Abstract
Canine bufavirus (CBuV) or Carnivore protoparvovirus-3, a nonenveloped DNA virus belonging to the genus Protoparvovirus, family Parvoviridae, has been identified in dogs with respiratory and enteric diseases. Although CBuV detection has been reported in multiple countries, descriptions of pathologic findings associated with infection have not yet been provided. In this study, the authors necropsied 14 dogs (12 puppies and 2 adult dogs) from a breeding colony that died during multiple outbreaks of respiratory diseases. Postmortem investigations revealed extensive bronchointerstitial pneumonia with segmental type II pneumocyte hyperplasia in all necropsied puppies but less severe lesions in adults. With negative results of common pathogen detection by ancillary testing, CBuV DNA was identified in all investigated dogs using a polymerase chain reaction (PCR). Quantitative PCR demonstrated CBuV DNA in several tissues, and in situ hybridization (ISH) indicated CBuV tissue localization in the lung, tracheobronchial lymph node, and spinal cord, suggesting hematogenous spread. Dual CBuV ISH and cellular-specific immunohistochemistry were used to determine the cellular tropism of the virus in the lung and tracheobronchial lymph node, demonstrating viral localization in various cell types, including B-cells, macrophages, and type II pneumocytes, but not T-cells. Three complete CBuV sequences were successfully characterized and revealed that they clustered with the CBuV sequences obtained from dogs with respiratory disease in Hungary. No additional cases were identified in small numbers of healthy dogs. Although association of the bufavirus with enteric disease remains to be determined, a contributory role of CBuV in canine respiratory disease is possible.
Bufaviruses (BuVs), which are nonenveloped, single-stranded DNA viruses that belong to the genus Protoparvovirus, family Parvoviridae, have approximately 4 kb genomes and contain 2 main open reading frames (ORFs), ORF1 and ORF2. ORF1 and ORF2 encode a nonstructural protein 1 (NS1) and viral capsid protein 2 (VP2), respectively.7,30,49 As the first identification of a BuV in a human showing gastroenteritis in 2012, 32 BuVs were subsequently identified not only in humans but also in wild and domestic animals, including nonhuman primates, wild shrews, bats, rats, pigs, seals, cats, and dogs.9,14,17,25,26,44,51 BuV in dogs, tentatively named canine bufavirus (CBuV) or carnivore protoparvovirus-3, was first identified in an outbreak of canine respiratory disease in a colony of mixed-breed puppies in Italy in 2016. 25 During the investigation, the CBuV DNA was commonly identified in stool samples of dogs with and without diarrhea; however, it was only detected in respiratory samples obtained from dogs with respiratory diseases, but not in healthy dogs. To date, the epidemiology of CBuV has been reported in Italy,10,25 Hungary, 25 China,22,48 India, 12 Türkiye, 1 and Canada, 4 suggesting its distribution of the virus over multiple continents. Although epidemiological investigations have suggested that CBuV is often found in cases of canine enteritis,22,25,48 neither descriptions of associated pathologic lesions nor association with diseases has been well defined.
Positive correlations have been found between the presence of CBuV DNA and diarrhea, and its genomic materials have also been detected in the serum of dogs with gastroenteritis. 22 However, this was challenged by studies that found the presence of CBuV was not significantly associated with diarrhea. 25 Similarly, the role of BuV infection in humans and other animals with enteric diseases remains uncertain.7,19,27,42,50 Notably, BuV DNA detection in the serum and spleen of infected wild shrews and monkeys,14,44 as well as in respiratory samples obtained from human patients with lower respiratory tract diseases, has also been reported, 51 suggesting either extraintestinal infections or hematogenous spreading of the virus. Recent studies exploring the genetic characterization of CBuV revealed genomic heterogenicity.4,48,53 Regarding the genotypic differences between CBuVs found in respiratory and enteric samples, there were 2 amino acid mutations present in the CBuV genome obtained from a respiratory origin. 25 Although the previous finding corroborated initial evidence that CBuV can be present in extraintestinal organs, further study is required to confirm the viral localization in tissues or organs that present significant lesions.
Although there have been anecdotal claims of CBuV affecting canine health,4,22,25,26,48,53 there have been no confirmed reports of CBuV as a possible etiological agent of fatal disease. Considering this concern, it is essential to determine the pathogenic potential of CBuV. Herein, this study describes episodes of recurrent fatal outbreaks associated with CBuV infection in multiple dog colonies on a breeding farm in Thailand and examines the pathological findings associated with the infection. In addition, full-length coding genome sequences of CBuV in each episode were characterized, and subsequently, the genetic relatedness was determined.
Materials and Methods
Outbreak Information
Four episodes of fatal disease outbreaks occurred in multiple colonies of 2 breeds of dogs, pit bulls (colonies A [cases 1–4 and 6–11] and C [cases 13–14]) and Pomeranians and a pit bull (colony B, cases 5 and 12) on a breeding farm in Thailand from May 2021 to April 2022 (Table 1). In the first episode of a fatal outbreak in pit bull colony A, 6, 35-day-old unvaccinated puppies, and 1 out of 2 completely vaccinated adult dogs developed respiratory distress. All the puppies presented with hemoptysis and died within 1 week of the clinical presentation. An adult female pit bull (dog No. 1) presented progressive pneumonia and then recovered after supportive treatments. Four dead puppies (cases 1–4) were submitted for necropsy. The second episode of the fatal outbreaks occurred 1 month later (June 2021) in Pomeranian colony B, in which 2 out of 5 adult dogs developed nasal discharge, followed by progressive pneumonia and hemoptysis, and died after 2 weeks of developing clinical signs. One Pomeranian (case No. 5) was submitted for necropsy. The third episode occurred in the same pit bull colony A in with August 2021. After delivery by dog No. 1, 6, 16-day-old puppies presented hemoptysis and died within 1–2 days. All 6 dead puppies (case Nos. 6–11) were submitted for necropsy. Subsequently, a 10-year-old male pit bull (case No. 12, colony B), who underwent treatment for chronic renal failure, died from acute respiratory distress and was later submitted for necropsy. The fourth episode occurred in April 2022 in pit bull breeding colony C that contained 1, adult female pit bull and 6, 47-day-old puppies. In this episode, all puppies were found dead with hemoptysis, but the adult female was clinically unremarkable. Two dead puppies (case Nos. 13–14) were submitted for necropsy.
Detection of canine bufavirus (CBuV) in outbreaks of dogs with respiratory disease.
