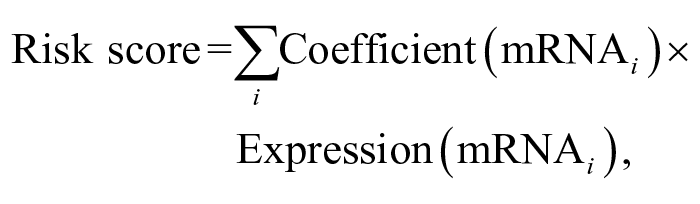Abstract
Canine diffuse large B-cell lymphoma (cDLBCL) is characterized by high mortality and clinical heterogeneity. Although chemo-immunotherapy improves outcome, treatment response remains mainly unpredictable. To identify a set of immune-related genes aberrantly regulated and impacting the prognosis, we explored the immune landscape of cDLBCL by NanoString. The immune gene expression profile of 48 fully clinically characterized cDLBCLs treated with chemo-immunotherapy was analyzed with the NanoString nCounter Canine IO Panel using RNA extracted from tumor tissue paraffin blocks. A Cox proportional-hazards model was used to design a prognostic gene signature. The Cox model identified a 6-gene signature (IL2RB, BCL6, TXK, C2, CDKN2B, ITK) strongly associated with lymphoma-specific survival, from which a risk score was calculated. Dogs were assigned to high-risk or low-risk groups according to the median score. Thirty-nine genes were differentially expressed between the 2 groups. Gene set analysis highlighted an upregulation of genes involved in complement activation, cytotoxicity, and antigen processing in low-risk dogs compared with high-risk dogs, whereas genes associated with cell cycle were downregulated in dogs with a lower risk. In line with these results, cell type profiling suggested the abundance of natural killer and CD8+ cells in low-risk dogs compared with high-risk dogs. Furthermore, the prognostic power of the risk score was validated in an independent cohort of cDLBCL. In conclusion, the 6-gene-derived risk score represents a robust biomarker in predicting the prognosis in cDLBCL. Moreover, our results suggest that enhanced tumor antigen recognition and cytotoxic activity are crucial in achieving a more effective response to chemo-immunotherapy.
Keywords
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most frequent lymphoma histotype in dogs, accounting for 50% to 60% of lymphoid neoplasms. Dogs typically present with systemic lymphadenomegaly that progresses to extranodal disease, primarily involving the peripheral blood (PB) and bone marrow (BM). 3 A definitive diagnosis requires lymphadenectomy, followed by morphological evaluation and immunophenotyping. Histologically, lymph node (LN) architecture is diffusely effaced by sheets of centroblasts and immunoblasts expressing B-cell markers (CD20, CD79, and PAX5) by immunohistochemistry (IHC). 3 Transcriptome studies demonstrated that canine DLBCL (cDLBCL) is a valid model for the human counterpart. In particular, the activated B-cell subtypes in dogs and human share similar pathways, such as nuclear factor kappa B (NF-kB) and B-cell receptor pathways.2,23,26 More recently, sequencing studies have identified the poor prognostic impact of TP53 mutations and reported recurrent mutations largely affecting the same signaling pathways and cellular processes in dogs as in human DLBCL, although the most recurrently mutated genes differ between the 2 species. 12
Despite advancements in cDLBCL treatment, curative therapy remains an unmet medical need. The introduction of an immunotherapeutic strategy (APAVAC) in addition to a CHOP-based chemotherapeutic protocol (vincristine, cyclophosphamide, doxorubicin, and prednisone) markedly improved the outcome of dogs with DLBCL. 12 However, prognosis remains scarcely predictable, and the available clinico-pathologic features are not always reliable in assessing treatment responses or selecting the most appropriate treatment. 18 Thus, a prognostic classification based on molecular rather than clinical features, which helps identify early cDLBCL cases that are potentially responsive to immunotherapy, is needed.
In a previous RNA-sequencing (RNA-Seq) study, dysregulation of immune-related genes, including genes involved in T-cell and macrophage regulation and immune response inhibition, was demonstrated in dogs with a poorer outcome; however, treatment was not standardized. 4 Similarly, the expression of the immune checkpoint–associated genes PD-1 and PD-L1 assessed by RNAscope was negatively associated with the outcome of dogs treated with chemo-immunotherapy.4,5 Taken together, these findings suggest a role of the immune landscape in cDLBCL, potentially driving immune surveillance and tumor defense mechanisms, and provide evidences that a broader understanding of the molecular basis beyond clinical response in dogs receiving immunotherapy is needed.
Here, we studied genes and pathways underlying the cancer-immune interaction in fully staged dogs with DLBCL using the NanoString nCounter Canine IO Panel. This technology allows the simultaneous analysis of hundreds of expressed immune-related genes using RNA extracted from formalin-fixed paraffin-embedded (FFPE) tissues.
