Abstract
Aspergillosis of gorgonian sea fans is a Caribbean-wide disease characterized by focal, annular purple pigmentation with central tissue loss. We applied a holistic diagnostic approach including histopathology and a combination of culture and direct molecular identification of fungi to evaluate these lesions with the goal of determining the diversity of associated micro-organisms and pathology. Biopsies were collected from 14 sea fans without gross lesions and 44 sea fans with lesions grossly consistent with aspergillosis in shallow fringing reefs of St. Kitts. Histologically, the tissue loss margin had exposure of the axis and amoebocyte encapsulation with abundant mixed micro-organisms. Polyp loss, gastrodermal necrosis, and coenenchymal amoebocytosis were at the lesion interface (purpled area transitioning to grossly normal tissue) with algae (n = 21), fungus-like hyphae (n = 20), ciliate protists (n = 16), cyanobacteria (n = 15), labyrinthulomycetes (n = 5), or no micro-organisms (n = 8). Slender, septate hyaline hyphae predominated over other morphological categories, but were confined to the axis with little host response other than periaxial melanization. Hyphae were absent in 6 lesioned sea fans and present in 5 control biopsies, questioning their pathogenicity and necessary role in lesion causation. From cultivation, different fungi were isolated and identified by sequencing of the nuclear ribosomal internal transcribed spacer region. In addition, 2 primer pairs were used in a nested format to increase the sensitivity for direct amplification and identification of fungi from lesions, thereby circumventing cultivation. Results suggest mixed and opportunistic infections in sea fans with these lesions, requiring longitudinal or experimental studies to better determine the pathogenesis.
Over the past 3 decades, Caribbean coral cover shrank to 10% as coral reefs face unprecedented environmental challenges tied to the human impact and an increase in coral disease epizootics.7,8 Despite the rapid emergence and impact of diseases and significant advances in methodology and knowledge base, the identification of proximate causes, pathology, and epizootiology remains poorly understood, 44 which hinders our ability to mitigate disease in affected scleractinian (hard) coral and octocoral (soft coral) populations. A major hurdle is the lack of clear diagnostic criteria and the limited ability to isolate and determine etiological agents. Descriptions of largely macroscopic lesion morphologies, such as the extent and pattern of tissue loss and abnormal coloration, form the basis for nomenclature and diagnosis. However, corals display few macroscopic signs that indicate a wide array of maladies and, consequently, similar signs may have multiple root causes. 46 Scleractinian coral declines increasingly restrict their sampling for scientific study (all are listed under Appendix II of the Convention on International Trade in Endangered Species). Octocorals are useful models for improving our understanding of anthozoan pathology because they are plentiful and their soft skeleton is more amenable to sample collection.
The octocoral disease aspergillosis is currently considered an endemic disease in the wider Caribbean region, 42 and one of the few coral diseases for which the Koch’s postulates were purportedly fulfilled.9,22,32 Aspergillosis particularly affects the sea fans Gorgonia ventalina and G. flabellum, and presents as (multi)focal circular tissue purpling, progressing to annular purpling with a central area of soft tissue (coenenchyme) loss, and exposure to and subsequent fouling of the axial skeleton by a variety of cyanobacteria, algae, sponges, hydrocorals, and other organisms. 42 This purple pigmentation results from an accumulation of sclerites that contain a carotenoid pigment, 18 and appears to be a generalized highly nonspecific response to contact with biotic agents.1,6 It is therefore unlikely to be pathognomonic for aspergillosis. Yet sea fans showing annular purple pigmentation with central tissue loss are typically given a field diagnosis of aspergillosis.14,15,19,41
Aspergillus sydowii, a saprophytic fungus ubiquitous in terrestrial and marine environments, 26 was identified as the putative causative agent based on Koch’s postulates, morphological description, and molecular identification targeting a non-translated region of the trpC gene. 9 Recent findings fuel the debate on whether this fungus is the primary pathogen or merely a secondary opportunistic invader and part of the yet largely undescribed mycoflora of gorgonians. In other animals, Aspergillus spp. are typically opportunistic pathogens. 30 Although different Aspergillus spp. have been isolated from both healthy and diseased sea fan colonies,2,47 A. sydowii is not always isolated from diseased tissue, 38 and experimental exposure does not consistently induce aspergillosis.21,36,40 Estimates of fungal diversity in sea fans are largely based on fungal cultivation, with subsequent isolation of culturable fungi and molecular identification based on amplification and sequencing the internal transcribed spacer (ITS) regions with ITS-1 and ITS-4 primers.38,43 Aspergillus spp. grow readily in culture under a wide range of conditions, and it is likely that culture-based studies may overstate the importance of Aspergillus relative to fungi that are less or nonculturable. Moreover, detailed and precise histological descriptions were lacking in the study addressing Koch’s postulates, 32 bringing uncertainty to comparisons with other studies histologically identifying fungi in sea fan tissues. 21
To this end, we analyzed lesions resembling sea fan aspergillosis with a holistic diagnostic approach, including detailed description of the histopathology, and a combination of culturing and direct molecular amplification of fungi. This enabled us to determine the variation of pathology and micro-organisms underlying these lesions, and thereby improve the understanding of their causation, pathogenesis, and diagnosis.
Materials and Methods
Sampling
Biopsies were collected from G. ventalina and G. flabellum sea fans affected with lesions consistent with aspergillosis. Study sea fans were from 4 reefs (Supplemental Table S1). An approximately 2 cm maximal width wedge-shaped biopsy was collected using titanium scissors to include the tissue loss margin (TLM), the zone of purple pigmentation, and the apparently healthy margin (AHM) of the lesion. For 25 diseased sea fans, an additional, approximately 2-cm wedge-shaped biopsy was also collected from the apparently healthy tissue at the periphery of the colony, which were termed lesioned controls. In addition, control biopsies were similarly collected from 14 sea fans lacking gross lesions (3 from Bird Rock Reef, 3 from Whitehouse Bay, 7 from Turtle Beach, and 1 from Reggae Beach). Sea fan biopsies were transferred to shore in Whirl-Pak bags. Once on shore, each biopsy was bisected with scissors. Half of the biopsy was immersed in Z-Fix (Anatech Ltd) diluted 1:5 with seawater. The other half of the sample was partitioned for fungal culture and molecular identification. The half used for mycological analysis was surface sterilized in ethanol 70% for 30 s and then washed with filtered autoclaved seawater for another 30 s to eliminate potential contaminant fungi present on the biopsy surface.
Histology
After 48 hours fixation, biopsies were partitioned into cassettes. Lesion biopsies were split into 2 cassettes: one with purple pigmented tissue and TLM, the other with purple pigmented tissue and AHM, (i.e., lesion interface; Fig. 1). Biopsies were then decalcified by immersion in Formical 2000 (StatLab, McKinney, TX) for 4–8 hours. Once decalcified, biopsies were processed routinely for histology and were embedded in a planar orientation in paraffin wax (i.e., oriented so that sections were parallel to the face of the sea fan). Samples were sectioned at a thickness of 4 µm and stained with hematoxylin and eosin. Ancillary histochemical stains, including Grocott’s methenamine silver, Periodic acid-Schiff, and Alcian blue, were used on a subset of sections containing micro-organisms.