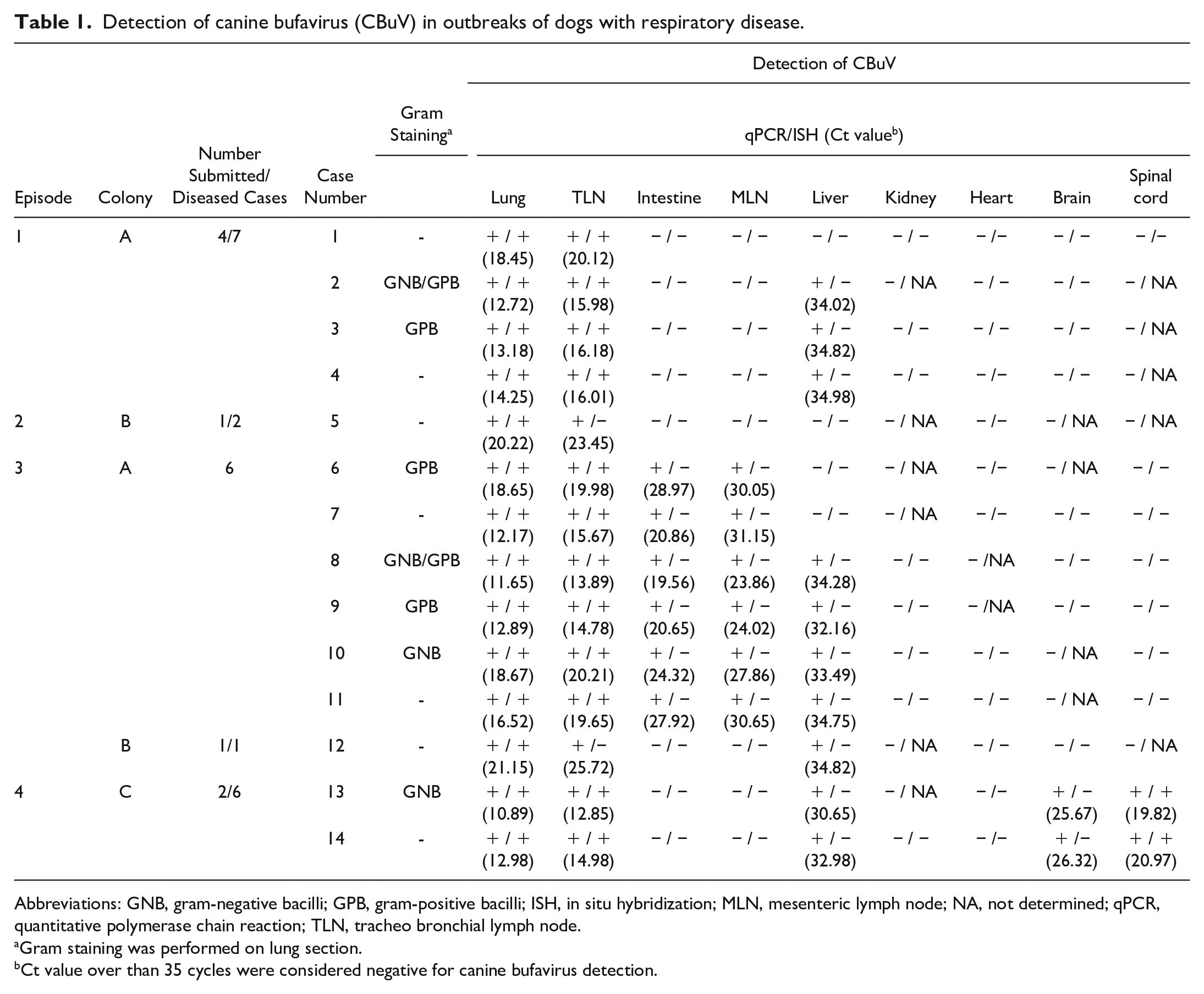
Abbreviations: GNB, gram-negative bacilli; GPB, gram-positive bacilli; ISH, in situ hybridization; MLN, mesenteric lymph node; NA, not determined; qPCR, quantitative polymerase chain reaction; TLN, tracheo bronchial lymph node.
Gram staining was performed on lung section.
Ct value over than 35 cycles were considered negative for canine bufavirus detection.
A total of 14 dead dogs were necropsied at Department of Pathology, Faculty of Veterinary Science, Chulalongkorn University, Bangkok, Thailand. Lung, liver, kidney, heart, intestine, tracheobronchial and mesenteric lymph nodes, brain, and spinal cord samples were obtained from all investigated dogs and were subsequently kept as fresh-frozen samples at −80°C for further use. The fresh-frozen lung and intestine samples were submitted for routine virologic polymerase chain reaction (PCR) testing. Lung samples of 4 dogs (cases 2, 5, 8, and 13) were further submitted for bacterial culture and identification. All major organs were also collected for histological examination. The collected samples were fixed in 10% neutral-buffered formalin for at least 48 hours and not more than 72 hours, and then processed for hematoxylin and eosin staining. Furthermore, the lung sections of all necropsied dogs were subjected for Gram staining. This study was approved by the Institutional Animal Care and Use Committee of the Faculty of Veterinary Science, Chulalongkorn University (IACUC No. 2231001), and all methods were carried out in accordance with relevant Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines and regulations.
Genomic Extraction and Routine Virologic Testing
All collected fresh tissue samples were subjected to viral genomic extraction. Briefly, the tissues were individually homogenized with an aseptic technique using an automatic homogenizer (Bead Ruptor 12; OMNI International, Kennesaw, Georgia). The homogenized tissue samples were further subjected to viral genomic extraction using the Viral Nucleic Acid Extraction II kit (Geneaid, Taipei, Taiwan) following the manufacturer’s protocols. The quality and quantity of the extracted nucleic acids were determined using a spectrophotometer (NanoDrop Lite, Thermo Fisher Scientific Inc., Waltham, Massachusetts). All extracted samples were kept at −80°C until used. Extracted lung samples were subsequently submitted to routine virologic testing using panels of conventional singleplex or multiplex PCR assays to detect canine influenza virus, canine parainfluenza virus, canine morbillivirus, canine respiratory coronavirus, canine herpesvirus-1, canine adenovirus-1 and canine adenovirus-2,36,37 and canine pneumovirus. 38 The extracted intestinal samples were also subjected to PCRs specific for carnivore protoparvovirus-1, 39 canine bocavirus, 40 canine enteric coronavirus, 21 and canine circovirus detection.8,33 PCRs targeting other novel viruses, including canine chapharmaparvovirus, 34 CBuV, 25 and severe acute respiratory syndrome coronavirus 2 were also performed on both lung and intestinal samples. 35 In addition, conventional polymerase chain reaction (cPCR) 29 to identify cytotoxic necrotizing factor 1 (CNF1) of Escherichia coli virulence was performed on the extracted lung samples of 13 necropsied cases (cases 1–13). Due to the limited lung sample obtained from case 14, CNF1 detection was not performed on this case.
