Abstract
Merkel cell carcinoma (MCC) is a cutaneous neuroendocrine tumor. We recently demonstrated that cats with MCC often have other proliferative cutaneous lesions, such as squamous cell carcinoma (SCC) and basal cell carcinoma (BCC). Based on this finding, we hypothesize that Felis catus papillomavirus (FcaPV) is involved in the development of MCC in cats, similar to SCC and BCC. To investigate this hypothesis, the presence of FcaPV nucleic acid and immunoreactivity for tumor suppressor proteins were examined in 21 feline MCC cases. Polymerase chain reaction using FcaPV type-specific primers detected FcaPV2 DNA in 20/21 samples of MCC. The complete FcaPV2 sequence was characterized in one case. In situ hybridization for FcaPV2 E7 revealed punctate nuclear signals within tumor cells in 19/21 MCC. Increased immunoreactivity for p16CDKN2A protein and decreased immunoreactivity for retinoblastoma (pRb) and p53 proteins were observed in 20/21 MCC. These results suggest that feline MCC cases are infected with FcaPV2 and the subsequent inhibition of pRb and p53 induced by integrated viral oncogenes is associated with feline MCC tumorigenesis, similar to other PV-induced proliferative cutaneous lesions. On the other hand, the single case of FcaPV2-negative MCC showed strong p53 immunoreactivity, suggesting mutations in p53 caused by cancer inducers other than FcaPV2 infection in this case. The present study suggests FcaPV2 as a cause of feline MCC.
Keywords
Merkel cell carcinoma (MCC) is a highly aggressive cutaneous neuroendocrine carcinoma showing epithelial and neuroendocrine differentiation. 4,10,19 MCC cells express epithelial markers such as cytokeratin (CK) 20, and neuroendocrine markers including synaptophysin and CD56 in human and cats. 4,10,19,23 More than 80% of human MCC cases are associated with Merkel cell polyomavirus (MCPyV) infection. 4,16 In these cases, virus genes are integrated into the tumor cell genome, and the viral oncoprotein, large T (LT) antigen, promotes cell division by suppressing the host retinoblastoma protein (pRb), a tumor suppressor protein that regulates cell cycle progression. On the other hand, ultraviolet (UV) light exposure is considered to be a major cause of MCPyV-negative MCC. 4,16
MCC was named after the phenotypic similarity between tumor cells and Merkel cells. Merkel cells (tactile epithelial cells) are cutaneous touch receptors distributed in skin regions with high tactile acuity, such as the fingertips, sinus hair follicles, and touch domes. 10,19,27,44 However, the gene expression profiling of human MCC suggested that MCC originates from epidermal stem cells, dermal fibroblasts, or immature B lymphocytes, and not likely from Merkel cells. 4,54 Recent studies suggested that epidermal stem cells and dermal fibroblasts are the cells of origin of MCPyV-negative MCC and MCPyV-positive MCC, respectively. 51
In animals, MCC has mostly been examined in cats. 3,10,19,41,50 Since a larger number of MCC cases has been reported in cats than in other animal species, cats appear to be more susceptible to developing MCC. In contrast to human MCC, our previous study on feline MCC showed the negative involvement of MCPyV genes and the LT antigen, and thus, the underlying cause of feline MCC has not yet been elucidated. 19 The immunohistochemical profiles of feline MCC and normal skin tissues have suggested that cutaneous stem cells in the hair follicle bulge are the cell of origin for feline MCC, 10,19 while a lymphoid origin was not demonstrated. 56 Other proliferative cutaneous lesions, including acanthosis with dysplastic changes, carcinoma in situ, squamous cell carcinoma (SCC), and basal cell carcinoma (BCC), were found in 80% of feline MCC cases, often adjacent to the MCC lesion. 19
In cats, UV exposure and Felis catus papillomavirus (FcaPV) infection are well-known factors involved in the development of these proliferative cutaneous lesions. 31,36,64,65 Actinic keratosis and SCC that develop at UV-exposed areas of the body, such as the pinna, nasal planum, and eyelids, are considered to be associated with chronic UV exposure. 31,38 On the other hand, papillomavirus (PV) infections have been confirmed in feline SCC that developed in UV-protected areas, such as haired or pigmented skin. 36 Furthermore, viral plaque and bowenoid in situ carcinoma (BISC) are recognized as PV-induced cutaneous lesions in cats. 31 Recent studies suggested that a molecular mechanism similar to that in human papillomavirus (HPV)-associated tumors contributes to the development of FcaPV-associated SCC in cats. 1,32,52 After viral genes are integrated into the host genome, the viral proteins E7 and E6 inhibit the tumor suppressor proteins pRb and p53, resulting in increased cell proliferation and survival, respectively. 5,42,45 Moreover, the inactivation of Rb causes aberrant increases in the tumor suppressor protein p16CDKN2A (p16), and thus, nuclear p16 immunoreactivity is considered as a surrogate marker for PV-associated tumorigenesis. 8,20,24
The high incidence of proliferative cutaneous lesions in cats with MCC suggests a common etiology. Therefore, we hypothesized that FcaPV infection is associated with the development of feline MCC. To elucidate the relationship between feline MCC and FcaPV, we examined the presence of FcaPV nucleic acid and immunoreactivity for the tumor suppressor proteins, p16, pRb, and p53 in feline MCC.
Materials and Methods
Samples
Biopsy samples from 21 cats that were histopathologically diagnosed with MCC were examined (Table 1). Samples were from the archives of the Laboratory of Veterinary Pathology, the University of Tokyo, and Diagnostic Laboratory, Patho-Labo, between 2008 and 2020. All cases were included in our previous study. 19 Feline SCC, BISC, and viral plaque tissues were additionally included as case controls.
Basic information on feline Merkel cell carcinoma cases.
