Abstract
Three cats, aged 2 to 11 years, presented to the University of Minnesota Veterinary Diagnostic Laboratory over a 3-year period following euthanasia or death due to respiratory distress. Thoracic radiographs revealed nodular, soft tissue opacities throughout the lung fields in all cases. On postmortem examination, approximately 60% to 80% of the lung parenchyma were expanded by multifocal to coalescing, well-demarcated, beige, semi-firm nodules. Histologically, large numbers of neutrophils, fewer macrophages, fibrin, and cellular and karyorrhectic debris effaced the pulmonary parenchyma. The inflammatory foci contained aggregates of gram-negative cocci. 16s rRNA Sanger sequencing and whole-genome sequencing identified the bacteria isolated from the lung of all cats under aerobic conditions as a novel Neisseria spp. Based on whole-genome sequence analysis, all 3 sequences shared 92.71% and 92.67% average nucleotide identity with closely related Neisseria animaloris NZ LR134440T and Neisseria animaloris GCA 002108605T, respectively. The in silico DNA-DNA hybridization identity compared to our isolates was 46.6% and 33.8% with strain DSM Neisseria zoodegmatis 21642 and strain DSM 21643, respectively. All 3 sequences have less than 95% average nucleotide identity and less than 70% DNA-DNA hybridization identity, suggesting that the 3 isolates are a novel species of the genus Neisseria. Infection with Neisseria spp. induces an embolic pneumonia in cats that radiographically and pathologically resembles a metastatic neoplastic process and should be considered among the etiologic differential diagnoses in cases of infectious pulmonary disease with a disseminated, nodular lung pattern.
Lower respiratory tract infection is an uncommon cause of morbidity and mortality in adult cats. These infections are difficult to diagnose clinically due to nonspecific or inconsistent clinical, hematologic, and radiographic findings and concomitant respiratory or systemic diseases.14,15,26 Diagnosis most commonly relies on a combination of signalment, history, physical examination, radiography, hematology, assessment and culture of bronchoalveolar lavage or pulmonary fine needle aspirates, and response to treatment, but there is no “gold standard” for diagnosis.14,15,26 Reported infectious causes associated with feline pneumonia include bacteria, viruses, fungi, protozoa, rickettsia, and parasites, with Bordetella bronchiseptica, Pasteurella spp., Mycoplasma spp., Streptococcus spp., and Escherichia coli considered the most common bacterial agents.9,14,15,26
Neisseria spp. are small, gram-negative cocci that are best characterized in humans as causative agents of gonorrhea (Neisseria gonorrhoeae) and bacterial meningitis (Neisseria meningitides). 39 They are also a reported cause of human infections secondary to dog and cat bite wounds (Neisseria animaloris and Neisseria zoodegmatis). These bacteria have been isolated as commensal flora from the oral cavity of healthy dogs and cats and have similarly been detected in these locations via molecular diagnostic techniques.14,16,18,23,33,34,39,41 Neisseria spp. infection is a seldom reported cause of feline bacterial pneumonia that was first described in 3 domestic cats in 1973, and sporadically thereafter in cases involving domestic cats, a tiger cub, a lion, and Chinese leopard cats.7,10,12,13,19,25,27,30,40 Previously known as “eugonic fermenter 4a and 4b” and biochemically characterized in 1974, the causative bacteria in previous cases of feline pneumonia were reclassified as Neisseria animaloris and Neisseria zoodegmatis, respectively, in 2006 following 16s ribosomal RNA (rRNA) gene sequence analysis and DNA-DNA hybridization (DDH) biochemical analysis. 34 Whole-genome sequencing of Neisseria spp. isolated from feline lungs with pneumonia has not been reported to the best of our knowledge.
Three privately owned, domestic short- and medium-haired cats were diagnosed with infectious pneumonia. The causative bacterium in each case was cultured. The isolates were analyzed using multiple molecular diagnostic techniques, including whole-genome sequencing. Results confirmed that all 3 instances of disease involved the same species of Neisseria and helped to characterize its genomic features, including protein makeup, relatedness to other Neisseria species, and potential antibiotic-resistance mechanisms. The results of the characterization suggest that the isolated bacteria are a novel Neisseria species.
Materials and Methods
Samples Source, Gross, and Histopathology
Three unrelated, privately owned, domestic short- and medium-haired cats from different households were submitted whole (2 cases) or as fresh and formalin-fixed tissues (1 case) to the University of Minnesota Veterinary Diagnostic Laboratory (UMN VDL) for postmortem examination between October 2018 and March 2020 following death or euthanasia due to severe respiratory compromise. Signalment, history, and clinical findings were compiled from submitted case paperwork and medical records, and discussions with individual practitioners involved with the cases.
Following submission, necropsy and/or tissue examinations were performed, and tissue samples were collected, fixed in 10% neutral-buffered formalin, and embedded in paraffin. Formalin-fixed, paraffin-embedded tissues were sectioned at a thickness of 4 µm and stained with hematoxylin and eosin, with subsequent Gram staining as needed.
