Abstract
The involvement of Felis catus papillomavirus type 2 (FcaPV2) in feline Merkel cell carcinoma (MCC) has been previously hypothesized. In this study, the expression and localization of FcaPV2 oncogene mRNA, the integration of FcaPV2 genes, and p53 mutations in feline MCC were examined by RNAscope in situ hybridization (ISH), whole genome sequencing (WGS), and Sanger DNA sequencing, respectively. Furthermore, the morphological and molecular characteristics of FcaPV2-positive (FMX-MCC01) and FcaPV2-negative (AS-MCC01) MCC cell lines were compared in vitro and in vivo using immunofluorescence, ISH, xenotransplantation into mice, and immunohistochemistry. ISH for FcaPV2 E6/E7 detected viral RNA in 18/21 FcaPV2-positive MCC and not in 1/1 FcaPV2-negative MCC. WGS of 2 FcaPV2-positive cases revealed the integration of FcaPV2 genes in both cases. In cultured cells and xenograft tissues of FMX-MCC01, most cells were positive for E6/E7 by ISH and p16CDKN2A, a few cells were positive for the retinoblastoma protein (pRb), and all cells were negative for p53. In cultured cells and xenograft tissues of AS-MCC01, all cells were negative for p16CDKN2A, most cells were positive for pRb, and some cells were positive for p53. Missense mutations in p53 were identified in 8/10 FcaPV2-positive and 1/1 FcaPV2-negative MCC. These results suggest that the expression of integrated FcaPV2 oncogenes might be associated with reduced expression of the tumor suppressor proteins pRb and p53 and might contribute to the development of feline MCC. On the other hand, p53 mutations may be involved in both FcaPV2-positive and FcaPV2-negative MCC tumorigenesis.
Keywords
Merkel cell carcinoma (MCC) is an uncommon and extremely aggressive neuroendocrine carcinoma of the skin.7,10 In humans, 2 distinct etiologies are involved in the pathogenesis of MCC: (1) Merkel cell polyomavirus (MCPyV) infection and (2) ultraviolet (UV) exposure. 10 Approximately 80% of human cases of MCC are caused by the integration of MCPyV DNA into the host genome, which leads to the persistent expression of viral oncogenes. 16 Conversely, MCPyV-negative MCC is triggered by high-frequency DNA mutations due to extensive UV exposure. 48
Papillomavirus (PV) has a double-stranded circular DNA genome and causes epithelial tumors in different species.12,36 PV is highly species-specific and often shows tropism for certain types of epithelium. To date, 6 types of Felis catus papillomavirus (FcaPV) have been identified in various cutaneous lesions of cats.37,46 Among these, FcaPV2 has been most frequently detected in viral plaques, Bowenoid in situ carcinoma (BISC), squamous cell carcinoma (SCC), and basal cell carcinoma (BCC).26,37,46 These are recognized as FcaPV-associated cutaneous lesions in cats. 37 Recent studies suggested that a molecular mechanism similar to that in high-risk human papillomavirus (HPV)-associated tumors contributes to the development of FcaPV-associated tumors.2,3,12,35,37 In HPV-induced tumorigenesis, after viral genes are integrated into the host genome, the viral oncoproteins E7 and E6 inhibit the host retinoblastoma (pRb) and p53 protein, respectively, resulting in increased cell proliferation and survival. 23 Moreover, the inactivation of pRb causes aberrant increases in the tumor suppressor protein p16CDKN2A (p16), which are the features of PV-induced tumors. 12 However, FcaPV2 infections are ubiquitous and asymptomatic in most of cats. 37 To distinguish between PV-induced or asymptomatic infection, immunohistochemistry (IHC) has been used to identify the molecular changes caused by PV described above. 35
Feline MCCs were first reported in 2003 with only around 25 of these rare neoplasms subsequently reported.5,43 Histologically, feline MCCs are located in the dermis and/or subcutis without showing continuity with the overlying epidermis.25,43 The tumor cells express epithelial markers such as cytokeratin (CK) 20 and CK18 and neuroendocrine markers such as synaptophysin and CD56.25,43 These findings are characteristic of MCC among cutaneous tumors and help distinguish it from SCC and BCC. We recently demonstrated the possible involvement of FcaPV2 in the tumorigenesis of feline MCC. 26 FcaPV2 DNA has been detected in 95% of feline MCC cases, and the concurrent FcaPV2-associated lesions, such as viral plaques, BISC, SCC, and BCC, have been observed in 62% of feline MCC cases. 26 To verify the involvement of FcaPV2 as a pathogen in feline MCC, we have immunohistochemically identified molecular changes caused by FcaPV2 in the tumor cells. 26 FcaPV2 DNA-positive MCC showed high immunoreactivity for p16 and low immunoreactivity for pRb and p53. In addition, DNA in situ hybridization (ISH) confirmed the presence of the FcaPV2 E7 DNA within the tumor cells of FcaPV2 DNA-positive MCC, 26 and its hybridization signal pattern implied the integration of FcaPV2 genes in tumor cells. 26 These results suggest that FcaPV2 may be an etiological agent of MCC in cats.
In the present study, to further understand the role of FcaPV2 in feline MCC, the integration of FcaPV2 DNA into the host genome and the expression of FcaPV2 E6 and E7 mRNA were investigated using tissue samples of feline MCC cases. Viral genomic integration is a critical step in PV-induced tumorigenesis, 12 and the active transcription of FcaPV2 oncogenes in MCC tumor cells would support the immunohistochemical results shown in the previous study. In vitro and in vivo analyses of cell lines derived from FcaPV2 DNA-positive MCC and FcaPV2 DNA-negative MCC were also performed. Furthermore, mutations in the p53 gene were evaluated.
Materials and Methods
Tumor Samples
Biopsy samples from 22 cats that were pathologically diagnosed with MCC were examined (Supplemental Table S1). Samples were from the archives of the Laboratory of Veterinary Pathology, the University of Tokyo, and Diagnostic Laboratory, Patho-Labo, Shizuoka, Japan, and were submitted between 2008 and 2021. Twenty-one cases were included in our previous studies.25,26 The FcaPV2 status of all cases was historically confirmed by polymerase chain reaction (PCR) in our previous study and are summarized in Supplemental Table S1.26,49 Concurrent FcaPV2-associated lesions were found in 14 cases (Supplemental Table S1). As for their locations, 11/12 viral plaques and 3/3 BISC were adjacent to MCC lesions, and 1/12 viral plaque, 1/1 SCC, and 1/1 BCC were distant from the MCC lesions.
RNAscope ISH
All tissues were routinely fixed in 10% phosphate-buffered formalin solution, embedded in paraffin, and sectioned at a thickness of 4 μm. Colorimetric ISH was performed using RNAscope 2.5 HD Reagent Kit-Red (Advanced Cell Diagnostics) according to the manufacturer’s instructions. An ISH probe (V-FPV-E6-E7) that targets the sequence between nucleotides 461 and 1154 (E6 and E7 genes) of FcaPV2 (GenBank EU796884.1) was applied in the hybridization step. This probe was designed and successfully detected FcaPV2 E6 and E7 mRNA in a previous study. 22 Probes for the feline gene peptidylprolyl isomerase B and the bacterial gene dihydrodipicolinate reductase served as positive and negative controls, respectively. ISH with an RNase pretreatment was performed on 3 cases (cases 9, 10, and 11) to investigate the contribution of viral DNA and mRNA to hybridization signals. After RNAscope protease digestion, slides were treated with RNase A solution (4 mg/mL; Promega) at 40°C for 30 minutes. Slides were then washed by distilled water 3 times before the addition of the targeting probe.
Next-Generation Sequencing
Genomic DNA was extracted from 2 freshly frozen tissue samples (cases 9 and 21) using the DNeasy Blood & Tissue Kit (QIAGEN). Genomic DNA was quantified by a Qubit fluorometer (Thermo Fisher Scientific), and its integrity was validated on a 1% agarose gel to confirm that the molecular weight of genomic DNA was larger than 10 kb and not markedly degraded. Sequence libraries were obtained using the TruSeq DNA PCR-Free Library Prep Kit (Illumina) and IDT for Illumina-TruSeq DNA UD Indexes (Illumina) following the manufacturer’s instructions. Genomic DNA was then randomly fragmented by Acoustic Solubilizer Covaris (Covaris). After end polishing and phosphorylation, fragment selection was performed using Agencourt AMPure XP (Beckman Coulter). The end repair of DNA fragments was performed, and an “A” base was added at the 3’-end of each strand. Sequencing libraries were constructed by ligated index adapters. The library preparation was automated using the Agilent XT-Auto system (Agilent Technologies). The quality of the libraries produced was verified by Agilent TapeStation (Agilent Technologies), and 150-bp paired-end sequencing was performed using the NovaSeq 6000 Sequencing System (Illumina). Real-Time Analysis v3.4.4 (Illumina) and bcl2fastq2 Conversion Software v2.20 (Illumina) were applied for base calling and data conversion to the FASTQ format, respectively.
Analysis of Whole Genome Sequencing (WGS) Data
The genome sequences of F. catus (GenBank assembly accession: GCA_000181335.4) and FcaPV2 (GenBank accession numbers: LC612600), which was identified in feline MCC in the previous study, 26 were set as the reference sequence and insertion sequence, respectively. Regions with high homology to the insertion sequence were searched in the reference sequence. Homologous regions were masked as an intrinsic sequence to avoid multi-mapping. Read mapping to the reference sequence and insertion sequence was then performed using the DRAGEN Bio-IT Platform (Illumina). To identify integration regions, discordant reads and split reads were extracted from mapping data using SAMtools and LUMPY.28,30 Discordant reads were defined as having one end of the paired-end read mapped to the insertion sequence and having its mate pair mapped to the reference sequence. Split reads were defined as having one end of the paired-end read spanning the integration junction and having its mate pair mapped to either the reference or insertion sequence. Chromosome structural variations, such as deletions, duplications, translocations, and inversions, were detected using the DRAGEN Bio-IT Platform (Illumina) and LUMPY. Structural variations associated with the integration were extracted, and the translocation sites of the insertion sequence into the reference sequence were identified as breakpoints. Breakpoints that were close to each other (<10 bp) in the reference sequence were defined as one integration region. The locations of the integration regions within annotated genes were then analyzed.