Materials and Methods
Recruitment of Dogs
Dogs with DLBCL of any World Health Organization (WHO) stage were prospectively enrolled at the Department of Veterinary Medical Sciences, University of Bologna. Dog owners were required to give written informed consent. This study was approved by the Ministero dell’Istruzione, dell’Università e della Ricerca Ethical Board (number RBSI14EDX9).
All dogs underwent a complete staging work-up, consisting of a history, physical examination, hematology, serum biochemistry (including serum lactate dehydrogenase [LDH]), thoracic radiographs, abdominal ultrasound, and cytological evaluation of a fine-needle aspirate of an enlarged peripheral LN, PB smear, and a BM aspirate. Flow cytometric (FC) immunophenotyping was performed on LN aspirates, PB, and BM aspirates. 11 The extent of PB and BM infiltration by large B cells was evaluated by FC and reported as the percentage of large (ie, forward scatter [FSC] > 400) CD21-positive cells out of the total CD45-positive cells (leukocytes and their precursors). 20
Finally, lymphadenectomy was performed on an enlarged peripheral LN, and histopathology was undertaken as a part of the initial staging. Histotype was confirmed by histological examination and IHC (CD3 and CD20).6,31
The following clinico-pathologic data were recorded: signalment, clinical stage, substage, FC assessment of LN, level of PB and BM infiltration, serum LDH level, pretreatment with steroids, and cause of death.12,18,20 -22 Finally, for each dog, the TP53 mutational status was assessed by whole-exome sequencing or Sanger sequencing, as previously reported. 12 Dogs received chemo-immunotherapy consisting of a CHOP-based protocol plus the autologous vaccine APAVAC. 18 Follow-up evaluation consisted of physical examination, including cytologic evaluation of peripheral LNs and imaging, if clinically indicated, on a monthly basis during the first year and every 3 months thereafter.
Response to treatment was classified as complete remission (CR), partial remission (PR), stable disease (SD), or progressive disease (PD) according to criteria published by the Veterinary Cooperative Oncology Group consensus statement. 30 Response was evaluated at each therapeutic session and was required to last for at least 28 days.
Dogs suffering from relapsed disease were offered CHOP-based chemotherapy without immunotherapy. Relapse was defined as the clinical reappearance and cytologic evidence of lymphoma with or without FC confirmation in any anatomical site in dogs having experienced CR. Time to progression (TTP) was calculated as the interval between initiation of treatment and relapse, whereas lymphoma-specific survival (LSS) was measured as the interval between initiation of treatment and lymphoma-related death. 30
RNA Extraction and NanoString nCounter Analysis
Before RNA extraction from FFPE LN tissues, the quality of tissue preservation and the adequacy of tumor representation were assessed by 3 pathologists (L.L., L.Mi., and L.A.) reviewing the histologic slides. 8 RNA was extracted from two 20-μm-thick FFPE tissue sections using RNeasy FFPE kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions. RNA concentration was quantified using Nanodrop 2000 Spectrophotometer (ThermoFisher Scientific, Waltham, Massachusetts), and DV200 values were obtained using Bioanalyzer 2100 (Agilent Technologies, Santa Clara, California). To be suitable for the following analysis, samples were required to have DV200 >30%. After this procedure, 3 DLBCLs were excluded (DV200 <30%), limiting the experiment to 48 tumors. NanoString nCounter (NanoString Technologies, Seattle, Washington) was used to quantify the expression of the immuno-oncology genes. Briefly, 300 ng of RNA was directly hybridized to reporter and capture probes from the nCounter Canine IO panel. This panel includes 780 genes involved in cytokines, chemokines, interferon and checkpoint signaling, complement cascade, immune cell abundance, tumor immunogenicity, inhibitory tumor mechanisms, stromal factors, and 20 internal reference genes. After hybridization, samples were automatically prepared on the nCounter Prep Station and the captured transcripts were immobilized on the cartridge. The cartridge was scanned using the nCounter Digital Analyzer, and barcodes of the reporter probes were counted.
Data Analysis
Normalization of raw data, quality control (QC), and functional analysis were performed using the nSolver Software v4.0 along with NanoString Advanced Analysis Module v2.0 plugin, following the manufacturer’s guidelines. Registered counts passing the QC parameters were used for further analysis, and the normalized data were scaled and transformed to log2. Functional analyses were conducted, including differential gene expression, gene set analysis (GSA), and cell type profiling. Genes were considered significantly differentially expressed when the false discovery rate (Benjamini-Yekutieli correction) was <0.05.
