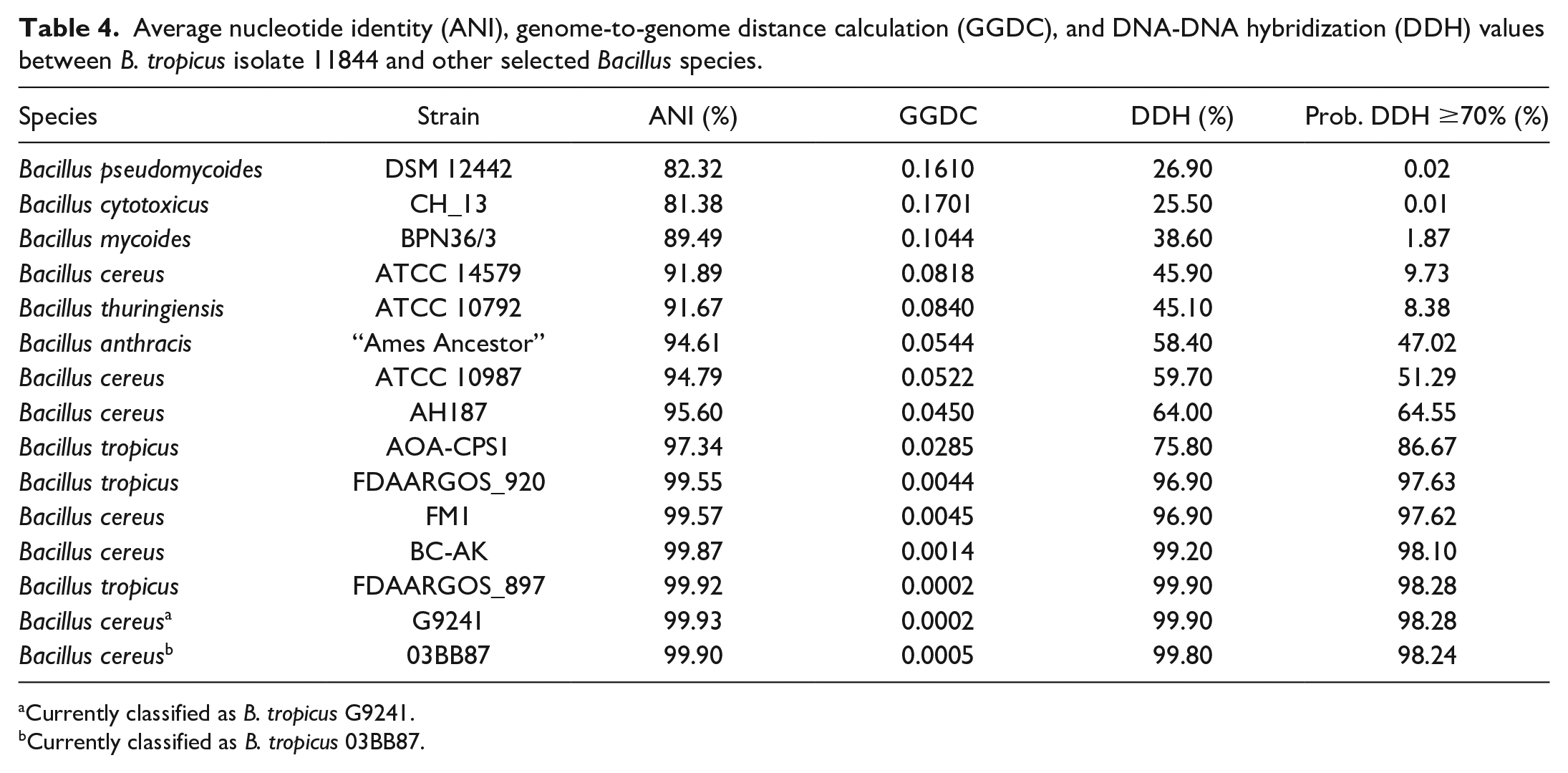
Currently classified as B. tropicus G9241.
Currently classified as B. tropicus 03BB87.
