Abstract
Chlamydia spp are reported to causes systemic disease in a variety of hosts worldwide including few reports in crocodilians. Disease presentations vary from asymptomatic to fulminant disease, some of which are zoonotic. The aim of this study was to describe the pathological, immunohistochemical, and molecular findings associated with the occurrence of a previously unreported Chlamydia sp infection causing a major mortality event in farmed American alligators (Alligator mississippiensis). The outbreak presented with sudden death in juvenile alligators mainly associated with necrotizing hepatitis and myocarditis, followed by the occurrence of conjunctivitis after the initial high mortality event. The widespread inflammatory lesions in multiple organs correlated with intralesional chlamydial organisms identified via immunohistochemistry and confirmed by 23S rRNA-specific real-time quantitative polymerase chain reaction (qPCR) for Chlamydiaceae bacteria. By sequencing and phylogenetic analysis of the OmpA gene, this uncultured Chlamydia sp grouped closely with Chlamydia poikilothermis recently described in snakes. This study highlights the significance of such outbreaks in farmed populations. Enhanced epidemiological monitoring is needed to gain further insight into the biology of Chlamydia sp in alligators, disease dynamics, risk factors, and role of carrier animals.
Bacteria within the order Chlamydiales are gram-negative, obligate intracellular pathogens of worldwide distribution that affect a wide range of host species including humans, domesticated and wild mammals, reptiles, amphibians, and fish.3,30 The Chlamydiales have a unique biphasic developmental cycle characterized by infectious, extracellular forms (elementary bodies, EB) measuring approximately 200 nm in diameter; and intracellular, metabolically active and replicative forms (reticulate bodies, RB), which can be visualized as intracytoplasmic inclusions within infected cells, measuring up to 1 µm in diameter. Reticulate bodies subsequently differentiate into EB, which exit from host cells via lysis or extrusion and infect other cells. 1 The strict dependency on host cell metabolites for survival and replication has a significant impact in the diagnostic approaches, which often include either isolation in susceptible cell lines or inoculation in embryonated chicken eggs. 3 Despite available culture systems, some Chlamydia species remain unculturable and, therefore, molecular diagnostics in combination with histopathology are frequently necessary to reach a diagnosis.3,22
Chlamydial infections affect a wide range of animal species, with some chlamydial species, such as Chlamydia trachomatis, Chlamydia pneumoniae, Chlamydia abortus, Chlamydia felis, Chlamydia psittaci, and Chlamydia pecorum, causing well-characterized diseases. 3 These infections mostly vary from asymptomatic to fulminant disease, with common manifestations including conjunctivitis, pneumonia, abortion, inflammation within the reproductive tract, encephalitis, enteritis, hepatitis, and polyarthritis. Asymptomatic infections are characterized by persistence of the microorganism and shedding with no clinical disease. In reptiles, chlamydiosis has been reported in both free-ranging and captive animals, with C. pneumoniae being the most frequently reported.2,3,11–16,19,20,25–29,31,32 Chlamydia-associated lesions reported in this group of vertebrates include necrotizing to granulomatous myocarditis, hepatitis, splenitis, enteritis, conjunctivitis, and pneumonia;3,12–15,19,26,27,32 however, C. pneumoniae and other potentially novel, poorly characterized Chlamydia species have also been identified in clinically normal snakes and turtles.16,20,25 Recently, 2 novel Chlamydia species, Chlamydia serpentis and Chlamydia poikilothermis (most closely related to C. pneumoniae and C. psittaci, and C. caviae and C. abortus, respectively), were isolated from captive snakes. 28 Regarding the occurrence of chlamydiosis in members of the order Crocodilia, outbreaks have been described in different species of crocodiles.13,14,26,32 Most recently, fulminant systemic disease was identified in an outbreak in juvenile Siamese crocodiles (Crocodylus siamensis) associated with a Chlamydia species closely related to C. caviae and C. psittaci. 32 The aim of this study was to describe the pathological, immunohistochemical, and molecular findings of a previously unreported, major mortality event associated with a Chlamydia sp infection in farmed American alligators in Louisiana.