Gene Signature Score Calculation
The association between LSS and gene expression was evaluated by univariable Cox proportional-hazards analysis. Genes whose expression significantly correlated with LSS (P < .001) were considered in the multivariable analysis. For each dog, a final risk score was calculated as the sum of the gene expression levels significantly associated with LSS from multivariable analysis (P < 0.05), multiplied by the corresponding regression coefficients, as follows: 10
where i is the selected gene from the gene signature.
Statistical and Survival Analyses
All the analyses were conducted in R environment (R Software v.4.1.2). The association between clinico-pathologic features and the risk score was investigated. Normality data distribution was assessed by the Shapiro-Wilk test. Associations were tested by Fisher’s exact test (categorical variables), Student’s t-test, or Wilcoxon rank-sum test (continuous variables, depending on the Shapiro-Wilk test results). Bonferroni correction was applied to take into account multiple testing. Survival analysis was conducted using the univariable Cox proportional-hazards model (survival and survminer packages) to test the impact of the following clinico-pathologic features on TTP and LSS: breed (pure vs mixed), sex (female vs male), age (<7 years vs ≥7 years), weight (<25 kg vs ≥25 kg), stage (III, IV, V), substage (a vs b), PB infiltration (%), BM infiltration (%), presence of BM infiltration (yes vs no), serum LDH levels (normal vs increased), pretreatment with steroids (yes vs no), TP53 mutational status (mutated vs wild type), and risk score (high vs low). A cutoff of P < .05 was used to screen the variables for multivariable analysis. Kaplan-Meier curves for categorical variables were drawn and compared by means of log-rank test (P < .05).
Six-Gene Signature Validation Cohort
RNA-Seq data of 50 dogs with DLBCL were available from a previous study and were used to validate the prognostic value of the risk score. 4 The gene expression measured as log counts per million (logCPM) reads was retrieved, and the risk score was calculated as previously described.
Results
Study Population and Clinical Characteristics
Complete patient demographics and clinical presentation of the 48 dogs with DLBCL included in the study are described in Supplemental Table S1. The most common breeds were mixed-breed dogs (n = 16; 33.3%), Golden Retriever (n = 6; 12.5%), German Shepherd Dog (n = 3; 6.3%) and Dobermann (n = 3; 6.3%). The population consisted of half females (n=24; 50%) and half males (n = 24; 50%). The median age was 7 years (range, 3-13 years) and the median weight was 25 kg (range, 5–43.3 kg). Before enrollment, 15 (31.3%) dogs received steroids. Thirty-one (64.7%) dogs had stage V disease, 16 (33.3%) had stage IV, and only 1 (2%) had stage III disease. Thirty-three (68.7%) dogs were asymptomatic at presentation (substage a), while 15 (31.3%) showed symptoms (substage b). Bone marrow was infiltrated in 15 (31.3%) dogs, with a median of 1.9% neoplastic cells (range, 0.6%–51.6%), while 16 (33.3%) dogs had PB infiltration with a median of 1.4% neoplastic cells (range, 0.1%–55.5%). At presentation, LDH was increased in 26 (54.2%) dogs. TP53 was mutated in 14 (29.2%) dogs and wild type in 34 (70.8%).
Treatment and Outcome
All dogs received chemo-immunotherapy: 39 (81.3%) dogs achieved CR, 6 (12.5%) had PR, 2 (4.2%) were stable, and 1 (2.0%) dog progressed. Overall, 45 (93.8%) dogs relapsed; among them, 34 (75.6%) received another round of CHOP chemotherapy. Median TTP and LSS were 190 and 332 days, respectively.
By multivariable Cox proportional-hazards analysis, TP53 mutation was significantly associated with shorter TTP and LSS (median TTP = 60 vs 245 days, P < .001; median LSS = 102 vs 407 days, P < .001) (Supplemental Table S2).
Six Immune-Related Genes Are Associated With Lymphoma-Specific Survival in cDLBCL
The immuno-oncology gene expression profile of the 48 cDLBCLs was explored by NanoString nCounter technology. A total of 615 of 800 genes passed QC and were further considered. To define the immune gene signature according to the outcome, an univariable Cox proportional-hazards analysis of each gene expression level was conducted, and 19 genes were found to be associated with LSS (P < .001) (Supplemental Table S3). These genes were then included in the multivariable analysis, eventually identifying 6 genes (P < .05) that represented the final signature (IL2RB, BCL6, TXK, C2, CDKN2B, ITK) (Table 1).
Multivariable Cox proportional-hazards analysis of the 19 candidate genes selected from univariable analysis (P < .001) associated with the LSS.
