Abstract
The amoeba Malpighamoeba mellificae is the etiologic agent of amoebic (amoeba) disease of Western honey bees (Apis mellifera). M. mellificae damages the Malpighian tubules, which is believed to weaken and kill the host bee. Here, the authors describe the detection of this organism in a honey bee colony in the Yukon Territory, Canada. The Malpighian tubules of 14% (7/50) of the adult worker bees were discolored dark brown. Fifteen bees screened using conventional polymerase chain reaction for the 18S gene of M. mellificae were positive for the pathogen. Histologically, the lumens of Malpighian tubules were packed with amoebae, causing dilation of the tubules and attenuation and loss of the tubular epithelium. This phylogenetic analysis places M. mellificae in a new clade, a sister group to the Entamoebidae. This work provides a foundation for further investigation into the distribution, prevalence, and pathology associated with M. mellificae infection.
Keywords
Western honey bees (Apis mellifera) are challenged by many pathogens that affect honey bee health and the sustainability of the beekeeping industry. 3 The amoeba Malpighamoeba mellificae has been associated with amoebic or amoeba disease in honey bees.5,14,17 This organism was discovered in the Malpighian tubules of honey bees by Maassen in 1916. 14
All castes of adult bees can become infected with M. mellificae, with the fecal-oral route likely being the predominant mode of transmission.1,8,10,14 Nosema spp is a microsporidian parasite that also affects the honey bee excretory tract. It causes clinical signs of dysentery and increased bee mortality. 14 M. mellificae is most often reported in conjunction with nosemosis.2,13,14 However, M. mellificae and Nosema spp are thought to infect different parts of the excretory system, leading to speculation about additive or synergistic interactions between the two parasites.11,12,14
Since the mid-20th century, little work has been done to describe this organism or its pathologic effects on honey bees, despite its worldwide distribution.3,14 M. mellificae is infrequently included in pathogen surveys, making its prevalence and distribution within countries uncertain. Herein, we present updated genomic information and light microscopic and histologic images to aid the diagnosis of amoebic disease.