Comparative Genome Analysis
Considering the phenotypic and pathogenic characteristics as well as the phylogenetic results, atypical B. cereus G9241 and B. anthracis “Ames Ancestor” were selected as representative strains for comparative genome analysis to provide insights into the virulence factors of the B. tropicus 11844 isolate. First, the BLASTn comparison of the genome sequences between B. tropicus 11844 and B. cereus G9241 strains using the CGView Comparison Tool confirmed that the isolate belongs to the same subspecies as B. cereus G9241 (Fig. 5). Next, the genome of the B. tropicus 11844 isolate was compared with those of the B. cereus G9241 and B. anthracis “Ames Ancestor” strains using the EDGAR pipeline. As shown in Fig. 6, a total of 5585 genes were found in the B. tropicus 11844 and B. cereus G9241 genomes, among which 5058 genes (90.6%) were common to both strains. There were 471 genes unique to the B. tropicus 11844 isolate, whereas 56 genes were specific to the B. cereus G9241 strain, as shown in the Venn diagram (Fig. 6). Similarly, the B. tropicus 11844 isolate shared 4363 genes (70.4%) with B. anthracis “Ames Ancestor” out of 6194 genes. The B. tropicus 11844 isolate possessed 665 unique genes, whereas the B. anthracis strain had 1166 genes specific to itself. Furthermore, the major virulence factors were identified and summarized in Table 5. Interestingly, B. tropicus 11844 and B. cereus G9241 carried the pBCXO1 and pBC210 plasmids that possess analogous