Polymerase Chain Reaction Detection of Canine Bufavirus and Viral Loads
The presence of CBuV DNA in the extracted samples was investigated by cPCR to detect the VP2 gene of CBuV following the protocols and newly modified primers of a previous study. 25 Briefly, 2 µl of extracted nucleic acids in each sample were used as a template in a combination with a cocktail of master mix buffer containing Taq DNA polymerase, deoxynucleotide triphosphates (dNTPs), and MgCl2, plus 10 pmol of each forward (5’-CTGGTTTAATCCAGCAGACT-3’) and reverse primer (5’-TGAAGCCCAAGGTAGTAGGT-3’). Thermal cycling amplification was performed at 94°C for 5 minutes as initial denaturation, followed by 45 cycles of 94°C for 30 seconds, 53°C for 30 seconds, 72°C for 30 seconds, and final extension at 72°C for 5 minutes. Amplification products were visualized using automated capillary–based electrophoresis (QIAxcel DNA Screening, Qiagen GmBH, Hilden, Germany) under procedures and settings previously described. 38 The presence of the 208-bp amplicons was considered positive, and it was subsequently confirmed by sequencing using a next-generation sequencing–based method (U2Bio, Incheon, South Korea).
CBuV viral loads in various organs were determined by performing quantitative polymerase chain reaction (qPCR) targeting the NS1 gene of CBuV according to the protocols described in Wang et al 53 with minor modifications. Briefly, 5 g of each collected tissue obtained from the CBuV cPCR-positive dogs were subjected to the genomic extraction described above and subsequently used as templates. The SYBR-based qPCR methods using the KAPA SYBR Fast qPCR Master Mix (2×) Universal kit (Kapa Biosystems, Sigma-Aldrich, South Africa) with the primers CBuV-NS1F (5’-AGGCAAAGGCAGCAAAACTA-3’) and CBuV-NS1R (5’-TGCATGGAATTTCCCAATCT-3’) were used. The qPCR was performed under a Rotor-Gene Q real-time PCR cycler (Qiagen GmbH) with thermal cycling conditions of 95°C for 2 minutes, followed by 40 cycles of denaturation at 95°C for 10 seconds and annealing at 60°C for 30 seconds. The presence of fluorescence signals with plateau curves that were gathered from the annealing step was determined by the cycle threshold (Ct) value. The obtained Ct value was used to determine the amount of CBuV load, and a Ct value over 35 cycles was considered negative. A nontemplate tube was used as a negative control. The qPCR was run in triplicate. Furthermore, extracted DNA samples derived from the retrospective lungs of 15 healthy dogs were investigated for the presence of CBuV DNA.
Genetic Characterization and Phylogenetic Analysis of Canine Bufavirus
To obtain a complete coding sequence of the identified CBuV in this study, multiple cPCR assays following specific primer pairs (Supplemental Table S1) that were newly designed based on the conserved regions of the nucleotide alignment of previously described CBuV sequence were performed. Samples that presented the highest viral loads obtained from dogs in each affected colony were subjected to this investigation. Briefly, the extracted nucleic acids obtained from the CBuV qPCR-positive organs that had the lowest Ct values were used as a template. The cPCR procedures were performed at 95°C for 5 minutes, followed by 45 cycles of denaturation at 95°C for 30 seconds, annealing at 50–53°C as the predicted annealing temperature in each primer pair for 1 minute, and extension at 72°C for 1 minute, followed by a final extension at 72°C for 10 minutes. The positive amplicons were further submitted to next-generation sequencing using the protocols described above. Partial CBuV contigs obtained from each cPCR assay were queried using Basic Local Alignment Search Tool (BLAST) in the National Center for Biotechnology Information (NCBI) database and aligned with the previous CBuV reference strains available in the GenBank database, and genomic sequences were constructed using BioEdit software v. 7.2. The constructed CBuV sequences were then phylogenetically analyzed using the maximum-likelihood model method with GTR+G+I as a best-fit model of nucleotide substitution according to the Bayesian information criterion embedded in the MEGA 7 software package. Bootstrap analysis was performed using 1000 replicates. The pairwise nucleotide similarity and deduced amino acids of the CBuV complete coding sequences were additionally analyzed by comparison with the previous CBuV strains via BioEdit software.
Canine Bufavirus In Situ Hybridization and Dual Labeling
Serial sections of formalin-fixed paraffin-embedded tissues were subjected to chromogenic ISH. An ISH DNA probe covering 208 bp of a partial VP2 gene was constructed using the PCR DIG Probe Synthesis Kit (Roche Diagnostics, Basel, Switzerland) according to the manufacturer’s protocols. The thermal cycling conditions and primer pairs targeting the VP2 gene described above were used, in addition to using the digoxigenin (DIG)-labeled oligonucleotide instead of normal deoxynucleotides. The success of the DNA probe construction was determined by size resolution on 1% (w/v) agarose gel electrophoresis, compared with a constructed control DNA. The ISH procedures were performed following previously described protocols with slight modifications. 41 After deparaffinization and dehydration, 4-µm thick sections were treated with citrate buffer (pH 6) at 95°C for 20 minutes. The sections were then permeabilized using 10 ng/ml of proteinase K solution at 37°C for 10 minutes and immediately postfixed with 0.4% cold formaldehyde for 5 minutes. The sections were triple washed with DNase/RNase-free water and pretreated with a prehybridization solution containing 50% (v/v) formamide in 4× saline-sodium citrate (SSC) solution at 37°C for at least 15 minutes. After incubation, the prehybridized sections were incubated with hybridization buffer containing 5× Denhardt’s solution, 0.5% (w/v) sodium dodecyl sulfate, 100 μg/ml of salmon sperm DNA, 5× SSC, and 10 ng/ml of the constructed CBuV DNA probe at 95°C for 5 minutes, followed by 50°C overnight in a humidified slide incubator. The incubated sections were stringency washed with a series of SSC solutions, and the nonspecific protein binding activity was blocked by immersing them in 2% (w/v) bovine serum albumin at 37°C for 1 hour. The CBuV hybridization was detected by incubation with anti-DIG-AP, Fab fragments (Roche, Basel, Switzerland) (1:200 in 1× bovine serum albumin) at room temperature for 30 minutes and visualized by an alkaline phosphatase (AP) system using a PermaRed/AP chromogen (Diagnostic BioSystems, Pleasanton, California). The CBuV DNA probe was replaced with a 214-bp canine bocavirus-2 VP1 probe for the ISH incubation, 40 and a CBuV qPCR-positive lung section treated with DNase enzyme prior to incubation with hybridization buffer containing the CBuV probe, were used as negative controls. Sections were then counterstained with hematoxylin, dried, and mounted with a coverslip. Red precipitates within cells were considered positive. A retrospective cohort of lung, tracheobronchial lymph node, and spinal cord sections obtained from 10 healthy dogs incubated with the CBuV probe were used as additional controls.