In fall 2021, an apiarist (Tardif) from the Yukon, Canada, contacted staff at the University of Florida Honey Bee Research and Extension Laboratory (UF HBREL) about increased bee mortality in a colony of honey bees. Since 2018, the apiarist had documented high individual bee mortality and Nosema spp infection overwinter in multiple colonies. Ten dead bees from the front of each hive were collected and placed into resealable bags labeled by colony to monitor for Nosema infection. Ten milliliters of distilled water was added to each bag, and then the contents were ground using a mortar and pestle to empty the bees’ abdomens. A drop of liquid from the bag was placed onto a microscope slide, cover-slipped, and viewed via light microscopy. Feces were collected from frames in the hives and diluted with warm distilled water, a drop of which was placed on a slide for microscopic review. In 2019, while performing fecal examinations to quantify the degree of infection with Nosema spp, the apiarist noted moderate numbers of approximately 8 to 10 µm diameter, spherical, colorless, translucent cysts (Fig. 1a) in samples of bees from one colony. Cysts were also observed filling Malpighian tubules (Fig. 1b).
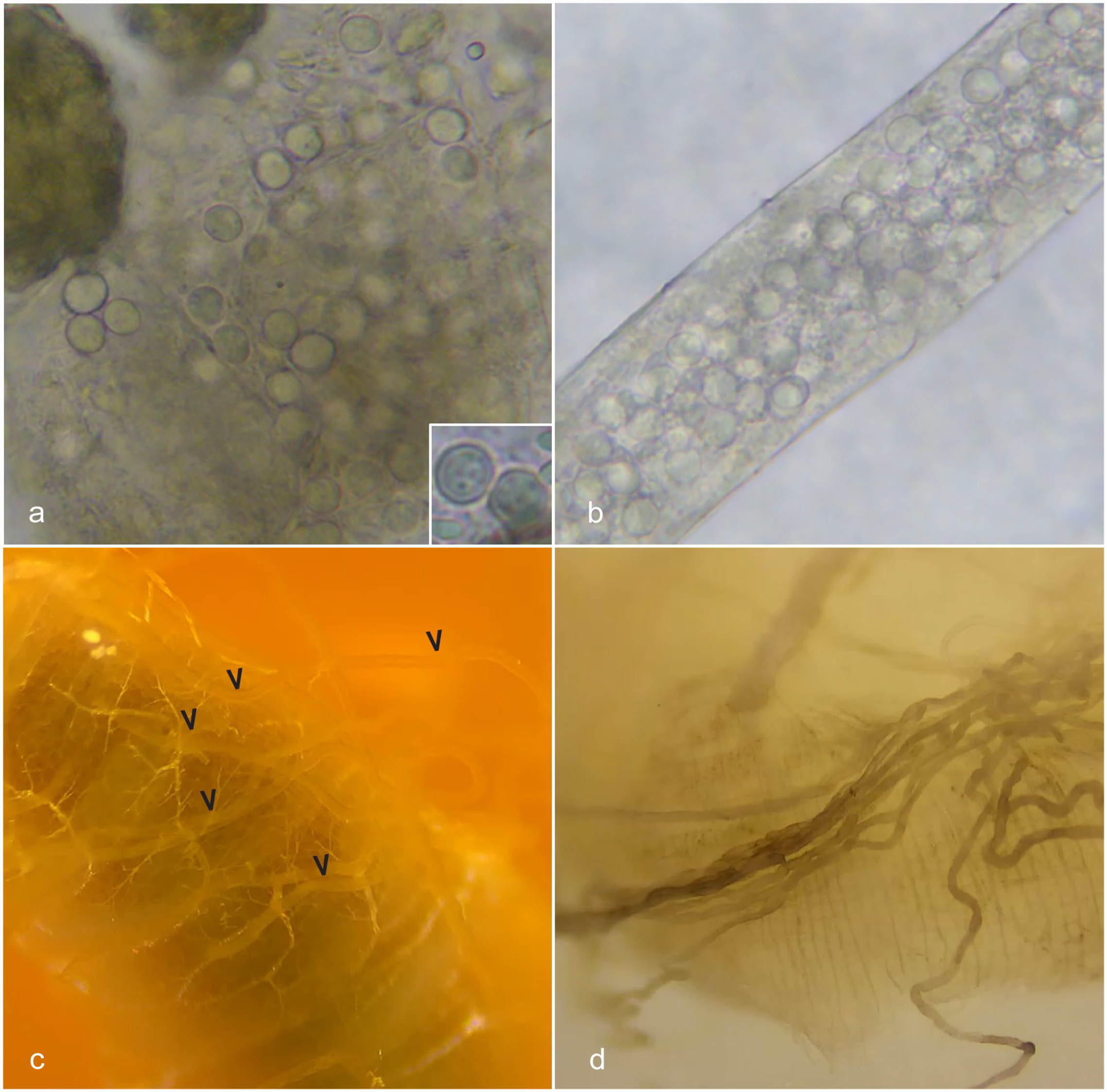
Amoebic disease (Malpighamoeba mellificae), alimentary canal, Western honey bee. (a) In fecal preparation, amoebic cysts were observed singly and in clumps and were approximately 8 to 10 µm in diameter, spherical, colorless, and translucent. Inset: higher magnification of amoebic cysts. (b) During dissection of honey bee guts, cysts were observed within Malpighian tubules using light microscopy. (c) Normal gross appearance of honey bee Malpighian tubules highlighted with arrowheads. (d) Malpighian tubules of a honey bee collected from a colony infected with Malpighamoeba mellificae were discolored, ranging from dark brown to black (interpreted as melanization).
To collect a sample of bees from the affected colony, the beekeeper closed the front of the hive and collected approximately 100 live bees from the entrance after approximately 12 hours. He transferred the bees to a conical tube, added 100% ethanol, and stored them in the freezer at 0°C until shipping. The colony sample was sent to the UF HBREL for further diagnostics. The colony continued to weaken and ultimately died over the 2021–2022 winter.