Materials and Methods
Animals Included in the Study and Clinical History
An outbreak associated with sudden mortalities of unknown cause was reported from an alligator farm in the state of Louisiana, USA during the spring months. Within 48 hours, a total of 40, 1.5-year-old American alligators (Alligator mississippiensis) suddenly died. Of these, 4 animals (Group 1) were submitted to the Louisiana Animal Disease Diagnostic Laboratory (LADDL), Louisiana State University School of Veterinary Medicine (LSU-SVM) for complete postmortem examination as part of the outbreak investigation. Approximately 10 days after the initial sudden mortalities were identified, 4 additional animals (Group 2) with gross evidence of conjunctivitis were submitted for postmortem examination (Table 1). The conjunctivitis was identified as a new clinical sign during the course of this outbreak, which was not evident earlier. Mortalities were still occurring in the population at this point. By day 10 of the outbreak, over 100 mortalities were estimated, with approximately 5000–10,000 animals at risk. No changes in husbandry were reported prior to the outbreak.
Cases examined at postmortem from an acute mortality event in American alligators.
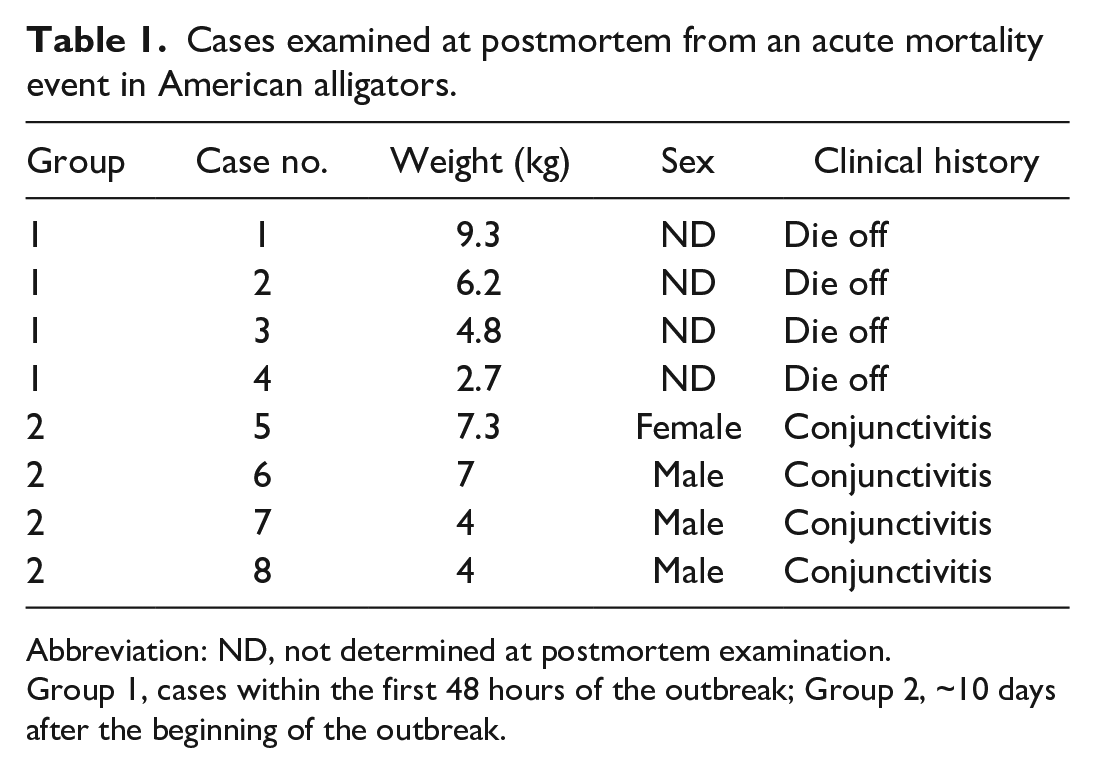
Abbreviation: ND, not determined at postmortem examination. Group 1, cases within the first 48 hours of the outbreak; Group 2, ~10 days after the beginning of the outbreak.
Postmortem Examination and Histopathology
Postmortem examinations were performed in 8 animals submitted to the LADDL (Table 1). Fresh pooled liver specimens from all 4 animals in Group 1 and individually from all 4 animals in Group 2 were aseptically collected and submitted for real-time quantitative polymerase chain reaction (qPCR) and bacterial (aerobic) culture. Conjunctival specimens from all individual alligators in Group 2 were submitted for qPCR. Samples from all organs were fixed in 10% neutral buffered formalin for 24–48 hours.