Bacterial Culture and Identification
Aseptically collected samples of fresh lung were inoculated on 5% sheep blood agar, MacConkey agar, and Columbia nalidixic agar plates as part of routine aerobic culture. The plates were incubated at 37°C with 5% CO2 for 24 hours. Isolated bacterial colonies were analyzed on Bruker MALDI-TOF Compass 4.1.80 for identification. Complementary triple sugar iron slants and Gram stains were used to support identification. The isolates were submitted to the Molecular Diagnostic Lab at the UMN VDL for bacterial identification by Sanger sequencing of the 16s rRNA gene.
Whole-Genome Sequencing and Genome Characterization
Bacterial isolates were processed for DNA extraction using the EZ1 DNA Tissue Kit on the EZ1 Advanced XL instrument (Qiagen). The extracted DNA was processed for library preparation using Nextera DNA Flex library prep kit (Illumina). Libraries were submitted to the University of Minnesota Genomics Center for NextSeq 500 mid output mode sequencing with 75 bp paired-end cycle. The fastq files obtained were processed for quality check, trimming, assembly, and annotation. The quality of the raw reads was determined with FastQC v0.11.9 software. 1 Pathosystems Resource Integration Center (PATRIC) web server (https://www.patricbrc.org) was used to analyze sequences. Assembly of reads was done using Unicycler v0.4.8, an assembly software for bacterial genomes at PATRIC. 42 Error correction of assembled genomes was done with Pilon version 1.23. 37 Annotation of assembled genomes was done using the RAST tool kit (RASTtk) through the PATRIC web server.5,38 To investigate function of proteins, proteins were mapped to enzyme commission, gene ontology, and KEGG pathways.2,21,32 Furthermore, PATRIC annotation was used to assign genus-specific protein families (PLFams), cross-genus protein families (PGFams), and subsystems.11,29 Profiling of antimicrobial resistance genes and their classification into different antimicrobial resistance mechanisms was done with a k-mer-based detection method, which utilizes PATRIC’s curated collection of representative antimicrobial resistance gene sequence variants (Supplemental Table S1). 38
The 16s rRNA gene sequence analysis was done with NCBI BLAST 16S ribosomal RNA sequence (bacteria and archaea) database. 20 In silico DDH analysis was performed at the Type Strain Genome Server DDH d4 value. 28
Results
Clinical Findings
The 3 affected cats were a 2-year-old neutered male, an 11-year-old spayed female, and a 2-year-old intact female (cases 1–3, respectively). Each cat was from a separate household, with the shortest distance between household areas measuring approximately 10 miles and the longest approximately 450 miles. The cats were housed indoors for months prior to clinical signs, with up to date or previously complete but lapsed vaccination histories and no abnormal clinical signs in cohabitating cats when present. History and clinical examination findings included lethargy, anorexia, and progressive dyspnea and/or tachypnea in all cats, with cases 2 and 3 exhibiting moderate to marked left submandibular lymph node enlargement and individual cases variably exhibiting pyrexia up to 103.2°F (normal temperature in cats is approximately 100.5°F–102.5°F), dehydration, ptyalism, eyelid swelling and ocular discharge, vomiting, and bradycardia. Cases 1 and 3 were negative on feline leukemia virus and feline immunodeficiency virus testing while the status of case 2 is unknown. Cases 1 and 3 had marked neutropenia of 0.30 × 103 cells/µL and 0.28 × 103 cells/µL (reference range 2.30–10.29 × 103 cells/µL), respectively, on complete blood count. Bloodwork was declined in case 2. A fine needle aspirate of the submandibular lymph node in case 3 revealed suppurative inflammation with mixed bacterial populations.
Thoracic radiographs interpreted by board-certified radiologists showed multiple, nodular, soft tissue opacities throughout the lung fields in cases 1 and 3 (Fig. 1a). Additional findings included pneumothorax, mild pleural effusion, and consolidation in case 1, and a mild, homogeneous increase in opacity of the caudal subsection of the left cranial lung lobe in case 3. Case 2 exhibited a diffuse, broncho-interstitial pattern, peribronchial markings in the lower airways, and patchy alveolar edema. Radiographic differential diagnoses favored pulmonary metastatic disease with a differential diagnosis of an infectious disease process caused by bacteria, fungi, or parasites.