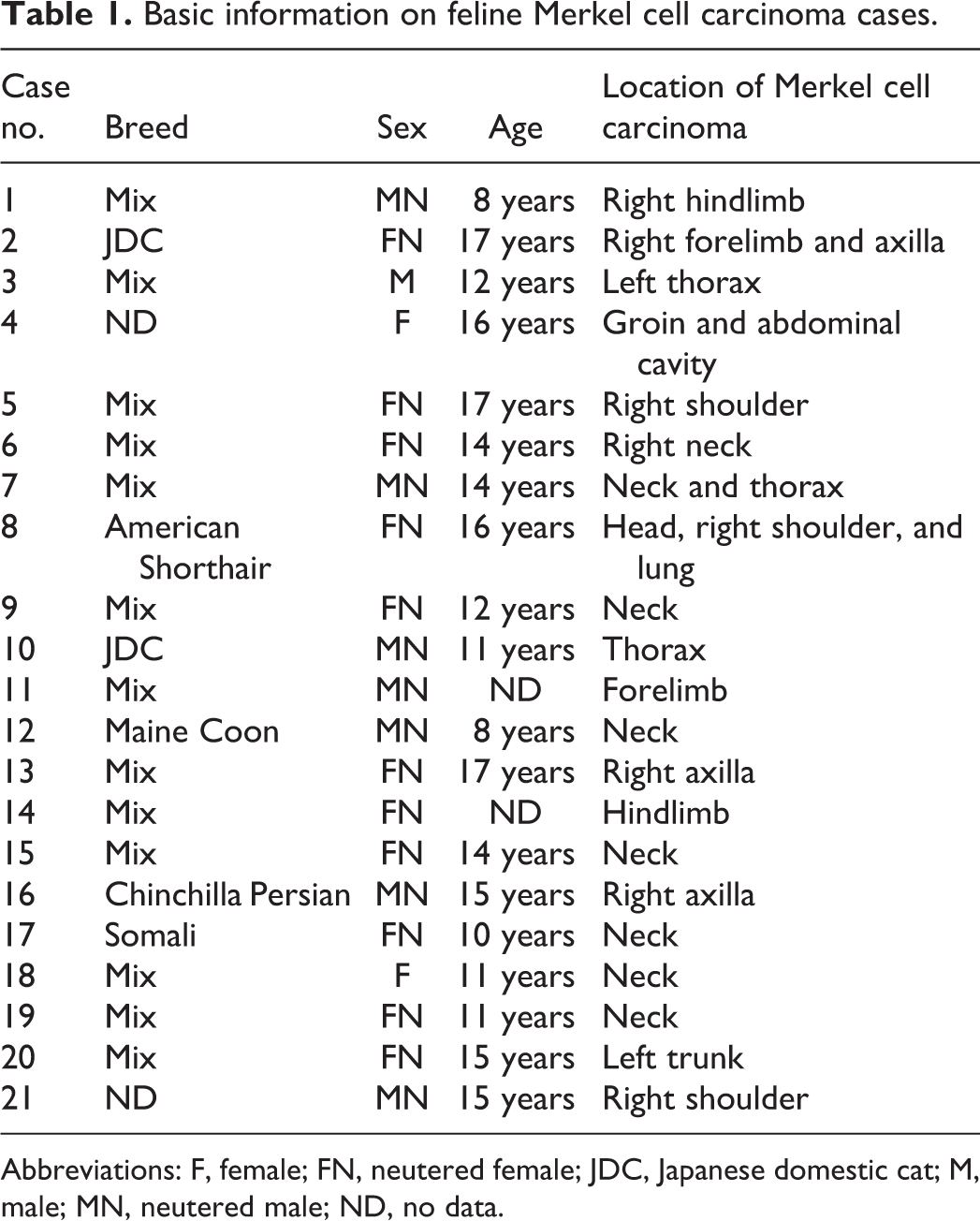
Abbreviations: F, female; FN, neutered female; JDC, Japanese domestic cat; M, male; MN, neutered male; ND, no data.
Histopathology and Immunohistochemistry
Specimens were routinely fixed in 10% phosphate-buffered formalin solution, embedded in paraffin, sectioned at a thickness of 4 μm, and stained with hematoxylin and eosin. Proliferative cutaneous lesions other than MCC in cats with MCC were also reviewed.
Immunohistochemistry (IHC) was performed using the primary antibodies listed in Supplemental Table S1. To detect PV antigens, IHC was performed using a cocktail of 2 monoclonal antibodies against the major capsid protein L1 of bovine PV type 1 and HPV type 16, which successfully detected FcaPV antigens in our previous study. 21 Deparaffinized tissue sections were immersed in 10% hydrogen peroxide (H2O2) in methanol at room temperature for 5 minutes and then incubated in 8% skim milk in Tris-buffered saline (TBS) at 37 °C for 40 minutes to avoid nonspecific reactions. Each tissue section was then treated with a primary antibody at 4 °C overnight. An anti-mouse IgG or anti-rabbit IgG polymer labeled with horseradish peroxidase (Envision, Agilent Technology) or the Optiview DAB detection kit (Roche Diagnostics) was applied at 37 °C for 40 minutes, and sections were then rinsed with TBS. Reactions were visualized with 0.05% 3,3′-diaminobenzidine and 0.03% H2O2 in Tris–hydrochloric acid buffer, followed by a counterstain with Mayer’s hematoxylin (Muto Pure Chemicals). The number of tumor cells positive for p16, pRb, and p53 were manually counted in 5 high-power fields (×400). The location of immunolabeling as nuclear or cytoplasmic was also assessed. Immunohistochemical positivity for p16 was evaluated based on the following parameters: percentage positivity, pattern, and intensity. 49 The pattern of p16 positivity was noted as nuclear, cytoplasmic, or both. The intensity was scored as negative, weak, intermediate, or strong. Lesions containing more than 20% of cells with intermediate or strong positivity in both nucleus and cytoplasm were considered to have increased p16 (+). Immunolabeling for pRb and p53 were scored as follows: 0, negative; 1, <20% positive cells with nuclear reactivity; 2, 20% to 50% positive cells with nuclear reactivity; 3, >50% positive cells with nuclear reactivity. Lesions containing less than 50% positive cells with nuclear reactivity were considered to be decreased pRb (−). Lesions containing less than 20% positive cells with nuclear reactivity were considered to be decreased p53 (−).
Polymerase Chain Reaction (PCR) Analysis
Genomic DNA was extracted from 21 formalin-fixed paraffin-embedded (FFPE) tissue samples using the QIAamp DNA FFPE Tissue Kit (QIAGEN). For one case (case 10), DNA was extracted from FFPE tissues and also from fresh-frozen tissue samples using the DNeasy Blood & Tissue Kit (QIAGEN). The success of DNA extraction from all samples was validated by conventional PCR using a primer pair targeting feline beta-actin named Feline ACTB 1 (F)/1 (R) (Suppl. Table S2). PCR was performed using KOD FX Neo DNA polymerase (TOYOBO) by preparing the following reaction mixture: 10 µL 2× PCR Buffer for KOD FX Neo, 4 µL dNTPs (2 mM), 0.4 µL forward primer (10 µM), 0.4 µL reverse primer (10 µM), and 0.4 µL KOD FX Neo (1 U/µL). Template DNA was added between 30 and 40 ng per PCR reaction followed by distilled water to a total volume of 20 µL. Distilled water instead of template DNA was applied for negative controls. The PCR cycle was performed using a 3-step cycling condition according to the manufacturer’s instructions as follows: pre-denaturation at 94 °C for 2 minutes followed by 39 cycles of denaturation at 98 °C for 10 seconds, annealing at 56 °C for 30 seconds, and extension at 68 °C for 15 seconds. PCR-amplified products were electrophoresed on 2% agarose gels and bands were visualized by UV exposure. To detect the FcaPV genome, type-specific primers targeting E1 of FcaPV2, FcaPV3, FcaPV4, and FcaPV5 were designed based on each reference sequence (GenBank accession numbers: EU796884.1, JX972168.1, KF147892.1, and KY853656.1, respectively; Suppl. Table S2). 64 An additional primer pair for FcaPV2 E7, named FcaPV2 E7 956 (F)/1130 (R), was designed to reconfirm the PCR result validated by the FcaPV2 E1 1769 (F)/1925 (R) primer pair (Suppl. Table S2). 64 Extracted DNA from a FcaPV2-positive feline BISC case was applied as the positive control for primers targeting FcaPV2 E1 and E7. Genomic DNA samples from previous studies were used as positive controls for the FcaPV3 E1 1165 (F)/1331 (R), FcaPV4 E1 1973 (F)/2115 (R), and FcaPV5 E1 1113 (F)/1269 (R) primers. 21,65 PCR reactions were performed using the 3-step cycling condition described above and by modifying the respective annealing temperatures listed in Supplemental Table S2. PCR results were confirmed by gel electrophoresis as described above.