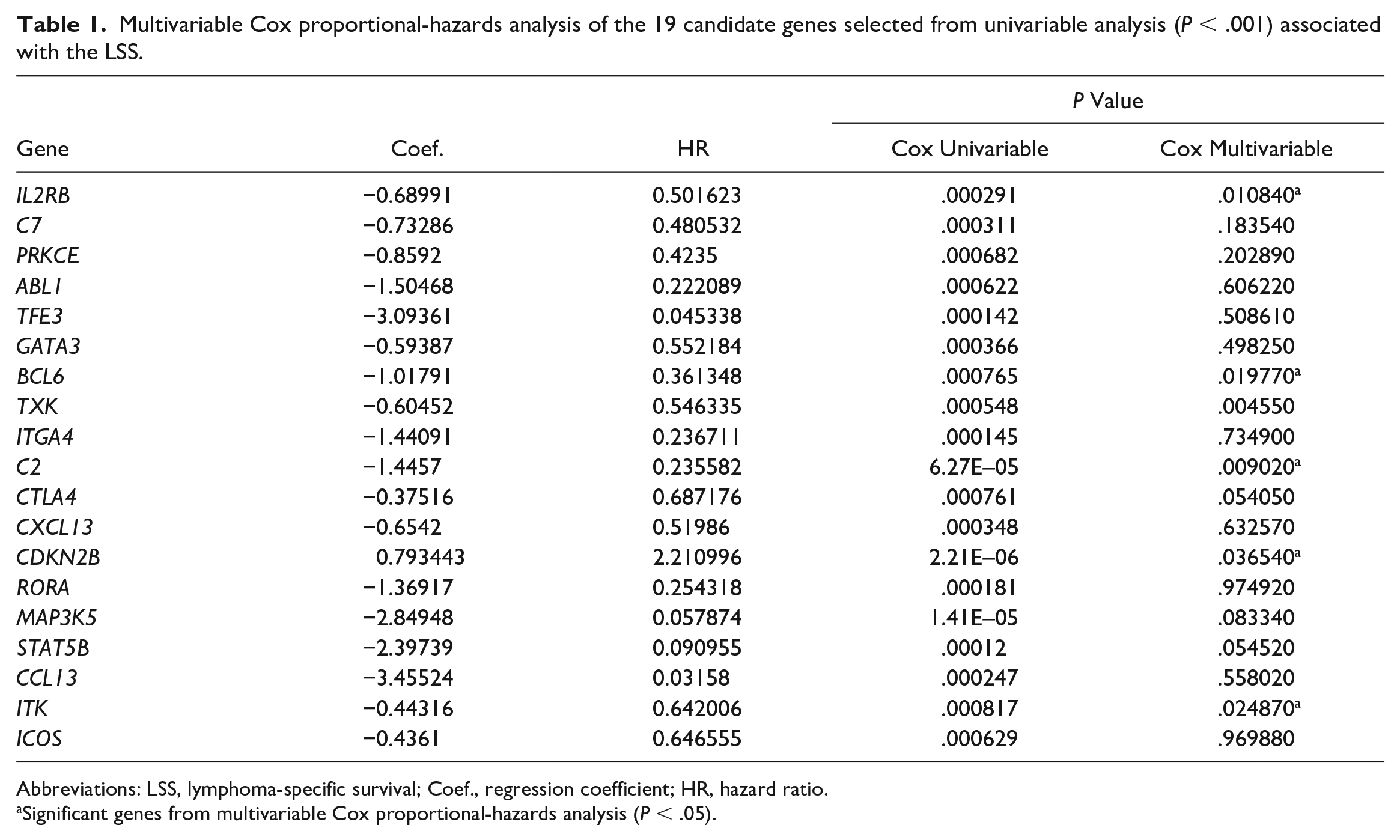
Abbreviations: LSS, lymphoma-specific survival; Coef., regression coefficient; HR, hazard ratio.
Significant genes from multivariable Cox proportional-hazards analysis (P < .05).
A 6-Gene-Derived Risk Score Predicts Outcome and Response to Chemo-Immunotherapy in cDLBCL
A risk score based on the 6 genes was calculated as follows: (−0.6899 × IL2RB expression level) + (−1.0179 × BLC6 expression level) + (−0.6045 × TXK expression level) + (−1.446 × C2 expression level) + (0.7934 × CDKN2B expression level) + (−0.4437 × ITK expression level). According to the median score (−24.9919), dogs were assigned to high-risk (score above the median) or low-risk group (score below the median). Survival analysis confirmed the prognostic significance of the 6-gene-derived risk score. Indeed, by multivariable Cox proportional-hazard analysis, dogs with high-risk score had significantly shorter TTP (median TTP = 64.5 vs 286.5 days, P = .002) and LSS (median LSS = 142 vs 672 days, P < .001) compared with dogs with low-risk score (Fig. 1; Supplemental Table S2). The risk score was also significantly associated with substage, with high-risk group having a higher percentage of dogs presenting symptoms (substage b) compared with the low-risk group (P = .001).
