Abstract
Cutaneous plaques and squamous cell carcinoma (SCC) are common in captive North American snow leopards (SLs) (Panthera uncia). Our objective was to determine whether these lesions are potentially associated with papillomavirus(es). Polymerase chain reaction (PCR) was performed on 3 cutaneous plaques using degenerate primers for papillomaviruses. A putatively novel papillomavirus was identified that shared 76% sequence identity to Felis catus papillomavirus 2. Specific PCR for this virus was performed on 5 cutaneous SCC samples and 7 normal skin samples, which were all positive. In situ hybridization for this putatively novel virus was performed, which revealed strong hybridization signals within hyperplastic cells in cutaneous plaques (n = 3) and within neoplastic cells in cutaneous SCC samples (n = 5). No hybridization signals were identified within normal skin. Ultimately, identification of a causal viral agent in the development of plaques and SCC in SLs will help guide therapeutic intervention and lay the foundation for development of prophylactic vaccines.
Snow leopards (SLs) (Panthera uncia) are iconic animals in many North American (NA) zoos and are currently listed as vulnerable by the International Union for Conservation of Nature. 7 Information gained about diseases that impact SLs will aid in the long-term management of the captive population and may have global implications for the conservation of this species.
In a 20-year mortality review of 241 captive NA SLs managed through the Association of Zoos and Aquariums SL species survival plan, malignant neoplasia affected approximately 46% of the population. 16 Squamous cell carcinoma (SCC) was the most common neoplasm, affecting 10% of the population. 16 The oral cavity, specifically the sublingual mucosa, and the skin were the most reported sites. 16 In domestic cats, papillomaviruses are associated with a subset of cutaneous SCC due to neoplastic transformation of viral plaques.9,11–14 Similar presumed viral sublingual and cutaneous plaques (CPs) occur frequently in the captive Association of Zoos and Aquariums species survival plan NA SL population affecting up to approximately 12% of animals (n = 14/124) (Timothy Georoff, SL species survival plan taxon advisory group, per personal communication). Panthera uncia papillomavirus 1 has been sequenced from oral plaques; however, a clear association between papillomavirus infection and the development of cutaneous neoplasia has yet to be established.5,8,22 Our objective was to determine whether papillomaviruses are associated with CPs and SCC in captive NA SLs.
Tissues from 15 adult captive SLs within the Association of Zoos and Aquariums species survival plan NA population were collected from archived biopsy and necropsy samples, which included 3 CP biopsies from 2 animals, 5 SCC biopsy and necropsy samples, and 7 control skin samples. Demographics for each sample are summarized in Table 1. In case 2, both CP and SCC were diagnosed on separate biopsies from different sites 1 year apart with histologic evidence of transition from CP to SCC. In the animals in which the cause of death was known, SCC was the cause of euthanasia in 1 case with metastasis to the lung, regional lymph node, and liver. Other known causes of death were generally unrelated to cutaneous lesions (Table 1). Control skin samples included follicular cysts, hypersensitivity dermatitis, dermal fibrosis, and normal skin.
Fifteen samples from 13 captive North American snow leopards.