To verify the FcaPV2 sequence amplified by the 2 FcaPV2 primer pairs, E1 1769 (F)/1925 (R) (case 10) and E7 956 (F)/1130 (R) (cases 2, 3, 4, 7, 9, 20, 21, and control cases), direct sequencing was performed. Each PCR-amplified product was purified using the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel) following the manufacturer’s instructions. Purified PCR products were bidirectionally sequenced with the BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems) and 3130xl genetic analyzer (Applied Biosystems). The sequences obtained were analyzed using Molecular Evolutionary Genetics Analysis Version X (MEGA X) software and the Basic Local Alignment Search Tool from the National Center for Biotechnology Information (NCBI; https://blast.ncbi.nlm.nih.gov/Blast.cgi). 22
Subcloning and Full-Genome Sequencing
To characterize the complete FcaPV2 sequence identified in the MCC lesion, extracted DNA from the fresh-frozen tissue-derived sample of case 10 was subjected to full-genome sequencing and subcloning. Three pairs of primers that divide the complete FcaPV2 genome into 3 amplicons, named FcaPV2 1732 (F)/4541 (R), FcaPV2 4474 (F)/7048 (R), and FcaPV2 6506 (F)/2700 (R), were applied (Suppl. Table S3). PCR was performed using KOD FX Neo DNA polymerase by setting the annealing temperature to 56 °C and the extension period to 4 minutes 30 seconds in the 3-step cycling condition described above. Amplified PCR products were electrophoresed on 1.2% agarose gels and each band was cut following the purification step using the NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel). Gel-purified products were cloned into the pCR XL-2 TOPO vector (Invitrogen). The complete FcaPV2 sequence was bidirectionally elucidated using sequencing primers (Suppl. Table S3) following the same sequencing methods as described. Sequence data were analyzed using MEGA X and the respective open reading frames (ORFs) were elucidated using the ORF Finder tool from NCBI (https://www.ncbi.nlm.nih.gov/orffinder/). 22 Pairwise nucleotide and amino acid identities were calculated using the EMBOSS Needle program (https://www.ebi.ac.uk/Tools/psa/emboss_needle/). 40
Sequencing Analysis of p16 Gene
To examine possible mutations of p16 in feline skin samples with MCC, 3 primer sets were designed to amplify each of the 3 exons (Suppl. Table S4). 30 Extracted DNA from the FFPE tissue-derived sample of cases 2, 3, and 16 was subjected to PCR. Extracted DNA from normal skin of a cat was applied as a control. PCR was performed by preparing the following reaction mixture: 12.5 µL KOD One PCR Master Mix-Blue- (TOYOBO), 0.75 µL forward primer (10 µM), 0.75 µL reverse primer (10 µM). Template DNA was added 30 ng per PCR reaction followed by distilled water to a total volume of 25 µL. The PCR cycle was performed using a 3-step cycling condition according to the manufacturer’s instructions as follows: pre-denaturation at 95 °C for 3 minutes followed by 45 cycles of denaturation at 98 °C for 10 seconds, annealing at 55 °C for 5 seconds, and extension at 68 °C for 1 second. After amplification, the product was confirmed by gel electrophoresis as described above. The sample was judged to have homozygous deletion of p16 if amplification of the p16 gene failed. For samples without evidence of overt homozygous deletion, the PCR products were subjected to a sequence analysis (FASMAC) and compared with the reference sequence (GenBank accession number: NM001290248.1).
In Situ Hybridization
Using extracted DNA from fresh-frozen tissue (case 10) as a template, a digoxigenin (DIG)-labeled DNA probe for FcaPV2 E7 gene was obtained by PCR amplification using the PCR DIG Probe Synthesis Kit (Roche Diagnostics) according to the manufacturer’s protocol. After amplification, the product was confirmed by gel electrophoresis as described above. Chromogenic in situ hybridization (CISH) was performed for all cases. Deparaffinized tissue sections were immersed in 10% H2O2 in methanol at room temperature for 5 minutes, followed by enzyme digestion with the Carezyme III: Pronase Kit (Biocare Medical LLC) at room temperature for 5 minutes. After an autoclave pretreatment at 105 °C in 10 mM citrate buffer, pH 6.0 for 15 minutes, DNA was denatured by heating at 95 °C for 5 minutes. Sections were hybridized with 1 ng/µL of a DIG-labeled DNA probe at 40 °C for 16 hours. After washing, sections were incubated in 1.5% Blocking Reagent (Roche Diagnostics) in TBS at room temperature for 30 minutes, and then incubated with a mouse anti-digoxigenin antibody at 37 °C for 1 hour. An anti-mouse IgG polymer labeled with horseradish peroxidase (Envision, Agilent Technology) was applied at 37 °C for 40 minutes and sections were then rinsed with TBS. Reactions were visualized with 0.05% 3,3′-diaminobenzidine and 0.03% H2O2 in Tris–hydrochloric acid buffer, followed by a counterstain with Mayer’s hematoxylin (Muto Pure Chemicals).
Fluorescence in situ hybridization (FISH) was performed for 7 cases (cases 1, 2, 3, 4, 9, 10, and 21). After hybridization, sections were incubated with 20 μg/mL DyLight 488-labeled anti-DIG-digoxin (Vector Laboratories) at 37 °C for 1 hour, counterstained with DAPI (Vectashield1; Vector Laboratory), and then observed under a Zeiss LSM 700 laser scanning confocal microscope (Carl Zeiss Meditec). Furthermore, to clarify the cellular distribution of hybridization signals, dual-fluorescence ISH and IHC was also performed for case 21. After incubation with DyLight 488-labeled anti-DIG-digoxin, anti-CK18 antibody (Suppl. Table S1), a marker of feline MCC, 19 was applied at 37 °C for 1 hour. The tissue section was subsequently incubated with anti-mouse Alexa 594 (1:100 dilution, Thermo Fisher Scientific) at 37 °C for 1 hour, counterstained with DAPI, and then observed under a Zeiss LSM 700 laser scanning confocal microscope.
Results
Histopathology and IHC
In all cases, MCC lesions were located in the dermis and/or subcutis and were not associated with the overlying epidermis (Fig. 1). Neoplastic foci consisted of tumor cells arranged in trabeculae and/or solid packets divided by a thin fibrous stroma (Fig. 2). Tumor cells were polygonal and had a moderate amount of an amphophilic cytoplasm and a round nucleus with dispersed chromatin.
The results of IHC for 21 cases are summarized in Table 2 and Supplemental Table S4. By immunohistochemistry, PV L1 antigens were not detected in the MCC tissue of any case (Fig. 3). Increased p16 immunoreactivity was confirmed in all MCC (Fig. 4), except for 1 case (case 8; Fig. 5). In these cases, more than 20% of MCC cells showed intermediate or strong immunoreactivity for p16 within both nucleus and cytoplasm. Immunoreactivity for pRb and p53 was confined to the nucleus of tumor cells in all MCC. Decreased pRb immunoreactivity was observed in 20/21 MCC (Fig. 6) while in the pRb-positive (+) case (case 8), most MCC cells showed strong pRb immunoreactivity (Fig. 7). In 20/21 MCC, p53 immunoreactivity was absent or weak (Fig. 8), and strong p53 immunoreactivity was observed in 1/21 MCC (case 8; Fig. 9). In normal cutaneous tissue around the tumor, immunolabeling for pRb was present in the epidermis, hair bulbs, epithelial root sheaths of hair follicles, apocrine glands, sebaceous glands, fibroblasts, and capillary endothelial cells, while all these cells were negative for p16 and p53 (Suppl. Figs. S1–S3).
Immunohistochemical analysis of Merkel cell carcinoma and other proliferative cutaneous lesions.