At the UF HBREL, the midguts and Malpighian tubules were dissected from 50 bees from the colony sample, pooled into a single sample (MEI1), and stored in phosphate-buffered saline at −80°C until DNA extraction. The Malpighian tubules of honey bees should be pale tan and translucent (Fig. 1c). Approximately 14% (7/50) of the honey bees from the affected colony had Malpighian tubules that were dark brown (Fig. 1d). We suspected this discoloration was due to melanization, as this is a common response to tissue damage in insects. 6 DNA was extracted from the Malpighian tubules using a DNeasy Blood & Tissue Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Subsequently, the extracted DNA was used to generate a DNA-sequencing library using a NEBNext Ultra II DNA Library Prep Kit (New England Biolabs, Ipswich, Massachusetts) according to the manufacturer’s protocol. The resulting library was sequenced using a v3 chemistry 600 cycle kit on a MiSeq (Illumina, San Diego, California). De novo assembly of the paired-end reads was performed using CLC Genomics Workbench v20.0.4 with default settings. The assembled contigs were subjected to BLASTN searches against the National Center for Biotechnology Information nonredundant nucleotide database using OmicsBox v1.2. A maximum likelihood (ML) phylogenetic tree was constructed based on the nucleotide alignments of partial small subunit (SSU) from 38 amoebae, including M. mellificae from this study, and 6 organisms from the Opisthokont clade as the outgroup (Supplemental Table S1). The alignment was performed using the MAFFT server (https://mafft.cbrc.jp/alignment/server/), and all columns containing gaps were removed in BioEdit version 7.0.5.3.7,9 The ML tree was constructed using IQ-TREE with 1000 nonparametric standard bootstraps performed to test the robustness of the clades. 4 Sample MEI1 (GenBank accession no. OP935659) resulted in a 1930-bp sequence with the highest nucleotide identity (97.46%) with the partial sequence of SSU RNA gene of M. mellificae isolate BK205-17. Phylogenetic analysis revealed that the M. mellificae sample MEI1 belongs to a new clade, a sister group to the Entamoebidae family, infraphylum Archamoebae (Supplemental Fig. S1).
Fifteen bees from the colony sample were individually homogenized by placing them into a microcentrifuge tube with 3 steel bashing beads and 750 µl of Bashing Bead Buffer (Zymo Research, Irvine, California) and running three, 20-second cycles at 6800 rpm, with a 15-second pause between cycles on a Precellys Evolution (Bertin Instruments, Basingstoke, England) tissue homogenizer. DNA was extracted from each sample using a Quick-DNA Tissue/Insect Miniprep Kit (Zymo Research) following the manufacturer’s protocol. The pooled sample of 50 bees that were positive for M. mellificae (using next-generation sequencing) was used as a positive control, and a nontemplate control was also included. Conventional polymerase chain reaction (PCR) was then performed following the protocol and using described primers. 18 All 15 (100%) honey bees were positive for M. mellificae (Supplemental Fig. S2).
Twenty bees from this colony were transferred to Davidson’s solution and fixed for 24 hours. Then, the bees were submitted to the UF Veterinary Teaching Hospital’s Histology Department for routine histologic processing and staining with hematoxylin and eosin (H&E), Giemsa, periodic acid-Schiff (PAS), and Fontana-Masson. A midline sagittal section was obtained for histopathologic review. Examples of normal microanatomy and negative controls are provided in Supplemental Fig. S3a, b, c.
Histologically, nosemosis was diagnosed in 100% (20/20) of the examined bees, and amoebic cysts were detected in 45% (9/20) of the bees. Amoebic cysts were observed distending the lumens of affected Malpighian tubules, within tubular epithelial cells, and in the lumen of the midgut and hindgut (Supplemental Fig. S5). The epithelium lining affected Malpighian tubules was degenerative, attenuated, or absent, presumably due to pressure induced by the amoebae in the lumen (Fig. 2b, c). The amoebic cysts in the Malpighian tubules were round to ovoid and 7 to 10 µm in diameter, with distinct mildly refractile walls and darkly eosinophilic to deeply basophilic cytoplasm (Fig. 2c, d). Melanin, highlighted with Fontana-Masson stain (Fig. 2e), was frequently associated with the basement membranes, hemocoelomic surfaces, and apical surfaces of enterocytes in areas of epithelial cell destruction secondary to nosemosis and amoebiasis. In the midgut, there was regionally extensive epithelial cell rupture and loss due to 4 to 6 µm in length, 2 to 4 µm in width, oval, intracellular Nosema spores (Fig. 2f). In 3 individuals (15%) with low numbers or no detectable cysts, structures similar to the trophozoite life stage of M. mellificae previously characterized by electron microscopy were rarely observed (Supplemental Fig. S4).8,11,12 These presumptive trophozoites were round, approximately 3 µm in diameter, with a magenta nucleus and wispy, deeply eosinophilic cytoplasm.