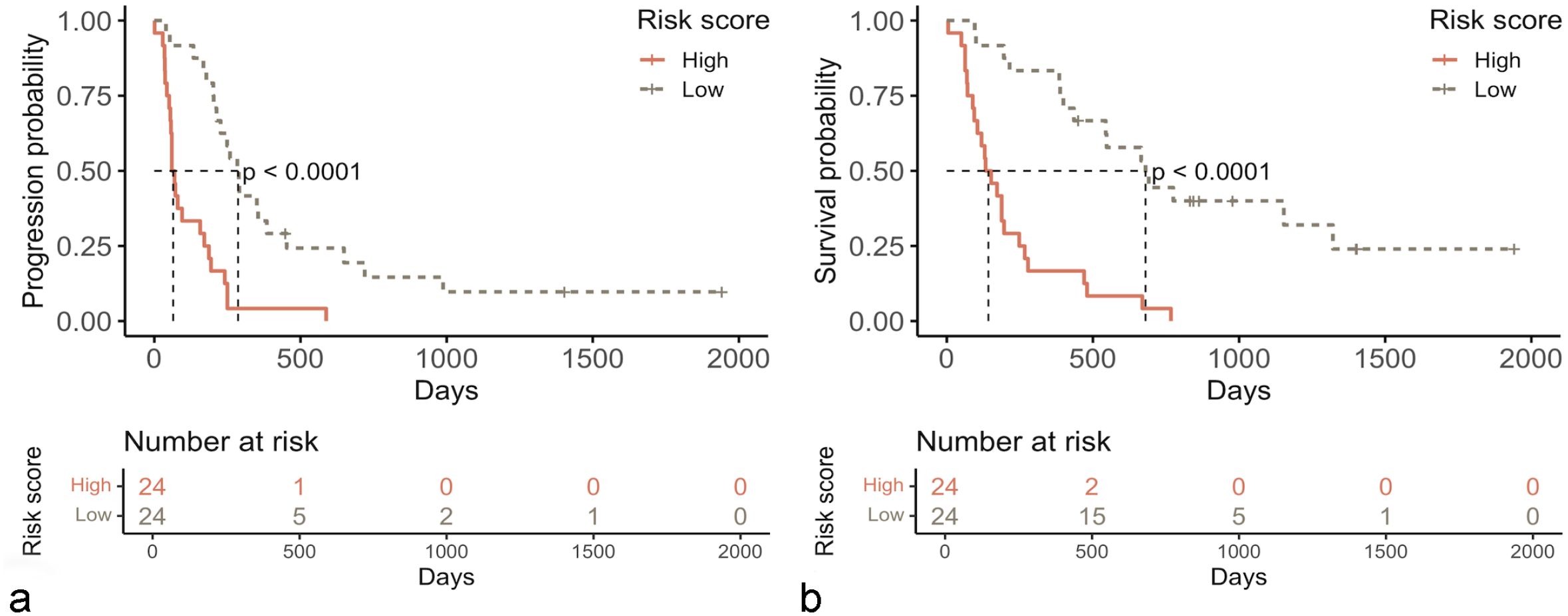
Kaplan-Meier curves for 48 dogs with diffuse large B-cell lymphoma according to the risk score, showing longer time to tumor progression (a) and lymphoma-specific survival (b) for low-risk dogs (tan dashed line) compared with high-risk dogs (red line) (P < .0001).
Low-Risk Dogs Have an Upregulation of Antigen Processing Signatures
To explore the immune profile of cDLBCL according to the risk score, the expression of immune-related genes was investigated by supervised analysis (low-risk vs high-risk dogs) (Supplemental Fig. S1). Thirty-nine genes were differentially expressed (DE), and in particular, all DE genes were upregulated in low-risk dogs (Fig. 2a; Supplemental Table S4). GSA highlighted an upregulation of genes involved in complement activation (C2), cytotoxicity (DLA-12), T-cell functions (TRBC, TRAC, CD5, MAF, LCK, CTLA4, IL4R, CD8A), and antigen processing (CD8A, DLA-12, B2M) in low-risk compared with high-risk dogs. Conversely, genes associated with the cell cycle (eg, TP63, CDKN2B, BCL2, MKI67, CDKN2C) were downregulated (Fig. 2b; Supplemental Table S5). Moreover, low-risk dogs were characterized by the upregulation of genes defining the signatures of T-cells (P < .001), tumor-infiltrating lymphocytes (P < .001), and cytotoxic lymphocytes (P = .003) (Fig. 2c–e).