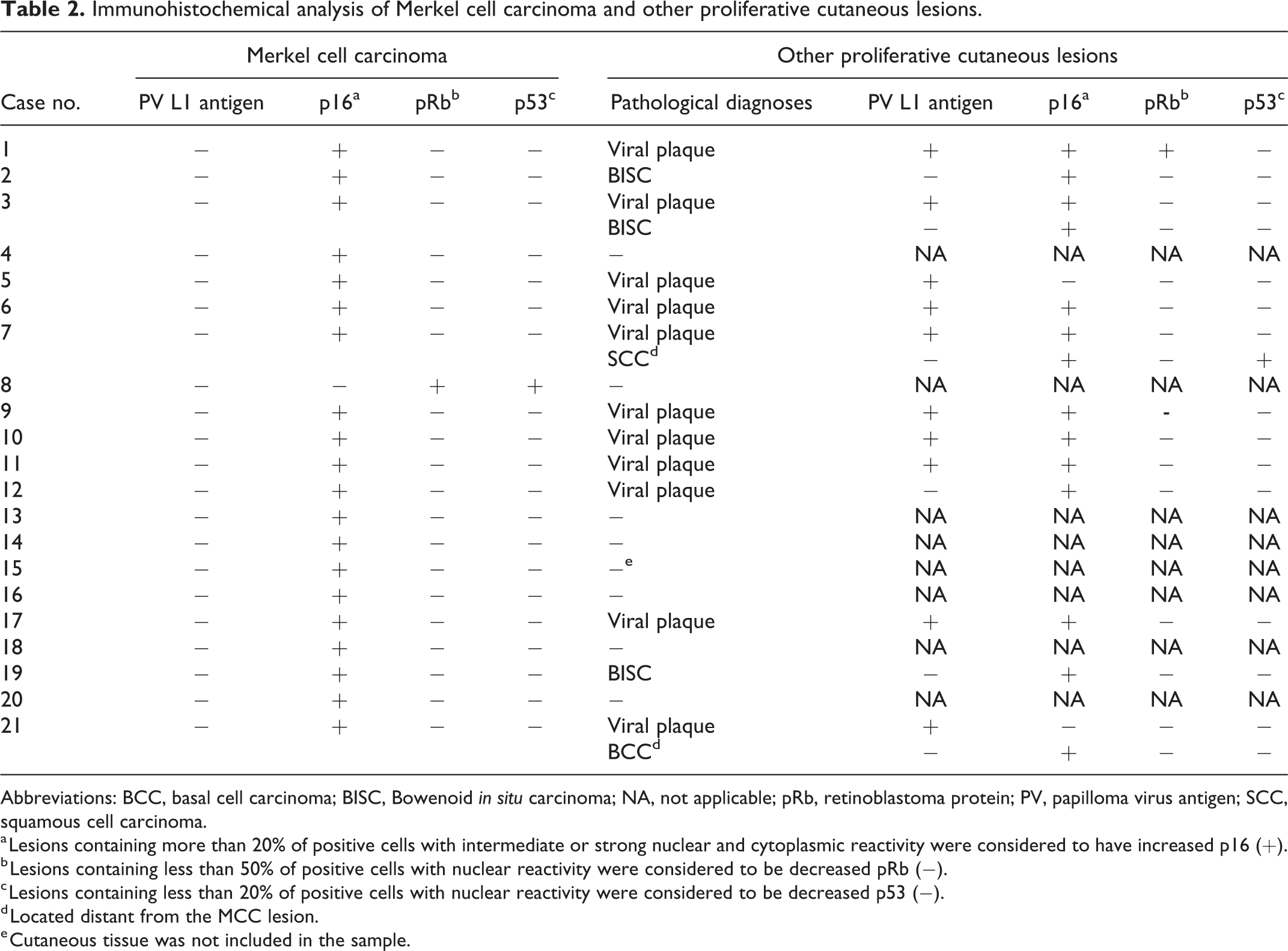
Abbreviations: BCC, basal cell carcinoma; BISC, Bowenoid in situ carcinoma; NA, not applicable; pRb, retinoblastoma protein; PV, papilloma virus antigen; SCC, squamous cell carcinoma.
a Lesions containing more than 20% of positive cells with intermediate or strong nuclear and cytoplasmic reactivity were considered to have increased p16 (+).
b Lesions containing less than 50% of positive cells with nuclear reactivity were considered to be decreased pRb (−).
c Lesions containing less than 20% of positive cells with nuclear reactivity were considered to be decreased p53 (−).
d Located distant from the MCC lesion.
e Cutaneous tissue was not included in the sample.
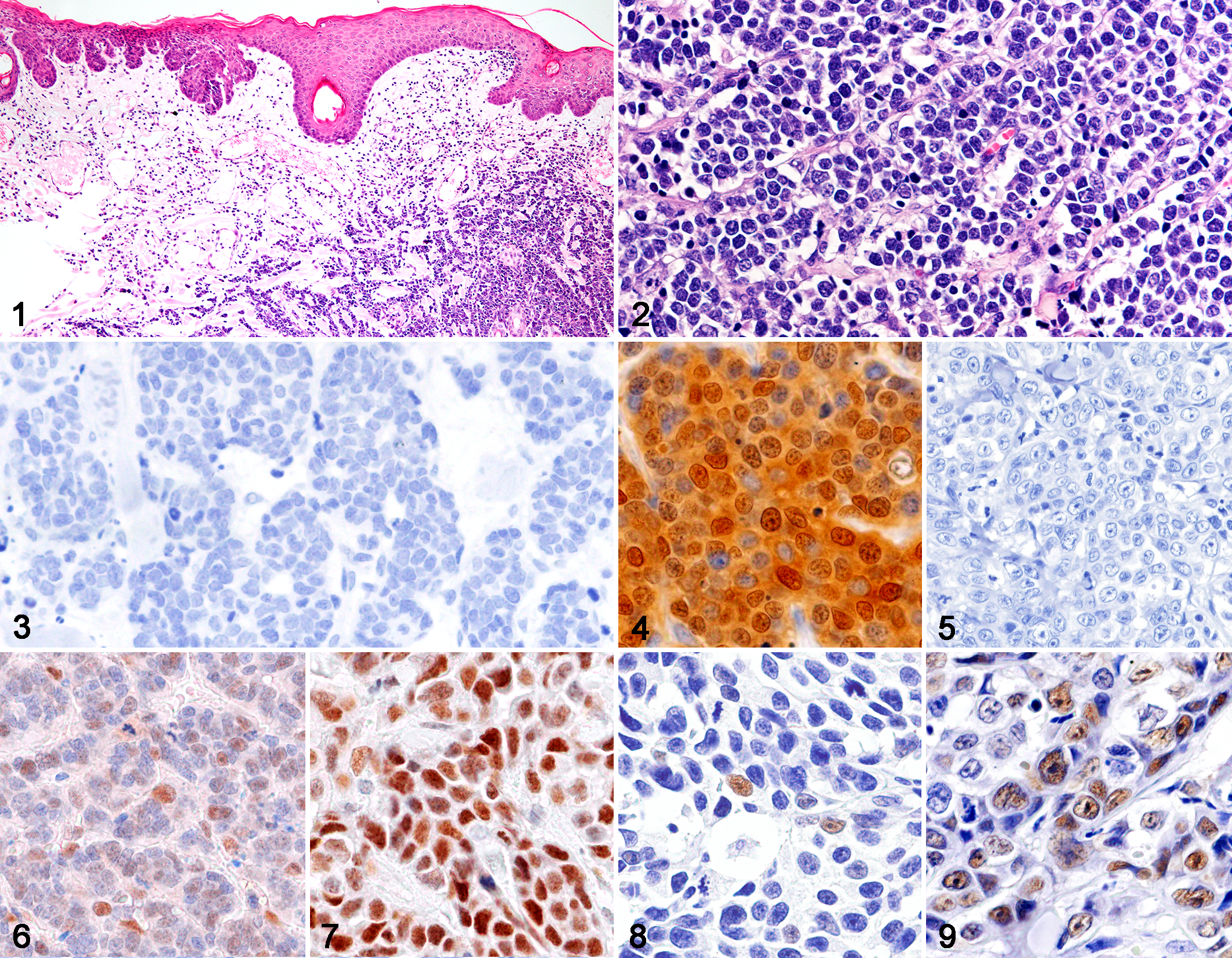
Merkel cell carcinoma, skin, cat.