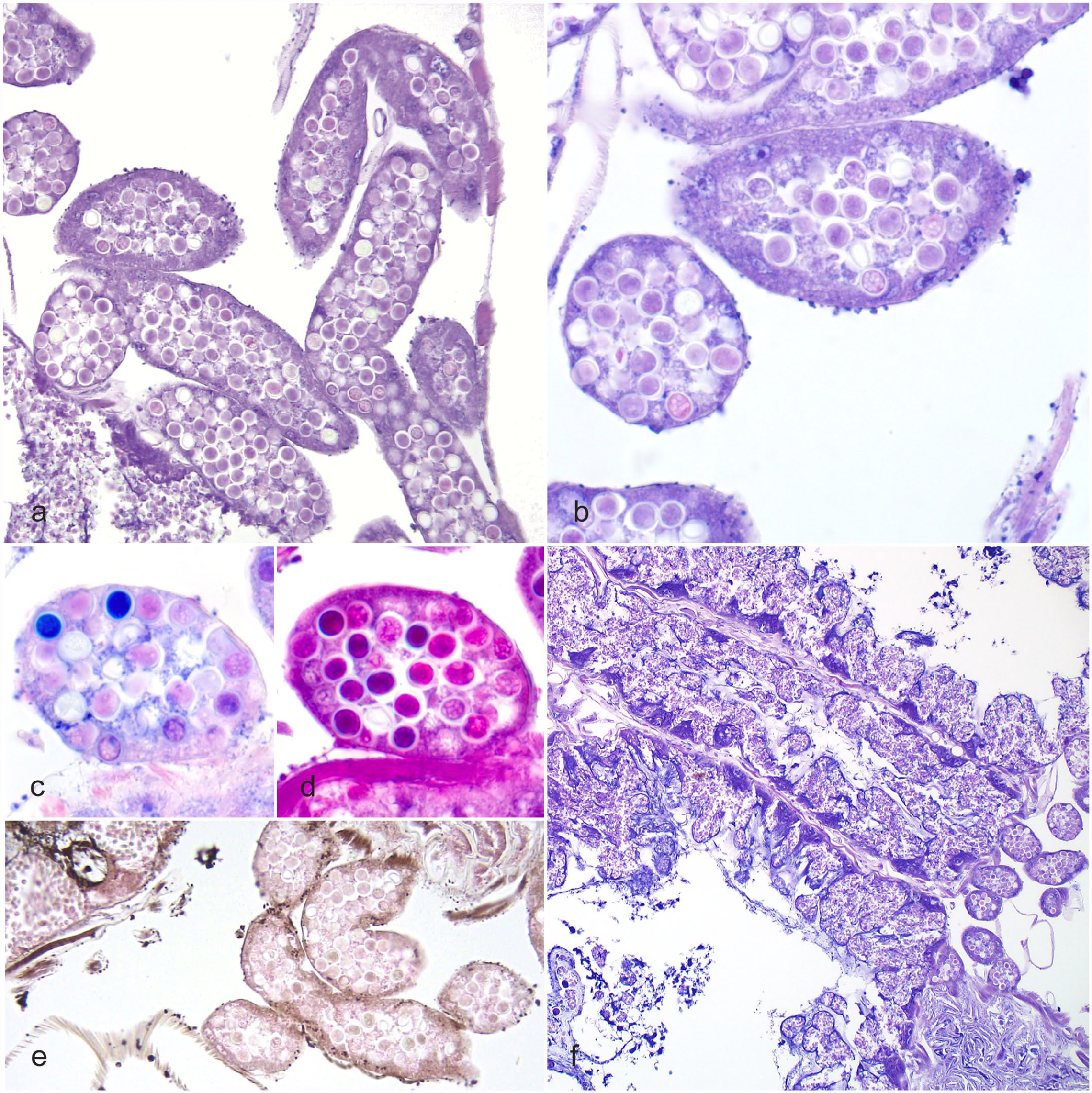
Amoebic disease (Malpighamoeba mellificae), alimentary canal, Western honey bee. HE. (a) Low magnification of Malpighian tubules severely distended by high numbers of amoebic cysts. HE. (b) High magnification of Malpighian tubules severely distended by amoebic cysts. Advanced cysts stain intensely with hematoxylin. HE. (c) PAS stain highlights the accumulation of PAS-positive material before encystment of trophozoites. (d) Toward the end of encystment, the cysts stain intensely blue/purple with Giemsa stain. (e) Fontana-Masson stain highlights melanin deposited in areas of tissue damage. (f) Destruction of midgut epithelium by Nosema spp and dilation of Malpighian tubules by amoebic cysts. HE. HE, hematoxylin and eosin; PAS, periodic acid-Schiff.