Cell Cultivation
Using fresh tumor tissue from a left shoulder mass in case 21, primary cell cultivation was conducted in the same manner as in our previous study. 25 The cell line was named FMX-MCC01 and characterized as described later. The previously established cell line AS-MCC01, which is derived from feline FcaPV2 DNA-negative MCC (case 22), 25 was also cultured and characterized as a comparison subject for in vitro/in vivo analyses.
Immunofluorescence in Cultured Cells
Using FMX-MCC01 and AS-MCC01 cells, immunofluorescence was conducted in the same manner as in our previous study. 25 The primary antibodies used in this experiment are listed in Table 1. Negative controls were performed by omitting the primary antibodies.
Primary antibodies used for immunofluorescence and immunohistochemistry.
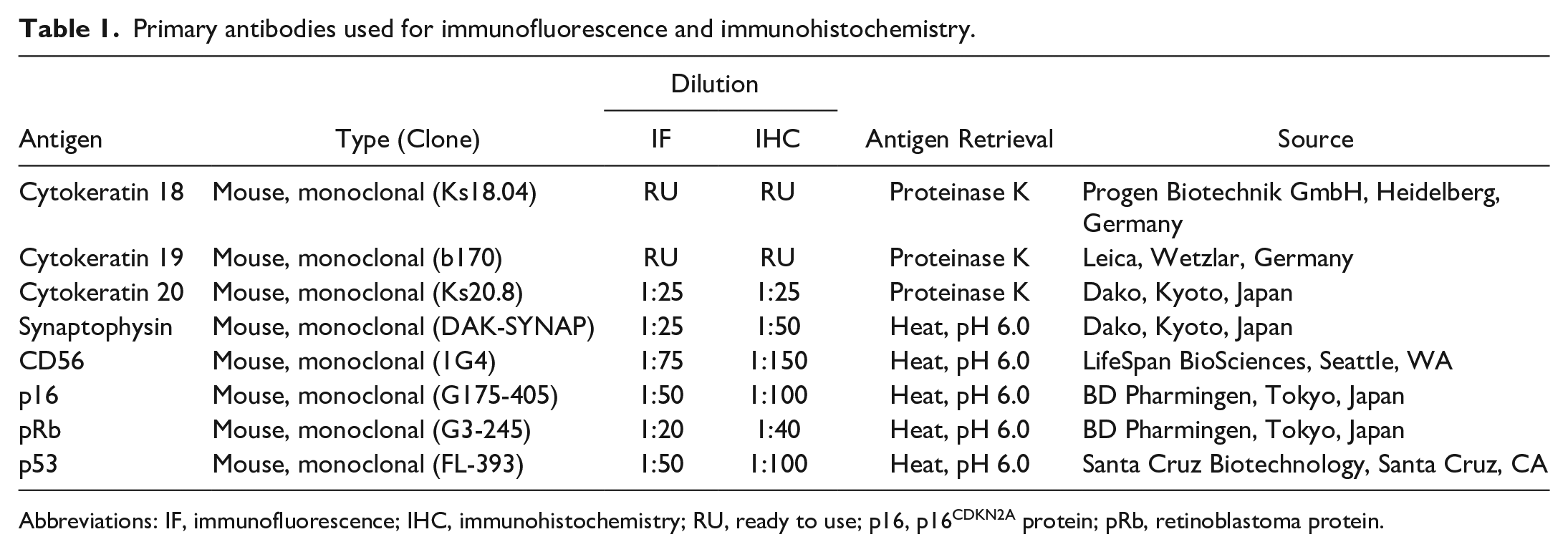
Abbreviations: IF, immunofluorescence; IHC, immunohistochemistry; RU, ready to use; p16, p16CDKN2A protein; pRb, retinoblastoma protein.
ISH in Cultured Cells
DNA fluorescence in situ hybridization (DNA-FISH) was performed for FMX-MCC01 cells. A digoxigenin (DIG)-labeled DNA probe for the FcaPV2 E7 gene designed in the previous study was used. 26 Cells were suspended in Roswell Park Memorial Institute 1640 (RPMI; Invitrogen) medium and centrifuged onto slides using CytoFuge 2 Cytocentrifuge (StatSpin Technologies). Cells were fixed with Carnoy’s solution at room temperature for 10 minutes and incubated at 70°C for 2 hours. After a pretreatment in 0.5 mg/mL pepsin and post-fixation in 1% paraformaldehyde, DNA was denatured in 70% formamide (pH 7.0) at 73°C for 3 minutes. Cells were hybridized with 1 ng/µL of the DIG-labeled DNA probe at 37°C for 16 hours. After washing, cells were further incubated with 20 µg/mL DyLight 488-labeled anti-DIG-digoxin (Vector Laboratories) at 37°C for 1 hour, counterstained with DAPI (Vectashield1; Vector Laboratories), and then observed under the Zeiss LSM 700 laser scanning confocal microscope (Carl Zeiss Meditec).
RNAscope ISH was performed for FMX-MCC01 cells using RNAscope 2.5 HD Reagent Kit-Red (Advanced Cell Diagnostics) and the ISH probe (V-FPV-E6-E7). Cells were suspended in RPMI medium and centrifuged onto slides using the CytoFuge 2 Cytocentrifuge (StatSpin Technologies). After fixation in 4% paraformaldehyde, ISH was conducted according to the manufacturer’s instructions.
Xenotransplantation Into Mice
A FMX-MCC01 cell suspension (2 × 107 cells) was prepared in 200 mL of Hank’s balanced salt solution (Invitrogen) and subcutaneously injected into the lower flank of 3 female severe combined immunodeficient (SCID) mice (FOX CHASE SCID C.B-17/Icr-scid/scidJcl; CLEA Japan). These mice were housed in a cage with free access to water and a standard purified rodent growth diet (AIN-93G, Oriental Yeast). All mice were euthanized with isoflurane 2 months after the injection or when the tumor volume exceeded 1000 mm3. Tumor masses were collected at necropsy, immediately fixed in 10% neutral-buffered formalin, and embedded in paraffin. Specimens were sectioned at a thickness of 4 µm and stained with hematoxylin and eosin. The mitotic count was calculated as the sum of mitoses in 10 consecutive high-power fields (HPF; diameter of the field of view = 0.55 mm; 10 HPF = 2.37 mm2; 40 × magnification; Olympus CX31 microscope, Tokyo, Japan) starting from the most mitotically active areas. 34 Fields with representative tumor cells were assessed. After counting mitoses in the first HPF, if no mitoses were found in the next 3 consecutive HPF, the process was repeated by moving to another mitotically active area. Experiments involving animals were approved by the Animal Care and Use Committee of the Graduate School of Agricultural and Life Sciences, the University of Tokyo (Approval # P20-083M02).
IHC was performed on FMX-MCC01 xenograft tissues and AS-MCC01 xenograft tissues obtained in the previous study. 25 Deparaffinized tissue sections were immersed in 10% hydrogen peroxide (H2O2) in methanol at room temperature for 5 minutes and then incubated in 8% skim milk in Tris-buffered saline (TBS) at 37°C for 40 minutes to avoid non-specific reactions. Sections were incubated at 4°C overnight with the primary antibodies (Table 1). An anti-mouse IgG polymer labeled with horseradish peroxidase (Envision, Agilent Technology) or the Optiview DAB detection kit (Roche Diagnostics) was applied at 37°C for 40 minutes, and sections were then rinsed with TBS. Reactions were visualized with 0.05% 3-3’-diaminobenzidine and 0.03% H2O2 in Tris–hydrochloric acid buffer, followed by a counterstain with Mayer’s hematoxylin (Muto Pure Chemicals). The numbers of xenograft cells positive for p16, pRb, and p53 were manually counted in 5 HPF (×400). The location of immunolabeling as nuclear or cytoplasmic was also assessed. Immunohistochemical positivity for p16 was evaluated based on the percentage positivity and pattern.26,35 Lesions containing more than 20% cells with both nuclear and cytoplasmic positivity for p16 were considered to have high p16 immunoreactivity (+). 28 Lesions containing less than 50% cells with nuclear positivity for pRb were considered to have low pRb immunoreactivity (−). 26 Lesions containing less than 20% cells with nuclear positivity for p53 were considered to have low p53 immunoreactivity (−). 26 Feline MCC tissues characterized in our previous studies and normal cutaneous tissues around the tumor were used as positive and negative controls.25,26
RNAscope ISH was also performed for FMX-MCC01 xenograft tissues using RNAscope 2.5 HD Reagent Kit-Red (Advanced Cell Diagnostics) and the ISH probe (V-FPV-E6-E7) as described above.
Transmission Electron Microscopy
Ultrathin sections were prepared from the targeted area of MCC identified in a hematoxylin and eosin-stained section derived from formalin-fixed paraffin-embedded tissue (case 20) after osmification and embedding in Epon blocks by the inverted beam capsule method.27,44 The section was stained with uranyl acetate and lead citrate and then examined with a transmission electron microscope (JEM-1400Plus; JEOL).
Sequencing Analysis of the p53 Gene
The statuses of p53 mutations in feline skin samples with MCC were assessed by a deep sequence analysis of exons 5 through 8. This region corresponds to the DNA binding domain and is the most common site of p53 mutations in tumor cells.6,39 Four primer pairs were designed to amplify each of the 4 exons (Supplemental Table S2). 39 Genomic DNA was extracted from 11 formalin-fixed paraffin-embedded (FFPE) tissue samples using the QIAamp DNA FFPE Tissue Kit (QIAGEN). Extracted DNA from skin samples with FcaPV2-associated SCC (case 7) and BCC (case 20) and 3 normal skin samples were applied as controls. PCR was performed by preparing the following reaction mixture: 12.5 µL KOD One PCR Master Mix-Blue (TOYOBO), 0.75 µL forward primer (10 µM), and 0.75 µL reverse primer (10 µM). Template DNA was added at 30 ng per PCR reaction followed by distilled water to a total volume of 25 µL. PCR cycling conditions referring to the manufacturer’s instructions were as follows: pre-denaturation at 95°C for 3 minutes followed by 45 cycles of denaturation at 98°C for 10 seconds, annealing at 59°C for 5 seconds, and extension at 68°C for 1 seconds. PCR-amplified products were confirmed by gel electrophoresis as described above, purified using the High Pure PCR product Purification Kit (Roche), and subjected to a sequence analysis (FASMAC). The sequences obtained were compared with the reference sequence (GenBank accession number: NM_001009294.1) using the Basic Local Alignment Search Tool from the National Center for Biotechnology Information (NCBI; https://blast.ncbi.nlm.nih.gov/Blast.cgi) and Unipro UGENE software.4,11,38 Cases with one or more alterations in the nucleotide sequence of the amplified exons leading to amino acid changes were classified as mutated. Furthermore, an in silico analysis using PolyPhen-2 predicted the possible impact of nucleotide alterations on the function and structure of the protein. 1 The predicted function of the variant was characterized as benign (score 0–0.5), possibly damaging (score >0.5–0.9), or probably damaging (score >0.9).