To indicate the cellular localization of the CBuV in lung tissue, serial formalin-fixed paraffin-embedded lung and mediastinal lymph node sections obtained from 4 dogs that were CBuV qPCR-positive with the lowest Ct values (cases 1, 5, 7, and 13) were evaluated. Immunohistochemistry (IHC) against CD20, CD3, ionized calcium-binding adapter molecule-1 (Iba-1), pancytokeratin (panCK), and thyroid transcriptional factor-1 (TTF-1) was first performed, followed by ISH targeting CBuV VP2. IHC was performed following the previously described protocols with some modifications. Briefly, after deparaffinization and dehydration, the sections were submitted to antigen retrieval by incubating with citrate buffer (pH 6) at 95°C for 20 minutes in a water bath. The sections were then blocked against endogenous enzyme activity using freshly prepared 3% (v/v) hydrogen peroxide (H2O2) in absolute methanol for 30 minutes at room temperature and following incubation with 2.5% (w/v) bovine serum albumin at 37°C for 30 minutes to block nonspecific binding. Subsequently, the sections were incubated with the primary antibodies, CD20 (1:300, ab27093, Abcam, Waltham, Massachusetts), CD3 (1:10, PF. Moore, California), Iba-1 (1:300; MABN92, Sigma-Aldrich, St. Louis, Missouri), panCK (1:200, AE1/AE3, Dako, Glostrup, Denmark), or TTF-1 (1:10, 8G7G3/1, Dako), at room temperature for 1.5 hours (CD20, CD3, panCK, Iba-1) or at 37°C for 32 minutes (TTF-1). Secondary antibodies were applied for 1 hour using immuno-AP polymer-conjugated anti-mouse antibody (Histofine simple stain AP [MULTI]; Nichirei, Japan) for CD20, Iba-1, panCK, and TTF-1 and ImmPRESS HRP, rat (Vector Laboratories Inc., Newark, California) for CD3. The visualization system for CD20, Iba-1, panCK, and TTF-1 was the Vector Blue substrate kit AP (Vector Laboratories Inc), whereas a DAB (Dako) was used as a substrate for the CD3 detection system. Subsequently, the slides were treated with 3% (v/v) H2O2 in absolute methanol for 30 minutes at room temperature to halt the remaining alkaline phosphatases/peroxidase activity, 52 and then subjected to the ISH procedures described above. The CBuV hybridization was visualized using a PermaRed/AP chromogen (Diagnostic BioSystems) as described above. Lung and tracheobronchial lymph node sections obtained from healthy dogs that suffered from traumatic disease were subjected to IHC controls.
Results
Postmortem Investigation
All investigated puppies (n = 12; cases 1–4, 6–11, and 13–14) revealed similar macroscopic and microscopic lesions, with slight differences in some organs. Macroscopically, they presented red-tinged discharge at the nostrils and diffuse consolidation in all lung lobes. The entire lungs were firm and red, similar to the livers (Fig. 1a). The tracheobronchial lymph nodes were swollen and congested in all affected cases. Severe congestion at the brain and liver was observed in 6 out of 12 cases. The intestines of most affected dogs had moderate to severe autolysis. In the 2 necropsied adult dogs (n = 2, cases 5 and 12), there was multifocal, dark red discoloration in the caudal lung lobes.
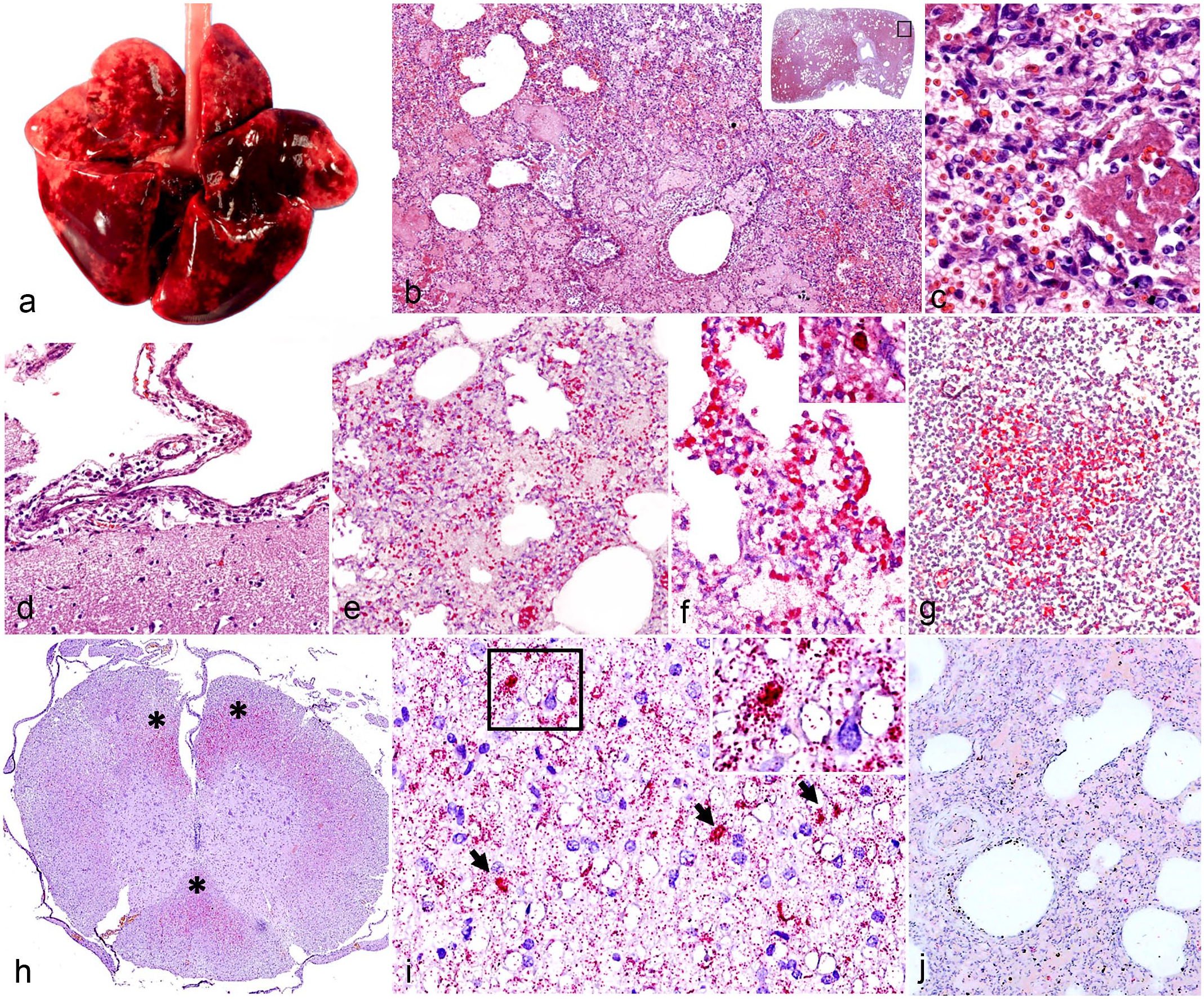
Canine bufavirus (CBuV) infection. (a–c) Lung. Case 7. (a) Severe generalized hemorrhagic pneumonia. (b) Pulmonary parenchyma is hypercellular. Inflammatory cells and fibrin are found in small bronchioles and alveolar lumens. Inset: lower magnification. The black frame corresponds to Fig. 1b. Hematoxylin and eosin (HE). (c) Abundant neutrophils and foamy macrophages are intermixed with organizing mats of fibrin-filled lumens of alveoli. HE. (d) Cerebrum. Mild mononuclear cell infiltrates are found within the meninges. Case 2. HE. (e–g) Detection of CBuV nucleic acid by in situ hybridization (ISH, red chromogen). Lung. Case 7. (e) CBuV nucleic acids are identified in the pulmonary parenchyma. (f) CBuV nucleic acids are identified within the pulmonary epithelia and infiltrating inflammatory cells. Inset: nuclear labeling of the CBuV nucleic acids. (g) Moderate amounts of CBuV nucleic acids are found in a follicle of the tracheobronchial lymph node. (h–i) Thoracic spinal cord. Case 2. (h). CBuV nucleic acids are found multifocally (asterisks) in the white matter of funiculi in cervical spinal cord. (i) CBuV nucleic acids are identified within the nucleus of glial cells (arrows). Inset: higher magnification of the area in the black frame (j). Lung. Case 7. There are minimal hybridization signals in the negative control.