Histologically, amoebic cysts were detected in the midgut, Malpighian tubules, and hindgut of infected bees. This supports the life cycle and fecal-oral route of transmission suggested by previous investigations using scanning electron microscopy (SEM).8,12 The primary lesions observed in the bees investigated in this study were consistent with nosemosis in the midgut and damage to the Malpighian tubules caused by the accumulation of M. mellificae cysts. Degeneration and attenuation of the Malpighian tubular epithelial cells in the presence of the amoeba were also observed in a previous study using SEM.11,12 The significance of this damage to the host needs to be elucidated further. When desert locusts were experimentally infected with Malpighamoeba locustae, the amoeba was shown to impact fluid secretion and P-glycoprotein-dependent detoxification by the Malpighian tubules. 16 Distension of the tubules by cysts in the lumen increases the surface area and fluid secretion rate, likely increasing water stress and energy costs in these infections. Similarly, increased surface area reduces net rhodamine extrusion, thus impairing P-glycoprotein-dependent detoxification. This may play an important role in removing xenobiotic substances from the hemolymph. 16 Considering these pathologic sequelae, it is likely that high burdens of M. locustae negatively impact the host, and the pathogenesis may be similar for M. mellificae in honey bees.
Melanization is a nonspecific immune response in insects and is often stimulated by trauma or in response to infection with pathologic organisms such as bacteria and fungi. 6 This response has also been detected in locusts infected with M. locustae. 8 In persistent infections, the Malpighian tubules of locusts became encapsulated by host hemocytes, forming aggregates and depositing melanin. 8 Melanization progressed throughout the interior of the locust, and increasing numbers of organs became involved in the encapsulated masses surrounding cysts freed from ruptured tubules. 8 This is presented as a contrast to what was observed in the honey bees in this case, where rupture of Malpighian tubules was not observed, and melanization was localized to the midgut and Malpighian tubules.
The colony in this study was lost due to high bee mortality and bee self-removal, likely due to the heavy burden of amoeba and Nosema spp. In prior years, the apiarist noted a high pathogen burden in the colony overwinter, which decreased following cleansing flights in spring. This winter buildup has also been observed when comparing the proportions of honey bees infected with M. mellificae and Nosema apis in affected colonies. 2 It was noted that honey bees long confined to hives, as occurs overwinter, contained more amoebae than bees able to perform cleansing flights. 2 Therefore, buildup overwinter may contribute to colony losses associated with amoebic infection.
Phylogenetic analysis placed M. mellificae in a new clade, a sister group to the Entamoebidae family, infraphylum Archamoebae. Entamoebidae is a predominantly endobiotic and aflagellated group representing a separate, deep-branching lineage of Archamoebae. 15 Entamoeba is a unique genus typified by the enteric pathogen of humans, Entamoeba histolytica. One of the unique aspects of this group is that its members, including E. histolytica, lack mitochondria and have a complex taxonomic relationship with other Archamoebae. In E. histolytica infections, transmission is fecal-oral, and excystation occurs in the intestinal lumen to produce trophozoites that invade host enterocytes. 19 The similarities between E. histolytica infection in humans and M. mellificae infection in honey bees are interesting considering their hosts’ broad taxonomic separation. This case, therefore, speaks to the strength of comparative pathology in elucidating disease syndromes in nontraditional species. More studies are needed to understand the relationship between Entamoeba and Malpighamoeba.
These data describe the infection of a honey bee colony with the amoeba M. mellificae. Based on these findings, and the historical reports of the widespread nature of this amoeba, it seems likely that amoebiasis is a common background infection of managed honey bees. Similarly, based on infrequent reports of disease associated with M. mellificae, severe illness may only occur when there is overwinter buildup of amoebae or a high concurrent burden with Nosema spp. This work also provides the phylogenetic description of this amoeba as a new clade and sister group to the Entamoebidae family and further helps to elucidate the intricate relationship of the amoeba group. Future investigation into amoebic disease requires surveys and infection experiments. The development of the PCR primers by Schäfer et al 18 and the genomic data generated in this study will aid in these future experiments.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231179956 – Supplemental material for Report of amoebic disease in a colony of Western honey bees (Apis mellifera)
Supplemental material, sj-pdf-1-vet-10.1177_03009858231179956 for Report of amoebic disease in a colony of Western honey bees (Apis mellifera) by Marley E. Iredale, Pedro H. O. Viadanna, Kuttichantran Subramaniam, Etienne Tardif, Bryony C. Bonning and James D. Ellis in Veterinary Pathology
Footnotes
Acknowledgements
We acknowledge the histotechnologists at the University of Florida College of Veterinary Medicine for the preparation of the tissues for histologic evaluation and Julia St. Amant (UF Honey Bee Research and Extension Laboratory) for her technical support for this project.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The funding was provided by Dave and Colleen Larsen, the University of Florida, and the USDA National Institute of Food and Agriculture, Multi-State Hatch Project 1019945.
Supplemental Material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