Results
RNAscope ISH
The cellular localization of the FcaPV2 E6 and E7 genes was identified by colorimetric ISH. Hybridization signal patterns were classified into 2 types: diffuse and dot-like (Fig. 1). Positive hybridization signals were detected in 18/22 MCC. All MCC with positive ISH signals were FcaPV2 DNA-positive cases. Hybridization signals were not detected in 3 FcaPV2 DNA-positive MCC (cases 5, 6, and 15) and 1 FcaPV2 DNA-negative MCC (case 22). Strong dot-like signals were observed within the nucleus and cytoplasm of most tumor cells (Figs. 2, 3). Pretreatment with RNase A resulted in the loss of cytoplasmic signal and the majority of nuclear signals; however, a few tumor cells contained faint dot-like signals within their nucleus (Fig. 4). Positive hybridization signals were also detected in concurrent FcaPV2-associated lesions. In 11/12 viral plaques, strong diffuse nuclear signals were consistently observed within the stratum granulosum and spinosum, and weak dot-like signals were rarely observed within the basal and suprabasal layers (Fig. 5). The stratum corneum also contained positive signals. Dot-like signals, similar to the pattern of MCC, were noted within the tumor cells of 2/3 BISC, 1/1 BCC, and 1/1 SCC (Figs. 6, 7). Diffuse nuclear signals were present in the central area of tumor nests in BCC. No hybridization signals were found in normal cutaneous tissues around the lesions.
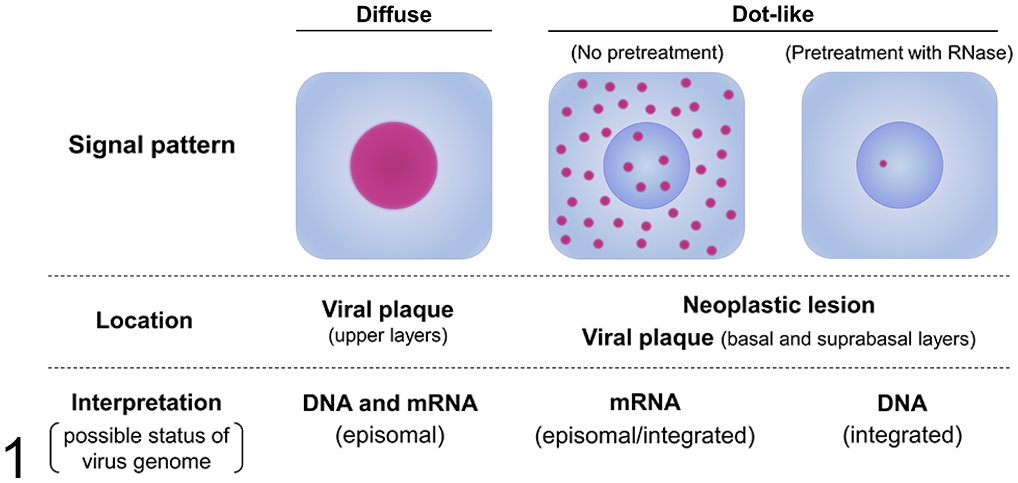
Signal patterns of RNAscope in situ hybridization for Felis catus papillomavirus type 2 (FcaPV2) E6 and E7. Diffuse nuclear signals in the upper layers of viral plaques represent the co-detection of mRNA and DNA of the episomal FcaPV2 genome. Dot-like nuclear and cytoplasmic signals in neoplastic lesions, such as Merkel cell carcinoma and squamous cell carcinoma, and the basal and suprabasal layers of viral plaques represent the mRNA of the episomal/integrated FcaPV2 genome. Dot-like nuclear signals remaining after a pretreatment with RNase represent the DNA of the integrated FcaPV2 genome.
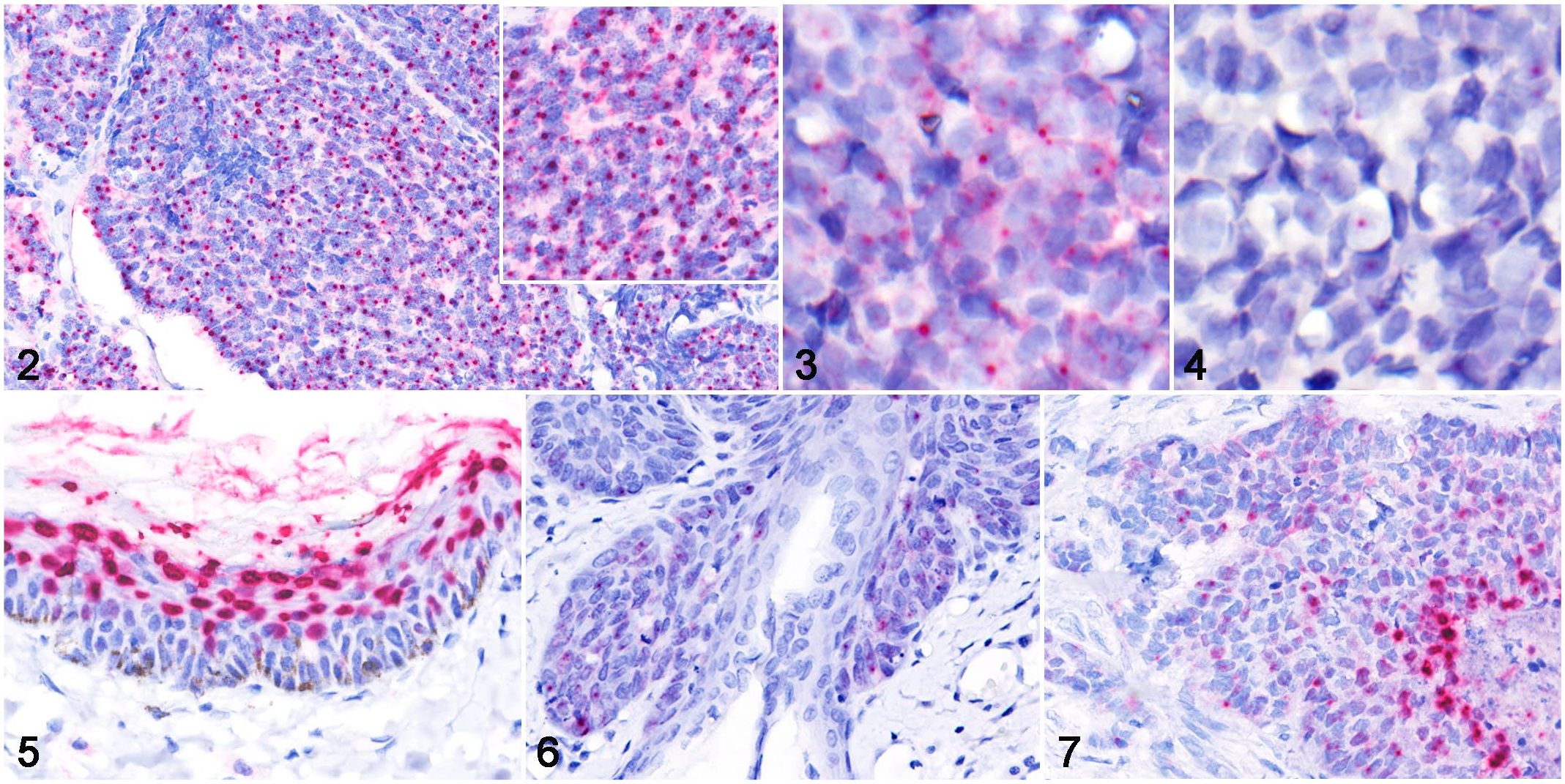
Merkel cell carcinoma, skin, cat. RNAscope in situ hybridization for Felis catus papillomavirus type 2 (FcaPV2) E6 and E7.
Exploring Viral Integration by WGS
The number of reads obtained from the sample of case 9 was 695,607,468 and the number of bases was 104,341,120,200 base pairs (bp), which produced sequencing coverage between a depth of 20–100× in 88% of the genomic regions. The number of reads obtained from the sample of case 21 was 720,226,102 and the number of bases was 108,033,915,300 bp, which produced sequencing coverage between a depth of 20–100× in 94% of the genomic regions. The numbers of chromosome structural variations detected in the mapping analysis are summarized in Supplemental Table S3.
WGS revealed putative FcaPV2 integration regions within the host (F. catus) genome in 2/2 cases, and the results obtained are summarized in Table 2. In case 9, integrations were identified in 7 regions (Regions 1–7): 2 regions on 1 chromosome (A2) and 1 region each on 5 chromosomes (B3, D4, E2, E3, and X) (Fig. 8). Three integration regions were intergenic; 2 were in the introns of AKT2 and PLD3, which have a shared intron, and ATP2B3; and 1 was upstream of PKMYT1. The sense strands of the FcaPV2 sequence were integrated in 3 regions and antisense strands were in 4 regions. Although 2 integration regions on chromosome A2 shared breakpoints in the host and FcaPV2 genomes, the strands of the FcaPV2 sequence were different (sense or antisense). In case 21, 5 integration regions were identified on chromosome D2 (Regions 1–5): 4 regions were intergenic and 1 was in the intron of CH25H (Fig. 8). Region 1 included 5 breakpoints and region 5 included 4 breakpoints in the host genome. Although 4 breakpoints in region 1 (breakpoints 2–5) and 3 breakpoints in region 5 (breakpoints 1–3) were located in the same positions in the host genome, the strands and breakpoints in the FcaPV2 genome were different. The sense strands of the FcaPV2 sequence were integrated into 8 breakpoints, and the antisense strands were integrated into 4 breakpoints in the host genome. Nineteen breakpoints in the FcaPV2 genome identified in the 2 samples were classified by the open reading frame (ORF) of FcaPV2: 1/19 in E1, 1/19 in E6, 1/19 in E7, 4/19 in L1, 4/19 in L2, and 8/19 in LCR (Fig. 9).
Whole genome sequencing analysis of Felis catus papillomavirus type 2 DNA integration in feline Merkel cell carcinoma samples.