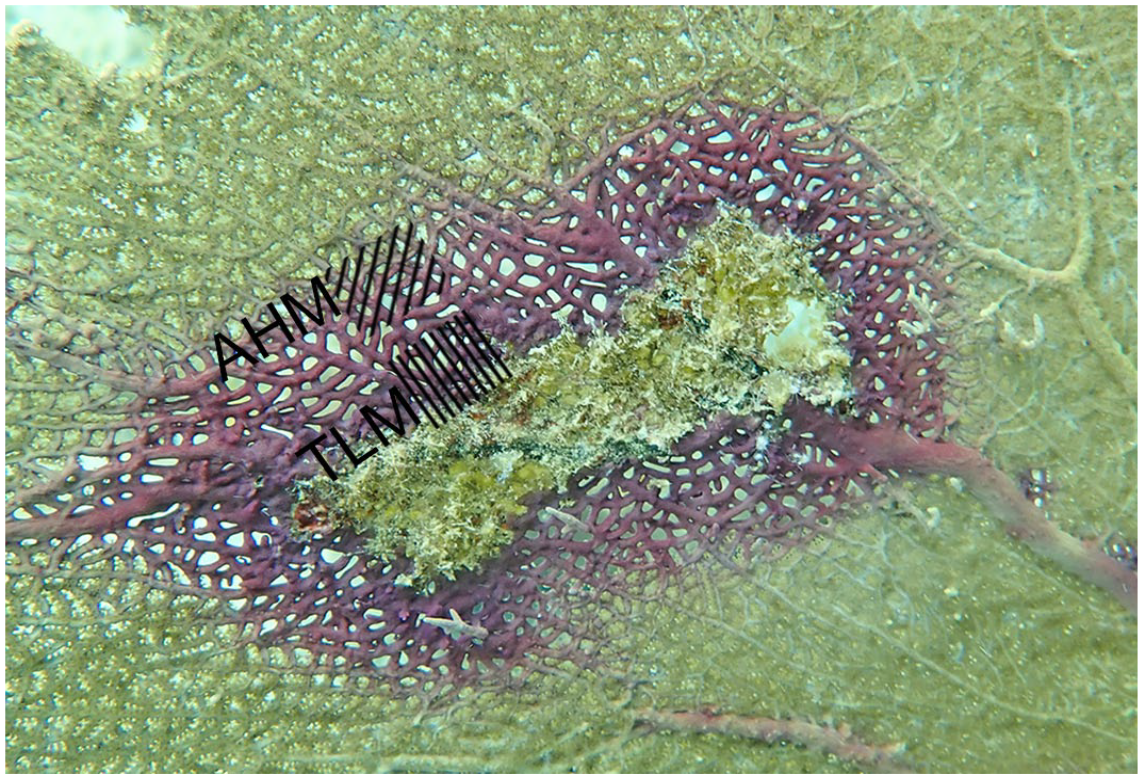
Purple annular pigmentation with central tissue loss, Gorgonia ventalina. Gross lesion appearance is typical of sea fan aspergillosis. The 2 lesion margins were histologically examined: tissue loss margin (TLM) and apparently healthy margin (AHM).
For histological analysis, microscopic features of lesions and associated micro-organisms were recorded separately for the 2 zones of the purpled lesion (TLM vs AHM), and for control samples. At the TLM, organisms were only recorded when they invaded the tissue (i.e., those within the area of tissue loss were not detailed). Organisms identified within sea fan tissues by histology were classified according to morphologic characteristics, using methods as described by Work et al. 45 Filaments or mosaics of cuboidal to rectangular cells with distinct cell walls were classified as algae. Filamentous structures without thick cell walls and internal cellular detail were classified as fungus-like hyphae. Filaments of concatenated trichomes (i.e., rows of densely stacked elongate cells) were classified as cyanobacteria. Ciliates were protists with cilia and macronuclei. An elongate (typically ~0.5-1 mm length in section) organism with a digestive gland, ganglia, subepidermal glands, and epidermis rich in mucous cells was classified as an unidentified metazoan. Hydroids were tubes exteriorly lined by epidermis containing cnidocytes, interiorly lined by gastrodermis containing granular gland cells, and encased within hyaline periderm. Ovoid magenta cells (1–10 µm) separated by clear space, suggestive of a mucous sheath, were classified as labyrinthulomycetes.5,39
Fungus-like hyphae were further subcategorized based on morphological features (Table 1).11,28 The abundance of hyaline hyphae was semi-quantitatively scored as follows: score 1—involved <10% of tissue in a section; score 2—involved 10–50% of tissue in a section; score 3—involved >50% of tissue in a section.
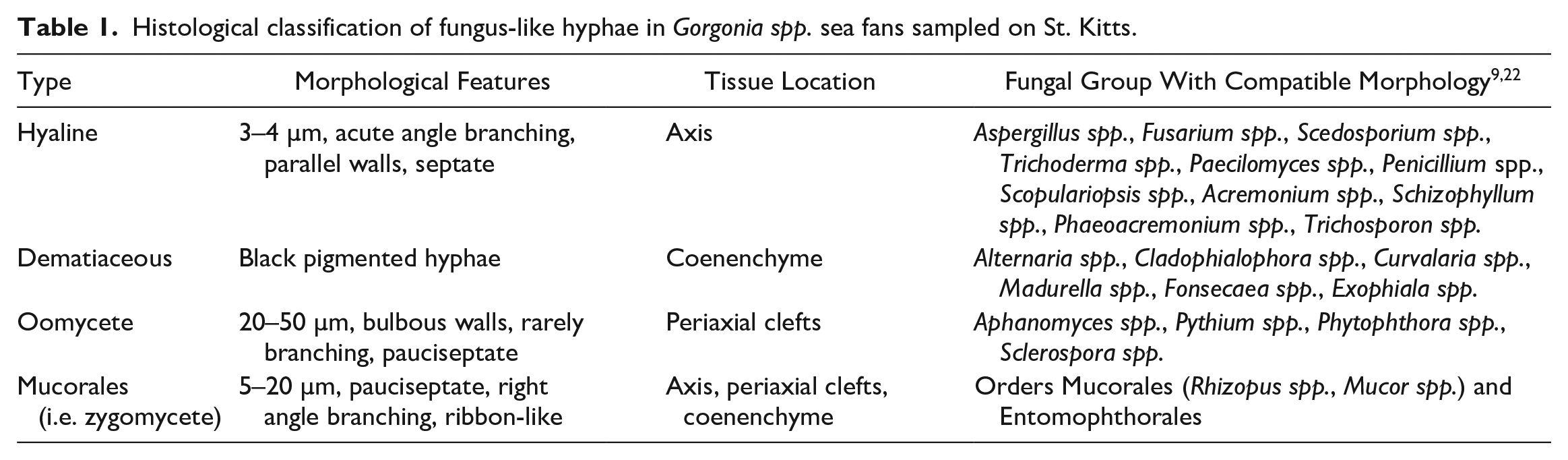
Histological classification of fungus-like hyphae in Gorgonia spp. sea fans sampled on St. Kitts.
CellSens Standard image analysis software (Olympus, Tokyo, Japan) was used to measure polyp diameter by taking the longest dimension of a polyp that appeared to be in full cross-section and representative of polyps across the section. Mean polyp diameter was determined from 3 polyp measurements per section. Endosymbiotic dinoflagellates (family Symbiodiniaceae) were similarly counted for a 100-µm length of superficial gastrodermis for 3 polyps, and a mean was recorded for each section. Amoebocytosis was subjectively scored according to approximate percentage of mesoglea occupied by the cells: mild = 15–25%; moderate 25–50%; severe >50%. Biopsies that contained polyps with identifiable gonads were categorized as reproductively active.