Microscopically, throughout the lungs of all dead puppies, alveoli were multifocally replaced by hemorrhage intermixed with variable organizing mats of eosinophilic fibrin, many lymphocytes, neutrophils, foamy macrophages, and eosinophilic granular to homogenous proteinaceous edema (Fig. 1b, c). The pulmonary interstitium was diffusely thickened by lymphocytes, neutrophils, and macrophages and lining type II pneumocytes with segmental plump, tombstoning hypertrophy. Similar aggregates of neutrophils and fibrin were also found within the lumens of occasional terminal airways and small bronchioles (Fig. 1c). Multifocally, there were clumps of short rod-shaped bacilli noted within the alveoli in 7 out of 12 cases (Table 1). Gram-positive large cadaver bacilli were identified in the alveoli of 4 out of 7 cases whereas Gram-negative bacilli were found in the other cases (Supplemental Fig. S1). The perivascular stroma of several pulmonary blood vessels was variably edematous. The tracheobronchial lymph node contained variable degrees of mildly to moderately hyperplastic lymphoid follicles with intact follicular polarity. The medullary and subcapsular sinuses were filled with pools of erythrocytes that were often engulfed by macrophages and few neutrophils. Variable numbers of plasma cells were scattered within the medullary cords. Regarding the 2 necropsied adult dogs, the alveolar spaces contained moderate amounts of homogenous proteinaceous substances with variable numbers of lymphocytes, neutrophils, and foamy macrophages. The pulmonary interstitium was multifocally thickening and contained lymphocytes and plasma cells. Within the cerebrum of case Nos. 2 and 13, there were minimal to mild interstitial and perivascular infiltrates of lymphocytes and plasma cells (Fig. 1d). The sections of the small and large intestines of the investigated dogs were autolyzed. All other organs of the investigated dogs were histologically unremarkable.
E. coli was cultivated from the lung samples of 3 out of 4 submitted cases (cases 2, 8, and 13); however, the CNF1 gene was not detected in 13 necropsied cases (Supplemental Fig. S2). None of the routine virologic PCR testing was positive in the submitted samples.
Detection of Canine Bufavirus, Viral Loads, and Localization
The most prominent lesion was found in the lung tissue in all submitted cases, and they were negative by ancillary testing, which prompted us to further investigate the possible cause using other available PCR assays targeting other viruses. Whereas others were negative, the CBuV-specific cPCR that was positive in the lung samples of all submitted cases, and sequence analysis of the partial VP2 gene of CBuV confirmed our findings. qPCR was used to determine the viral distribution and its loads in other organs. CBuV DNA loads were mostly detected in the lung tissues (average Ct = 14.42), followed by the tracheobronchial lymph node (average Ct = 15.35) in all cases. Interestingly, the intestines and mesenteric lymph nodes were positive only in samples obtained from 6 dead puppies (from colony A, second episode), with average Ct values of 23.71 and 27.93, respectively. CBuV DNA was also detected in the liver (9/14, Ct < 35), brain (2/14, Ct < 27), and spinal cord (2/14, Ct < 21) of some cases (Table 1). No CBuV DNA was found in the additional retrospective cohort samples obtained from healthy dogs.
ISH identified large amounts of CBuV DNA within the lung tissues of all puppies, but rarely detected it in the 2 adult dogs. In the lung sections, CBuV DNA was identified diffusely in pulmonary parenchyma and occasional cellular aggregates within the lumen of small blood vessels (Fig. 1e). Infiltrative mononuclear cells in the described interstitial pneumonia were also strongly positive for CBuV on ISH. The hybridization signals indicated nuclear localization of the CBuV in the affected cells (Fig. 1f). The tracheobronchial lymph node was also focally positive for CBuV by ISH in the center of a lymphoid follicle (Fig. 1g). Interestingly, the CBuV hybridization signal was also detected in the nuclei of glial cells located in the white matter of the funiculus in the cervical spinal cords of the 2 qPCR-positive spinal cord sections (Fig. 1h, i). All other organs examined were negative for CBuV by ISH. Minimal hybridization signals were observed in the negative controls (Fig. 1j).
In the dual IHC/ISH assays, CBuV DNA was positive within the cells labeled with the TTF-1 antibody (Fig. 2a). Coexpression of CD20 and CBuV DNA was diffusely observed throughout the lung and lymphoid follicles (Fig. 2b, c), whereas it was detected in Iba-1-positive cells (Fig. 2d) and rare panCK-positive cells. By contrast, CD3-positive cells were not found to have CBuV DNA colabeling. No ISH signal was found in any investigated tissue sections from healthy dogs (Supplemental Fig. S3).