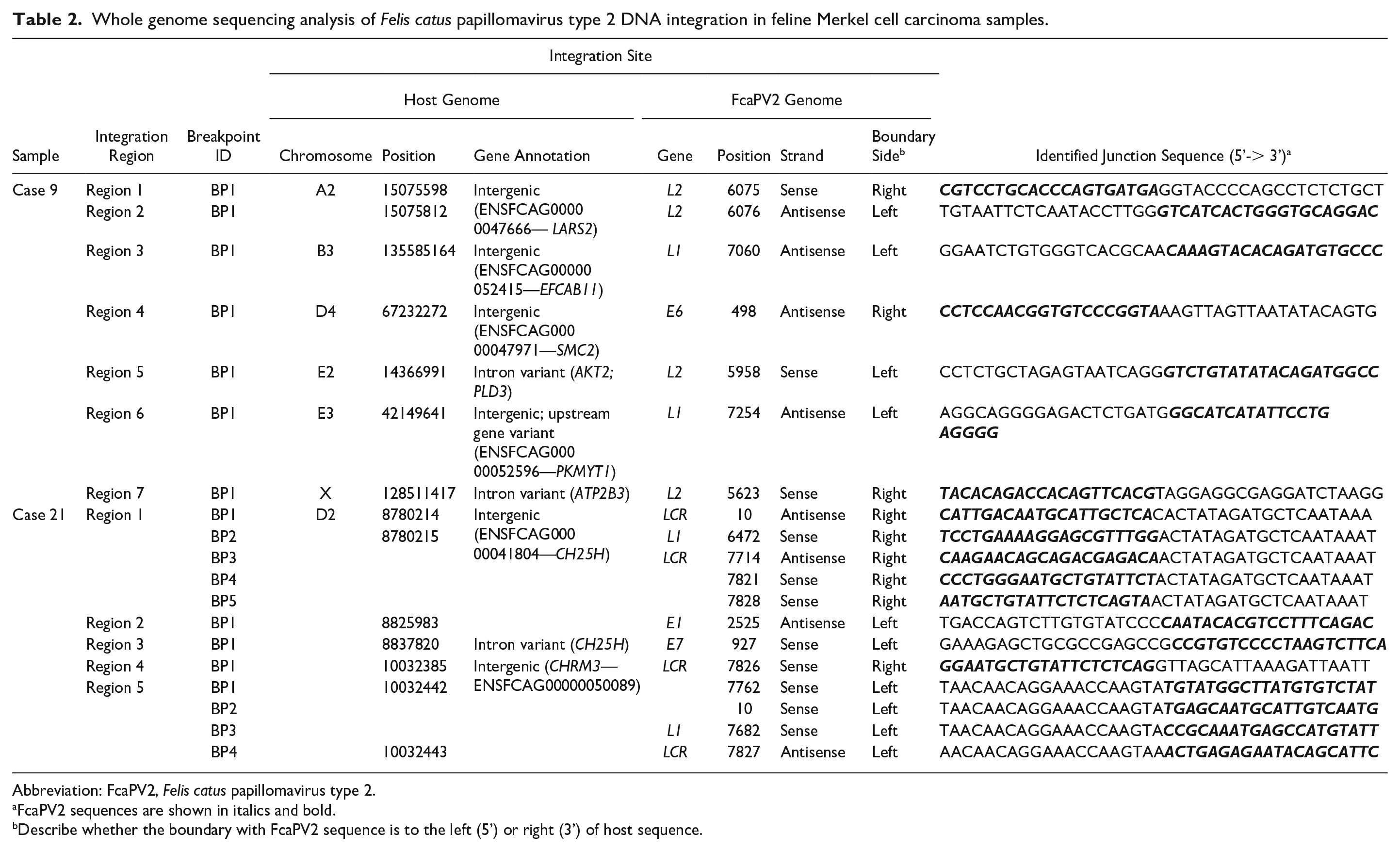
Abbreviation: FcaPV2, Felis catus papillomavirus type 2.
FcaPV2 sequences are shown in italics and bold.
Describe whether the boundary with FcaPV2 sequence is to the left (5’) or right (3’) of host sequence.
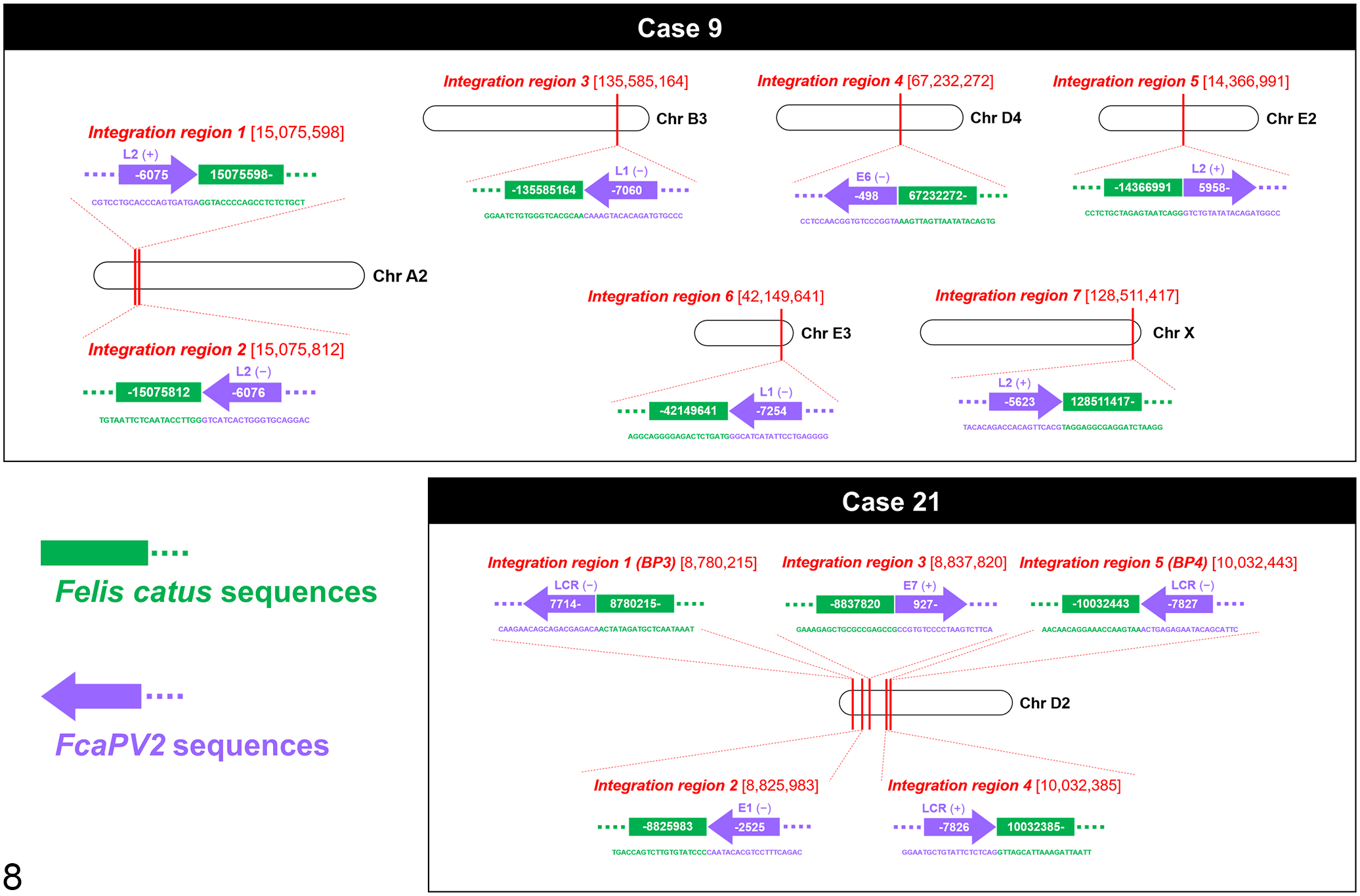
Integration regions of Felis catus papillomavirus type 2 (FcaPV2) genes in host genomes of Merkel cell carcinoma cases 9 and 21. Nucleic acid sequences are described as 5’ to 3’, and the direction of purple arrows indicates strands of the FcaPV2 sequence (sense or antisense). In case 9, 7 integration regions are present in 6 chromosomes (A2, B3, D4, E2, E3, and X). Chromosome A2 includes 2 integration regions. In case 21, 5 integration regions are present in 1 chromosome (D2).
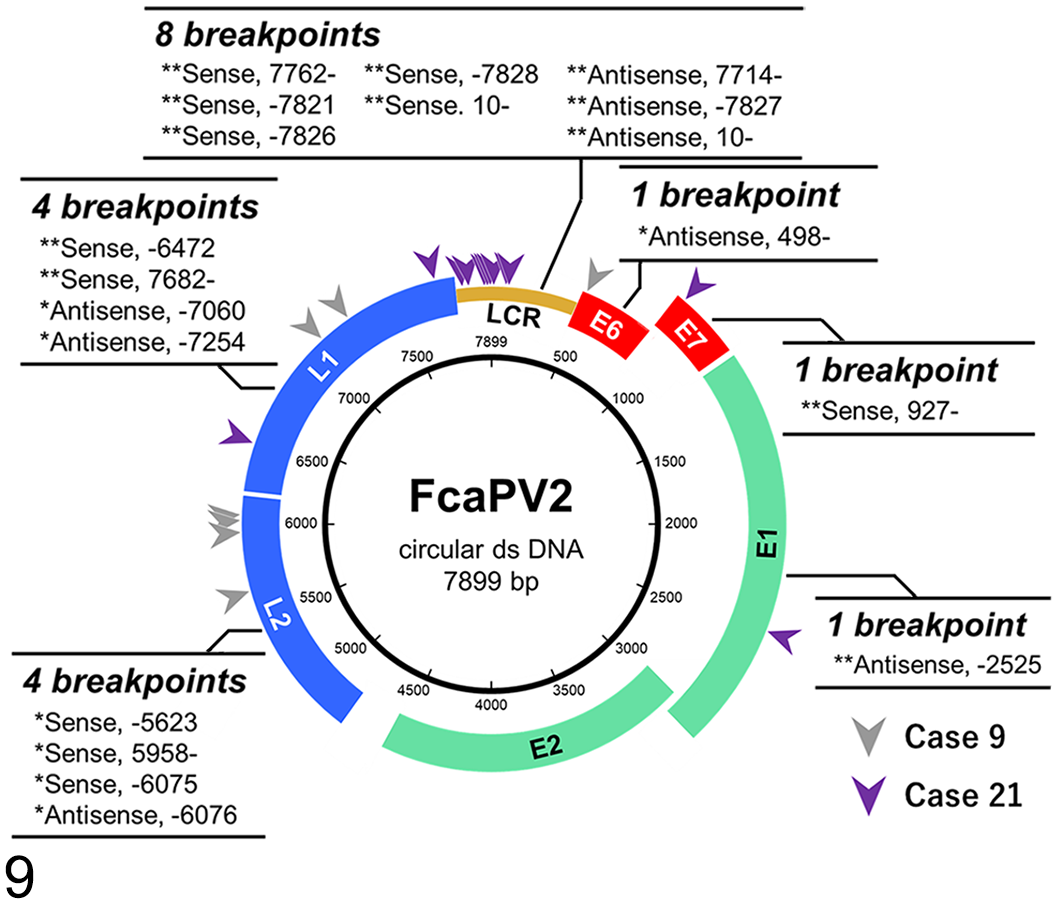
Breakpoints in the Felis catus papillomavirus type 2 (FacPV2) genome identified from FcaPV2 DNA-positive Merkel cell carcinoma cases 9 and 21. Single asterisks (*) and gray arrowheads indicate breakpoints identified in case 9, and double asterisks (**) and purple arrowheads indicate breakpoints identified in case 21.