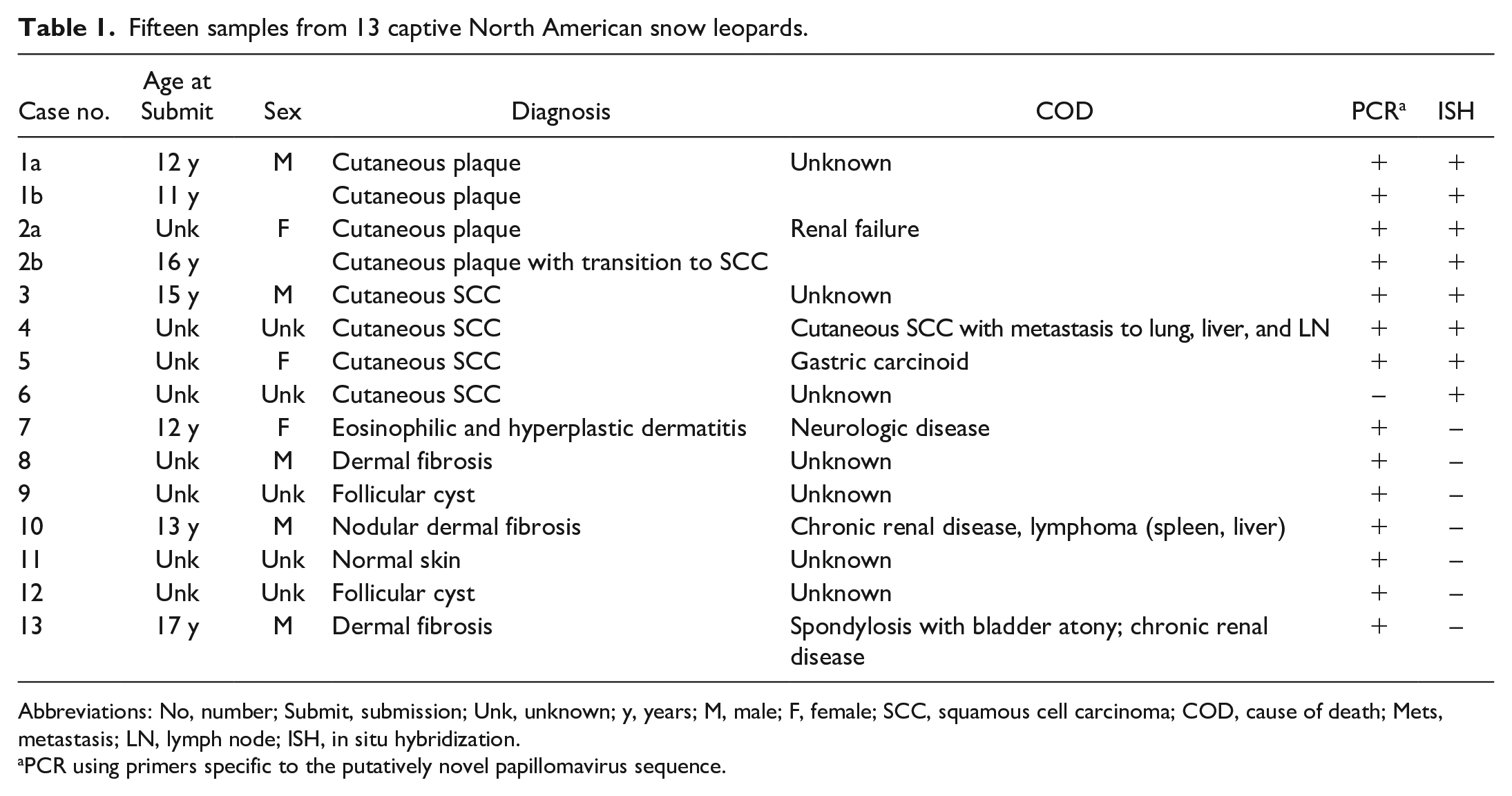
Abbreviations: No, number; Submit, submission; Unk, unknown; y, years; M, male; F, female; SCC, squamous cell carcinoma; COD, cause of death; Mets, metastasis; LN, lymph node; ISH, in situ hybridization.
PCR using primers specific to the putatively novel papillomavirus sequence.