functions to the pXO1 and pXO2 plasmids of B. anthracis. The anthrax toxins (protective antigen, lethal factor, and edema factor) were observed in all 3 Bacillus strains. In addition, other important components for virulence, such as polyglutamate capsule in the B. anthracis “Ames Ancestor” genome, as well as hyaluronic acid and exopolysaccharide capsules present in the B. tropicus 11844 and B. cereus G9241 genomes were found to be associated with anthrax-like diseases. Besides, the B. tropicus 11844 and B. cereus G9241 strains contained certhrax toxin, which was absent in the B. anthracis genome. Several virulence factors of typical B. cereus, including nonhemolytic enterotoxin, hemolysin BL, and cytotoxin K, were also observed in both B. tropicus 11844 and B. cereus G9241 genomes.
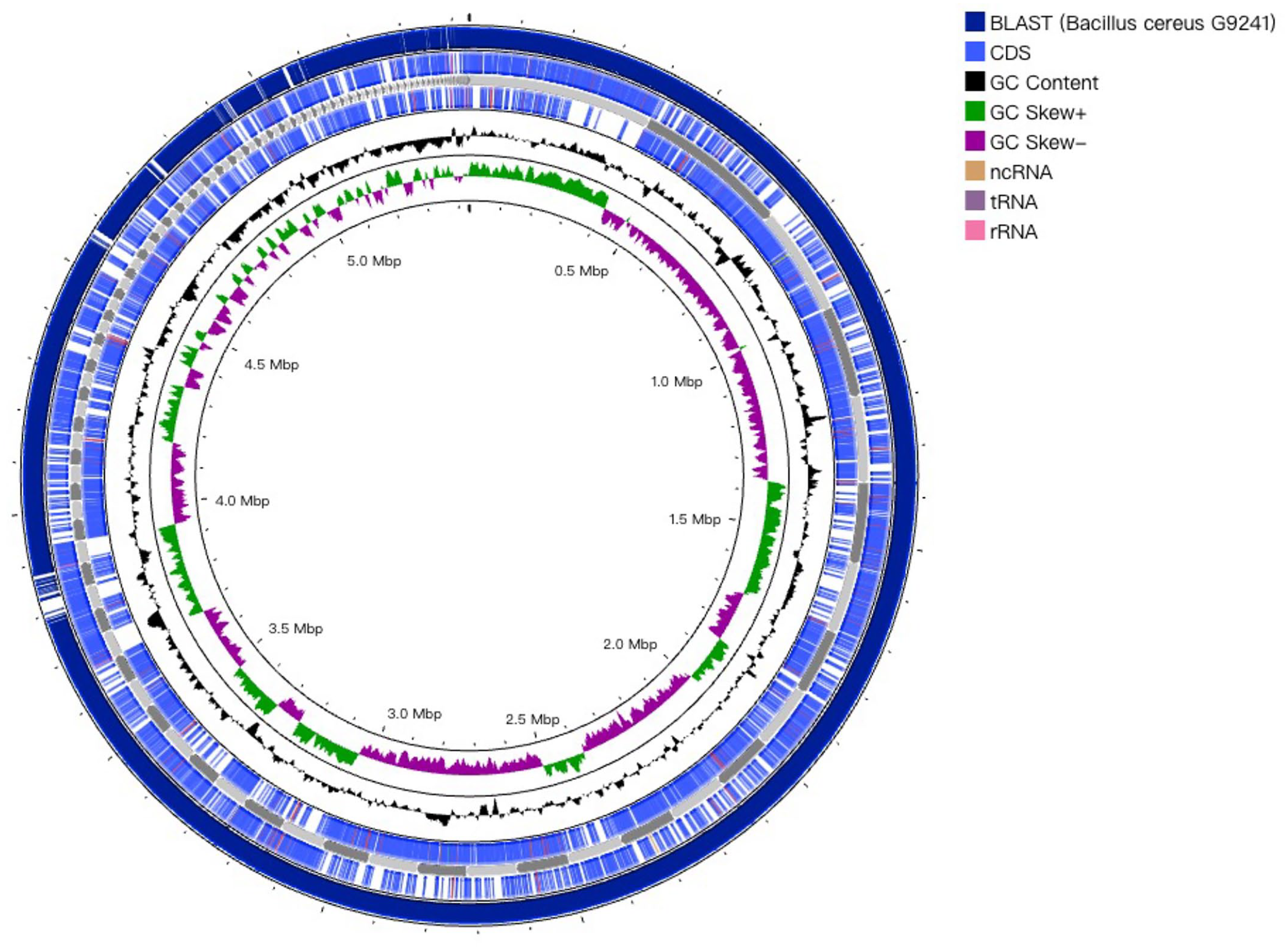
Circular map of the genome of Bacillus tropicus 11844 generated using the CGView Server. From the outside to the center: BLASTn vs Bacillus cereus G9241 (currently, B. tropicus G9241), coding sequences (CDSs) in the positive strand, CDSs in the reverse strand, GC content, and GC skew.
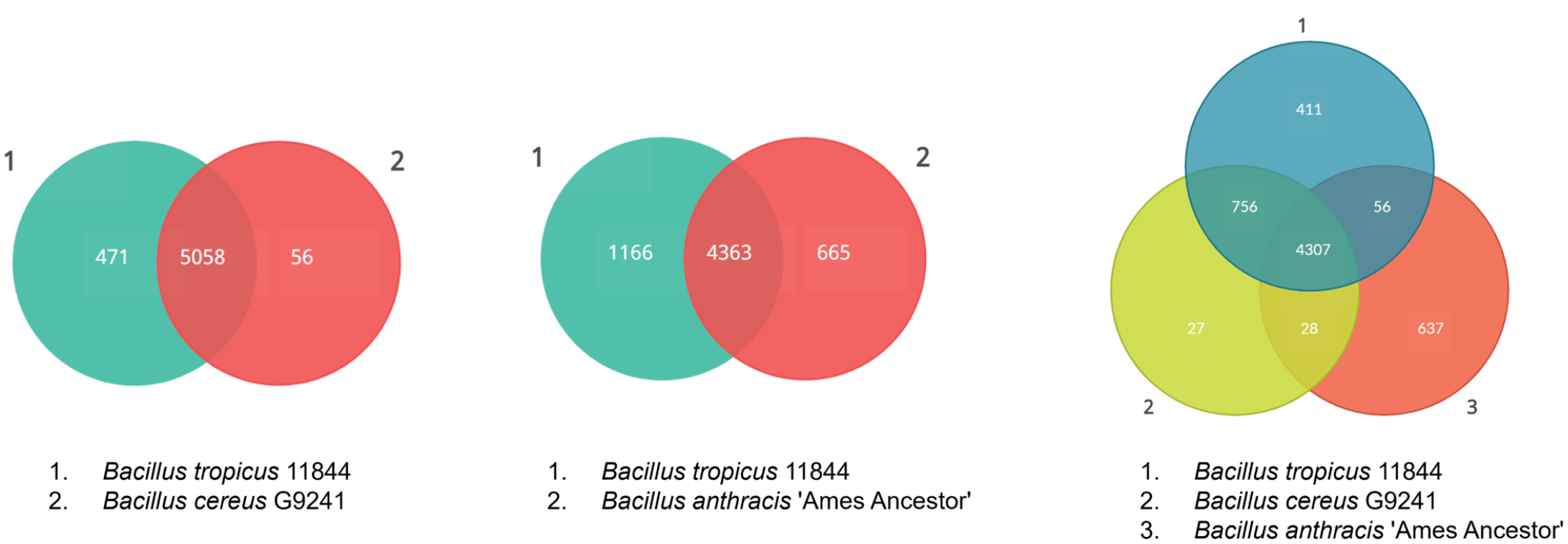
Major phenotypic differences and virulence characteristics of Bacillus tropicus 11844 isolate, atypical B. cereus G9241 strain, and B. anthracis “Ames Ancestor” genomes. The numbers indicated in the Venn diagrams represent the number of common and unique genes identified in each Bacillus sp. genome (each genome being depicted with a circle).