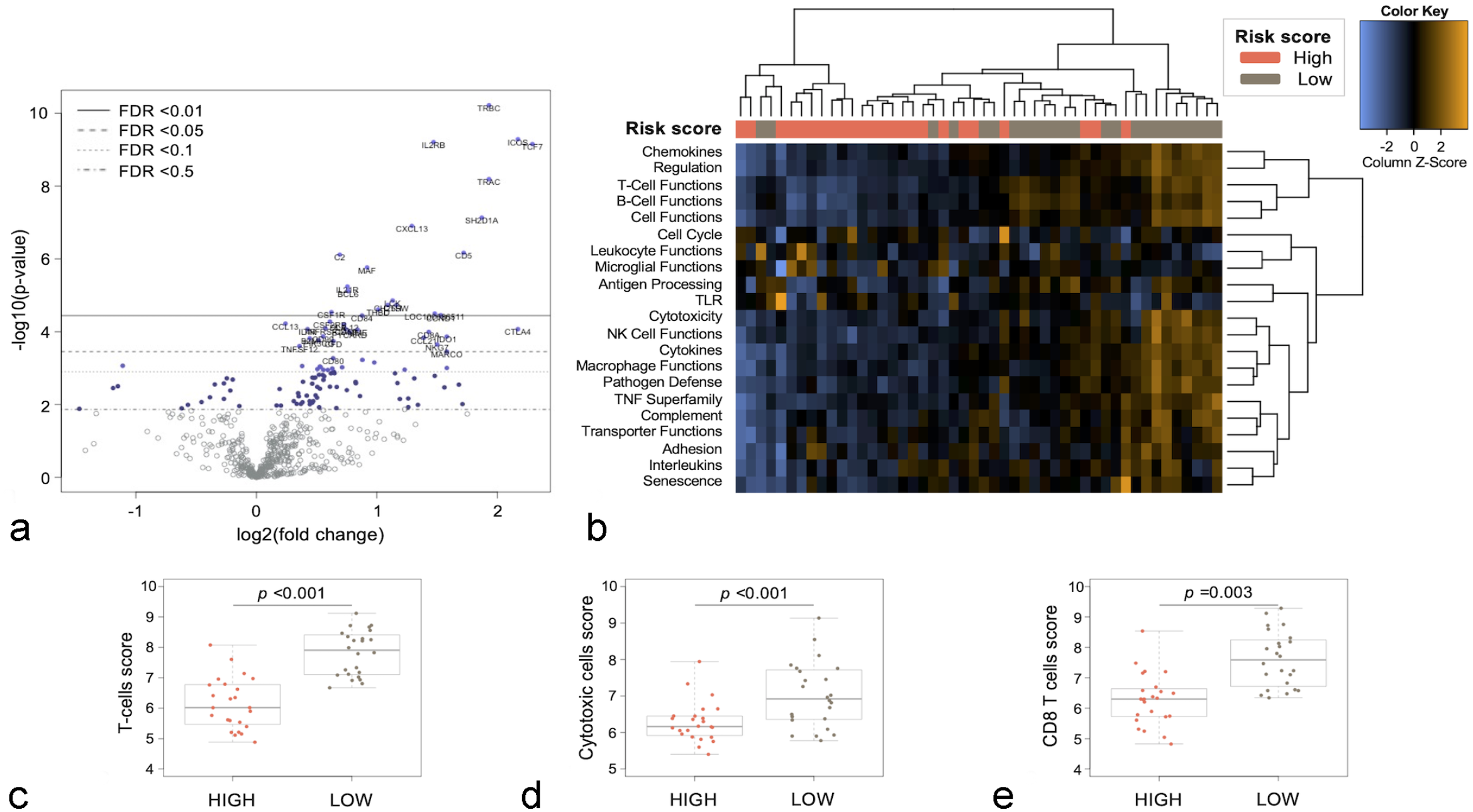
Immune landscape of canine diffuse large B-cell lymphoma according to the risk score. (a) Volcano plot showing differentially expressed genes. Thirty-nine genes were significantly upregulated in low-risk versus high-risk dogs (false discovery rate [FDR] <0.05). (b) Heatmap of pathway scores. Scores were scaled via a Z-transformation; orange indicates high scores; blue indicates low scores. (c–e) Box-plots showing cell-type scores for T-cells (c), cytotoxic cells (d), and CD8 T-cells (e) in low-risk and high-risk dogs. The difference between low-risk and high-risk dogs was assessed by means of Wilcoxon rank-sum test (P < .05).