Complete FcaPV2 genomes were also sequenced by WGS. The complete genome in case 9 was 7899 bp in size and showed 100% nucleotide sequence identity to the reference FcaPV2 sequence (GenBank accession number: LC612600), which was identified in the same case in the previous study. 26 The complete genome in case 21 was 7899 bp in size and showed 99.9% nucleotide sequence identity, 7889/7899 nucleotide (nt) to the reference FcaPV2 sequence (GenBank accession number: LC612600). Nucleotide sequence identities based on each ORF were E1: 99.9% (1807/1809 nt); E2: 99.9% (1592/1593 nt); E6: 100% (417/417 nt); E7: 100% (288/288 nt); L1: 99.9% (1513/1515 nt); L2: 99.9% (1503/1506 nt); and LCR: 99.9% (666/667 nt). The predicted amino acid sequence was 100% identical to the reference FcaPV2, except for E2 (amino acid substitutions: A260T and A380T) and L2 (amino acid substitutions: Q99H, T167A, V168L). One nucleotide substitution in LCR was detected (c. 401T>C).
Characterization of the FcaPV2 DNA-Positive MCC Cell Line
Cultured FMX-MCC01 cells floated in the medium and formed clusters (i.e., spheroids) of small round cells (Fig. 10). Immunofluorescence revealed that cultured cells were positive for CK18, CK19, CK20, synaptophysin, and CD56 (Figs. 11, 12). After the subcutaneous injection of FMX-MCC01 cells, tumor masses developed at the injection site (flank) of all SCID mice. Macroscopically, all tumors were dome-shaped, and their cut surfaces were pink-gray with central necrosis and hemorrhage. Metastatic lesions were not found in any mice. Xenograft tissues were histologically composed of polygonal or round cells with scant basophilic cytoplasm arranged in solid packets separated by a fibrovascular stroma (Fig. 13). The mean mitotic count was 44.0 per 10 HPF. Xenograft cells were immunohistochemically positive for CK20, CK19, CK18, synaptophysin, and CD56 (Figs. 14, 15). The cellular localization of FcaPV2 E6 and E7 in FMC-MCC01 cells was confirmed by ISH. DNA-FISH for E7 detected dot-like nuclear signals within a small number of cells (Fig. 16). RNAscope ISH for E6 and E7 detected dot-like cytoplasmic signals within cultured cells (Fig. 17). Positive hybridization signals were also detected within the tumor cells of xenograft tissues by RNAscope ISH (Fig. 18).
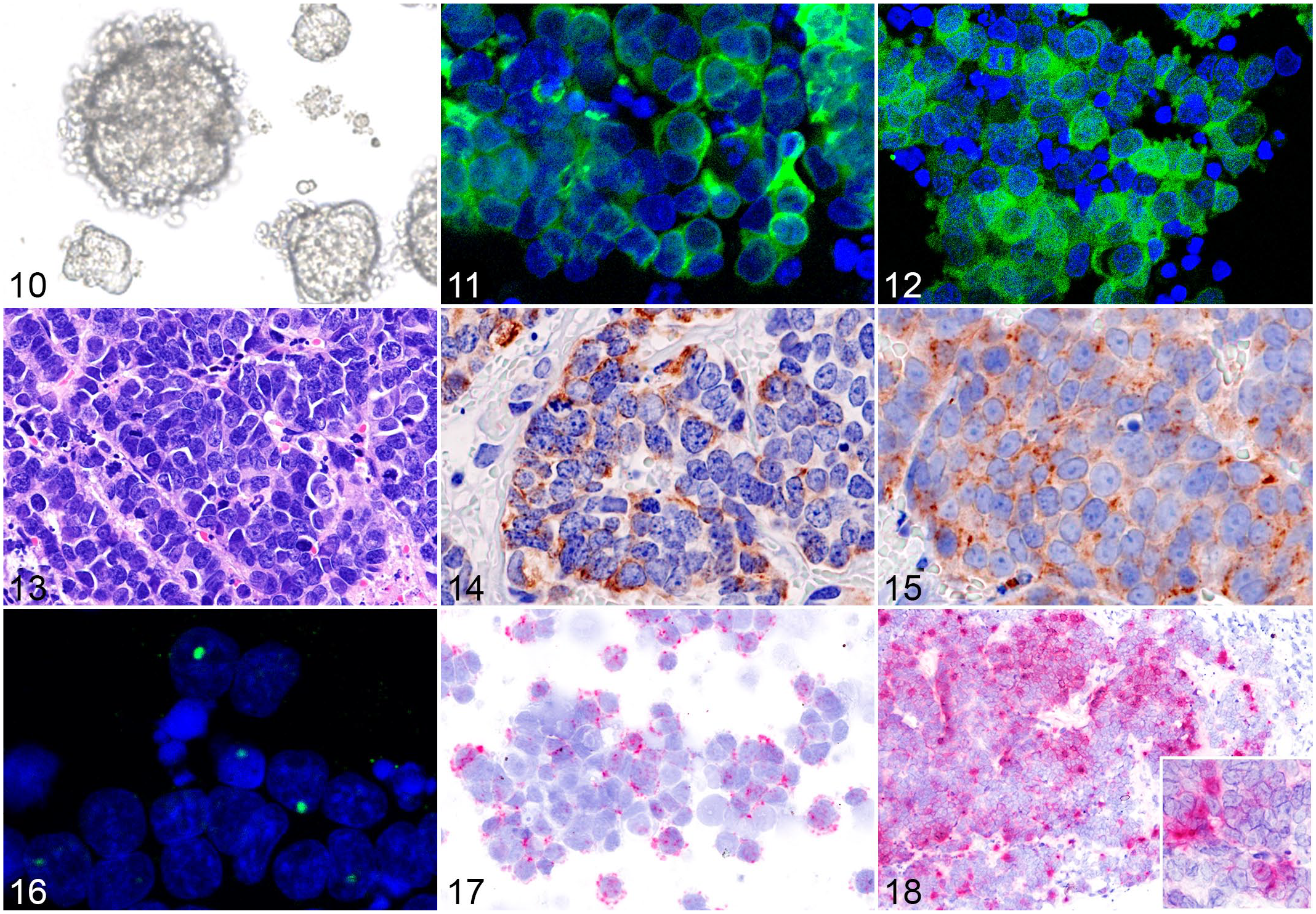
A cell line derived from Felis catus papillomavirus type 2 (FcaPV2)-positive Merkel cell carcinoma (FMX-MCC01).
Comparison of FcaPV2 DNA-Positive and FcaPV2 DNA-Negative MCC Cell Lines
Comparisons between FcaPV2 DNA-positive and FcaPV2 DNA-negative MCC cell lines are summarized in Table 3. In contrast to FMX-MCC01 cells, AS-MCC01 cells formed an adherent monolayer of polygonal- or spindle-shaped cells in culture dishes, as described in the previous study. 25 Immunofluorescence showed that FMX-MCC01 cells were positive for p16 (Figs. 19–21) and negative for pRb and p53, while AS-MCC01 cells were negative for p16 and positive for pRb and p53 (Figs. 22–27). IHC on xenograft tissues revealed high p16 immunoreactivity and low pRb and no p53 immunoreactivity in FMX-MCC01 (Figs. 28–30). Nuclear and cytoplasmic immunolabeling for p16 was observed in most xenograft cells, and nuclear immunolabeling for pRb was observed in a small number of xenograft cells. Xenograft cells were negative for p53. Conversely, the AS-MCC01 cells of xenograft tissues were negative for p16 and positive for pRb and p53 (Figs. 31–33). Most xenograft cells showed nuclear immunolabeling for pRb, and xenograft cells with marked cellular pleomorphism, such as giant- and multi-nucleated cells, showed nuclear immunolabeling for p53.
Comparison of Felis catus papillomavirus type 2-positive and -negative Merkel cell carcinoma cell lines.