Other proliferative cutaneous lesions were observed in 13/21 cases, including 11 viral plaques, 3 BISC, 1 SCC, and 1 BCC (Figs. 10–12). A combination of 2 different types of lesions was observed in 3 cases (cases 3, 7, and 21). The viral plaques and BISC were adjacent to MCC lesions; that is, they were found in the same tissue section. SCC and BCC were distant; that is, they were found in a different body location. The tissue sample of case 15 was collected from the subcutis and did not contain dermal tissue. Viral plaque was characterized by focal irregular epidermal hyperplasia, keratinocytes with vacuolated cytoplasm and shrunken nuclei known as “koilocytes,” clumped keratohyalin granules, and orthokeratotic hyperkeratosis (Fig. 10). The histopathological features of BISC were characterized by the focal proliferation of keratinocytes involving the epidermis and hair follicle infundibulum without invasion of the basement membrane, and dysplastic changes, such as an irregular cell arrangement, anisokaryosis, and the loss of nuclear polarity (Fig. 11). SCC and BCC infiltrated the deep dermis (Fig. 12). By immunohistochemistry, PV L1 antigens were detected in 10/11 viral plaques. Immunoreactivity for PV L1 antigens was observed in the nucleus of keratinocytes in the stratum granulosum and in keratin of the stratum corneum (Fig. 13). PV antigens were not detected in the neoplastic foci of BISC, SCC (Fig. 14), or BCC. Immunoreactivity for p16 was observed in all lesions, while the normal epidermis was negative for p16. Increased p16 immunoreactivity was confirmed in 9/11 viral plaques (Fig. 15), 3/3 BISC, 1/1 SCC (Fig. 16), and 1/1 BCC. The cellular positivity for p16 varied in viral plaques: <20% in 2/11, 20% to 50% in 5/11, and >50% in 4/11. Intermediate or strong p16 immunoreactivity within both nucleus and cytoplasm was observed in 9/11 viral plaques, 3/3 BISC, 1/1 SCC, and 1/1 BCC, and weak cytoplasmic immunoreactivity was observed in 2/11 viral plaques with less than 20% positive cells. Weak or intermediate pRb immunoreactivity was observed in 10/11 viral plaques (Fig. 17), 3/3 BISC, 1/1 SCC, and 1/1 BCC (Fig. 18). Decreased p53 immunoreactivity was observed in 11/11 viral plaques (Fig. 19), 3/3 BISC, and 1/1 BCC (Fig. 20).
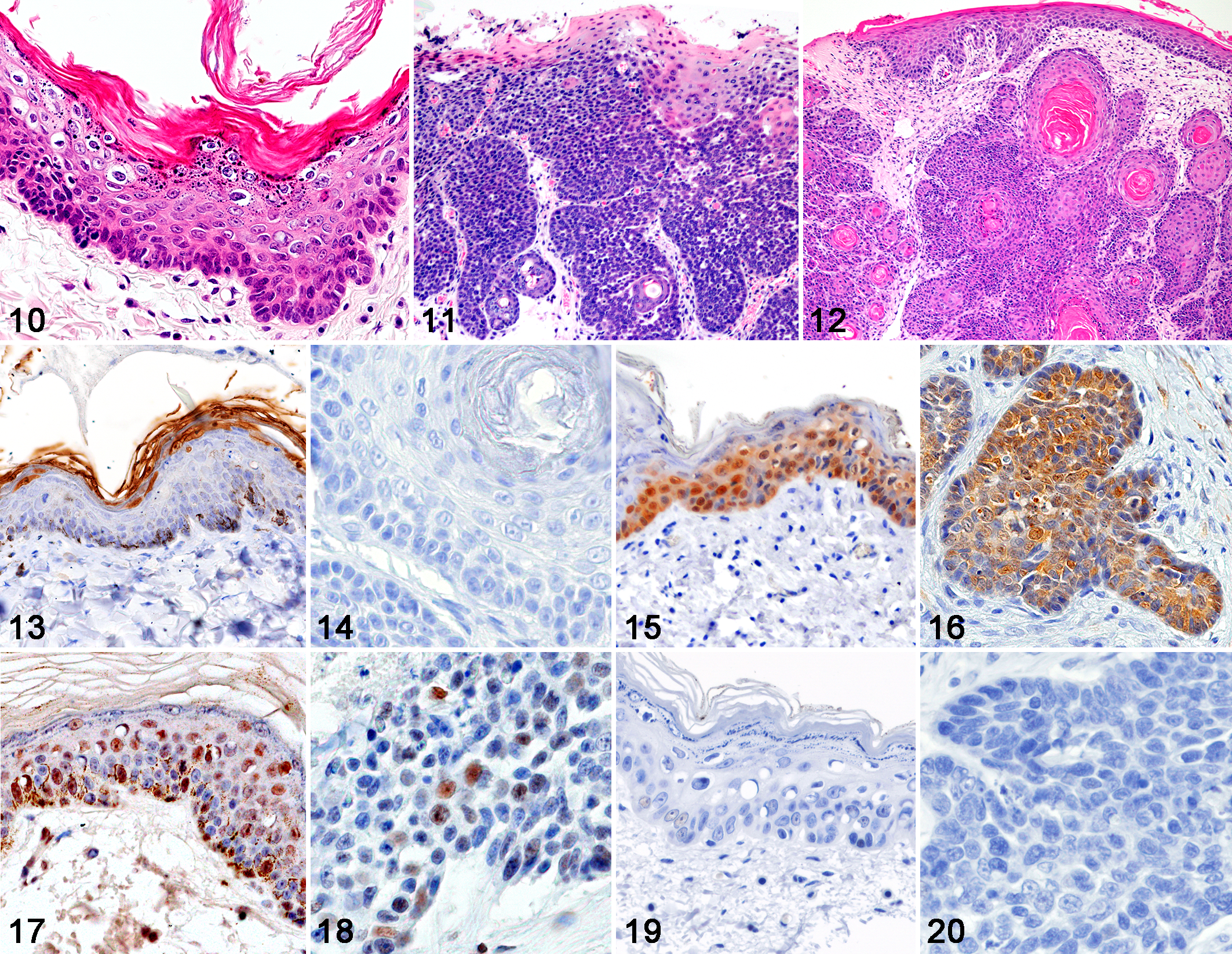
Other proliferative lesions associated with Merkel cell carcinoma, skin, cat.
PCR Detection and Full-Genome Sequencing of FcaPV2
In 20/21 samples, papillomaviral DNA was detected with 2 primer sets for FcaPV2 (Table 3, Fig. 21), and no positive results were obtained with the primer sets for FcaPV1, 3, 4, and 5. No amplification product was obtained from 1/21 sample (case 8) with any of the primer sets used in the experiment.
Detection of Felis catus papillomavirus by PCR and in situ hybridization.