The proportion of biopsies with histologically observed microbial colonization and reproductively active polyps were compared among paired sampling groups (TLM, AHM, and lesioned controls) using contingency tables and a McNemar’s chi-square test. The proportion of biopsies with histologically observed microbial colonization and reproductively active polyps were compared with control biopsies (lesioned vs non-lesioned sea fans) using a Pearson chi-square test. A Wilcoxon Signed Rank test was used to compare median hyphae abundance score of TLM with AHM. A Bonferroni correction to allow for multiple comparisons adjusted the significance level from P ≤ .05 to ≤ .01. 25 Statistical analyses were performed using SPSS version 28.0.0.0 (IBM Corporation, New York).
Fungal Culture and Identification
Small fragments (1–2 cm2) from the lesion margin, including the zone of purple pigmentation, and from apparently healthy tissue in 18 diseased sea fan colonies were plated (1 fragment per plate) on Sabouraud dextrose agar (SDA) and glucose peptone yeast agar (GPYA). The latter is a standard medium for marine fungi (3 g glucose, 0.3 g yeast extract, 0.3 g peptone, and 20 g agar l-1 0.2-μm membrane-filtered seawater).4,35 In parallel, small fragments (1–2 cm2) from 6 sea fan colonies without gross lesions were also cultivated. Plates were incubated at 28°C and observed for fungal growth every 2 days for 1 month. For identification, mycelial mass observed on the top of the coral tissue was isolated and re-cultured on SDA or GPYA to obtain pure isolates and (where possible) identified using microscopic and macroscopic characteristics of the mycelium and reproductive structures.33,34 Some colonies did not sporulate and morphology-based identification was not possible. Thereafter, all isolates were transferred to liquid potato-dextrose (PD) medium made with filtered, autoclaved seawater for DNA extraction. One week after transfer to liquid PD, fungal mats were removed from the surface of the agar with a sterile scalpel, and DNA extracted using MO BIO’s PowerSoil® DNA Isolation Kit. The nuclear ribosomal ITS regions were amplified using the primers ITS-1/ITS-4 with PCR conditions described by White et al. 43 PCR amplicons of the expected size of approximately 700 bp were sent to an external laboratory for bidirectional sequencing. The resulting sequences were identified as having a fungal origin using BLAST searches in GenBank. Identifications based on BLAST searches were confirmed by microscopic observations.
Direct Fungal Amplification
Small fragments from the lesion margin from 25 sea fans were used for DNA extraction that was performed using a tissue DNA purification kit (EurX Poland). DNA concentrations were quantified using a NanoPhotometer (Implen) and used as templates for subsequent PCR. Initial PCR methods followed White et al, 43 using the universal fungal primers ITS-1, ITS-2, ITS-3 and ITS-4; however, due to the consistent amplification of host and other non-target DNA, typically Symbiodiniaceae, with some combinations of these primers, new more targeted fungal primers were designed by eye from alignments of multiple fungal isolates, predominantly ascomycetes, from GenBank in ClustalX; the objective being to aid amplification of a broad range of fungal taxa in a given DNA sample. Two primer pairs were designed that could be used in a nested format, to increase sensitivity. The external primers Fun-1490fwd 5′ agrgccagcgagtacatcac and 618-rev 5′ atccctacctgatccgaggtc amplify approximately 800 bp including the 3′ end of the 18S, full ITS1 and 5.8S and partial ITS2. For the nested primers, an internal forward primer, still within the 18S, Fun-1659fwd 5′ tactaccgattgaatggct was used with the reverse primer ITS-2. 43 The nested primers were used on samples where no, or poor, amplification was observed with the external primer pair and amplified approximately 400 bp including partial 18S, complete ITS-1, and partial 5.8S genes. PCR conditions were as described by White et al, 43 with the exception of 20-µl PCR volumes and an annealing temperature of 60°C for the external primers and 55°C for the internal pair.
PCR products were resolved on 1% agarose gels with molecular weight ladders, allowing amplicons of the expected sizes to be identified, purified, and sent to an external laboratory for bidirectional sequencing. The resulting sequences were identified as having a fungal origin using BLAST searches in GenBank and contigs constructed using Bioedit and ClustalX.
Results
Histopathology
Two main lesion areas were compared with each other and to control biopsies from lesioned and non-lesioned sea fans (Table 2). The purple lesion adjacent to the TLM presented a consistent general picture of opportunistic microbial overgrowth of exposed axis (Fig. 2a, b), variably bordered by amoebocyte encapsulation (Fig. 2c), gastrodermal necrosis (Fig. 2d, e), polyp loss (Fig. 2f), and coenenchymal amoebocytosis. Gastrodermal necrosis was characterized by dissociated clusters of gastrodermal epithelial cells and associated endosymbionts within gastrovascular canals. Polyp loss was evident as stretches of coenenchyme devoid of polyps. Coenenchymal amoebocytosis consisted of increased numbers of amoebocytes within the mesoglea. Coenenchymal amoebocytosis at the TLM was mild (n = 8, 22%), moderate (n = 16, 44%), or severe (n = 12, 33%). Peri-axial clefting was also consistently present, and was indicated by a cavitated space between the axis and axial epithelial cells with subepithelial laminar deposition of melanin (Figs. 2e, f). Melanin deposits were golden brown, as previously described in sea fans. 21 Peri-axial clefts were filled with basophilic pleomorphic debris, a mixture of micro-organisms (typically cyanobacteria or algae), and clear areas suggestive of hard contents artefactually lost during sectioning.
Microscopic pathology and microorganisms observed in Caribbean sea fan (Gorgonia spp.) biopsies of lesions grossly resembling aspergillosis, with comparison among the purpled tissue adjacent to the TLM, the AHM, and control biopsies.