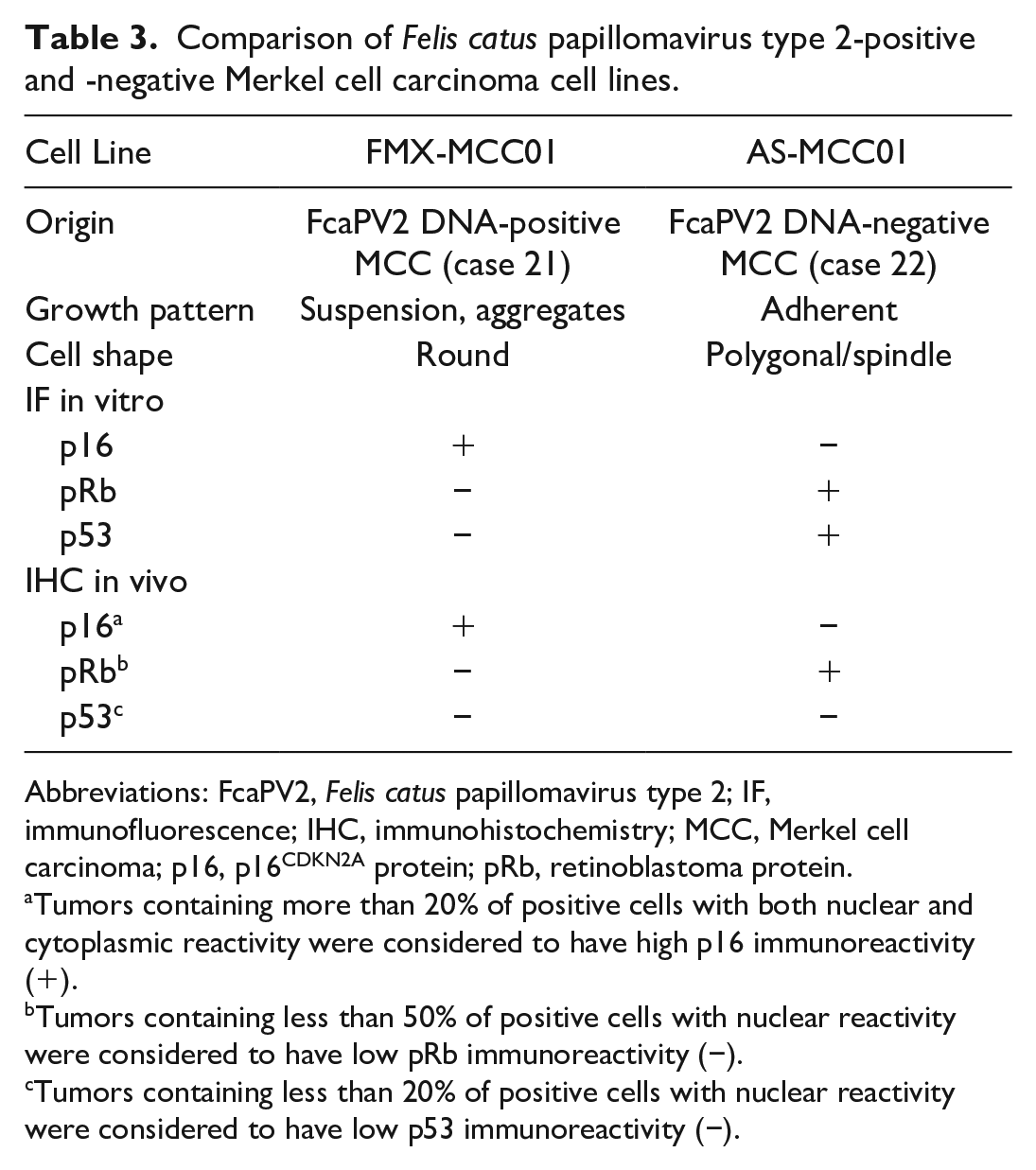
Abbreviations: FcaPV2, Felis catus papillomavirus type 2; IF, immunofluorescence; IHC, immunohistochemistry; MCC, Merkel cell carcinoma; p16, p16CDKN2A protein; pRb, retinoblastoma protein.
Tumors containing more than 20% of positive cells with both nuclear and cytoplasmic reactivity were considered to have high p16 immunoreactivity (+).
Tumors containing less than 50% of positive cells with nuclear reactivity were considered to have low pRb immunoreactivity (−).
Tumors containing less than 20% of positive cells with nuclear reactivity were considered to have low p53 immunoreactivity (−).
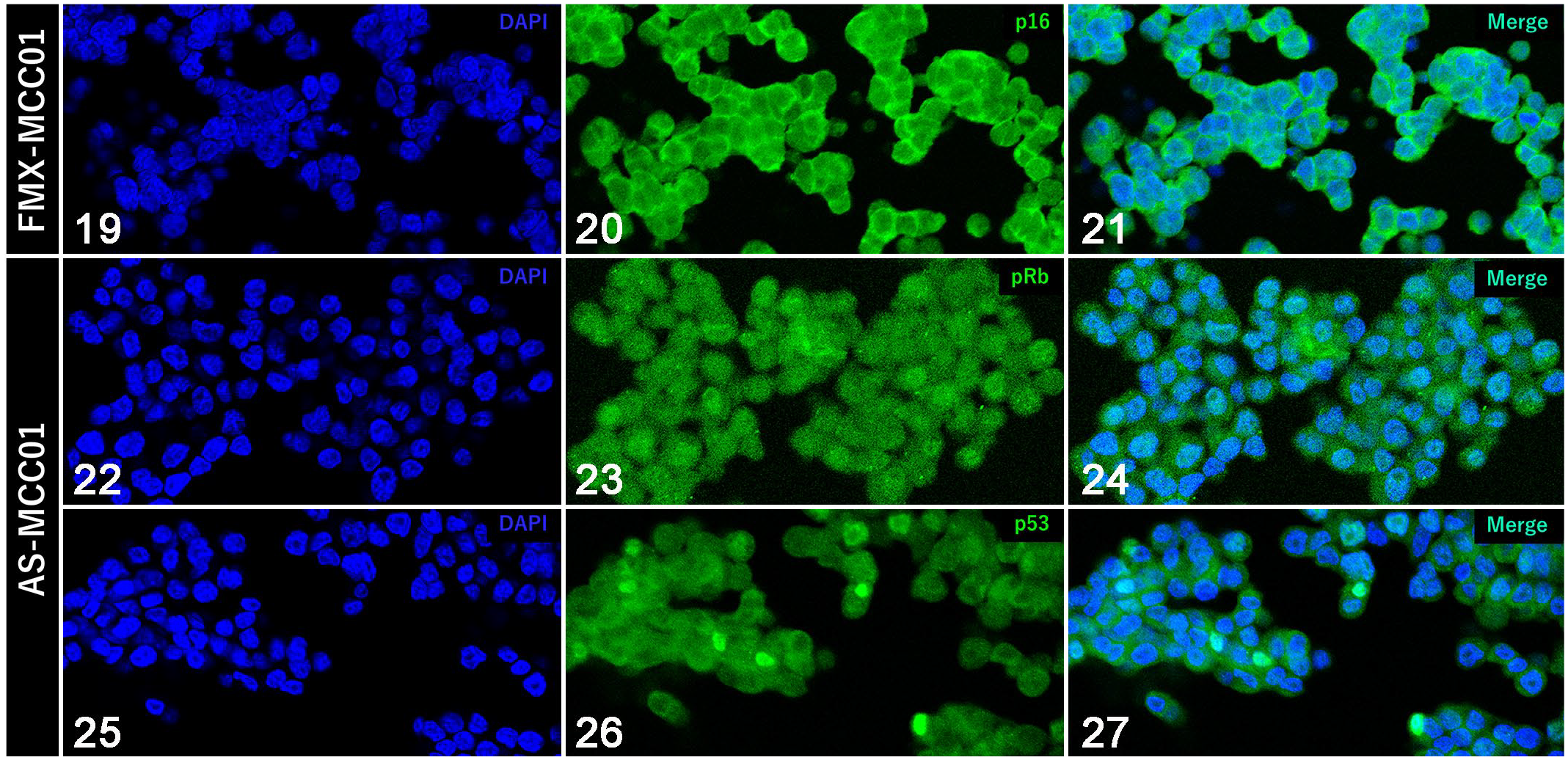
Immunofluorescence for tumor suppressor proteins (in vitro).
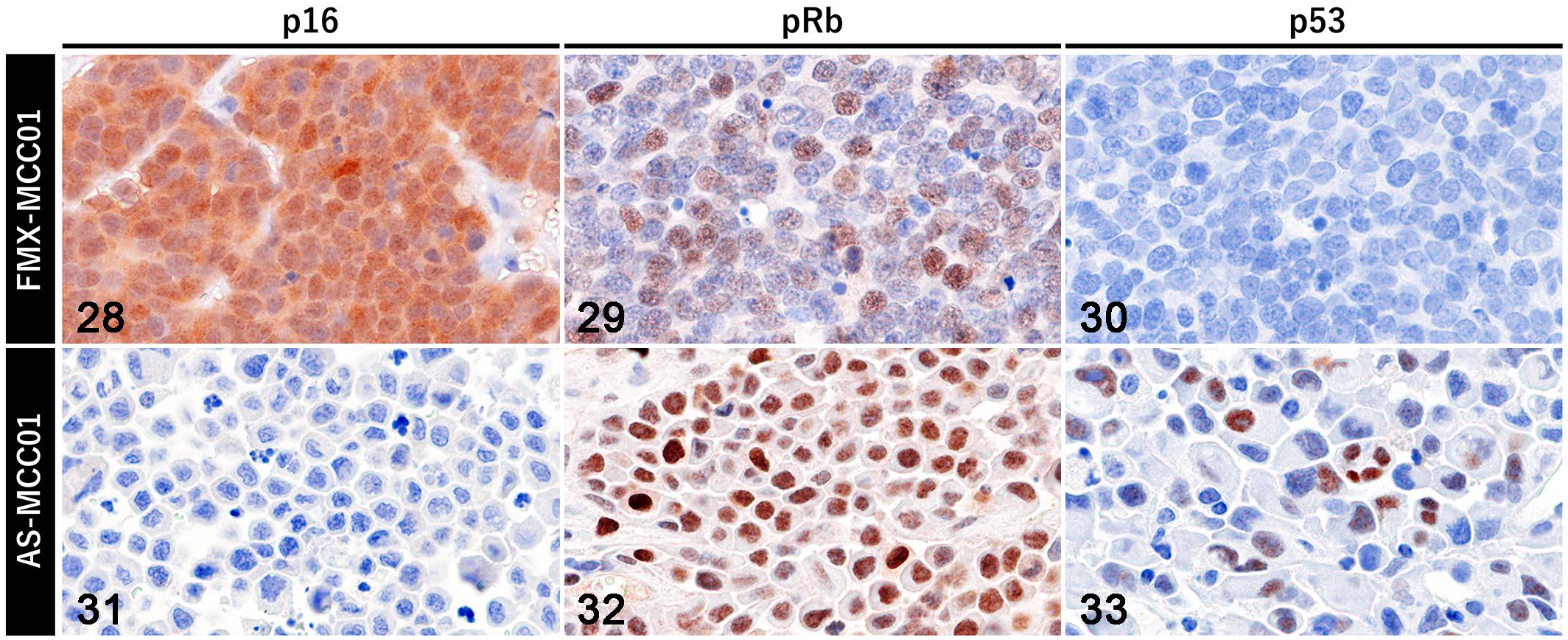
Immunohistochemistry for tumor suppressor proteins in xenograft tumors.