Hematoxylin and eosin-stained slides were evaluated to confirm the diagnoses. CPs were characterized by irregular epidermal hyperplasia forming short exophytic ridges overlain by orthokeratotic hyperkeratosis. The epidermis had varying degrees of spongiosis, hypergranulosis, and koilocytosis (viral cytopathic change) with variable superficial dermal inflammation (Fig. 1A). In case 1, in both examined biopsy samples (a/b) submitted a year apart, epithelial cells had normal differentiation with orderly maturation. In case 2a, epithelial cells had mild disorderly maturation and dysplasia. In a second sample from the same animal (case 2b), submitted a year prior, there was histologic evidence of a CP with transition to SCC (Fig. 1B). In this case, and in 3 other cases of SCC, the neoplasm was characterized by anastomosing islands and cords of atypical stratified squamous epithelium supported by varying amounts of fibrovascular stroma (Fig. 1C). Neoplastic cells showed disorganized maturation and prominent intercellular bridging with centralized concentric lamellae of keratin (keratin pearls). Neoplastic cells were polygonal with a moderate amount of eosinophilic cytoplasm and round nuclei with vesiculated chromatin and prominent magenta nucleoli (Fig. 1C). In 1 animal (case 4) that was euthanized due to cutaneous SCC, the skin lesions were not available for review, but a biopsy of the right popliteal lymph node had evidence of metastatic SCC (Fig. 1D).
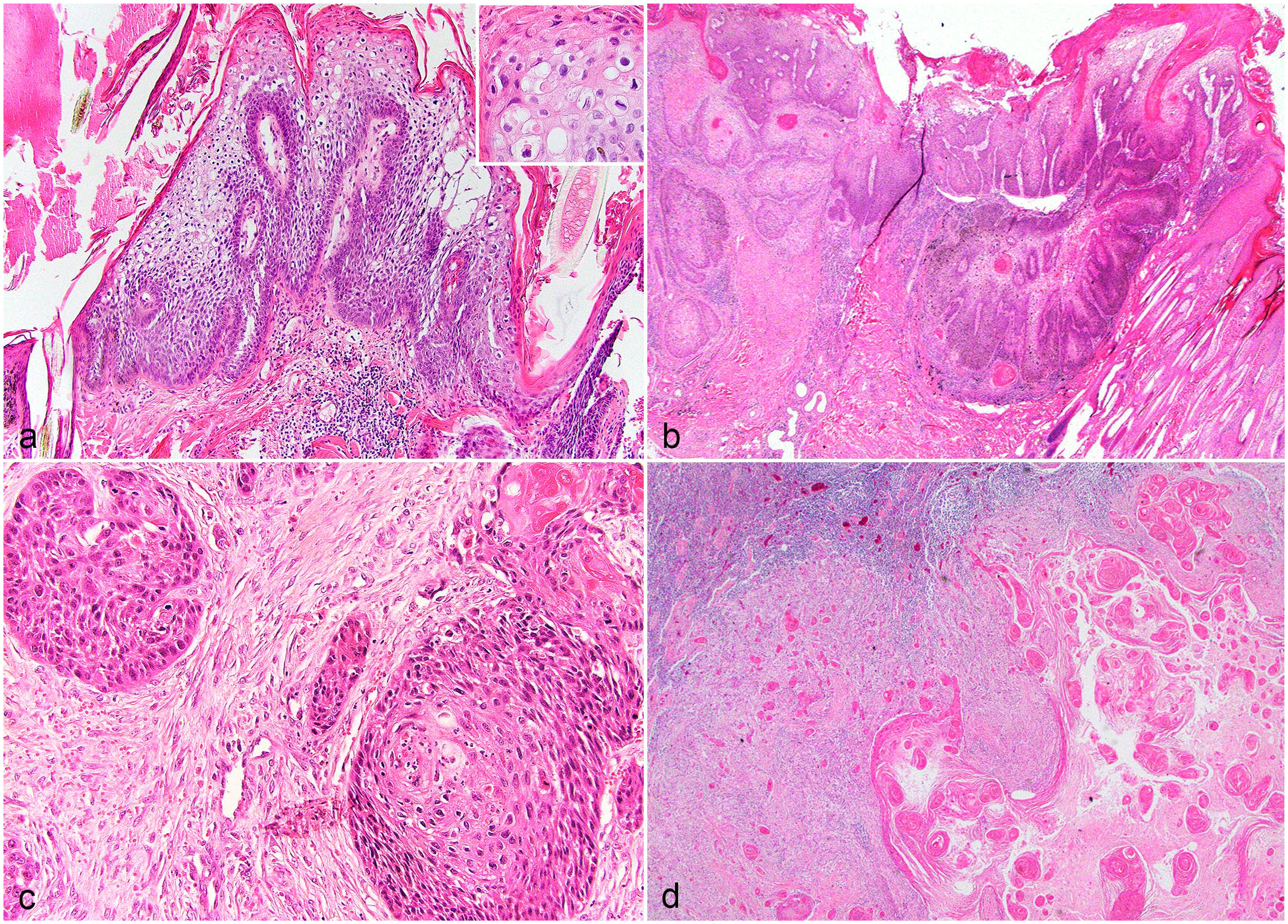
Tissues from captive North American snow leopards. Hematoxylin and eosin. (a) Haired skin. Cutaneous plaque with viral cytopathic effect. The epithelium is thickened by irregular epidermal hyperplasia forming short exophytic ridges overlain by orthokeratotic hyperkeratosis and is supported by inflamed fibrovascular stroma. The epidermis exhibits viral cytopathic effect (inset) with koilocytosis, clumping of keratin tonofilaments, and spongiosis. Case 2a. (b) Haired skin. Cutaneous plaque with transition to squamous cell carcinoma. Viral plaque exhibits epithelial cell dysplasia with transition to invasive squamous cell carcinoma as dysplastic epithelial