Major phenotypic differences and virulence characteristics of Bacillus isolate 11844, B. cereus G9241 (currently B. tropicus G9241), and B. anthracis “Ames Ancestor” genomes.
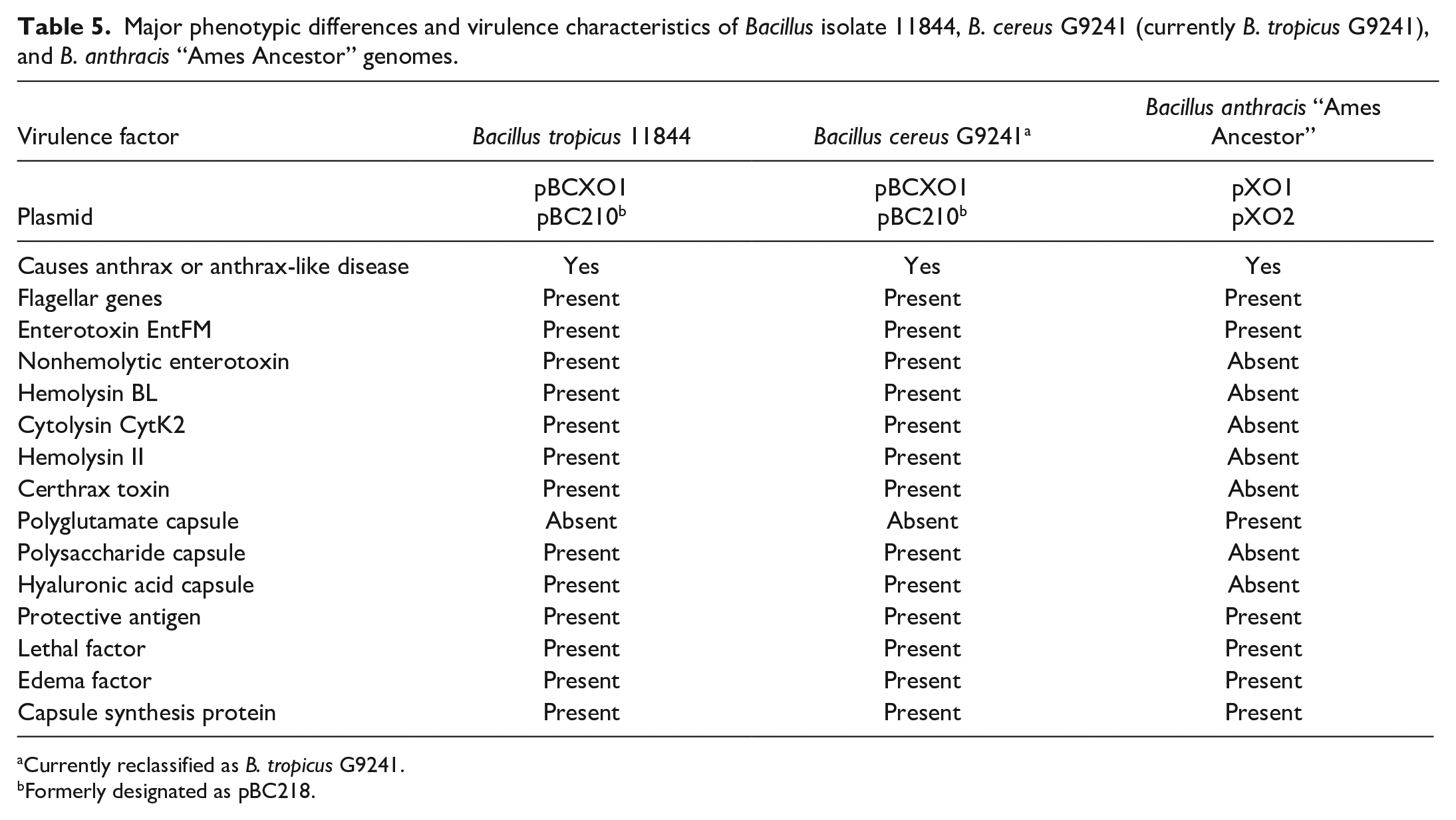
Currently reclassified as B. tropicus G9241.