Ultrastructural Analysis
One FcaPV2 DNA-positive MCC was ultrastructurally evaluated by transmission electron microscopy. MCC tumor cells had oval nuclei with coarse chromatin and distinct nucleoli (Supplemental Figs. S1, S2). Some tumor cells had a nucleus with a higher density of chromatin, obscuring the fine structure of nuclei. The cytoplasm contained numerous, approximately 100-nm diameter dense-core vesicles (i.e., neuroendocrine granules). No virus particles were observed within tumor cells.
Sequencing Analysis of the p53 Gene
Extracted genomic DNA from all samples was successfully amplified. DNA sequencing identified point mutations in 8/10 FcaPV2 DNA-positive and 1/1 FcaPV2 DNA-negative MCC samples (Table 4). Two mutations were detected in case 8, 5 in case 19, and 1 in each of the 7 other cases. The same mutation (c. 631C>T) was found in cases 1 and 19. Thirteen different mutations were identified in 4 exons: 4 in exon 5, 4 in exon 6, 3 in exon 7, and 2 in exon 8, and all were missense mutations. Based on sequencing chromatograms, 13 9/13 mutations were detected as heterozygous and 4/13 mutations as homozygous (Figs. 34–37). An in silico analysis using PolyPhen-2 predicted a negative impact on protein function in 10/13 mutations. No mutations, insertions, or deletions were detected in FcaPV2 DNA-positive SCC (1/1) or BCC (1/1) samples.
Statuses of p53 gene in feline Merkel cell carcinoma, squamous cell carcinoma, and basal cell carcinoma samples.
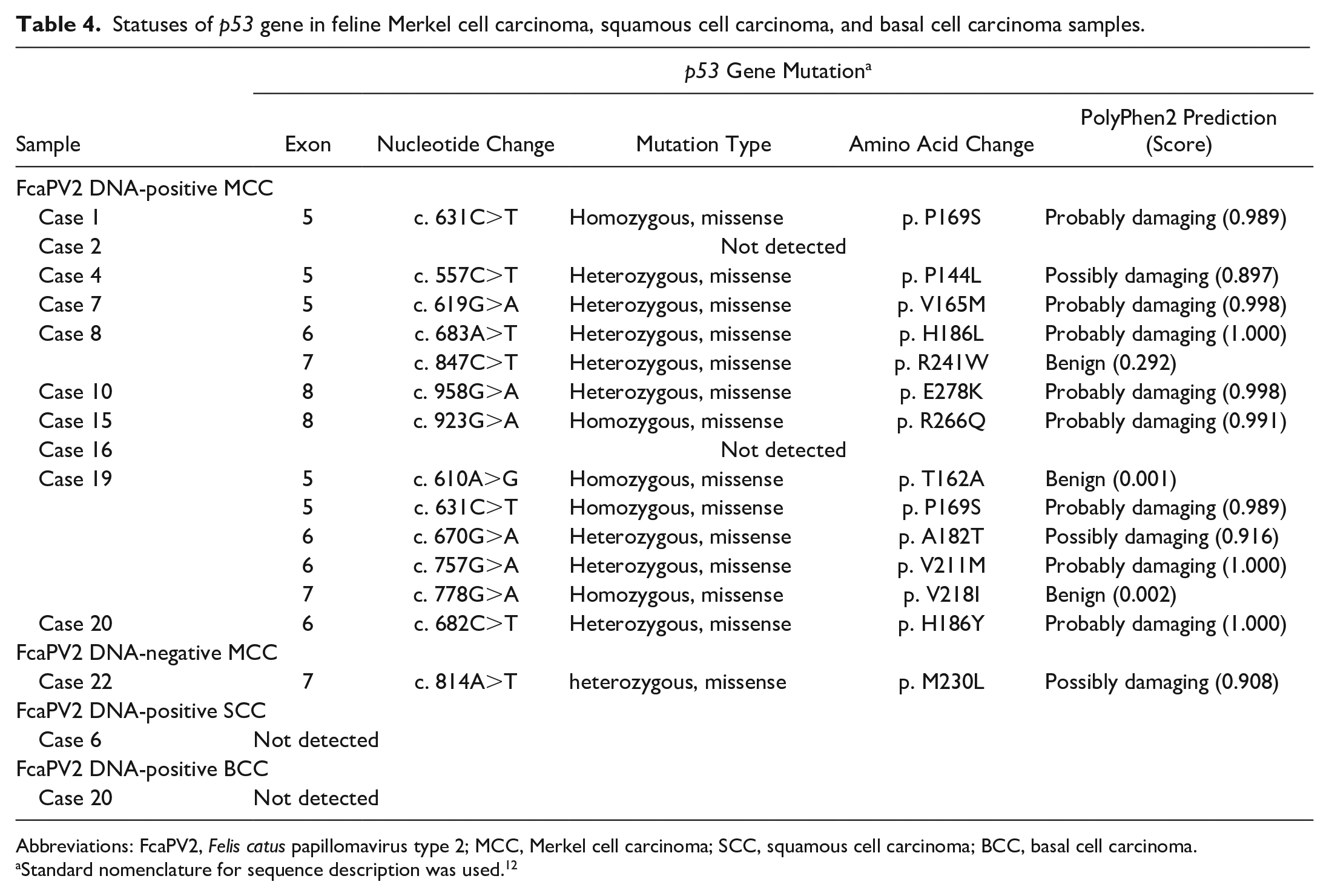
Abbreviations: FcaPV2, Felis catus papillomavirus type 2; MCC, Merkel cell carcinoma; SCC, squamous cell carcinoma; BCC, basal cell carcinoma.
Standard nomenclature for sequence description was used. 12
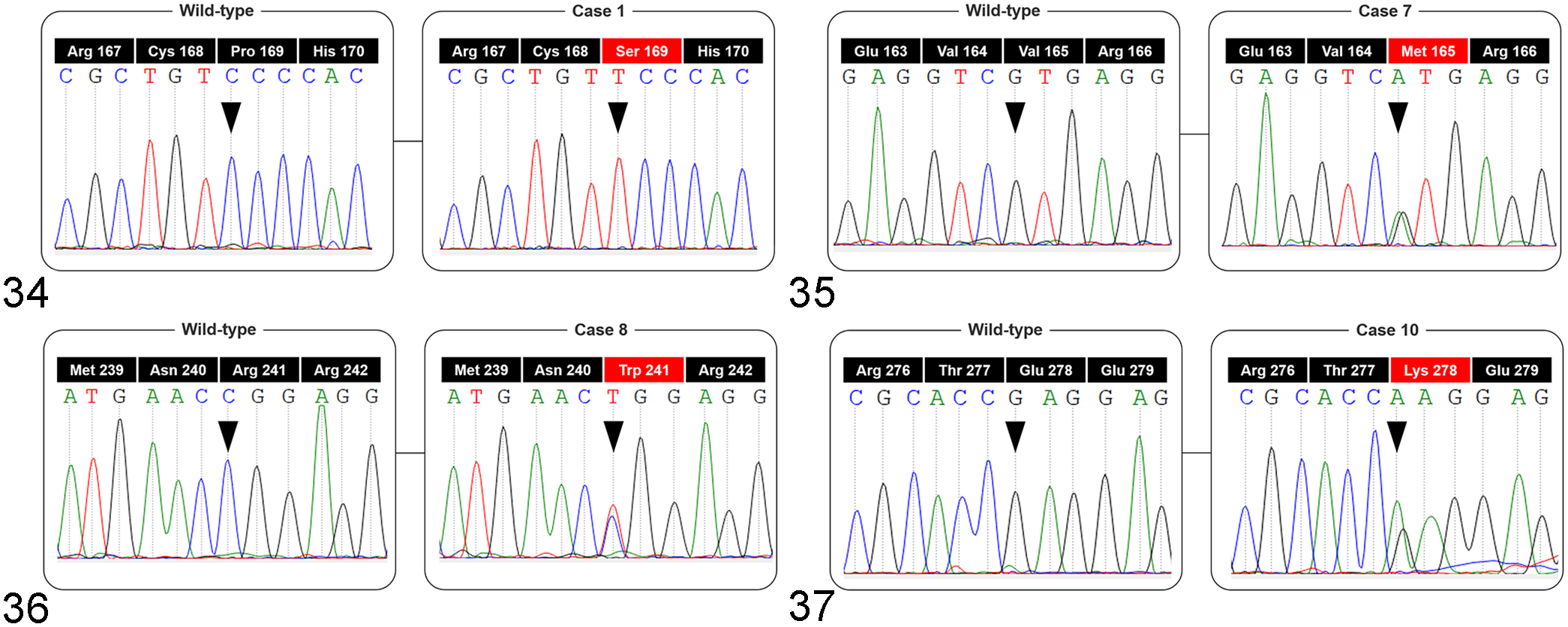
Sequence chromatograms of the p53 gene obtained from feline Merkel cell carcinoma samples.