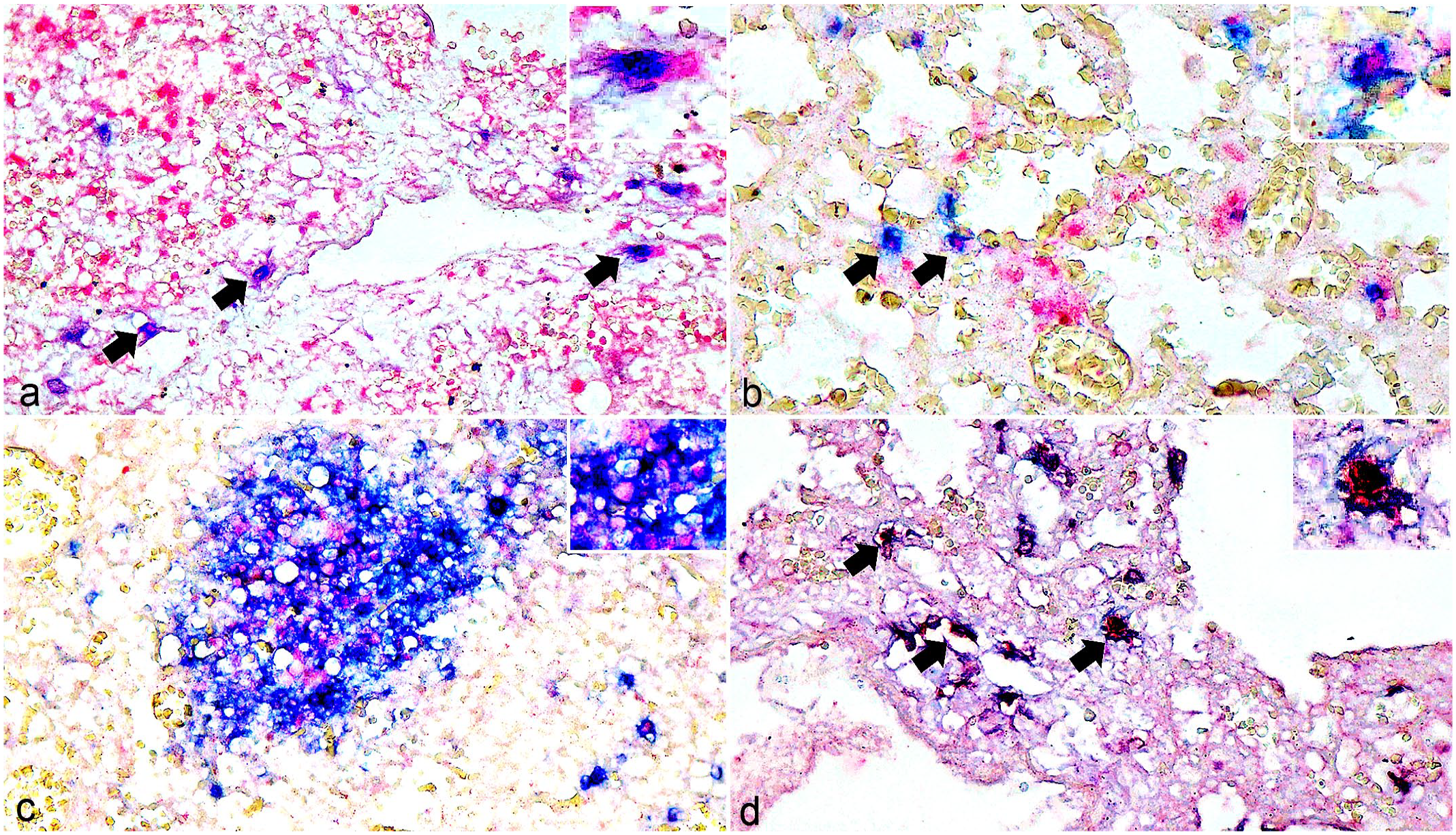
Canine bufavirus (CBuV) infection. Dual labeling of CBuV nucleic acids by in situ hybridization (ISH, red chromogen) and cellular-specific antigens by immunohistochemistry (IHC, blue chromogen). Lung and tracheobronchial lymph node. Case 2. (a) CBuV nucleic acids are identified in the nuclei of cells with nuclear thyroid transcription factor-1 (TTF-1) labeling (arrows). Inset: higher magnification of a positive cell. (b) Nuclear labeling of CBuV DNA within infiltrating B-cells with cytoplasmic CD20 labeling (arrows). Inset: higher magnification of a positive cell. (c) CBuV nucleic acid labeling within the cells with cytoplasmic CD20 labeling in the follicle of the tracheobronchial lymph node. Inset: higher magnification of the positive cells. (d) Rare nuclear CBuV labeling within the pulmonary alveolar macrophages that are labeled with ionized calcium-binding adapter molecule-1 (Iba-1) (arrows). Inset: higher magnification of a positive cell.
Genome Characterization and Phylogenetic Analysis
Genomic analysis data obtained from the PCR assay amplifying the CBuV VP2 gene identified the most nucleotide similarity among CBuV-positive cases (99.0%–100.0%) in each affected colony. The representative samples in the 3 colonies were subsequently subjected to CBuV complete coding sequence analysis. Three 4219-bp sequences were successfully characterized, tentatively named CBuV-CP21R187C TH/2021-colonyA, CBuV-CP21R204C TH/2021-colonyB, and CBuV-CP22R73C TH/2022-colonyC and were submitted to the GenBank database under accession numbers OR004240–OR004242, respectively. The identified CBuV genomes contained the 2 main ORFs 1–2. ORF1 encoded the NS, NS1 (nt 1–1917). ORF2 encoded the overlapping structural proteins (VP), VP1 (nt 1904–1931 and nt 2115–4219) and VP2 (nt 2513–4219). Phylogenetic analyses of the obtained CBuV genomes revealed that they clustered together and presented a monophyletic clade, genetically close to the CBuV sequences identified in dogs with respiratory disease in Hungary since 2012 (MF198245 and MF198246, 99.6% pairwise nucleotide identity) (Fig. 3). In addition to the unique phylogenetic clade of CBuV in dogs, they were divergent from the previously described CBuV genomes found in foxes (OM640104–OM640105) and the human BuV sequences, significantly supported by the 100% bootstrap value. Pairwise nucleotide analysis revealed that CBuV-THs had a genetic similarity of ~70.0% and 57.7% to the fox-CBuV and human BuV, respectively.