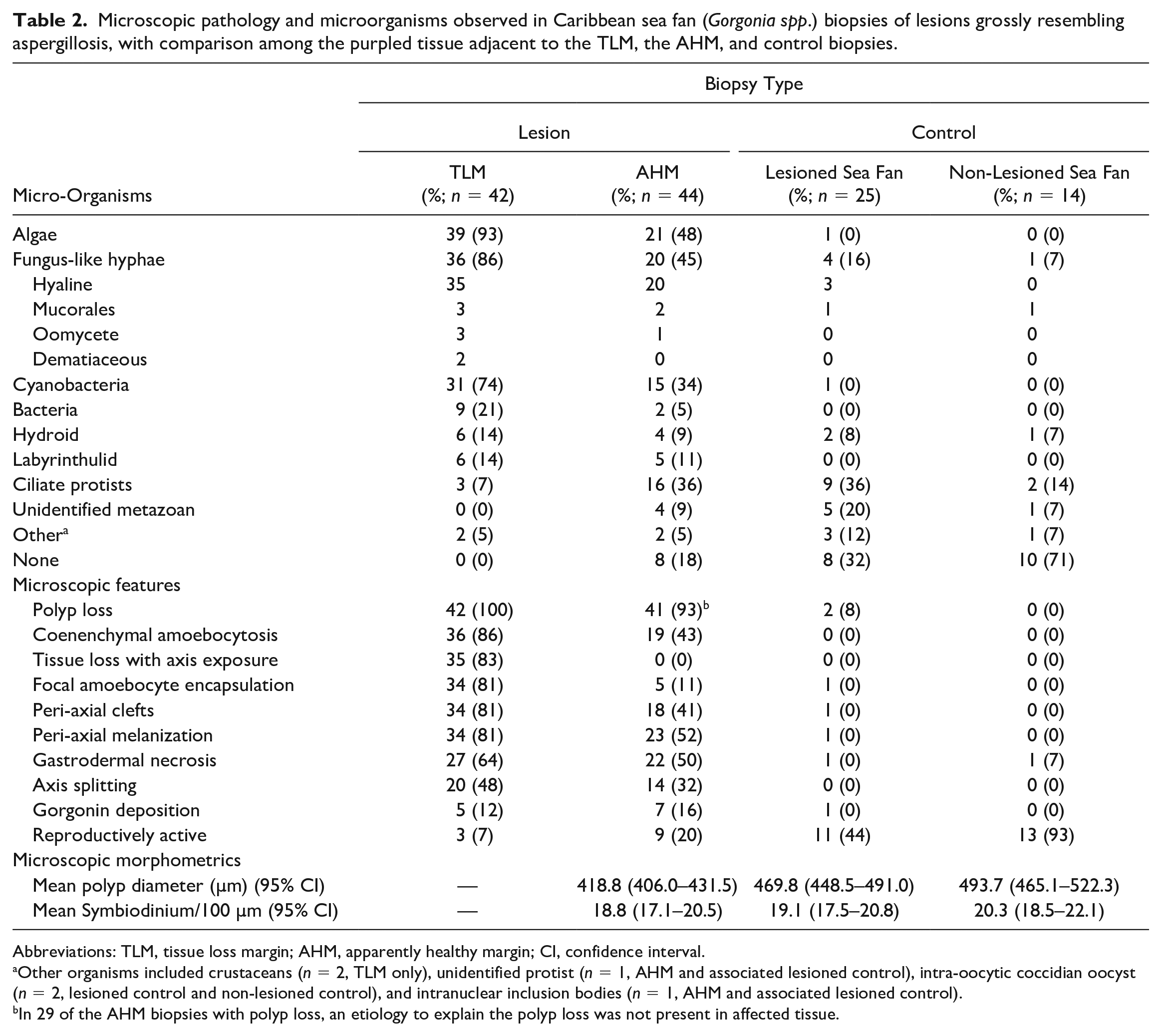
Abbreviations: TLM, tissue loss margin; AHM, apparently healthy margin; CI, confidence interval.
Other organisms included crustaceans (n = 2, TLM only), unidentified protist (n = 1, AHM and associated lesioned control), intra-oocytic coccidian oocyst (n = 2, lesioned control and non-lesioned control), and intranuclear inclusion bodies (n = 1, AHM and associated lesioned control).
In 29 of the AHM biopsies with polyp loss, an etiology to explain the polyp loss was not present in affected tissue.
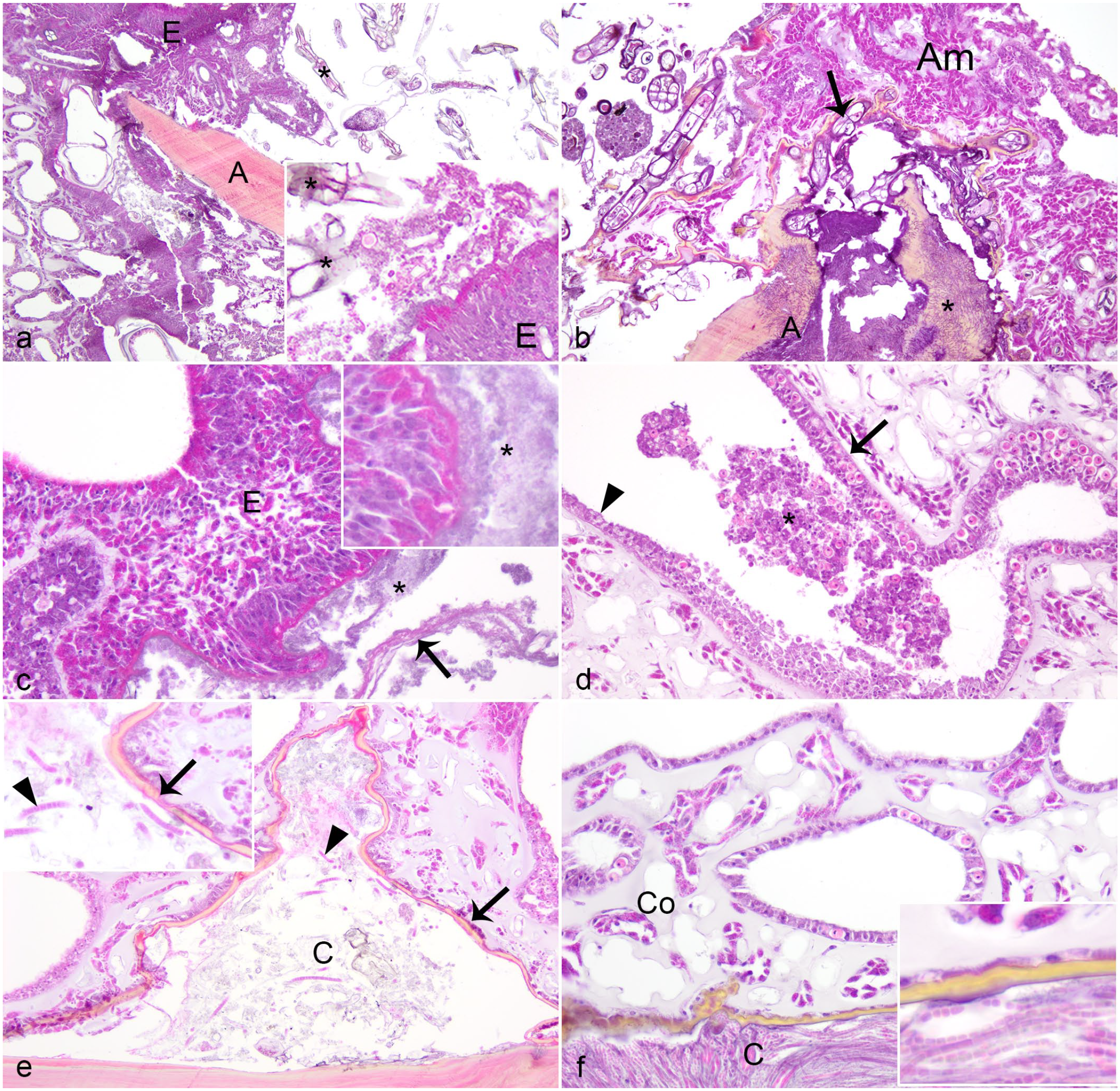
Tissue loss margin of Caribbean sea fans with gross lesions suspicious for aspergillosis. Hematoxylin and eosin. (a) Tissue loss exposing the axis (A) with amoebocyte encapsulation (E, inset) bordering infiltrates of macroalgae (asterisk, inset). (b) Severe amoebocytosis (Am) adjacent to tissue loss with axis exposure (A), with infiltrates of algae encapsulated by melanin (arrow), and the presence of abundant hyaline hyphae (asterisk) within the axis (A). (c) Amoebocyte encapsulation (E, inset) of bacteria (asterisks) within coenenchyme; note hyaline membrane formation (arrow). (d) Gastrodermal necrosis. Clumps of necrotic sloughed cells (asterisk) are within the lumen of a gastrovascular canal, which is segmentally lined by normal-appearing (arrow) and attenuated (arrowhead) gastrodermis. (e) Subepithelial periaxial cleft (C) bordered by laminar melanin deposition (arrow, inset) and containing debris and cyanobacteria (arrowhead, inset). (f) Periaxial cleft (C) filled with cyanobacteria (inset) and bordered by laminar melanin deposition, with polyp loss in overlying coenenchyme (Co).