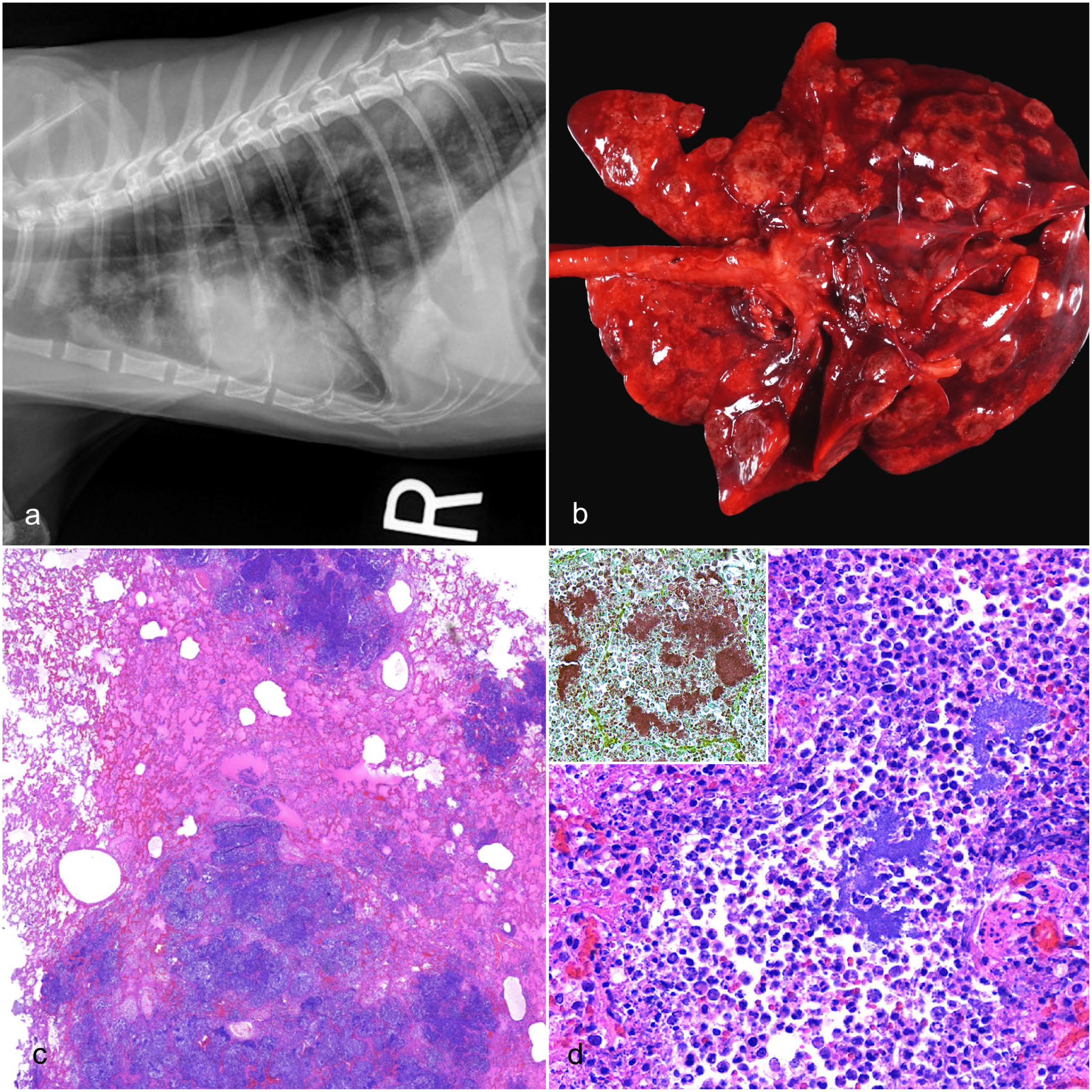
Neisseria sp. pneumonia, lung, cat. (a) On antemortem thoracic radiographs, all cats exhibited widely disseminated, nodular, soft tissue opacities in all lung lobes. (b) The lung lobes in all cats were expanded by numerous, widely disseminated, well-demarcated, multifocal to coalescing, tan to beige, semi-firm to firm, circular to irregular nodules ranging in size from 0.5 to 5.0 cm in diameter. (c) Histologically, the lungs were characterized by multifocal to coalescing, randomly distributed, well-demarcated inflammatory nodules that infiltrated and effaced the parenchyma. Hematoxylin and eosin (HE). (d) The alveolar spaces and interstitium were infiltrated, expanded, and effaced by degenerate and nondegenerate neutrophils and foamy macrophages with fibrin accumulation. Throughout the affected areas, pulmonary architecture, including alveolar septa, airways, and vasculature, exhibited coagulative to lytic necrosis. Clusters of extracellular and occasionally intra-neutrophilic and intra-histiocytic basophilic, approximately 1 µm diameter, coccoid bacteria were present in the center of the nodules. HE. Inset: coccoid bacteria were gram-negative. Gram stain.
Case 1 was euthanized due to continued decline in clinical signs despite antibiotic therapy with intravenous ampicillin. Unfixed tissue samples, including lung, were submitted to the UMN VDL after the gross postmortem examination was performed at the veterinary clinic. Case 2 died overnight following the clinical examination and initiation of subcutaneous fluids and amoxicillin/clavulanate potassium antibiotic therapy. Case 3 was euthanized prior to treatment due to a guarded prognosis following the clinical examination. The carcasses of cases 2 and 3 were submitted to the UMN VDL for postmortem examination.
Gross Lesions
Grossly, all cases were characterized by numerous, multifocal to coalescing, raised, tan to beige, well-demarcated, semi-firm to firm, circular to irregular nodules widely disseminated throughout all lung lobes (Fig. 1b). Occasionally, the exposed, raised surface of the nodules was centrally discolored dark brown to black. Nodules varied in size from up to 0.5 cm (case 1) to 2.0–5.0 cm (case 2) in diameter. The estimated percentage of lung parenchyma affected ranged from 60% (case 3) to 80% (case 2). Affected sections of the lung parenchyma sank in formalin. The remaining lung parenchyma was mottled pink to red, wet, heavy, slightly voluminous, and oozed translucent, red, watery fluid, and pink-tinged stable foam on cut section (interpreted as congestion and edema). Individual findings included enlargement of the left submandibular lymph node, enlargement of the left retropharyngeal lymph nodes, nasal turbinate reddening, hydrothorax with small amounts of hemorrhage and fibrin, and fibrinous pleural adhesions. There was no evidence of bite wounds or external trauma on any of the cats.