cells invade through the basement membrane forming islands of neoplastic cells within the superficial dermis. Case 2b. (c) Haired skin. Squamous cell carcinoma. Islands and lobules of dysplastic epithelial cells invade the dermis, exhibit disorderly maturation, pleomorphism, and increased mitotic count. Case 4. (d) Lymph node. Cutaneous squamous cell carcinoma with metastasis to the regional lymph node. The lymph node is partially effaced by neoplastic squamous epithelial cells that form concentric lamellated layers of keratin (keratin pearls). Case 3.
To determine whether there is an association with papillomavirus, genomic DNA from each CP and SCC sample was extracted from two 25-m scrolls of formalin-fixed, paraffin-embedded tissue blocks using a commercially available kit (DNeasy Blood and Tissue Kit, Qiagen, Hilden, Germany), as previously described. 5 In order to develop primers specific to a SL reference gene, a polymerase chain reaction (PCR) assay was run on SL samples using primers specific for canine ribosomal protein L13a (RPL13A), K9RPL13A-For 5’-TGG GCC GGA AGG TTG TAG TCG T-3’ and K9RPL13A-Rev 5’-TTG CGG AGG AAG GCC AGG TAA TTC A-3’. Using the amplified sequence, specific primers were designed using Primer3 (v. 0.4.0) design software (https://bioinfo.ut.ee/primer3-0.4.0/) and are as follows: SL-RPL13-For, 5’-CAAGTGTAAGTTC GGACGTG-3’ and SL-RPL13-Ref, 5’-CCCCAGACACACAA ACATC-3’, which amplifies a 156 bp product. RPL13A was amplified from all samples except 1 cutaneous SCC (case 6). This sample was excluded from further PCR assays. An initial PCR was performed on genomic DNA extracted from 3 CP samples and 1 control tissue using 2 degenerate primer pairs, FAP64/6085 and MY09/MY11, which amplify a highly conserved region of the papillomavirus L1 gene, following previously published protocols.6,44 The positive control consisted of genomic DNA from a canine CP that contained papillomavirus previously shown to react with these primers. A sample without template DNA was used as the negative control. All PCR products were electrophoresed through 1% agarose, stained with SYBR Safe DNA gel stain (ThermoFisher