Typically, the deep periaxial coenenchyme and axis bordering the TLM were highly colonized by a variety of microorganisms, particularly algae, fungi, and cyanobacteria (Fig. 3). The location of fungi depended on their morphology (Table 1), but hyaline hyphae within the axis predominated. There was a large degree of variation in the amount of hyaline hyphae colonization at the TLM, including abundance scores of 1 (n = 13, 37%), 2 (n = 12, 34%), and 3 (n = 10, 29%). In contrast to fungi, algae and cyanobacteria were typically within the coenenchyme, associated with melanization or amoebocyte encapsulation, or within periaxial clefts (Fig. 2e). Algae were also within the axial skeleton in many instances (13/21 instances of algae at TLM; Fig. 3d, f), sometimes co-localized with fungi (10/21; Fig. 3f). Intra-axial organisms (fungi, labyrinthulids, and algae) were associated with periaxial melanin deposition (Figs. 2e, f, 3a-f). Intra-axial labyrinthulids and algae were associated with axial splitting (Figs. 3d, f, g), whereas fungi were not, unless they were co-localized with other micro-organisms.
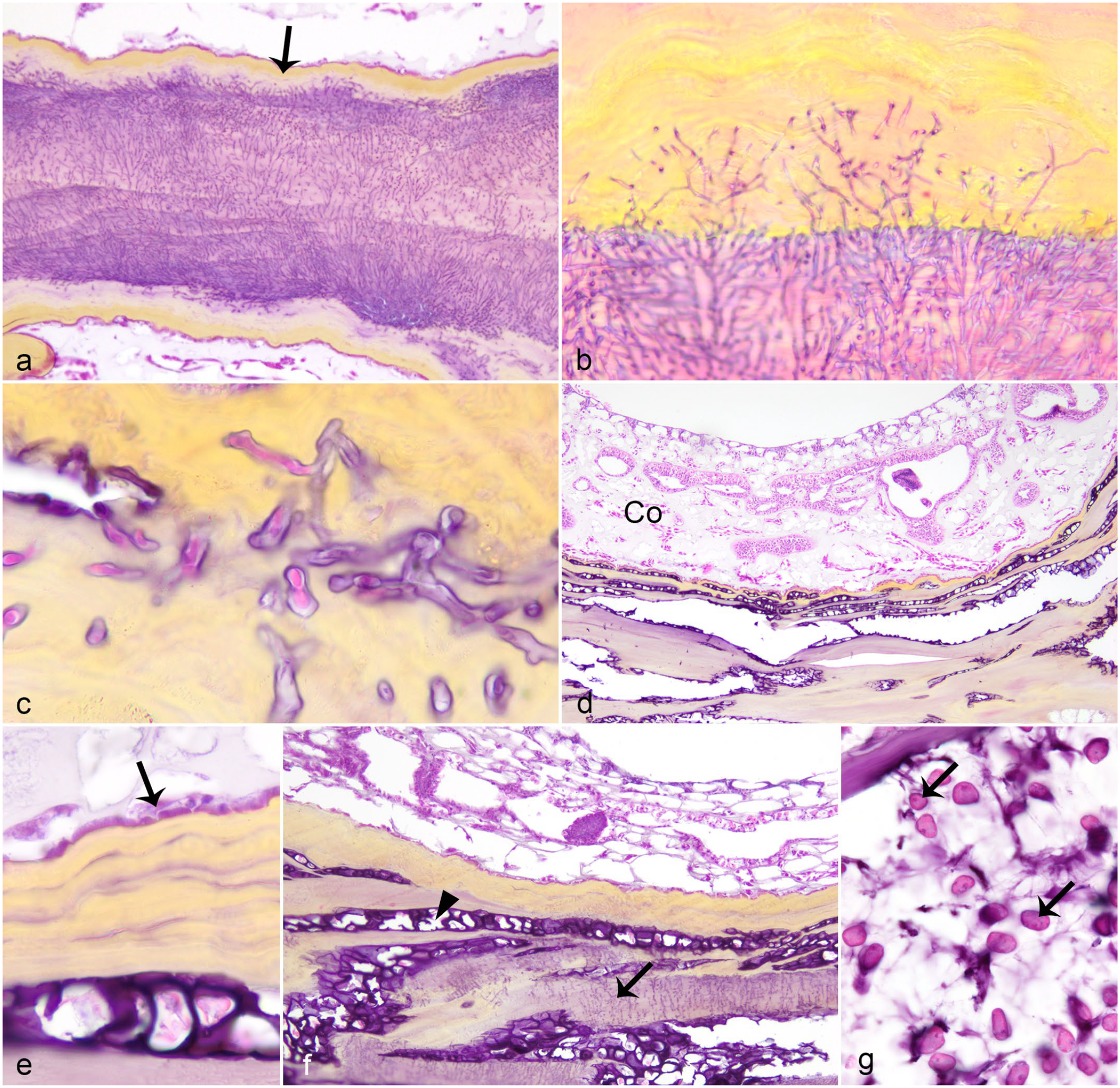
Common micro-organisms of Caribbean sea fans with gross lesions suspicious for aspergillosis. Hematoxylin and eosin. (a) Hyaline fungi within the axis, bordered by melanin deposition (arrow). (b) Higher magnification of axis hyaline fungi; note extension into adjacent melanin deposition. (c) Mucorales fungi within periaxial melanin deposition (same magnification as Fig. 3b). (d) Infiltrates into the axis of filamentous algae with prominent cell walls associated with axis splitting and absence of polyps in overlying coenenchyme (Co). (e) Higher magnification of algae within the axis showing thick cell wall and subepithelial (arrow) periaxial location associated with melanin deposition. (f) Combined axial hyaline fungi (arrow) and algae (arrowhead); note axis splitting associated with algae. (g) Labyrinthulomycete (arrows) within rarified axis.
Tissue loss with axis exposure and amoebocyte encapsulation were absent and infrequent, respectively, in the area of the purple lesion adjacent to the AHM. However, polyp loss, gastrodermal necrosis, and coenenchymal amoebocytosis were present (Fig. 4a-c). In 29/41 (71%) instances of polyp loss at the AHM, an explanatory etiologic agent was not evident in affected areas of the section. At the AHM, coenenchymal amoebocytosis was mild (n = 15, 79%), moderate (n = 3, 16%), or severe (n = 1, 5%). Reproductively active polyps were observed in TLM, AHM, and control biopsies of lesioned sea fans (Fig. 4d, Table 2), but differences in proportion of biopsies with reproductively polyps were not statistically significant (TLM relative to AHM: McNemar χ2 = 4.2, P = .041; AHM relative to lesioned control: McNemar χ2 = 2.3, P = .131). The proportion of biopsies with reproductively active polyps was lower in control biopsies from lesioned sea fans relative to non-lesioned sea fans (Pearson χ2 = 9.05, P = .003).