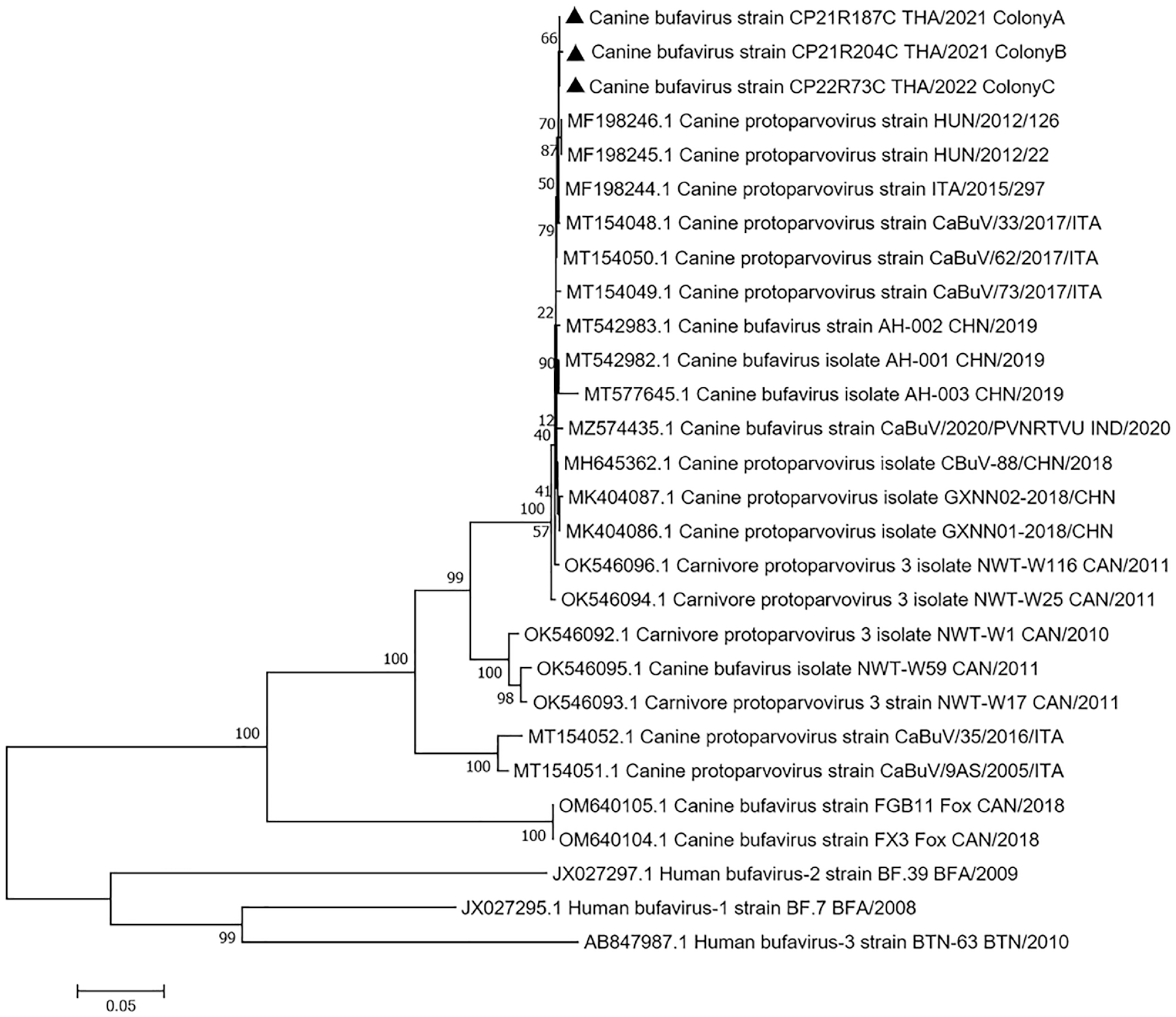

Phylogenetic topology construction of canine bufavirus (CBuV) complete coding sequences using a maximum-likelihood model with 1000 bootstrap replicates. The CBuV sequences found in each outbreak are identified by black triangles, and they are clustered together by presenting a unique topologic clade. The obtained CBuV sequences in this study are genetically close to the CBuV sequences identified in dogs with respiratory disease in Hungary. Taxon names are presented as GenBank accession numbers, and the names of viral strains are presented in parentheses. The human bufavirus sequences are used as an outgroup. The scale bar indicates nucleotide substitution per site.
Discussion
Although BuVs have been identified in various hosts,17,25,26,42,44,50 whether there is a pathogenic role of this virus or they only serve as a commensal agent in the susceptible hosts has not been fully established. Gathering relevant information obtained from observational studies through epidemiological investigations of BuVs may facilitate the understanding of whether BuVs may act as a pathogen. Although recent studies found that CBuV has often presented in dogs with diarrhea,12,22,48,53 our study found that CBuV was present in cases of respiratory disease. The association between CBuV infection and diseases has not been completely determined because there is no experimental model, and the virus could be identified in the feces of healthy dogs.10,48 Although a positive correlation of the presence of CBuV with enteric diseases was revealed in a study conducted in dog populations in China, 22 2 other studies indicated no correlation in dogs showing diarrhea and suggested that CBuV was common in enteric samples obtained from dogs both with and without diarrhea.10,48,53
Although CBuV DNA was frequently identified in fecal samples, a positive correlation between the presence of CBuV DNA and respiratory diseases was also indicated. 25 As the first detection of CBuV in dogs using a metagenomic study, the viral DNA was identified in the oropharyngeal samples obtained from puppies of an unknown respiratory disease outbreak in a dog litter in Italy. This previous finding also suggested the potential role of CBuV in a contributory cause of respiratory diseases in puppies. 25 Since then, CBuV genomic characterization has been widely studied;1,4,10,12,22,26,48,53 however, neither evidence nor findings supporting the CBuV tropism in infected cases have been investigated. Here, we report a recurrent outbreak of CBuV infection in multiple colonies of puppies in Thailand. Regarding the significant respiratory diseases presented in all affected dogs, CBuV DNA was identified in respiratory samples using PCR, which, together with the large amounts of CBuV DNA localized in the lung sections with histologic lesions of bronchointerstitial pneumonia, suggests the contribution of this virus to the clinical cause for dyspnea in these outbreaks. Furthermore, CBuV was not found in the samples obtained from healthy dogs, which may further support the role of CBuV in association with respiratory disease. Consistent results between qPCR and ISH showed that the CBuV viral loads were much higher in the lung and tracheobronchial lymph node than in other organs. The CBuV PCR-positive/ISH-negative result in other organ samples is, therefore, more likely due to hematogenous spreading. Although recent studies have focused on the association of CBuV with enteric disease because this virus belongs to the Parvoviridae family,7,30 together with speculation of other BuV infections in association with diarrhea in other susceptible hosts,42,44,48,50 lower amounts of CBuV DNA were identified in the intestine by the qPCR in some affected puppies. Unfortunately, histological findings of CBuV infection associated with intestinal lesions could not be determined in this study due to tissue autolysis. Although CBuV DNA was detected in the intestinal sample that was similar to the identification of CBuV in fecal samples found in previous studies,10,12,22,25,53 viral localization in the intestinal tissues could not be determined in this study, which may result from either tissue autolysis or viral exocytosis from the infected cells. The exact mechanism to explain the findings could not be determined based on this study, and further investigations are needed.