Formerly designated as pBC218.
Discussion
B. cereus is a gram-positive, aerobic-to-facultative, spore-forming rod that is widely distributed in the environment and bears close phenotypic and genetic relationships to several other Bacillus species, especially B. anthracis. 2 Two variants of B. cereus cause anthrax-like disease: (1) atypical strains such as B. cereus strains G9241, FL2013, and 03BB102, which are now classified as B. tropicus and (2) B. cereus biovar anthracis strains such as CA and CI. 22 Most of the atypical strains of B. cereus isolated in human anthrax-like cases in the United States are distinct from the anthrax-like disease isolated in mammals in Africa, which are classified as B. cereus biovar anthracis strains.1,18 Atypical B. cereus strains (currently, B. tropicus) possess virulence plasmids BCXO1 and pBC210 (formerly pBC218) that are primarily involved in causing anthrax-like disease and that are similar to anthrax virulence plasmids pXO1 and pXO2.4,20 B. cereus biovar anthracis strains can be differentiated from atypical strains of B. cereus based on the absence of pBC210 and the presence of pBCXO2, which is analogous to pXO2 from B. anthracis. The Bacillus sp. identified in this study (11844) is classified as a B. tropicus strain, as it is highly similar to the atypical strain G9241 of B. cereus described as the cause of anthrax-like disease in humans in the United States 20 and possesses both the pBCXO1 and the pBC210 virulence plasmids. At the same time, our isolate exhibits several phenotypic differences with other atypical B. cereus strains, such as strain ATCC 10987, which is not associated with anthrax-like disease, and lacks toxins such as hemolysin BL, lethal factor, edema factor, and hemolysin II.
In contrast to B. cereus and B. thuringiensis, B. anthracis is non-motile, lacks beta-hemolytic activity, is sensitive to penicillin G, is sensitive to lysis by the gamma phage, and can produce a capsule in vivo and in vitro under appropriate conditions. 39 Atypical B. cereus strains, including G9241 and other B. cereus strains commonly found in the food chain, are typically penicillin-resistant.20,31 Our isolate presented resistance to cephalothin (cephalosporins), polymyxin B (polypeptides), and amoxicillin/clavulanic acid. The beta-lactam resistance observed in B. tropicus 11844 is likely due to the presence of beta-lactamase (bla) genes, which hydrolyze beta-lactams, conferring resistance to penicillins and cephalosporins. 13 The bla1 and bla2 genes in B. tropicus 11844 are identical to those found in the penicillin-resistant B. cereus G9241 strain. In addition, the regulatory elements associated with these bla genes, such as penicillin-binding proteins and sigma factors, which may influence beta-lactamase gene expression, are conserved in B. tropicus 11844.7,32 Therefore, the beta-lactam resistance in B. tropicus 11844 is likely due to these conserved bla genes and their associated regulatory elements.
Interestingly, 8 previously described atypical B. cereus strains from human cases in the United States and the one in this study were identified in the southern states, including Louisiana, Texas, and Florida.3,4,20,28 B. cereus BC-AK, also highly similar to our isolate, was presumptively identified in a kangaroo in China in 2016; however, no information regarding disease or clinical course is available. 14 Our isolate is highly similar to B. cereus G9241, which was identified in a human immunocompetent patient of Louisiana with pulmonary anthrax-like disease. Our isolate and B. cereus G9241 had a 99.9% DDH value, demonstrating the high similarity between strains. Studies have demonstrated that the atypical B. cereus G9241 strain is capable of causing fatal anthrax-like disease in immunocompromised and immunocompetent mice, while being avirulent in New Zealand white rabbits. 40 However, the study also demonstrated that for full virulence of B. cereus G9241 and development of anthrax-like disease in mice, both plasmids pBCXO1 and pBC218 must be present. 40 B. anthracis has 2 circular, extrachromosomal DNA plasmids, pXO1 and pXO2, which carry the major virulence factors required for pathogenesis. pXO1 carries the genes that encode 3 important anthrax toxin proteins, including protective antigen, lethal factor, and edema factor.2,20 The strain of B. cereus obtained in our study also contained these toxin proteins encoded within the pBCXO1, which are likely associated with the development of acute sepsis and anthrax-like disease culminating in the death of the examined kangaroos.