Histopathology
Within representative sections of lung tissue from all 3 cats, widely disseminated throughout 50% to 80% of the lung parenchyma were variably well-demarcated nodules with infiltration of the alveolar spaces and interstitium by numerous degenerate and nondegenerate neutrophils, foamy macrophages, and moderate to abundant amounts of fibrin (Fig. 1c). Throughout the affected areas, the lung parenchyma, including alveolar septa, airways, and vasculature, exhibited coagulative to lytic necrosis, with multifocal hemorrhage and numerous clusters of extracellular and occasionally intra-neutrophilic and intrahistiocytic colonies of basophilic, approximately 1 µm diameter, gram-negative cocci (Fig. 1d). There was occasional fibrinosuppurative vasculitis with rare fibrinoid necrosis of vascular walls, occasional partially to fully occlusive thrombi, and rare intraluminal rafts of cocci. Remaining intact airways occasionally contained disorganized epithelium piled up to 5 cells thick, infiltrated by neutrophils and macrophages, with patchy sloughing into airway lumina. Airway lumina multifocally contained variable amounts of fibrin, degenerate epithelial cells, neutrophils, macrophages, necrotic cellular debris, and cocci. The pleural mesothelium was plump and reactive. It was multifocally thickened by fibrin, variable numbers of degenerate neutrophils, and smaller amounts of necrotic cellular debris and macrophages. The surrounding, less affected pulmonary tissue had congested septal microvasculature, and alveolar spaces were filled with homogeneous, slightly eosinophilic material (interpreted as edema fluid), small amounts of fibrin, and moderate numbers of neutrophils and foamy macrophages. Case 2 was diagnosed with a concurrent disseminated mast cell tumor (systemic mastocytosis) affecting lymph nodes and the spleen.
Bacterial Culture and Identification
Bacteria grew on sheep blood agar but not on MacConkey agar or Columbia nalidixic agar plates (Supplemental Figure S1). Microflex LT MALDI-TOF mass spectrometer (Bruker Daltonics) calibrated prior to sample testing using the bacterial test standard, as per the manufacturer’s recommendations, identified the samples as Neisseria animaloris. The triple sugar iron slant and Gram stain were consistent with Neisseria species. The 16s rRNA Sanger sequencing analysis also confirmed the isolates as Neisseria species.
Whole-Genome Sequence Analysis and Characterization