Following 24–48 hours of fixation, tissues (Group 1 and Group 2: lung, liver, spleen, kidneys, heart, stomach, intestines, cloaca, and brain; additional tissues in Group 2: eye and conjunctiva) were embedded in paraffin and sections of formalin-fixed paraffin-embedded tissues (4 μm) were stained with hematoxylin and eosin, Gimenez, Machiavello, acid-fast, and Gomori’s methenamine silver stains according to standard laboratory procedures prior to histological evaluation.
Bacterial Culture
Samples from the liver were inoculated onto blood agar (tryptic soy agar with 5% sheep blood), chocolate agar, and MacConkey agar plates (Remel, San Diego, Calif.). Inoculated blood and chocolate agar plates were aerobically incubated at 37°C with 5% CO2 for the first 24 hours and without CO2 for the next 48 hours. Inoculated MacConkey agar plates were incubated at 37°C for up to 72 hours. Salmonella-specific culture from the intestines of Group 1 alligators was performed following standard procedures. Salmonella serotyping was performed at the National Veterinary Services Laboratory, US Department of Agriculture, Ames, Iowa. For Chlamydia sp culture, a 10% tissue homogenate was prepared in 1× phosphate-buffered saline pH 7.4 and inoculated via yolk sac route in 5- to 12-day-old, specific pathogen-free embryonated chicken eggs following standard laboratory procedures. Inoculated eggs were incubated for 7 days at 37°C and subjected to a second blind passage.
Immunohistochemistry
Sections from the lung, liver, spleen, kidneys, heart, stomach, intestines, cloaca, encephalon, eye, and conjunctiva were evaluated by immunohistochemistry (IHC) using a genus-specific mouse monoclonal antibody to C. trachomatis, C. pneumoniae and C. psittaci lipopolysaccharide (LPS; 1681, ViroStat Inc., Portland, Maine). Briefly, 4-μm sections of formalin-fixed paraffin-embedded tissues were mounted on positively charged Superfrost Plus slides (VWR, Radnor, Pa.) and subjected to IHC using the automated BOND-MAX and the Polymer Refine Detection kit (Leica Biosystems, Buffalo Grove, Ill.). Following automated deparaffinization, heat-induced epitope retrieval (HIER) was performed using a ready-to-use EDTA-based buffer (pH 9.0; Leica Biosystems) at 100°C for 20 minutes before incubation with anti-Chlamydia LPS monoclonal antibody diluted 1:50 in Primary Antibody Diluent (Leica Biosystems). Sections were then incubated with the primary antibody for 30 minutes at room temperature, followed by an anti-mouse IgG coupled with horseradish peroxidase (Leica Biosystems) for 20 minutes at room temperature. 3,3′-Diaminobenzidine tetrahydrochloride (DAB) was used as the chromogen (10 minutes), and counterstaining was performed with hematoxylin. Slides were mounted with a permanent mounting medium (Micromount®, Leica Biosystems). A C. psittaci-infected liver specimen was used as a positive control and tissues incubated without the primary antibody were used as negative controls. Antigen abundance was semiquantitatively scored as – (none), 1 (sporadic positive target cells per 40× field examined), 2 (approximately 10%–20% positive target cells per 40× field examined), 3 (approximately 20%–50% of positive target cells per 40× field examined), and 4 (>50% positive cells per 40× field examined).
qPCR for Chlamydiaceae and West Nile Virus
A total of 75 mg of liver (pooled for Group 1 alligators and individually for Group 2 alligators) or conjunctiva (individually only for Group 2 alligators) were homogenized in 240 μl of 1× phosphate-buffered saline using a TissueLyser II (Qiagen, Valencia, Calif.). DNA from tissue homogenates was extracted using the QIAamp DNA Mini kit (Qiagen) in a QIAcube (Qiagen) following the manufacturer’s instructions. Elutions (200 μl) were stored at −80°C until processed.