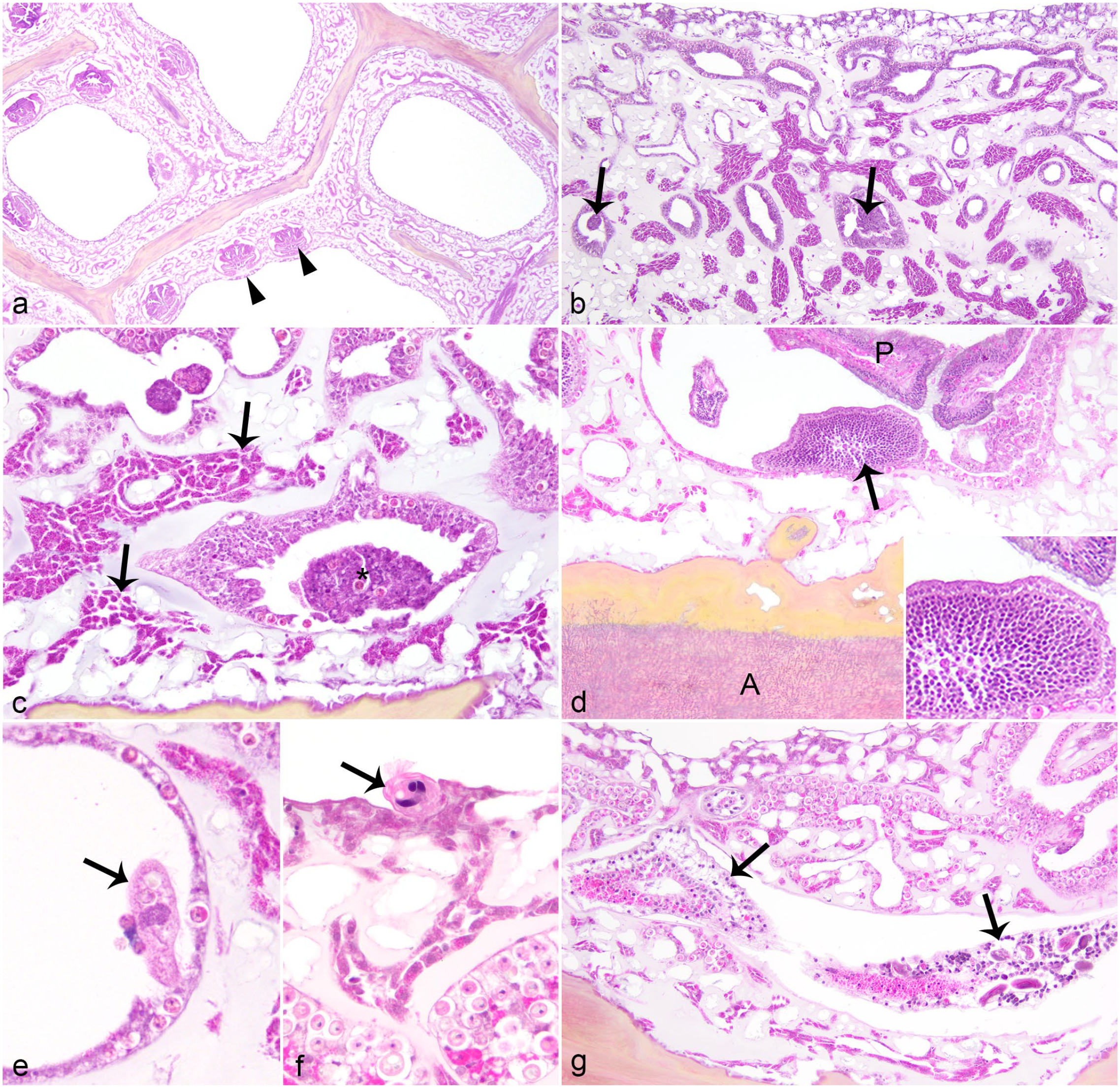
Apparently healthy margin of Caribbean sea fans with gross lesions suspicious for aspergillosis. Hematoxylin and eosin. (a) Polyp (arrowhead) loss at the transition from grossly purpled (right) to normal (left) coloration. Note absence of micro-organisms. (b) Gastrodermal necrosis (arrows) with moderate ameobocytosis. Note absence of micro-organisms. (c) Higher magnification of gastrodermal necrosis (asterisk) with ameobocytosis (arrows designating enlarged aggregates of amoebocytes). (d) Male gametes (arrow, inset) indicative of reproductive activity in a polyp (P) within an area with axial hyaline fungi (A) and periaxial melanin deposition. (e) Ciliate protist (arrow) within gastrovascular canal. (f) Ciliate protist (arrow) associated with surface epidermis. (g) Hydroid (arrows) within coenenchyme.
The proportion of biopsies with histologically identifiable micro-organisms varied among sampling groups (Table 2), but these differences were not statically significant (TLM relative to AHM: McNemar χ2 = 5.1, P = .023; AHM relative to lesioned control: McNemar χ2 = 0.44, P = .51; lesioned relative to non-lesioned control: Pearson χ2 = 5.61, P = .018). Ciliates were always sparse at the AHM, morphologically diverse; associated with epidermal surface, within superficial coenenchyme or gastrovascular canals; and, in some instances, were histophagous (Fig. 4e, f). Fungal colonization of the AHM was less frequently identified than the TLM, and had hyaline hyphae abundance scores of 1 (n = 7, 35%), 2 (n = 6, 17%), and 3 (n = 7, 20%). The median hyaline hyphae abundance score was higher in the TLM (median = 1.6) relative to AHM biopsies (median = 0.89; Wilcoxon Signed Rank test, Z = -2.829; p< 0.005).
Fungus-like hyphae were assigned to the category (Table 1) that most closely approximated the combination of morphological features observed histologically. Some of these features, particularly angle of branching and presence of septation, could be difficult to discern in histology sections, especially when hyphae were intra-axial. The walls of both algae and fungus-like hyphae predominantly stained black with GMS, blue with Alcian blue, and pink with PAS stains. GMS and Alcian blue stains were often equivocal when attempted on intra-axial organisms because of heavy concurrent staining of axis surrounding organisms.
Mycology: Cultivation and Direct Molecular Identification
From 18 diseased sea fan colonies, 12 fragments from the lesion margin plated on SDA or GPYA resulted in fungal growth, though the majority did not sporulate to allow reliable identification solely based on colony or conidial morphology. However, 11 out of 12 cultured isolates could be further identified using White’s ITS-1/ITS-4 primers (Table 3).
Histological fungal morphology, fungal cultivation and direct fungal amplification in Caribbean sea fan (Gorgonia spp.) biopsies of lesions grossly resembling aspergillosis.