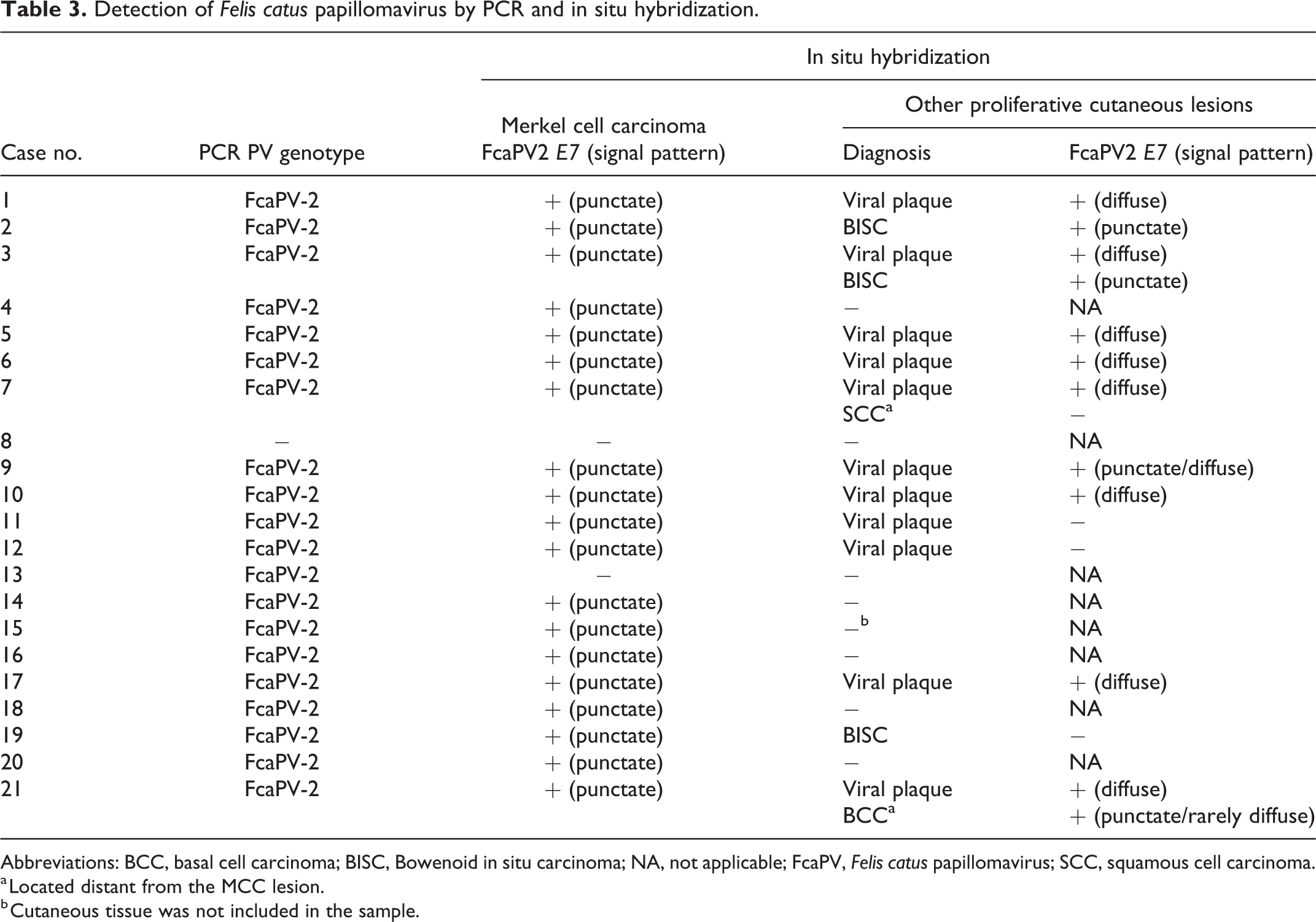
Abbreviations: BCC, basal cell carcinoma; BISC, Bowenoid in situ carcinoma; NA, not applicable; FcaPV, Felis catus papillomavirus; SCC, squamous cell carcinoma.
a Located distant from the MCC lesion.
b Cutaneous tissue was not included in the sample.
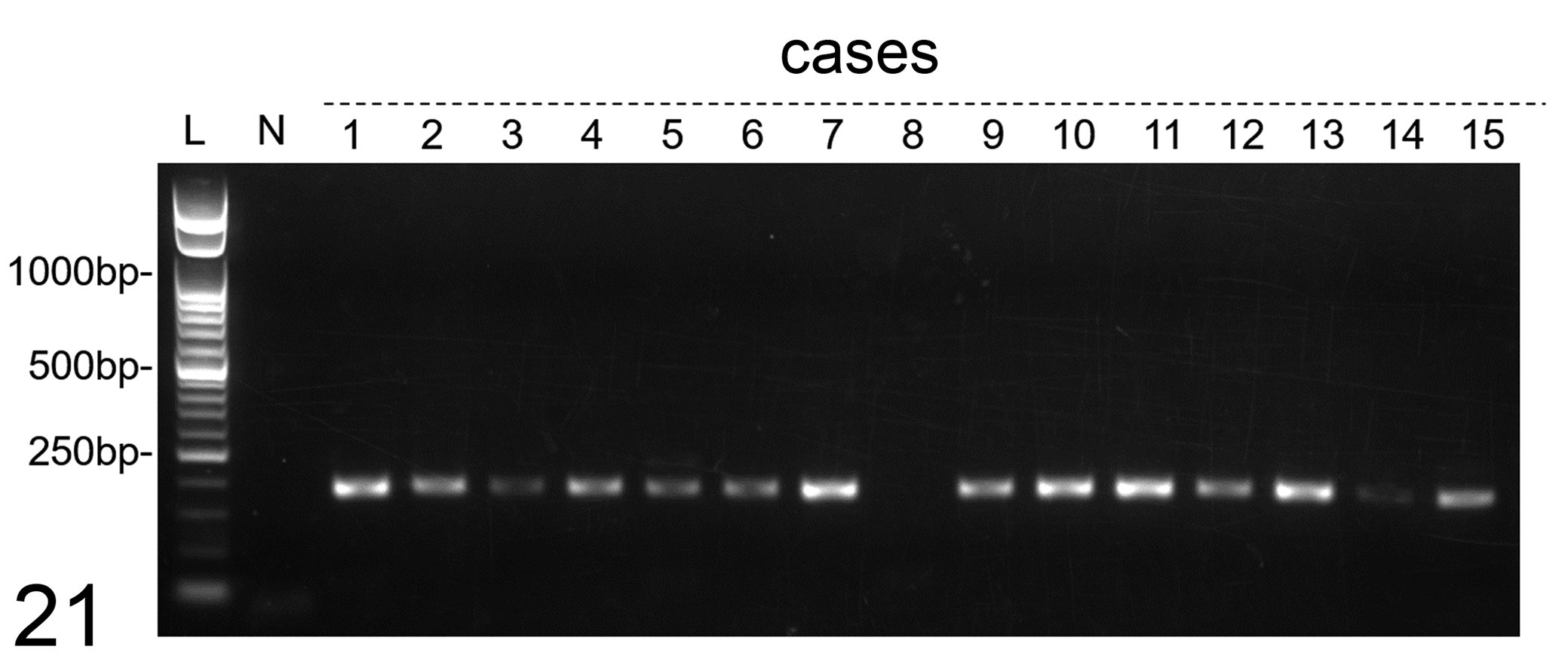
Gel electrophoresis of PCR amplicons for Felis catus papillomavirus type 2 E7. PCR-positive bands in cases 1 to 7 and 9 to 15, but not in case 8. Abbreviations: bp, base pairs; L, 50 bp ladder; N, negative control.