CBuV was detected in oropharyngeal samples of dogs with respiratory diseases, similar to our findings in this study. 25 In addition to CBuV DNA detection by PCR, we consistently identified the viral DNA in lesions presenting interstitial pneumonia in all selected cases. Evidence of parvovirus infection associated with respiratory diseases has been documented in dogs and humans, such as in an experimental study of canine bocavirus-1 (formerly known as minute virus of canines) infection in dogs and human bocavirus infection in children showing respiratory distress.5,18,23,24,46,47 CBuV infection in association with respiratory diseases in dogs, therefore, could not be neglected.
Pulmonary changes associated with extraintestinal pathogenic E. coli infection in dogs have been previously described.3,6,15 Histologic evidence of bacterial clumping was found in conjunction with the described pulmonary lesions, and the E. coli was cultured from the lung samples of some cases, which prompted us to perform the Gram stain and also investigate the CNF1 virulence factor of the extraintestinal pathogenic E. coli as a playing contributory role in these outbreaks. Although the gram-negative bacteria were found to be intralesional in some cases, CNF1 was not detected in the lung samples of necropsied cases. Given the character of the gross and histologic changes, the presence of intralesional clumps and gram-negative bacilli and the isolation of E. coli from the lung of cases 2, 8, and 13, negative CNF1 PCR does not completely exclude the potential role of extraintestinal pathogenic E. coli as a contributory pathogen in the development of the lesions in at least these 3 cases. Although CNF1 has been the main virulence factor identified in cases of canine extraintestinal pathogenic E. coli–associated necrohemorrhagic pneumonia, other virulence factors include CNF2, adhesins, iron acquisition factors, lipopolysaccharides, polysaccharide capsules, and invasins, which are usually encoded on pathogenicity islands (PAIs), plasmids, and other mobile genetic elements. 43
We also found that the type II pneumocytes harbored the CBuV, as it was found colabeling in TTF-1 and panCK-positive cells, and some infected cells were proliferating. Desquamated or reactive type II pneumocytes have been found in pulmonary manifestation of parvovirus B-19 infection in humans and mink infected with Aleutian mink disease parvovirus;2,13 however, the significance of this finding is unknown. As CBuV has also been found to be colocalized with B-cells and macrophages as it was colabeled with CD20 and Iba-1 IHC, respectively, we speculate that CBuV may harbor these cells in the lung and tracheobronchial lymph node, and the virus may simply replicate in lymphoid tissues, as evident by several studies indicating parvovirus replication.11,16,20,28 A recent study predicted that the CBuV genome contains the B-cell epitope, 53 which is commonly known as it is recognized by the B-cell receptor on B-cells. We consistently found that CBuV preferred the B-cells, and we also speculate that CBuV may use the B-cells for trafficking in the lung or other organs. This finding offers evidence for CBuV in B-cells, which may be useful for further studies. In addition, CBuV DNA was identified in glial cells in the spinal cord in a case that presented minimal lymphoplasmacytic meningitis; however, the significance of this finding is unknown and could not be determined from the current information. Parvovirus localization in the central nervous system, where it was associated with minimal to no inflammation, has been reported in many infected cases, but the pathogenesis of this phenomenon is still questioned.31,40,41,45,54
According to genomic analysis, these detected CBuV Thai strains were genetically close to the CBuV strains found in Hungary. 25 A previous study speculated that there are 2 amino acid differences between the CBuV strains obtained from respiratory and enteric origins; however, we did not find such amino acid changes in our sequences. This finding reflects that these changes may not be associated with the clinical significance of this virus.
Overall, this study identified CBuV as a possible causative agent in multiple outbreaks of breeding colonies showing fatal respiratory disease. Naturally infected puppies revealed histopathological pulmonary diseases, in which there were the highest viral loads. The virus was also detected in glial cells in the spinal cords of some infected cases. Using dual labeling, CBuV localization was verified; it is harbored in B-cell lymphocytes, macrophages, and type II pneumocytes. Although our findings corroborate evidence that CBuV is associated with respiratory diseases, the pathogenesis of infection is still lacking, and further studies are needed to address this issue.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231198000 – Supplemental material for Canine bufavirus (Carnivore protoparvovirus-3) infection in dogs with respiratory disease
Supplemental material, sj-pdf-1-vet-10.1177_03009858231198000 for Canine bufavirus (Carnivore protoparvovirus-3) infection in dogs with respiratory disease by Chutchai Piewbang, Panida Poonsin, Pattiya Lohavicharn, Tin Van Nguyen, Sitthichok Lacharoje, Tanit Kasantikul and Somporn Techangamsuwan in Veterinary Pathology
Footnotes
Supplemental material for this article is available online.
Author Contributions
CP designed the experiments. CP, PP, PL, TVN, and SL performed experiment. CP, TK, and ST performed histological evaluation. CP wrote the first draft. ST proved the finalized manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: CP was supported by the Ratchadapisek Somphot Fund for Postdoctoral Fellowship, Chulalongkorn University. PP received a grant from Thailand Research Fund through the Royal Golden Jubilee Ph.D. Program (Grant No. NRCT5-RGJ63001-013) and The Second Century Fund (C2F), Chulalongkorn University. PL received a grant from The National Research Council of Thailand (NRCT): (N41A640189) and The Second Century Fund (C2F), Chulalongkorn University. TVN was granted from The Second Century Fund (C2F), Chulalongkorn University. ST was partly supported by the National Research Council of Thailand (NRCT): R. Thanawongnuwech NRCT Senior Scholar 2022 #N42A650553. This research was funded by Animal Virome and Diagnostic Development Research Unit, Faculty of Veterinary Science, Chulalongkorn University.
Ethical Approval
All experimental protocols were approved by the Chulalongkorn University Animal Care and Use Committee (No. 2231001). All procedures were done in accordance with the ARRIVE guidelines and regulations.
Data Availability Statement
All the data supporting our findings are contained within the manuscript. Three full-length coding sequences of the CBuV have been deposited in NCBI GenBank under accession numbers OR004240–OR004242.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