The whole-genome sequencing of the isolated bacteria was performed using the Illumina (short read) sequencing platform and approximately 8 million reads were added to each isolate. The sequencing reads were assembled de novo to yield a draft genome. The resulting sequence consisted of a 2,436,300 bp genome length with G+C content of 50.06% (Table 1). The sequences from all 3 isolates were identical and related to the same species/subspecies as indicated by the percent identity (>99.99%), DDH Score (99.9), and GC% differences (<0.01%). This genome has 2675 protein coding sequences, 47 transfer RNA genes, and 3 rRNA genes. The annotated genome contained 901 hypothetical proteins and 1774 proteins with functional assignments (Table 2 and Supplemental Figure S2).
Statistics of the assembled sequence of novel Neisseria species.
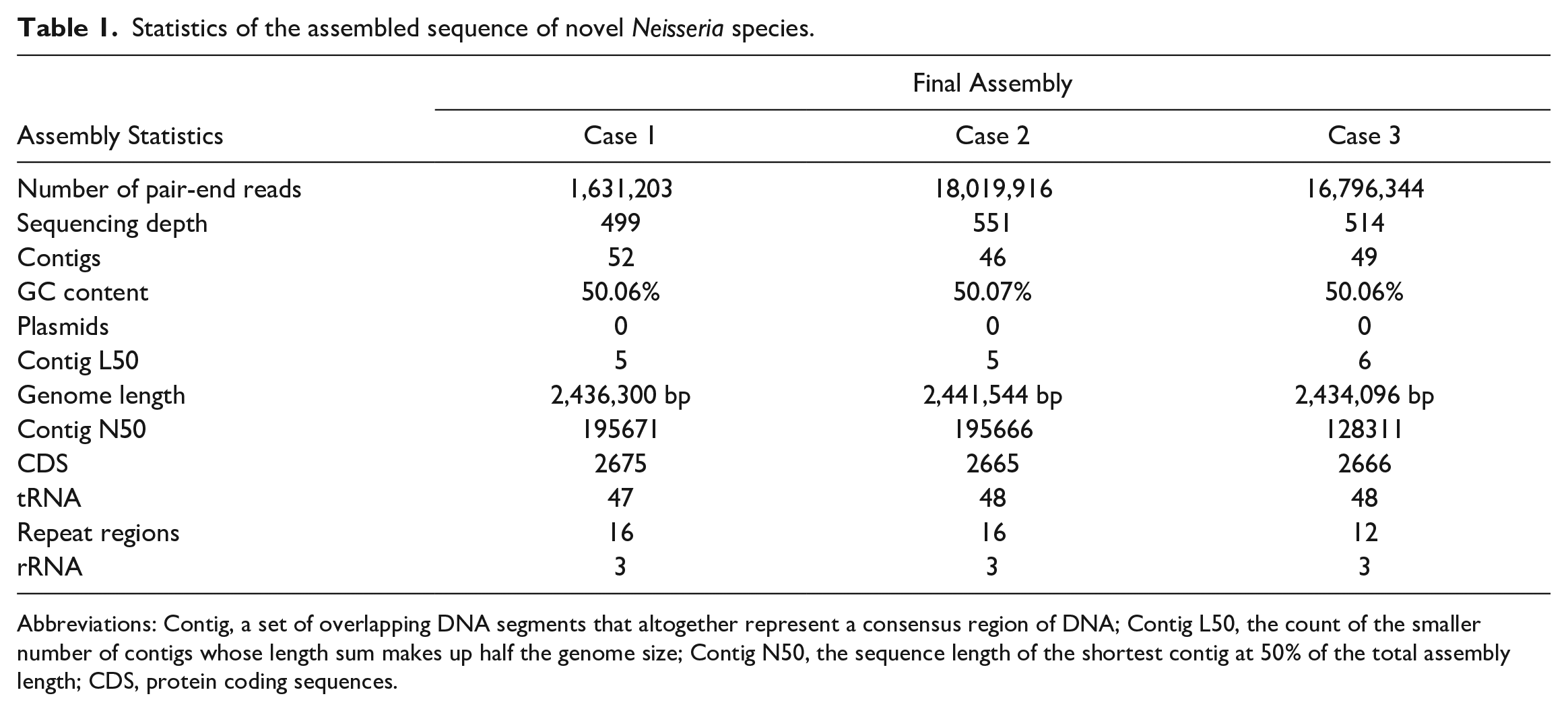
Abbreviations: Contig, a set of overlapping DNA segments that altogether represent a consensus region of DNA; Contig L50, the count of the smaller number of contigs whose length sum makes up half the genome size; Contig N50, the sequence length of the shortest contig at 50% of the total assembly length; CDS, protein coding sequences.
Annotation of the novel Neisseria species with a breakdown of protein features.
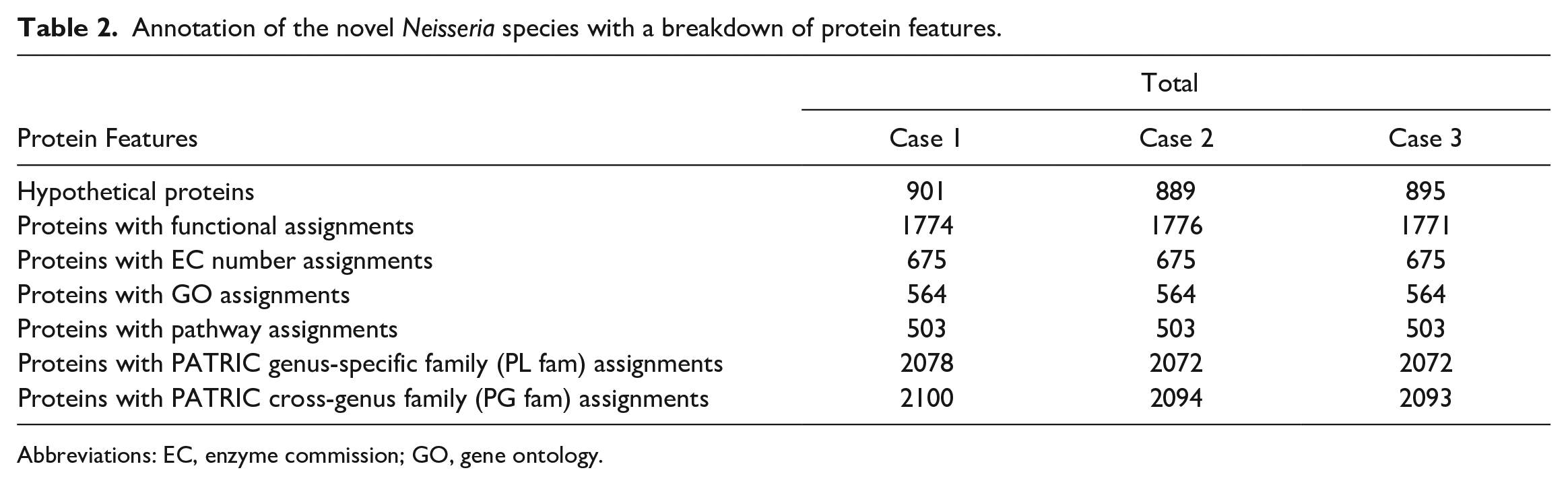
Abbreviations: EC, enzyme commission; GO, gene ontology.
A BLAST analysis of the 16S rRNA gene from the Neisseria sp. genome against the NCBI 16S rRNA gene sequence database for bacteria and archaea indicated that it has a unique 16S rRNA sequence with 97.21% to 97.88% nucleotide identity to the most closely related Neisseria species such as N. zalophi, N. canis, N. zoodegmatis, and N. animaloris. In silico DDH was performed using the Type Strain Genome Server to compare Neisseria sp. to the type strains in the Type Strain Genome Server database. The bacteria with the highest predicted DDH to the isolated Neisseria sp. were Neisseria animaloris strain DSM 21642 (d4 of 46.6%), Neisseria zoodegmatis strain DSM 21643 (d4 values of 33.8%), and Neisseria weaveri strain LMG 5135 (d4 values of 31.8%). An approximate average nucleotide identity (ANI) of Neisseria sp. calculated using the Microbial Genome Atlas (http://microbial-genomes.org) online server found the closest relatives within the database to be Neisseria animaloris NZ LR134440T (92.71% ANI) and Neisseria animaloris GCA 002108605T (92.67% ANI) (Table 3). Together, the 16S rRNA gene sequencing, DDH, and ANI results indicate that the isolated bacterium is a novel Neisseria species and its closest related species is Neisseria animaloris (Figs. 2, 3). The sequences were submitted in GenBank under the BioProject IDs PRJNA993729, PRJNA993730, and PRJNA993732 and accession numbers JAUKWG000000000, JAUKWH0 00000000, and JAUKWI000000000 for 41461, 42950, and 10259 sequences, respectively.