Gel electrophoresis results for the PCR products amplified with the FcaPV2 1732 (F)/4541 (R), FcaPV2 4474 (F)/7048 (R), and FcaPV2 6506 (F)/2700 (R) primer pairs confirmed the successful PCR amplification of the complete FcaPV2 genome in case 10 (data not shown). The full genome was 7899 bp in size and showed 99.8% nucleotide sequence homology (7880/7899 nt) to the reference FcaPV2 sequence (GenBank accession number: EU796884.1). Nucleotide sequence homologies based on each ORF were E1: 99.8% (1806/1809 nt); E2: 100% (1591/1593 nt); E6: 100% (417/417 nt); E7: 100% (288/288 nt); L1: 99.8% (1512/1515 nt); L2: 99.5% (1499/1506 nt); and LCR: 99.4% (663/667 nt). The predicted amino acid sequence was 100% identical to the reference FcaPV2, except for E2 (amino acid substitutions: T260A and A380T) and L2 (amino acid substitutions: M67T, L168V, I301F, W369F, and R491K). Four nucleotide substitutions in LCR were detected (G282A, G418A, C596T, and C608T). The full-genome sequencing results of the FcaPV2 clones of case 10 were 100% identical to those of the direct sequencing results. The complete sequence of FcaPV2 identified from MCC case 10 was deposited in the DDBJ/EMBL/GenBank database under accession number LC612600.
Sequencing Analysis of p16 Gene
In order to examine mutations of p16 in feline skin samples with MCC, 3 exons were amplified separately by PCR in 3 cases with increased p16 immunoreactivity, as well as in normal feline skin. PCR amplification of all 3 exons was successful, and subsequent sequencing analysis revealed no mutations or deletions in MCC cases compared to normal feline skin. When compared with feline p16 reference sequence (GenBank accession number: NM001290248.1), single nucleotide substitutions (c.108G>C, 131C>T) were found in the 3 MCC cases and normal feline skin, 9 which did not result in amino acid substitution.
In Situ Hybridization
The cellular localization of the FcaPV2 E7 gene was examined by CISH and FISH and the results obtained are summarized in Table 3. Positive hybridization signals were detected in 19/21 MCC by CISH. The hybridization signal pattern was characterized by punctate nuclear signals within MCC cells (Fig. 22). All 19 MCC with positive hybridization signals were also PCR-positive for FcaPV2. No hybridization signals were detected one PCR-positive MCC (case 13). The one PCR-negative MCC (case 8) was also negative by ISH. Punctate signals in MCC cells were also detected by FISH in 7/7 PCR-positive cases. Dual-fluorescence ISH and IHC revealed punctate hybridization signals (green) within the nuclei (blue) in MCC cells (Fig. 23). Positive hybridization signals were observed in other proliferative cutaneous lesions. Diffuse nuclear hybridization signals in keratinocytes of the stratum spinosum and stratum granulosum were observed in 8/11 viral plaques. Punctate signals were also detected in one of these cases (case 9; Fig. 24). Punctate nuclear signals, similar to the pattern of MCC, were noted in 2/3 BISC and 1/1 BCC (Fig. 25). No hybridization signals were found in normal skin around the lesions.
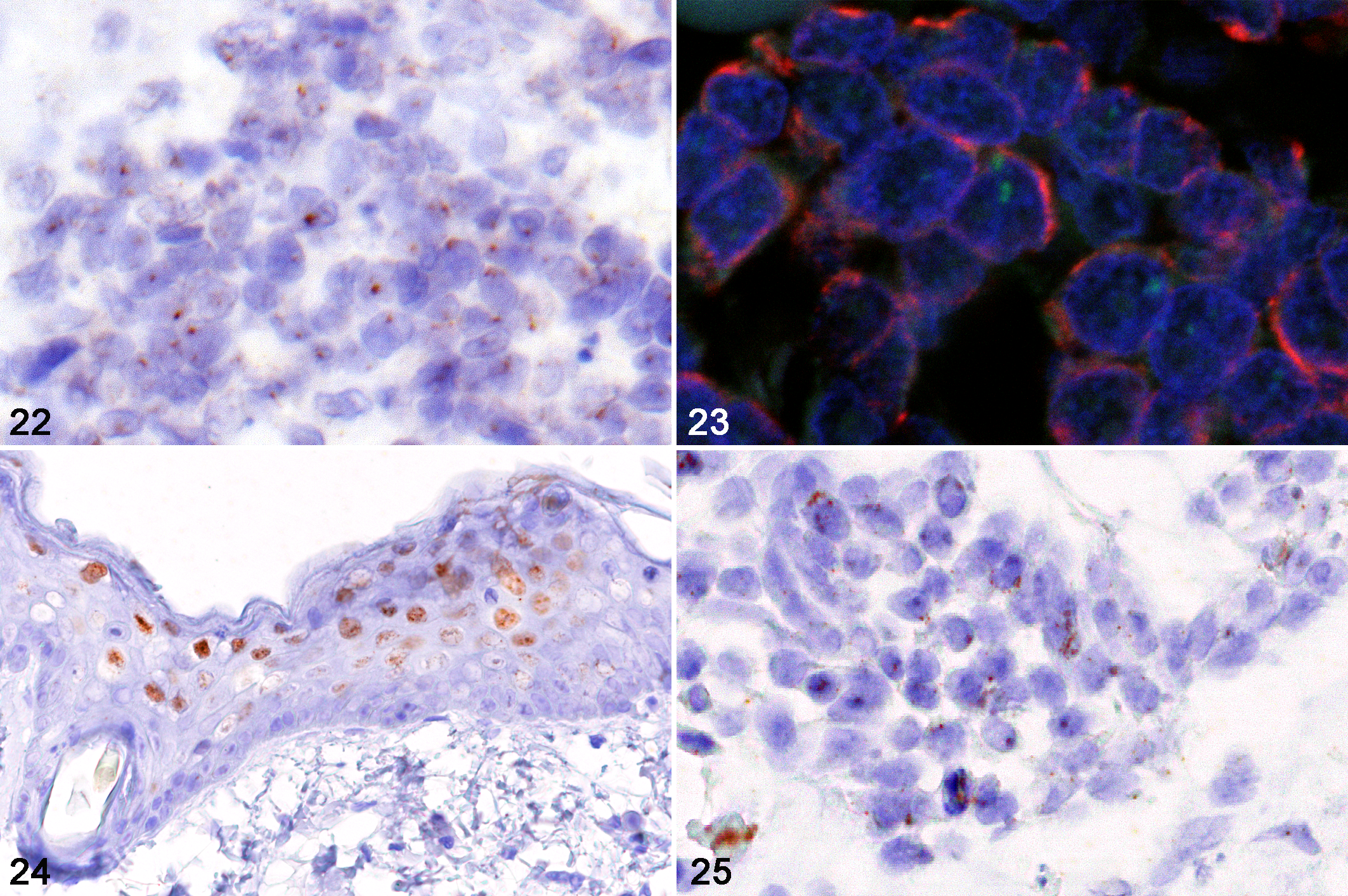
Skin, cat, in situ hybridization (ISH) for Felis catus papillomavirus type 2 E7.
Discussion
PV are highly species-specific and classified by their DNA sequence identities of the major capsid protein, L1 ORF. 57 To date, 6 types of FcaPV have been detected in various epithelial skin tumors of cats, but not in MCC. 6,35 Among these, FcaPV2 has been most frequently detected in viral plaque, BISC, and cutaneous SCC. 28,31 In the present study, FcaPV2 genes were detected in 20/21 (95%) MCC-containing feline skin samples by PCR. However, the detection of FcaPV genes in skin tumors does not necessarily mean that the virus infection is involved in tumorigenesis because FcaPV has also been detected in asymptomatic cats. 34,37 In a previous study, FcaPV2 DNA was detected from 39% of normal feline skin. 37 IHC has been used to identify molecular changes caused by PVs in tumor cells, 26 such as reduced pRb immunoreactivity due to the inactivation and degradation of pRb; 5,32,42 the absence of p53 immunoreactivity due to the degradation of p53; 1,45 and increased p16 immunoreactivity due to negative feedback secondary to pRb degradation. 13,24,32,33 In the present study, FcaPV-positive MCC, as well as other neoplastic lesions, exhibited these IHC features, suggesting that PV infection may be involved in their etiology.