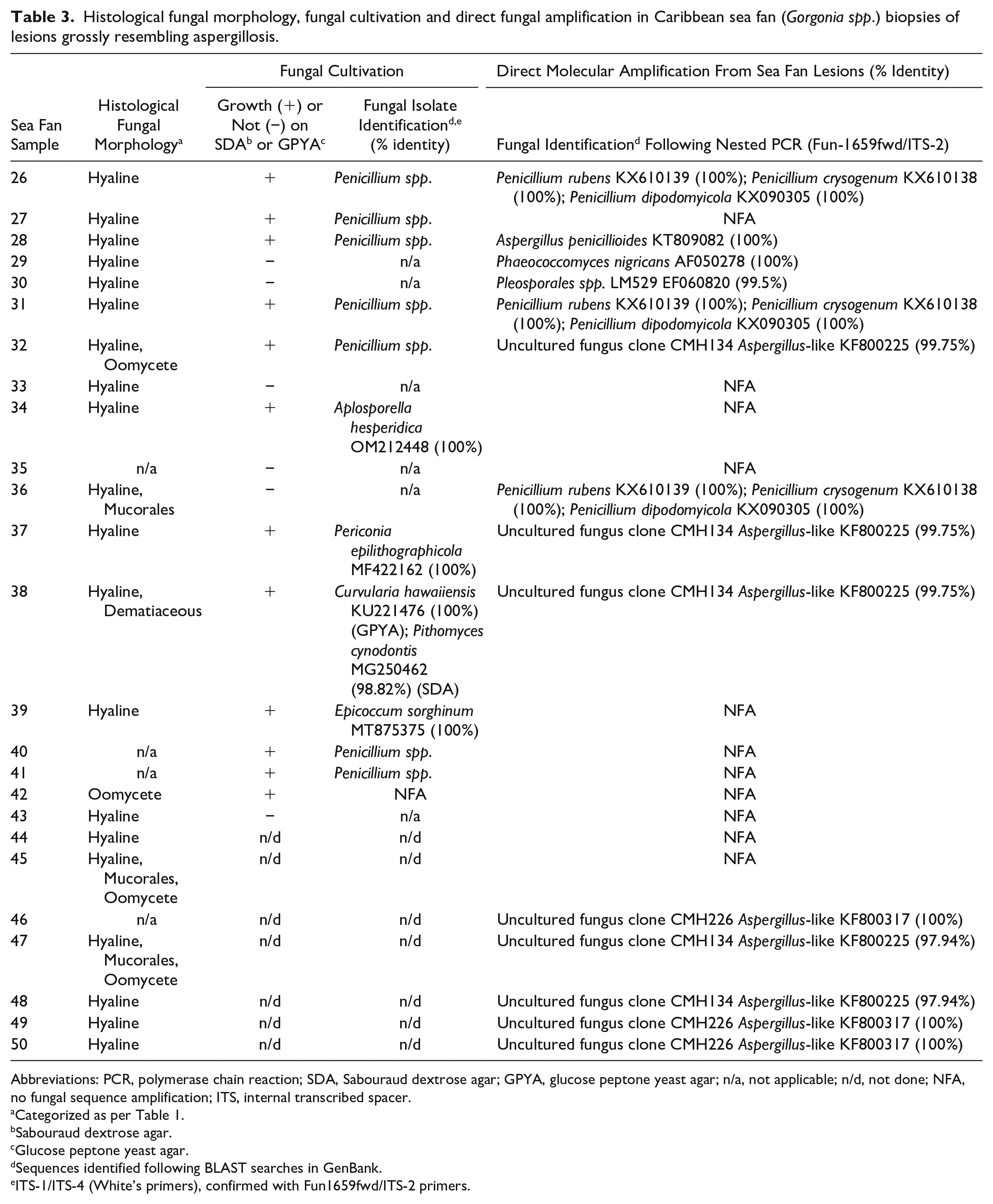
Abbreviations: PCR, polymerase chain reaction; SDA, Sabouraud dextrose agar; GPYA, glucose peptone yeast agar; n/a, not applicable; n/d, not done; NFA, no fungal sequence amplification; ITS, internal transcribed spacer.
Categorized as per Table 1.
Sabouraud dextrose agar.
Glucose peptone yeast agar.
Sequences identified following BLAST searches in GenBank.
ITS-1/ITS-4 (White’s primers), confirmed with Fun1659fwd/ITS-2 primers.
More specific primers were needed to amplify and sequence fungal DNA from mixed template DNA taken directly from the sea fan lesions. Indeed, using the aforementioned nested PCR proved to be the most successful method to amplify and sequence fungal DNA from sea fan lesions. Fourteen fungal contigs were generated from DNA extracted directly from the sea fan lesions. Of these, 7 different fungal sequences were identified, some from multiple sea fan lesions with others being unique to 1 sea fan (Table 3). These 7 sequences have been assigned the accession number OQ676388-OQ676495 in GenBank. Discrepancies between culture and DNA sequencing-based results showed an increased relative abundances of Penicillium, Cladosporium, and Aspergillus spp. in culture compared to DNA sequencing.10,31 Interestingly, the histopathological observation of hyaline hyphae in the biopsies was not consistently correlated with fungal cultivation or direct molecular identification, underpinning the need for a combined diagnostic approach to lesions resembling aspergillosis in sea fans.
Discussion
Sea fan aspergillosis is often considered among the best characterized of all coral diseases; however, by using a standardized biomedical approach to disease diagnosis that includes histology, culture, and molecular techniques, this study underpins the complexity of sea fan aspergillosis and the limitations of each one of these methods. It is surprising that no such combined diagnostic procedure has been attempted for sea fan aspergillosis given aspergilloses in many wild and domestic animals are diagnosed through a combined diagnostic procedure. 30
This study conclusively shows for the first time that gross lesions of annular purple pigmentation with central tissue loss are not specific to sea fan aspergillosis, or even fungal infection more broadly. In our study, microscopic examination demonstrated that algae were at least equally as common as fungus-like hyphae in these lesions. This contrasts with the initial description of sea fan aspergillosis, where colonization by other organisms was limited to cyanobacteria along the exposed axis (described as “recessed rind”) and was considered secondary to tissue loss caused by the fungus. 32 In the present study, fungi were not clearly causing tissue loss (Fig. 2a). Fungal infections were mild at the tissue loss margin in many instances, and fungus-like hyphae were not observed at all in the biopsies of 6/44 lesioned sea fans, indicating that fungal infection is not a necessary cause for this lesion. In addition, fungus-like hyphae were not unique to the lesioned tissue, as they were histologically identified in 7–16% of control biopsies, consistent with other studies that demonstrate axial fungi in grossly non-lesioned samples.6,16 These findings challenge Koch’s postulates, which require an organism to be present in all cases of the disease, but not in healthy individuals.
Histologically, more than 1 morphological type of fungus-like hyphae were identified in lesions suspected to have sea fan aspergillosis. Other studies have similarly documented multiple fungal morphologies in purpled lesions of sea fans.6,20 The thin, septate, dichotomously branching hyaline hyphae which predominated in this study and others,16,21,23 could represent Aspergillus species as well as numerous other fungi. 11 Histological identification of fungi can be problematic due to overlapping morphological features and challenges in accurately assessing morphology in 2-dimensional tissue sections. 27 Structures convincingly resembling conidiophores were never seen within tissue sections to allow more specific histological identification of fungus-like organisms. Consistent with the histological findings, microbiological and molecular techniques identified several different fungi associated with these lesions.
Fungal cultures are considered the complementary gold standard tool for laboratory diagnosis,11,27 but can be time consuming and technically challenging. As demonstrated by our study, a major challenge for marine mycology is the inability to easily culture the microbially diverse fungal populations, with plating methods favoring the isolation of faster growing generalist and osmotolerant genera, and many fungi not sporulating in culture, making visual identification challenging. 10 Sea fan tissue fragment size and processing can also have direct impact on the fungal yield and different genera retrieved from cultivation 35 which further explains the discordant results between histopathology and cultivation. With the continued growth of the molecular diagnostic field, sequence-based identification should also be used in parallel for a reliable identification. Moreover, the complexities in fungal identification reflected in our study further scrutinize prior conclusions that relied on colony and cellular morphology of fungal isolates retrieved on Hektoen enteric agar, a selective differential medium for gram negative bacteria, to determine if Aspergillus spp. is the cause of Caribbean sea fan mortalities. 31 In addition, the study implicating A. sydowii as the cause of these lesions based on experimental infection did not histologically confirm the presence of fungi, 8 thereby highlighting the importance of our findings.