Average nucleotide identity (ANI Value).

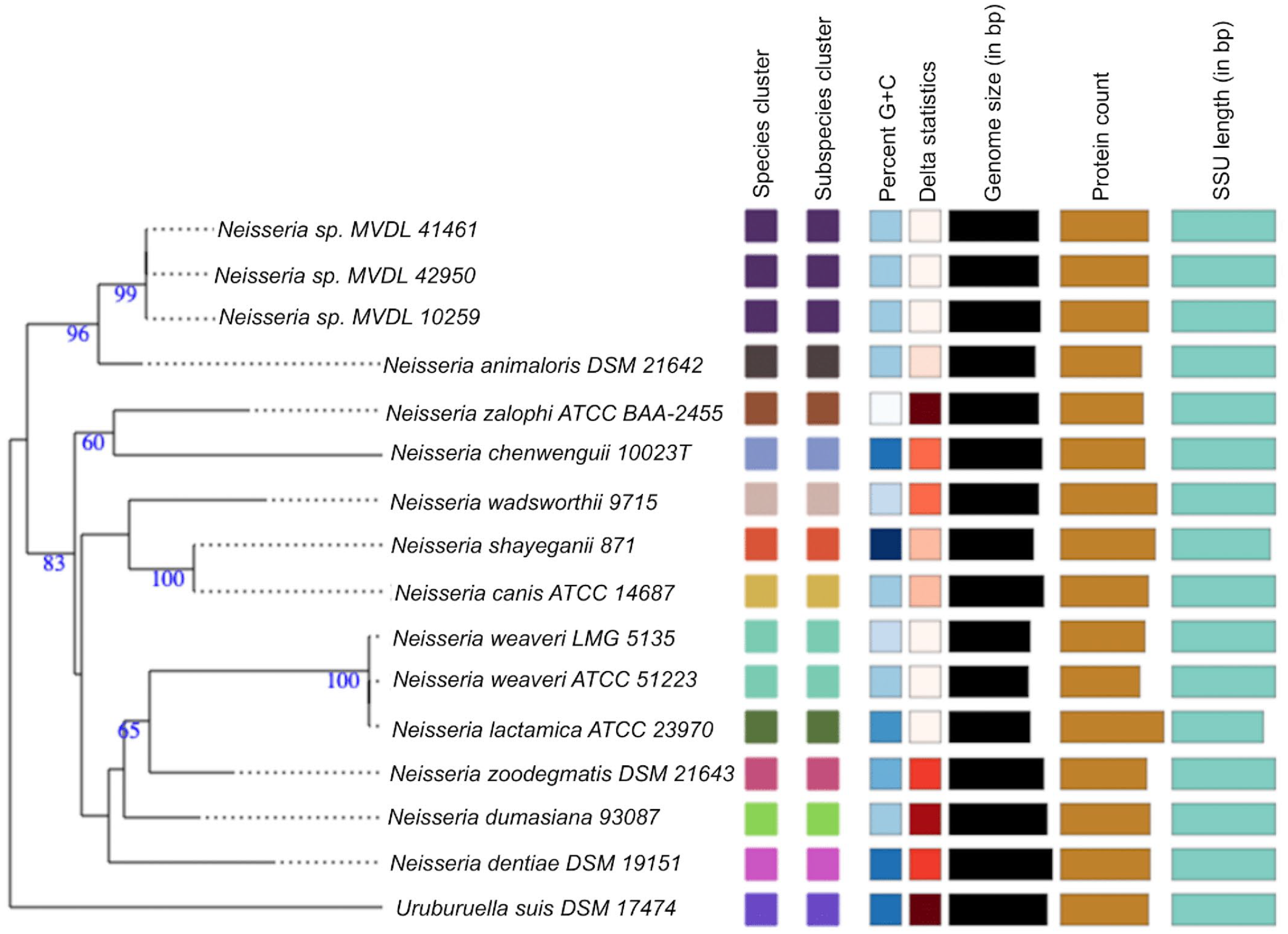
Neisseria species and subspecies delineation based on the Genome BLAST Distance Phylogeny (GBDP) phylogenetic analysis retrieved from the Type (Strain) Genome Server (TYGS) website based on whole-genome sequence. The tree was inferred with FastME from GBDP distances calculated from genome sequences. The branch lengths are scaled in terms of GBDP distance formula d5. The numbers above branches are GBDP pseudo-bootstrap support values >60% from 100 replications. Color coding represents similar or dissimilar features/characteristics among species in tree, while bar length represents comparative measure for respective features/characteristics in columns. Delta statistics allow for assessment of phylogenetic accuracy in terms of treelikeness, with lower delta value representing higher accuracy. G+C, guanine and cytosine; bp, base pair; SSU, small subunit rRNA.