Scientific, Waltham, Massachusetts) and visualized using a BioRad ChemiDoc Imaging System. Using both sets of degenerate primers, papillomavirus DNA was identified in 1/3 CPs (case 1a). Positive and negative controls reacted appropriately. The PCR amplicons were purified using a commercially available kit (QIAquick PCR purification kit, Qiagen) following the manufacturer’s protocol and submitted to a commercial DNA sequencing laboratory (Eton Biosciences, Research Triangle Park, North Carolina). The MY09/11 and FAP6085/64 primer sets amplify different regions of the L1 gene. To obtain a slightly larger sequence of the L1 gene, specific primers were designed that spanned the region between the amplicons generated from the MY09/11 and the FAP6085/64 primer set. These primers are as follows: PuPV-3316-For 5’-TCCAATTGAACT CGTTGCTA-3’ and PuPV-3316-Rev 5’-GCGTCATAGGAC TCAGGTTT-3’. The PCR was then performed using these specific primers on the 3 CPs samples with the following reaction conditions: 95°C for 15 minutes followed by 45 cycles of 95°C for 1 minute, 57°C for 1 minute, and 72°C for 1 minute, followed by 72°C for 10 minutes. Using these primers, papillomavirus DNA was amplified from all 3 CP samples. Amplicons from all 3 samples were purified and sequenced as above, which yielded an identical sequence from all 3 samples. The 3 amplified sequences from case 1a from the 3 different primer sets were aligned and combined to yield a 1115 bp nucleotide sequence. This sequence has been deposited in GenBank, accession number OR355483. This 1115 bp sequence was aligned to known papillomavirus sequences using the National Center for Biotechnology Information Basic Local Alignment Search Tool (BLAST) and was most closely related to Felis catus papillomavirus 2, which shared 76% sequence identity (Fig. 2A). This supports identification of a novel SL papillomavirus sequence. 9 A phylogenetic tree was constructed using Geneious Prime software based upon alignment of the L1 gene from 48 different papillomaviruses representing 17 different Genera, and the partial L1 sequence obtained from this study (Fig. 2A).
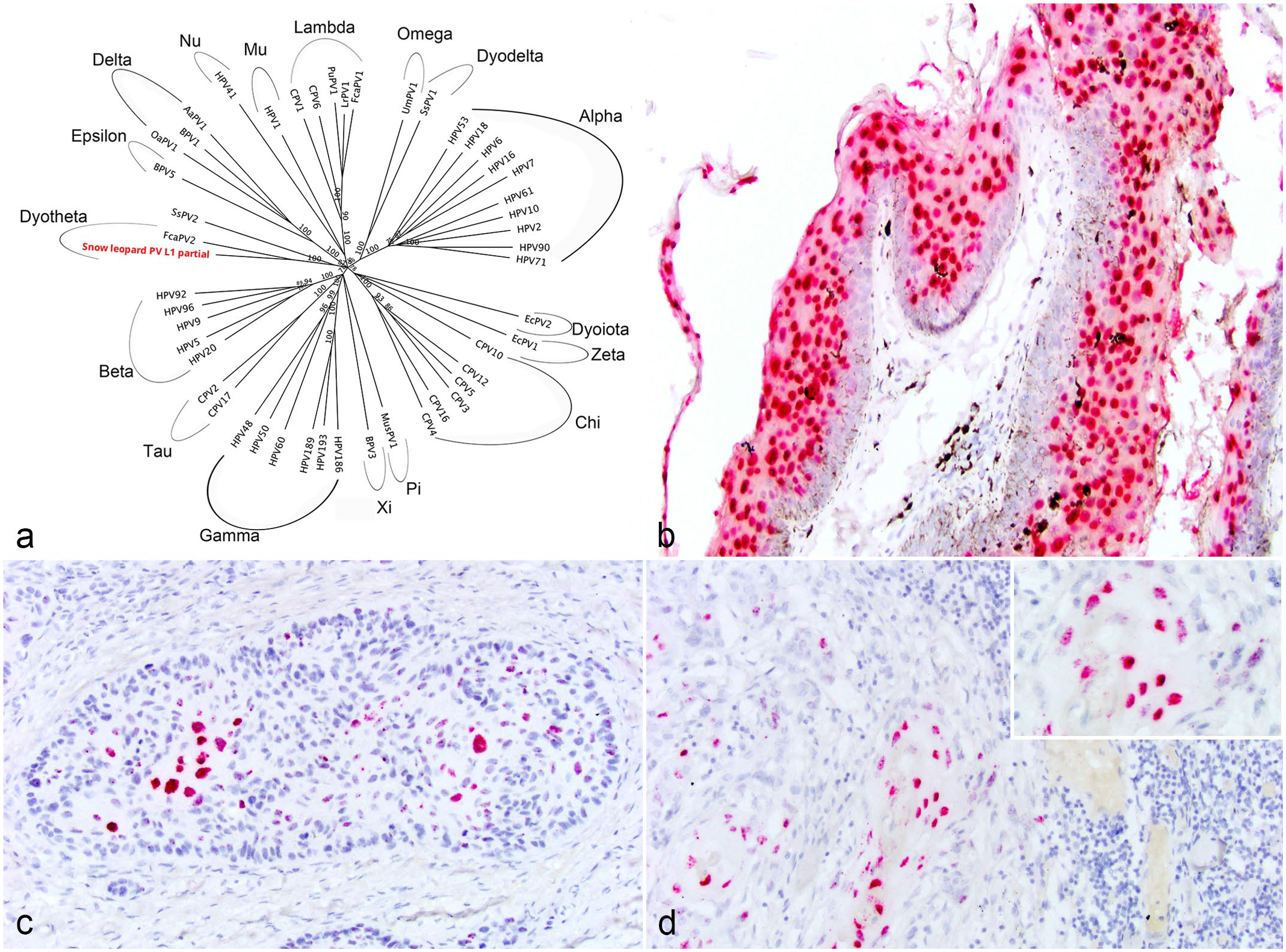