Validation of the 6-Gene Signature
The prognostic value of the 6-gene-derived risk score was then validated in a second cohort of 50 dogs with DLBCL. The complete clinical data of these dogs are described elsewhere. 4 Gene expression levels of the 6 genes included in the signature, obtained by RNA-Seq as logCPM, were used to calculate the risk score. 4 Dogs were dichotomized according to their risk score into high-risk and low-risk groups. Univariable Cox proportional-hazards analysis confirmed that prognosis was significantly associated with the risk score. Indeed, dogs with a high-risk score had shorter TTP (78 vs 202 days, P = .05) and LSS (150 vs 296 days, P = .047) compared with low-risk dogs (Supplemental Fig. S2).
Discussion
In human medicine, advances in genomic analysis have led to the identification of the tumor microenvironment as a critical cancer hallmark. In this study, we characterized the immune landscape of 48 dogs with DLBCL treated with immunotherapy and identified a novel gene signature predicting the survival.
Active immunotherapy has emerged as one among the most promising treatment options for canine B-cell lymphoma. 18 However, an outstanding response only occurs in a subset of dogs, reflecting the need for an improved understanding of molecular mechanisms beyond tumor resistance to properly select the most effective treatment and predict clinical response. Recent genetic and transcriptome studies have shed light on the heterogeneity of cDLBCL, uncovering subsets of tumors which show unfavorable prognosis. Few genetic markers were also correlated to LSS and TTP, opening new scenarios for genetic analysis in the clinical setting. Conversely, the immune landscape of cDLBCL is scarcely known. To date, the impact of T-cell microenvironment on cDLBCL pathogenesis and clinical outcome was specifically investigated by RNAscope, and the expression of RNA encoding 2 well-known immune checkpoint molecules, PD-1 and PD-L1, was associated with shorter survival time in dogs treated with chemo-immunotherapy.4,5 Also, the likelihood of relapse was significantly increased in dogs with DLBCL characterized by a high tumor CD4+/CD8+ ratio based on FC. 22
Here, we investigated the immune landscape of cDLBCL by NanoString nCounter Technology to clarify the interaction mechanisms between tumor and immune system driving response to chemo-immunotherapy and to identify potential predictive biomarkers. A robust performance of the NanoString Canine IO panel using FFPE cDLBCL biopsy samples was obtained overall. Sufficient RNA yield and gene expression quality were largely gained in all runs, and none of the samples with a DV200 >30% was excluded. More than 78% of the panel genes passed QC and were used for the downstream analyses.
Deregulation in genes clustered in interferon, cytokine, and chemokine signaling and lymphoid compartments was identified. The main aim of the study was to identify a subset of genes whose expression may contribute to predict outcome. To this end, we processed the data in a stepwise approach and obtained 19 genes associated with LSS by univariable analysis. Then, we considered only genes that remained significant at multivariable analysis, ultimately leading to the identification of 6 genes. To confirm the clinical relevance of the 6-gene signature, a second cohort of dogs with DLBCL was tested, confirming its utility in discriminating between dogs with better and poorer outcome.
The 6-gene signature was largely defined by genes controlling the adaptive immune response, including TXK, ITK, IL2RB, and C2. Both TXK and ITK encode for members of the Tec family of tyrosine kinases and regulate T-cell receptor–mediated T-cell activation and T-helper cell differentiation. 29 Downregulation or the absence of ITK and TXK alters the development of CD4+ and CD8+ T-cells, reduces T-cell differentiation toward the Th2 lineage, and decreases interleukin-2 (IL-2) production. 29 Dogs having a lower gene expression level of both TEC kinases exhibited a more aggressive phenotype with shorter survival. Our hypothesis is that ITK and TXK overexpression may favor the activation of tumor-specific cytotoxic T-cells to attack cancer cells by boosting the presentation of tumor-associated antigens after APAVAC administration. IL2RB encodes 1 of the 3 subunits of IL-2 receptor and is expressed on T-cells and natural killer cells. IL-2/IL-2 receptor signaling conveys essential signals for immune cells, including T-cells and natural killer cells. Therefore, IL2 receptor has been recently considered as a promising drug target to activate and empower immune therapy in several human cancers. 14 In head and neck cancers, overexpression of IL2RB has been associated with longer survival, and the intratumoral addition of recombinant IL-2 increased therapeutic antitumor activity and the magnitude of the antitumor immune response, and enhanced the avidity of antigen-specific CD8+ T-cells.13,33 Importantly, toxicities were not recorded. Based on our findings, a similar approach might represent a promising therapeutic strategy in dogs with DLBCL before APAVAC administration. Furthermore, recent studies demonstrated that IL2RB is positively linked to the abundance of tumor-infiltrating lymphocytes and neoantigens in several tumors. 14 Indirectly, this biological mechanism might also occur in dogs with DLBCL, as increasing numbers of CD3+ T-cells were associated with better outcome in cDLBCL. 4
Finally, C2 encodes for complement component 2, which is part of the classical pathway of the complement system and whose activation is known to be detrimental to cancer cells. 1 Indeed, transcriptomic data from several human cancers revealed that complement activation has antitumor activity, even if most of these studies were focused on complement system components other than C2. 27 In one study, higher C2 expression was associated with better prognosis in patients with hepatocellular carcinoma, suggesting an influence of C2 on intratumoral CD4+ T-cell proliferation and macrophage M0 cells, supporting our findings. 24 Although this association was not demonstrated here, it deserves further investigation.