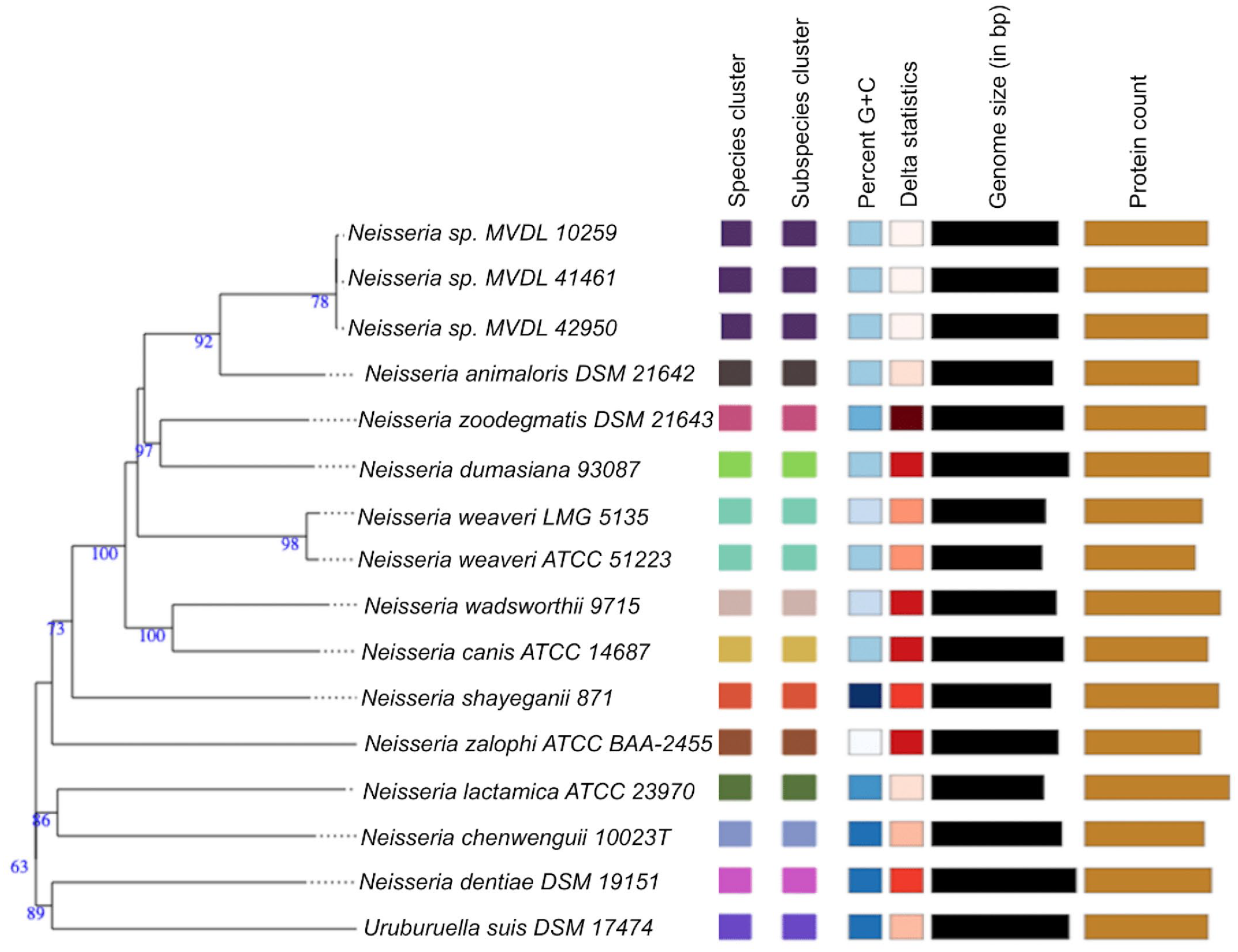
Neisseria species and subspecies delineation based on the Genome BLAST Distance Phylogeny (GBDP) phylogenetic analyses retrieved from the Type (Strain) Genome Server (TYGS) website based on 16S rRNA. The tree was inferred with FastME from GBDP distances calculated from genome sequences. The branch lengths are scaled in terms of GBDP distance formula d5. The numbers above branches are GBDP pseudo-bootstrap support values >60% from 100 replications. Color coding represents similar or dissimilar features/characteristics among species in tree, while bar length represents comparative measure for respective features/characteristics in columns. Delta statistics allow for assessment of phylogenetic accuracy in terms of treelikeness, with lower delta value representing higher accuracy. G+C, guanine and cytosine; bp, base pair.