(a) A phylogenetic tree constructed based upon an alignment of the L1 gene from 48 different papillomaviruses representing 17 different genera and the partial L1 sequence obtained from this study (snow leopard PV L1 partial). The putatively novel papillomavirus shares 76% sequence identity to Felis catus papillomavirus 2. (b-d) Tissue from captive North American snow leopards. In situ hybridization specific for the putatively novel snow leopard papillomavirus. (b) Haired skin, cutaneous plaque. Epithelial cells exhibit strong nuclear hybridization signal to probes against the putatively novel snow leopard papillomavirus. Case 1a. (c) Haired skin, squamous cell carcinoma. Neoplastic epithelial cells exhibit strong nuclear hybridization signal to probes against the putatively novel snow leopard papillomavirus. Case 4. (d) Lymph node, metastatic squamous cell carcinoma. Neoplastic epithelial cells within a metastatic regional lymph node exhibit strong nuclear hybridization signal to probes against the putatively novel snow leopard papillomavirus. Inset: higher magnification image highlighting strong nuclear hybridization. Case 3.
As genomic DNA from formalin-fixed paraffin-embedded samples may be degraded and not able to amplify fragments larger than 300 bp, we designed an additional specific primer set that amplified a smaller 234 bp product using the identified putatively novel papillomavirus DNA sequence. This specific primer pair was designed using Primer3 (v. 0.4.0) design software (https://bioinfo.ut.ee/primer3-0.4.0/). The primer set included PuPV-1179-For 5’-CTTTCCCCAATGCTCGCCTA-3’ and PuPV-1179-Rev 5’-TGACAAGGAACGGTTGGTGT-3’. The PCR reaction conditions were as above for the PuPV-3316 primer set. This specific papillomavirus PCR was performed on CPs (n = 3), SCCs (n = 4), and controls (n = 7) and identified putatively novel papillomavirus DNA in 3/3 (100%) CPs, 4/4 (100%) SCCs, including a metastasis to the popliteal lymph node, and 7/7 (100%) control skin samples. These data suggest that this putatively novel papillomavirus is ubiquitous in this population, being present in all samples and control tissues. The no-template DNA control was negative.