Discussion
The following criteria have been presented to support the causal role of HPV in the development of oropharyngeal SCC: (1) the presence of viral DNA within tumor samples, (2) the interaction of viral oncoproteins with host tumor suppressor proteins, and (3) the active transcription of the viral oncogenes E6 and E7 in tumor cells. 8 E6 and E7 are key factors in the tumorigenesis of high-risk alpha HPV. 23 The major tumorigenic pathway is the inactivation of the host tumor suppressor proteins p53 and pRb by E6 and E7, respectively. E6 promotes the proteasomal degradation of p53 by forming a complex with ubiquitin ligase E6AP and p53, which inhibits p53-dependent cell cycle arrest and apoptotic pathways. E7 binds to pRb and directs it toward proteasomal degradation, leading to unscheduled cell cycle progression. In HPV-induced tumors, these 2 viral oncogenes are continuously expressed to maintain the growth and malignant phenotype of tumor cells. 23 Recent studies demonstrated that similar molecular mechanisms to HPV contribute to the development of FcaPV-associated SCC in cats.2,3,35,37 In the present study, RNAscope ISH detected FcaPV2 E6 and E7 mRNA within the tumor cells of FcaPV2 DNA-positive MCC (86%). Three FcaPV2 DNA-positive cases without ISH signals may have represented latent infections, in which viral DNA could be detected by PCR, but no viral mRNA was present, or the low-level expression of viral oncogenes not detectable by ISH. Dot-like ISH signals similar to those observed in MCC have been confirmed in HPV-positive cervical carcinoma and FcaPV2-positive SCC.14,22 It has been reported that more dot-like signals were observed as the lesions progressed from premalignant to invasive. The disruption of E2, the viral regulatory gene of E6 and E7, due to neoplastic transformation has been suggested to increase the transcriptional activity of oncogenes. 40 Previous studies using RNAscope ISH reported intense diffuse nuclear signals within HPV-positive intraepithelial neoplasia (CIN) I/II and epidermal hyperplasia adjacent to FcaPV2-positive SCC that persisted even after a pretreatment with RNase A.14,22 Since unzipped single-stranded PV DNA is produced during up-regulated episome synthesis in the productive phase of the PV life cycle, these diffuse signals are considered to represent the co-detection of viral mRNA and DNA.14,22 RNAscope ISH in the present study also showed similar diffuse signals within the upper layers of concurrent viral plaques, indicating that the productive infection of FcaPV2 occurred in the epidermis around FcaPV2 DNA-positive MCC in cats. Weak dot-like signals within the basal and suprabasal layers of viral plaques may represent the expression of early viral genes in the basal layer during proliferative infection. On the other hand, after the pretreatment with RNase A, dot-like nuclear signals remained within FcaPV2 DNA-positive MCC cells. These signals were similar to those detected within FcaPV2 DNA-positive MCC cells by DNA ISH for FcaPV2 E7 in the present and previous studies. 26 This signal pattern represents integrated PV DNA in tumor cells. 9
In the productive phase of HPV life cycle, the viral genome is maintained in the host epithelial cell as circular DNA (i.e., episomal form). 12 On the other hand, in HPV-associated tumors, the viral DNA is found as fragmented sequences in the host genome (i.e., integrated form). Viral genomic integration is a terminal event for the life cycle and is tightly linked to tumor progression in HPV-associated cervical lesions. 12 HPV is present in the episomal form within low-grade lesions, such as CIN I/II, and the integrated form within CIN III and invasive carcinoma.9,47 Therefore, the integration of HPV DNA into the host genome may occur in the late stage of tumorigenesis. 47 For example, the viral integration was confirmed in 100% of HPV18-positive tumors and 80% of HPV16-positive tumors. 47 In the present study, WGS analyses revealed the integration of FcaPV2 in 2 cases of FcaPV2 DNA-positive MCC in cats. The present results are consistent with the findings of ISH. This is the first study to demonstrate the integration of FcaPV in feline neoplastic diseases by WGS. Although HPV DNA is generally integrated into the non-coding region of the human genome, accumulating evidence suggests several hot spots, including POU5F1B, FHIT, KLF5–KLF12, and HMGA2, as well as important cancer-related genes at or around integration regions.18,24 HPV DNA is known to be integrated into multiple human chromosomes in 80% of tumors with HPV integration, and recurrent integration may occur in close regions in the same chromosome. 18 HPV breakpoints are randomly distributed throughout the viral genome.18,24 In the present study, half of the integration regions in the F. catus genome were found in intergenic regions and the other half in introns and upstream of genes. Among these genes, AKT2 participates in cell proliferation and epithelial-to-mesenchymal transition (EMT) in pulmonary carcinoma, 31 PKMYT1 in cell proliferation and apoptosis resistance in gastric carcinoma, 51 and CH25H in tumor vascularization in various tumors. 32 In the present study, various integrations into multiple chromosomes (case 9) and into one chromosome (case 21) were confirmed. The possible causes of these complicated integration statuses are as follows: (1) viral DNA broken at different sites was integrated into different regions of the host genome and/or (2) structural mutations, such as translocation and deletion, occurred at or around the integration sites during cell division. 18 Since WGS performed in the present study comprehensively analyzed the genomes of cells comprising tumor tissues, it is important to consider the possibility that tumor cells with different genomic statuses were examined. Besides, in this study, WGS analyses were performed on only 2 cases. Further studies with larger sample size are needed to support the association between viral integration and MCC in cats.
Regarding breakpoints in the FcaPV2 genome, most breakpoints were found in late viral genes and LCR (84%). Since ISH revealed the expression of FcaPV2 oncogenes in MCC cells, FcaPV2 DNA was considered to be broken at late viral genes or subsequent regions and then integrated into the host genome while retaining E6 and E7. The changes associated with tumor progression, including viral integration, abrogate the replicative life cycle of PV. 20 In PV-induced tumors, the expression of viral capsid proteins is lost and progeny virions cannot be produced. 20 In the present study, virions were not observed in the tumor cells of FcaPV2 DNA-positive MCC by electron microscopy. Furthermore, the L1 antigen, the major capsid protein of PV, was not immunohistochemically detected within tumor cells in our previous study. 26 These results indicate that the life cycle of FcaPV2 is impaired by viral integration in FcaPV2 DNA-positive MCC.
In our previous study, 21/21 cases of FcaPV2 DNA-positive MCC were negative or weakly positive for p53 and 1/1 case of FcaPV2 DNA-negative MCC was strongly positive for p53 by IHC. 26 Since mutant p53 prevents normal p53 functions and accumulates in the nucleus, creating a stable target for immunohistochemical detection, strong and diffuse p53 immunopositivity indicates mutations within the p53 gene. 17 In addition, the complete lack of p53 immunopositivity has been associated with null mutations within p53. 50 The sequencing analysis performed in the present study revealed a p53 missense mutation with a negative impact on protein function in the FcaPV2 DNA-negative MCC sample (case 22), which may be related to the p53 immunopositivity exhibited by a large number of tumor cells in this case. 26 High p53 expression and p53 mutations are common features of MCPyV-negative MCC and HPV-negative SCC in humans.19,33,42 A high mutational burden in these tumors is induced by UV exposure and associated with their tumorigenesis. Similarly, UV-induced DNA damage including p53 mutations may contribute to the development of FcaPV2 DNA-negative MCC. On the other hand, p53 mutations were also detected in 80% of FcaPV2 DNA-positive MCC samples, but not in FcaPV2 DNA-positive SCC and BCC samples. In general, p53 mutations are rarely found in MCPyV-positive MCC and HPV-positive SCC.10,33 The negative results for SCC and BCC were consistent with these findings. Possible reasons why FcaPV2 DNA-positive MCC with p53 mutations were immunohistochemically negative or weakly positive for p53 are as follows: (1) the degradation of mutant and normal p53 by FcaPV2 E6 and (2) the loss of p53 expression due to null mutations in other p53 exons not examined in the present study.
ISH revealed the presence and expression of FcaPV2 oncogenes in the FcaPV2 DNA-positive MCC cell line. Besides, high immunoreactivity for p16 and low immunoreactivity for pRb and p53 were confirmed in the cultured cells and xenograft tissues of the FcaPV2 DNA-positive MCC cell line. These results were similar to those observed in primary tumor tissue. 26 Therefore, these associations in vitro and in vivo suggest that the integrated FcaPV2 oncogenes might contribute to the proliferation of FcaPV2 DNA-positive MCC by inhibiting tumor suppressor proteins. In contrast, the FcaPV2 DNA-negative MCC cell line was negative for p16 and positive for pRb and p53. Although the majority of tumor cells were positive for p53 in the primary tumor tissues of FcaPV2 DNA-negative MCC, only a few cells were positive for p53 in vitro and in vivo. Genomic alterations during subculturing and selective cloning may be associated with the loss of the p53 protein. Since xenograft cells with marked cellular pleomorphism showed p53 immunolabeling, these cells were considered to contain p53 mutations and survived as a result of the inhibition of apoptosis.
In humans, a number of cell lines have been established from MCPyV-positive and MCPyV-negative MCC.21,29,41,45 Human MCC cell lines are classified into classical and variant MCC according to their growth characteristics in vitro. 21 Classical MCC cells have a neuroendocrine growth pattern, namely, they grow in suspensions as loose or compact spheroids. Variant MCC cells grow in adherent monolayers. Although most of the frequently used classical MCC cell lines are positive for MCPyV, most of the variant MCC cell lines are negative for MCPyV and have a high mutational burden associated with UV exposure. 48 Recent studies demonstrated that classical MCC highly expressed atonal bHLH transcription factor 1 (ATOH1), which is a pro-neural transcription factor and essential for Merkel cell development, and variant MCC highly expressed EMT-related genes.15,21 Since forced expression of ATOH1 in variant MCC caused a change in the growth pattern from adherent to suspension, variant MCC is considered to be an intermediate state between neuroendocrine and epithelial differentiation.15,21 In the present study, the growth patterns of FcaPV2 DNA-positive and FcaPV2 DNA-negative MCC cell lines were the same as those of human classical and variant MCC, respectively. These results suggest that, similar to human MCC, the involvement of an oncogenic virus and/or neuroendocrine differentiation in tumorigenesis are associated with the growth characteristics of feline MCC in vitro. FcaPV2 DNA-positive and FcaPV2 DNA-negative MCC cell lines are useful for detailed analyses of tumorigenesis and therapeutic development for feline MCC.
The present study demonstrated the integration of FcaPV2 DNA and the active transcription of FcaPV2 oncogenes in feline MCC. The expression of integrated viral oncogenes might participate in the inhibition of the host tumor suppressor proteins, which might contribute to the development and progression of feline MCC. These findings support the possibility that FcaPV2 infection is an etiological agent of MCC in cats. However, the current study did not directly reveal the interaction of FcaPV2 oncoprotein with host tumor suppressor proteins. We have only examined the consequent changes in the tumor suppressor proteins using limited methods and materials. Further studies are needed to provide mechanistic support for a causal role of FcaPV2 in feline MCC tumorigenesis, and the cell line established in this study and genetic recombination technology would be useful.
Conclusion
The present study suggests that the tumorigenesis of feline MCC may be caused by FcaPV2 infection: FcaPV2 DNA was integrated into the host genome, and the expression of the integrated viral oncogenes is associated with loss of host tumor suppressor proteins, potentially leading to the development of feline MCC. Our results provide additional evidence for FcaPV2 being a major cause of MCC in cats.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858221139197 – Supplemental material for Genomic integration and expression of Felis catus papillomavirus type 2 oncogenes in feline Merkel cell carcinoma
Supplemental material, sj-pdf-1-vet-10.1177_03009858221139197 for Genomic integration and expression of Felis catus papillomavirus type 2 oncogenes in feline Merkel cell carcinoma by Soma Ito, James K. Chambers, Ayumi Sumi, Tetsuo Omachi, Makoto Haritani, Hiroyuki Nakayama and Kazuyuki Uchida in Veterinary Pathology
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