Discussion
Neisseria sp. pneumonia has been sporadically reported in cats since 1973, with all cases exhibiting similar, relatively unique radiographic, gross, and histologic findings.7,10,12,13,19,25,27,30,40 Cases are most commonly reported in young adult cats.7,10,12,14,19,27 Antemortem diagnostic results are variable, with the most consistent finding being discrete, multifocal nodular densities and/or lung consolidation on thoracic radiographs with leukopenia and degenerative left shift on bloodwork.14,19 Gross lesions consist of widespread, white to tan nodules throughout the lung parenchyma, with microscopic changes of multifocal, necrosuppurative, and histiocytic embolic pneumonia with intralesional gram-negative cocci.7,10,12,19 While relatively unique, the reported radiographic and gross findings are not pathognomonic for Neisseria sp. pneumonia. Similar patterns have been reported in neoplasia and rarely other infectious causes, including Yersinia pestis, Cryptococcus neoformans, Toxoplasma gondii, Mycoplasma spp., Mycobacterium spp., Rhodococcus equi, and fungal, parasitic, or other sporadic bacterial infections.6,14,15 Impression smears and cytology on the necropsy floor may allow for the initial differentiation as a coccal bacterial pneumonia rather than fungal, neoplastic, or other disease process, with aerobic culture allowing further speciation.
The pathogenesis of feline Neisseria sp. pneumonia is not clear, nor is the reason for the acute and rapid nature of deterioration.7,26,30 However, the pneumonia is likely due to hematogenous spread, potentially from the oral cavity given its commensal nature. A review of 6 cats with Neisseria sp. pneumonia included 5 cats that had clinically reported gingivitis prior to respiratory infection. 26 One proposed mechanism is that chronic infection circumvents the host’s immunity, leading to a period of asymptomatic bacteremia and hematogenous spread to locations that favor survival and growth, subsequently leading to severe, acute on chronic exacerbation of disease. 4
Immunosuppression has been hypothesized as a potential causative factor for the pathogenesis in previous case reports due to histories of “lost” animals or feline leukemia virus/feline immunodeficiency virus positivity, with a single case report of an epizootic outbreak of disease in a colony with enzootic upper respiratory viral disease and a high rate of feline leukemia virus infection.7,14,40 However, immunosuppression was not a consistent finding in previous reports, and while 2 out of the 3 cats submitted to the UMN VDL exhibited marked neutropenia, this is a nonspecific finding and may have arisen either secondary to chronicity of disease, secondary to factors within the bacterium itself, or due to a different cause prior to infection or exacerbation of Neisseria sp.-related disease. Neisseria spp. infections have been reported to affect the immune system of the host in a variety of ways, including through disruption of the complement system and dampening of the adaptive immune response in N. meningitidis and N. gonorrhoeae, with potential widespread conservation of genes required for persistence between Neisseria species.22,24,31
To the authors’ knowledge, the cases at the UMN VDL represent the first cases of novel feline Neisseria sp. with whole-genome sequence-based characterization. The 16s rRNA gene sequence analysis showed 97.21% to 97.88% nucleotide identity to the closest relatives, such as N. zalophi, N. canis, N. zoodegmatis, and N. animaloris. A similar identity was shown by previous reported novel Neisseria species such as Neisseria chenwenguii sp. nov. isolated from the rectal contents of a plateau pika, Neisseria zalophi sp. nov., and Neisseria montereyensis sp. nov. isolated from the oral cavity/oropharynx of a California sea lion.35,36,43 Based on whole-genome comparisons, our feline cases were all caused by the same species of Neisseria, with all 3 sequences being 99.9% identical to one another. The predicted DDH (d4) value of ≤70% and ANI of ≤95% have been accepted as species-defined thresholds for describing novel prokaryotic species at the genomic level.3,8,17,28 In this study, the nearest Neisseria species has less than 95% ANI and less than 70% DDH identity compared to our isolates, suggesting that the 3 isolates are a novel species of the genus Neisseria.8,35,36 Similar criteria have been used to classify novel Neisseria zalophi and Neisseria montereyensis isolated from a sea lion.35,36 The Neisseria isolates from previously reported cases of feline pneumonia were known as eugonic fermenter 4a and 4b, before they were reclassified to Neisseria animaloris and Neisseria zoodegmatis in 2006 following 16s rRNA gene sequence analysis and DDH analysis. 34 The relationship and comparison between the novel Neisseria sp. in these cases with those previously classified as N. animaloris via other techniques is not known. Further phenotypic analysis will be helpful in better understanding of this novel Neisseria sp.
Unfortunately, the prognosis of feline Neisseria sp. pneumonia is poor, with decline and death generally within hours to days of the onset of clinical signs, and only a single case report of successful treatment following antemortem diagnosis via lung fine needle aspirate. 14 Neisseria sp. pneumonia should be included in the listed differentials for a pulmonary nodular radiographic pattern in cats and in cases of nodular, necrosuppurative pneumonia in feline patients.
Supplemental Material
sj-docx-1-vet-10.1177_03009858241235392 – Supplemental material for Embolic necrosuppurative pneumonia in domestic cats induced by a novel Neisseria species
Supplemental material, sj-docx-1-vet-10.1177_03009858241235392 for Embolic necrosuppurative pneumonia in domestic cats induced by a novel Neisseria species by Christopher R. Bolt, Vikash K. Singh, Arno Wünschmann, Hallie C. Richards, Kelly L. Gehlhaus and Sunil K. Mor in Veterinary Pathology
Footnotes
Acknowledgements
We thank Drs Christina Foutz and Emma Torii for their help in initial gross and histologic examination of the cases, as well as the necropsy, histology, bacteriology, and molecular diagnostic laboratory staff for their help in diagnosis and preparation of materials. In addition, we thank Dr Tim Johnson for his invaluable help in critical revision of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