To establish a more specific association of this putatively novel papillomavirus with the development of CPs and SCC, colorimetric in situ hybridization (ISH) was performed on all formalin-fixed, paraffin-embedded tissue samples using the RNAscope ISH method (Advanced Cellular Diagnostics, Inc, Newark, California). A 7-ZZ probed named V-PuPV-1179-L1 was designed and synthesized by Advanced Cellular Diagnostics. A probe to the bacterial gene dihydrodipicolinate reductase served as the negative control and a predesigned probe to the reference gene ubiquitin C was used to assess the presence of adequate RNA within each sample (Advanced Cellular Diagnostics, Inc). ISH was performed on 5-µm-thick, formalin-fixed paraffin-embedded sections using the RNAscope 2.5 RED assay kit following the manufacturer’s protocols. Strong hybridization signals for this novel papillomavirus were observed within hyperplastic epithelial cells of 3/3 CPs (Fig. 2B) and neoplastic epithelial cells in 5/5 SCCs, including the popliteal lymph node metastasis (Fig. 2C, D). No hybridization signals were observed in adjacent epithelial and stromal cells. No hybridization signals were identified in any of the control tissues (n = 7) or using the negative control probe. Hybridization signals for ubiquitin C were identified within all samples.
The specific localization of viral nucleic acid within both hyperplastic and neoplastic epithelial cells of CPs and SCCs combined with the histologic evidence within 1 sample of a transition between the 2 lesion types suggests a strong association between this putatively novel papillomavirus and the development of hyperplastic and neoplastic cutaneous lesions in captive NA SLs; however, fulfillment of Koch’s postulates is needed to establish a more definitive causation. Given the ubiquitous PCR prevalence even in control skin samples, it is also possible that papillomavirus may replicate in spontaneously occurring CPs without direct causation. Over the past several decades, a viral association with CPs and SCC has been speculated within the captive NA SL population and has been name Panthera uncia papillomavirus 2 in some reports; however, the full L1 sequence would be necessary for this virus to be officially named.5,22
In a recent report evaluating free-ranging SL fecal samples for DNA viruses, a sequence most similar to Felis catus papillomavirus 2 was identified, sharing 74% sequence identity to the E1 gene. 1 We speculate that this virus may be the same as the putatively novel papillomavirus we identified in captive NA SLs, suggesting that this papillomavirus may occur in free-ranging populations. This information combined with the presence of DNA from this putatively novel papillomavirus within control skin samples also suggests that this virus is common in the captive population and can be present without causing disease, reinforcing the importance of using ISH to demonstrate presence of viral nucleic acid within hyperplastic or neoplastic lesions, and to rule out a latent infection or surface contaminant.
In other species, including humans and domestic cats, papillomaviruses are known to express the oncoproteins E6 and E7.9,11,13 E6 activates telomerase, impairing cell cycle regulation while also inactivating the transformation-related protein 53.9,11,13 E7 promotes cell cycling by inactivating retinoblastoma tumor suppression protein.9,11,13 Future studies to evaluate potential oncoprotein expression in these cutaneous hyperplastic and neoplastic lesions in SLs are warranted. Ultimately, identifying a viral association with cutaneous lesions in captive NA SLs will provide guidance for the medical management of this populations and aid in the future development of prophylactic vaccines.
Footnotes
Acknowledgements
The authors are grateful to the snow leopard species survival plan taxon veterinary advisory committee Jay Tetzloff, D. McAloose, Timothy Georoff, Nancy Carpenter, and Kelly Helmick for their support with this project. The authors thank the Santa Barbora Zoo, Woodland Park Zoo, Northeastern Lincoln University, Brookfield Zoo, Los Angeles Zoo, Oregan Zoo, Zoo New England, San Franciso Zoo, and Rosamond Griffon Zoo and well as the NCSU CVM histology laboratory, for their contribution to the project. The authors also thank Cathy Minogue of Northwest ZooPath for data retrieval.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research reported in this publication was supported by the Office of Research Infrastructure Programs of the NIH under award number K01 OD123219-03.