The other 2 dysregulated genes in the prognostic signature were BCL6 and CDKN2B. BCL6 is widely recognized in human DLBCL pathogenesis and acts as a transcriptional repressor of the BTB/Zinc finger family of transcription factors. It negatively regulates several cell programs, such as cell-cycle arrest, response to DNA damage, terminal differentiation, and apoptosis. 25 Chromosomal translocations and somatic mutations occurring in the 5′ noncoding region of the BCL6 gene are the most frequent genetic abnormalities associated with human B-cell lymphomas, leading to a constitutive expression of this gene.15,32 So far, similar translocations have not been identified in dogs with DLBCL, follicular lymphoma, or marginal zone lymphoma. 7 However, the positive correlation between mRNA levels and survival obtained here mirrors the findings in human DLBCL.16,17 Finally, CDKN2B encodes for the cyclin-dependent kinase inhibitor p15 and its expression negatively correlated with LSS, reflecting the abundance of various transcripts related to cell cycle in dogs with poor outcome.
To gain further information from the 6-gene signature, a risk score was calculated, clustering dogs into low-risk and high-risk. Different survival and treatment responses were obtained when comparing these groups, confirming the biological relevance of the 6-gene panel. Also, a total of 39 genes were found to be significantly upregulated in dogs showing a better outcome compared with those having a worse prognosis. Interestingly, low-risk dogs expressed genes associated with increased antigen processing machinery and cytotoxic activity, and were characterized by an enrichment of genes involved in T-cells, cytotoxic cells, and tumor-infiltrating lymphocyte signatures, suggesting an abundance of these cytotypes within the tumor itself.
Recently, several immunotherapeutic approaches have been developed to improve the release and presentation of tumor-associated antigens to the immune system. APAVAC is an autologous vaccine consisting of hydroxyapatite ceramic powder and heat shock proteins purified from the tumor itself. 19 Heat shock proteins act as tumor-associated antigens carriers, thereby facilitating antigen presentation on major histocompatibility complex class I and activating CD8+ cytotoxic T-cell response, which results in an antitumor immune response.9,28 Based on the current findings, it is suggested that the presence of cytotoxic T-cells within the tumor may be associated with a better response to immunotherapy. 22
Unlike previous results, PD-1 and PD-L1 mRNA expression was not correlated to outcome by multiple comparison analysis.4,5 This discrepancy may be due to the different enrollment criteria in previous studies, including lack of treatment standardization and different laboratory techniques, such as RNA-Seq and RNAscope.
In conclusion, the measurement of transcripts and pathways related to the cancer-immune interaction can predict the response to chemo-immunotherapy in dogs with DLBCL. In addition, the NanoString nCounter technology can be successfully used to analyze RNA extracted from FFPE samples also in veterinary oncology. Future studies are in the pipeline to further validate the current findings by incorporating the 6-gene signature assay in the clinical setting and replicate the experiment in dogs treated with CHOP only to compare the cDLBCL immune landscape with the clinical response to chemotherapy alone.
Supplemental Material
sj-pdf-6-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-pdf-6-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Supplemental Material
sj-xlsx-1-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-xlsx-1-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Supplemental Material
sj-xlsx-2-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-xlsx-2-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Supplemental Material
sj-xlsx-3-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-xlsx-3-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Supplemental Material
sj-xlsx-4-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-xlsx-4-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Supplemental Material
sj-xlsx-5-vet-10.1177_03009858231162209 – Supplemental material for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature
Supplemental material, sj-xlsx-5-vet-10.1177_03009858231162209 for Predicting outcome in dogs with diffuse large B-cell lymphoma with a novel immune landscape signature by Luca Licenziato, Lucia Minoli, Ugo Ala, Laura Marconato, Antonella Fanelli, Diana Giannuzzi, Raffaella De Maria, Selina Iussich, Giulia Orlando, Francesco Bertoni and Luca Aresu in Veterinary Pathology
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the MyLav laboratory (Milan, Italy).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.