However, amplification and sequencing of DNA from fungal isolates using the universal primers ITS-1/ITS-4 was not consistent, and DNA extracted directly from sea fan lesions never resulted in fungal amplification with this primer set and typically targeted more abundant templates such as the endosymbiont (Symbiodiniaceae) or the sea fan host. It has been previously stated that some fungal primers, ITS-1 included, would amplify non-fungal DNA 43 and other researchers have noted that due to the diverse nature of fungi, ITS primers need to be carefully chosen, or new primers designed to target a certain region of the ITS within the rRNA gene. 3 These primers may reflect the specific target group, basidiomycetes or ascomycetes and the sample type being analyzed, such as environmental samples or those with a clinical application. 3 Therefore, alternative primers were designed in this study to directly amplify fungi from lesions, thereby circumventing cultivation. We found the ITS-2 primer to be specific enough for fungal amplification, so we utilized this primer when designing the nested PCR. The nested cycle of the new PCR yielded the best amplification and subsequent sequence reads with 100% bidirectional agreement from direct sequencing. The nested amplicon includes the entire ITS-1 region of the rRNA gene, which is well represented in the genetic databases and hence can be used to identify fungal pathogens in complex mixtures of host or environmental DNA. The ITS region is the official DNA barcoding marker for identification of fungi; although certain limitations have been identified, it is still considered superior for species-level identification than using other gene regions or morphology alone.17,24,29
Amplification of fungal DNA from the first round of the PCR was not considered sufficient for direct sequencing, with faint bands only sometimes being observed in gel electrophoresis; hence the need for the nested cycle. This is most likely due to the low amount of fungal DNA in the templates extracted from the sea fan lesions. No mixed fungal sequences were generated with the nested PCR which was somewhat surprising as more than 1 fungal morphology was confirmed histologically and the primers have been designed to amplify a wide range of fungal DNAs. This may suggest that each lesion has a dominant fungal type or that some form of preferential amplification is occurring during the PCR; however, a wide range of fungi have been amplified with these primers from various tissues and hosts (unpublished data). Next-generation amplicon sequencing, using suitable primers or the ones described here, could potentially yield important information on the number and relative abundance of fungal taxa that are associated with sea fan lesions. Such molecular techniques have been widely applied in scientific research but less so in clinical pathogen detection, with fungi receiving far less attention than bacteria and viruses. The development of such methodologies will assist in the identification of fungi causing infections in numerous clinical and environmental scenarios. 13
The micro-organisms within areas of annular purple pigmentation away from the tissue loss margin were mainly localized within the axis, the pliable proteinaceous skeleton of the octocoral, or peri-axial clefts. The pathogenicity of micro-organisms in this location is uncertain as they are fairly well confined by melanin deposition and bordered by coenenchymal amoebocytosis putatively representing a cell-mediated immune response, as observed in this and other studies.20,21,23 Indeed, the pathogenicity of organisms which reside in the skeleton (i.e. endoliths) is similarly debated in scleractinian corals. 12 A mechanism whereby axial fungi or algae can injure the polyp, resulting in the gastrodermal necrosis and polyp loss documented in this study is undetermined. Moreover, intra-axial organisms were often present without gross or histological changes (other than melanin deposition), even with the host gorgonian showing active gonadogenesis in some instances, suggesting that their presence was of little consequence to neighboring coenenchyme and polyps. The purpling of sea fan tissues reflects accumulation of a carotenoid pigment within sclerites, 18 calcified deposits within coenenchyme hypothesized to provide a physiochemical barrier to infection. 37 While sclerites are dissolved during histologic processing, and thus were not assessed by this study, it may be that the clear spaces within peri-axial clefts represents a site of pigmented sclerite accumulation.
Detailed histologic descriptions of the lesion interface (i.e., AHM, where the lesion transitions to normal tissue) were lacking from previous histological descriptions of sea fan aspergillosis. 32 This is remarkable because histological assessment is often directed toward the lesion interface as it provides the most uncomplicated view of the earliest or primary pathological features, whereas a lesion center (such as the central area of tissue loss in these sea fan lesions) is often obscured by secondary processes such as necrosis and opportunistic infection. Surprisingly, micro-organisms were not consistently present at the lesion interface where earliest lesion features are observed, suggesting they do not instigate the lesion and that infection is a secondary change (i.e. opportunistic infection). Unexplained polyp loss or gastrodermal necrosis, instances of surface-associated metazoans, ciliate infestations, an unidentified protist, and intranuclear inclusion bodies were all observed at this lesion interface, raising the possibility that a number of primary injurious processes can precede fungal or other infections of the axis. Moreover, the reduced proportion of reproductively active biopsies from control areas of lesioned sea fans compared to healthy sea fans, suggests affected sea fan colonies may be debilitated, possibly predisposing to various infections. Reproduction was similarly suppressed in fungus-infected sea fans in the Florida Keys, but this was attributed to systemic suppressive effects of the fungal infection on reproduction. 23
Previous studies induced tissue loss in small sea fan colonies maintained in aquaria by direct inoculation with A. sydowii. 9 However, the initial health status of those sea fans was not histologically verified prior to inoculation, nor was an identical disease state histologically verified post-inoculation. It is possible that opportunistic fungal infection or many other mechanisms of tissue loss were experimentally induced in that study, which would explain inconsistent results in reproducing sea fan aspergillosis.36,40 Our study demonstrates that these lesions are more complex than initially reported and questions the pathogenicity of the diverse range of associated fungi. Experimental infection trials substantiating the role of fungi in these lesions will require a similar multi-modal approach.
In conclusion, annular purple pigmentation in sea fans reflects multiple infections, predominantly algae and hyaline fungi, among others. There is a growing body of literature that contests the fulfillment of Koch’s postulates by A. sydowii. Longitudinal or experimental studies are required in order to disentangle the etiologies and sequence of pathological events currently described as aspergillosis in Caribbean sea fans. A combined diagnostic approach employing multiple techniques will likely yield the best results. Accepted limitations to fungal identification using histology and the poor sporulation of marine fungi in culture suggest that increasingly efficient and affordable molecular techniques can play an important role in supporting traditional diagnostic methods.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231173355 – Supplemental material for A combined diagnostic approach for the investigation of lesions resembling aspergillosis in Caribbean sea fans (Gorgonia spp.)
Supplemental material, sj-pdf-1-vet-10.1177_03009858231173355 for A combined diagnostic approach for the investigation of lesions resembling aspergillosis in Caribbean sea fans (Gorgonia spp.) by Anne A. M.J. Becker, Mark. A. Freeman and Michelle M. Dennis in Veterinary Pathology
Footnotes
Acknowledgements
The authors thank Gina Zeitlin, Nicole Atherley, Veronica Chavis, Kathleen Gilbert, Alexandra Gutierrez, Anna Pleto, David Papanu, Louis-Pierre Rich, Abigail Michelson and Victoria Vaughan for assistance with coral sampling or sample processing, and David Hilchie for performing histological processing.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Center for Conservation Medicine and Ecosystem Health, Ross University School of Veterinary Medicine, St. Kitts, West Indies.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