Since p16 is typically upregulated in the nucleus following PV infection, nuclear p16 labeling with or without cytoplasmic labeling is of potential importance in predicting HPV infection. In human head and neck tumors, HPV-positive tumors have stronger p16 labeling in both the nucleus and cytoplasm than HPV-negative tumors. 66 In present study, FcaPV-positive MCC showed both nuclear and cytoplasmic immunoreactivity for p16 and were considered to have increased p16. Furthermore, sequence analysis of p16 revealed wild-type status in 3 cases with increased p16.
The life cycle of PV is intimately tied to the process of epithelial differentiation. 11,12,14,48 PV infects the basal cells of the epidermis and mucosa, and as these cells divide, they produce daughter cells that are pushed outwards toward the epithelial surface. In the parabasal and suprabasal layers, early viral genes (E1–E7) reprogram the cells to reenter the cell cycle and promote cell division. In the upper epithelial layers, the cells leave the cell cycle and late viral genes (L1–L2) are expressed to form viral capsid proteins. Newly formed infectious virions are eventually released in the stratum corneum during desquamation. The integration of viral genes into the host genome is a terminal event for the viral life cycle and a critical step in the tumorigenesis of PV-induced neoplasms. 14,43,61 Integration results in the increased expression and stability of transcripts encoding the viral oncogenes E6 and E7. The E6 and E7 proteins inactivate and/or degrade p53 and pRb, respectively, resulting in neoplastic progression. In humans, ISH signal patterns for HPV-16 DNA in cervical intraepithelial neoplasia (CIN) and SCC have been grouped into 3 types and are associated with the physical status of HPV in infected cells, that is, episomal or integrated forms. 7,18 A type 1 signal is diffuse and present throughout the nucleus; a type 2 signal is punctate; and type 3 is a combination of types 1 and 2. A type 1 signal represents an episomal virus, type 2 an integrated virus, and type 3 a combination of both. A previous study reported that CIN I lesions exhibited type 1 signals, while CIN II/III and SCC lesions showed type 2 and rarely type 3 signals. 15 In the present study, ISH for FcaPV2 E7 revealed type 2 signals within the tumor cells of MCC, BISC, and BCC, and type 1 signals within the keratinocytes of viral plaque. These results implicate the integration of FcaPV2 genes in feline MCC.
The full-genome sequencing of FcaPV2 in case 10 revealed 99.8% identity with the reference FcaPV2 (GenBank accession number: EU796884.1). Five amino acid substitutions were detected in L2 (M67T, L168V, I301F, W369F, and R491K). In HPV16, L2 amino acid residues from 64 to 73 (SGTGGRTGYI) are one of the neutralization epitopes, 39,59 and L2 residues from 301 to 400 are required for the downregulation of E2-dependent transcriptional activation. 59 Further molecular studies are needed to clarify the functional difference associated with the amino acid substitutions of FcaPV2 L2 in feline MCC.
The viral L1 antigen is a major component of the capsomeres of PVs. Expression of L1 protein depends on transcriptional factors that are expressed during the maturation process of PV-infected basal epithelial cells to superficial epithelial cells. 2,11,12,29 It has been demonstrated that L1 expression is decreased in conjunction with viral integration in HPV-positive cervical carcinoma and FcaPV-positive SCC, and negative immunoreactivity of PV capsid antigen is associated with malignant progression. 63,65 In present study, PV L1 antigen was immunohistochemically detected in the stratum granulosum and stratum corneum of viral plaque. These results suggest that viral DNA encapsidation and the release of newly formed infectious virions occur in feline viral plaque, but not in neoplastic lesions, including MCC, SCC, BCC, and BISC. Furthermore, FcaPV2 DNA was detected by ISH in MCC, suggesting possible FcaPV gene integration.
Among the 21 MCC cases examined, the results of case 8 differed from those of the other cases as follows: (1) no proliferative lesions besides MCC, (2) immunohistochemically negative for p16 and strongly positive for pRb and p53, and (3) PCR- and ISH-negative for FcaPV genes. Negative p16 and strong pRb immunoreactivities are strong indicators of HPV negativity in human SCC. 13 Therefore, it is unlikely that FcaPV infection was associated with the development of MCC in this case (case 8). These results also indicate that there are more than one molecular pathway for the tumorigenesis of MCC in cats. Mutations within the p53 gene prevent normal p53 function and result in stable p53 that is immunohistochemically detectable. 17 In humans, p53 mutation and increased p53 are common features of HPV-negative SCC and MCPyV-negative MCC. 25,47,53 Since MCPyV-negative MCC exhibits a high mutation burden in the Rb1 and p53 genes, which are associated with a UV-induced DNA damage signature, UV exposure is considered to be the major cause of MCPyV-negative MCC. 4,62 These findings from human MCC suggest that mutations in p53 due to UV exposure may have contributed to the development of FcaPV-negative MCC in case 8.
There has been substantial debate in the literature about the cells of origin of human MCC. Merkel cells are unlikely to be the origin of MCC because MCC lesions are located in the dermis and/or subcutis, while Merkel cells are present in the epidermis. 4,46,51 Moreover, Merkel cells appear to be terminally differentiated with limited proliferative potential and a slow turnover. 58 Feline MCC developed in the dermis and/or subcutis, and not in the epidermis. Therefore, intact Merkel cells in the epidermis are not likely the origin of MCC. 19 Accumulating evidence suggests that the PV infection of epithelial stem cells in the skin or mucosa is necessary for the development of PV-induced neoplasms. 11,14 The bulge region of the hair follicle is a well-characterized region at which epithelial stem cells reside. Virions of HPV are considered to gain access to epithelial stem cells through either a wound or the hair follicle. 14 The integration of FcaPV genes and their involvement in the tumorigenesis of feline MCC cells strongly support the cutaneous stem cells residing in hair follicles as the likely origin of feline MCC, which has been suggested based on the immunohistochemical characteristics of tumor cells. 19 However, the mechanisms by which the cells of origin of feline MCC acquire neuroendocrine phenotypes in the process of neoplastic transformation are unclear. In human MCC, the inhibition of pRb by the MCPyV LT antigen or pRb and p53 mutations by UV irradiation are considered to induce neuroendocrine differentiation in MCPyV-positive MCC and MCPyV-negative MCC, respectively. 51,55,60
Conclusion
The present study suggests FcaPV2 as a cause in many cases of feline MCC. Similar to other PV-induced tumors, viral genome integration and the inhibition of pRb and p53 by viral oncogenes participate in the tumorigenesis of feline MCC. The relationship between feline MCC and FcaPV may explain why MCC is more likely to occur in cats than in other animals.
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211045440 - Involvement of Felis catus papillomavirus type 2 in the tumorigenesis of feline Merkel cell carcinoma
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211045440 for Involvement of Felis catus papillomavirus type 2 in the tumorigenesis of feline Merkel cell carcinoma by Soma Ito, James K. Chambers, Ayumi Sumi, Nanako Yamashita-Kawanishi, Tetsuo Omachi, Takeshi Haga, Hiroyuki Nakayama and Kazuyuki Uchida in Veterinary Pathology
Footnotes
Declaration of Conflicting Interests
The author(s) declares no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