In previous reports, anthrax-like diseases were usually associated with inhalation of the bacterium and were described as cases of pulmonary anthrax-like disease.3,20 In addition, 2 cases of cutaneous anthrax-like disease have been observed in humans with features of anthrax eschar.4,28 In our study, the route of infection in the affected animals remains unclear. However, it is hypothesized that in the affected kangaroos this likely occurred through ingestion of the bacterium, as they had evidence of enteritis (cases 1 and 2) or concurrent amoebic (ulcerative) gastritis (case 3), which could have served as a port of entry. In case 3, a percutaneous route of infection could also be speculated, as this animal had evident excoriations on the chest accompanied by cellulitis with numerous bacilli. Although the source of infection could not be confirmed, ingestion of spoiled food and percutaneous environmental contamination were speculated to be the route of infection in this mob of kangaroos.
This outbreak raises awareness regarding the zoonotic potential of infectious agents affecting wildlife. Since such potential is more commonly encountered with wild animal populations, it is, therefore, important for pathologists and diagnostic facilities where postmortem examinations of wildlife are performed to implement and strictly adhere to biosafety practices and biosecurity measures following appropriate risk assessments. Among biosafety practices, the use of personal protective equipment is the most important component and should include safety elements to protect mucosal surfaces from exposure to infectious droplets, such as safety glasses/goggles and face masks or face shields. In addition, the use of cut-resistant gloves for additional protection against accidental cuts is highly recommended and should be strongly considered as well as the use of other primary containment devices such as biosafety cabinets for dissections whenever the animal size permits. Ideally, postmortem remains should be transported securely and destroyed on-site using appropriate disposal methods such as incineration or alkaline digestion, and postmortem examinations should be followed by effective and standardized decontamination protocols. In terms of biosecurity, enhancement of facility security and implementation of policies that prohibit transfer of specimens potentially bearing zoonotic disease(s) to taxidermists or other collectors should be strongly considered in order to minimize the risk of inadvertent exposure of humans and other animals.
In conclusion, this study describes the occurrence of a fatal outbreak caused by an anthrax-like, B. tropicus strain affecting red kangaroos in a wild animal preserve. This outbreak reported is the first to demonstrate the ability of atypical B. cereus strains to cause severe systemic infections in marsupials and is the first report of anthrax-like disease in animals in the United States. This outbreak highlights the importance of ensuring high sanitary standards and microbiological quality controls for feeds and supplements used in animal feeding. Enhanced surveillance is needed to gain further insight into specific predisposing factors, bacterial dynamics, and distribution in different animal species. Furthermore, this infection carries high relevance from a public health standpoint associated with its potential zoonotic transmission.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858241306399 – Supplemental material for Pathologic and genomic characterization of an outbreak of anthrax-like disease caused by Bacillus tropicus (formerly atypical Bacillus cereus) in red kangaroos (Macropus rufus)
Supplemental material, sj-pdf-1-vet-10.1177_03009858241306399 for Pathologic and genomic characterization of an outbreak of anthrax-like disease caused by Bacillus tropicus (formerly atypical Bacillus cereus) in red kangaroos (Macropus rufus) by Bianca Santana de Cecco, Naomi Grace Falconnier, Weiyi Chen, Yun Young Go, Laura Peak, Emi Sasaki, Christine Walsh, Maria S. Mitchell, Mariano Carossino and Fabio Del Piero in Veterinary Pathology
Footnotes
Acknowledgements
The authors acknowledge the members of the Histology and Immunohistochemistry section at the Louisiana Animal Disease Diagnostic Laboratory, School of Veterinary Medicine, Louisiana State University for their assistance.
Author Contributions
BSdC performed postmortem and histologic examinations and wrote the manuscript. NGF performed postmortem and histologic examinations and edited the manuscript. WC and YYG performed whole-genome sequence analysis and edited the manuscript. LP performed next-generation sequencing on submitted specimens and edited the manuscript. ES postmortem and histologic examinations and edited the manuscript. CW and MM performed bacteriologic culture and identification and edited the manuscript. MC performed histologic evaluations, coordinated next-generation sequencing, coordinated whole-genome sequence analysis with WC and YYG, contributed to data interpretation, and reviewed/edited the manuscript. FDP performed histologic evaluations, coordinated follow-up on all cases, and reviewed/edited the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Louisiana Animal Disease Diagnostic Laboratory, Louisiana State University School of Veterinary Medicine.
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