For qPCR, a primer and probe set targeting the 23S rRNA gene and specific for all known Chlamydiaceae 10 was used (Table 2). The reaction was set up using the Path-ID™ Multiplex One-Step RT-PCR kit (ThermoFisher Scientific) following the manufacturer’s recommendations. The 25 μl reaction contained 12.5 μl of 2× Multiplex RT-PCR Buffer, 2.5 μl of Multiplex Enzyme Mix, 200 nM TaqMan® fluorogenic probe, 1 μM of each primer, 3.5 μl of nuclease-free water, and 5 μl of template DNA. Amplification was carried out in a Cepheid Smartcycler® (Cepheid, Sunnyvale, Calif.). The program included 20 minutes at 48°C, 10 minutes at 95°C, followed by 40 cycles at 95°C for 15 seconds (denaturation) and 58°C for 1 minute (combined annealing/extension). For West Nile virus (WNV), a primer-probe set targeting the envelope gene (E) was used as described previously. 18 The 25 μl reaction contained 12.5 μl of 2× Multiplex RT-PCR buffer, 2.5 μl of Multiplex Enzyme Mix, 200 nM TaqMan® fluorogenic probe, 1 μM of each primer, 3.5 μl of nuclease-free water, and 5 μl of template DNA. Amplification was carried out in a Cepheid Smartcycler® (Cepheid). The program included 20 minutes at 48°C, 10 minutes at 95°C, followed by 40 cycles at 95°C for 15 seconds (denaturation) and 60°C for 1 minute (combined annealing/extension).
Primers and probe used for Chlamydiaceae 23S rRNA qPCR and primers used for standard PCR and sequencing of Chlamydia sp 16S rRNA and OmpA genes.
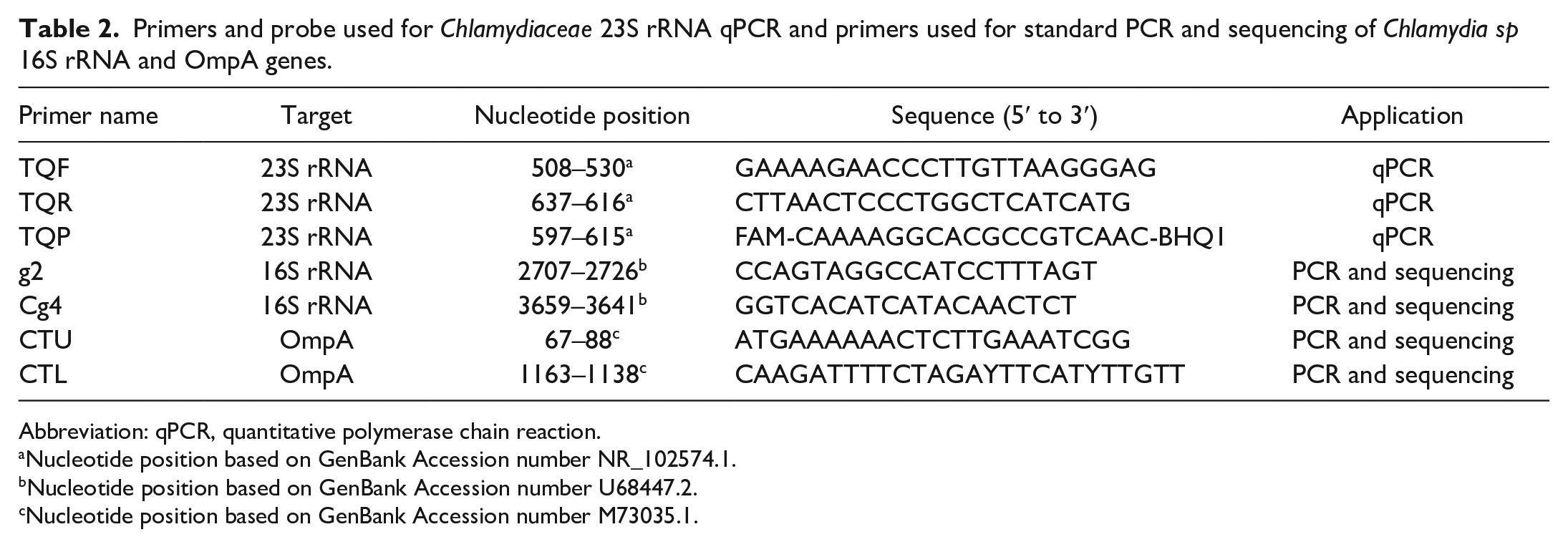
Abbreviation: qPCR, quantitative polymerase chain reaction.
Nucleotide position based on GenBank Accession number NR_102574.1.
Nucleotide position based on GenBank Accession number U68447.2.
Nucleotide position based on GenBank Accession number M73035.1.
16S Ribosomal and Outer Membrane Protein A (OmpA) Gene Sequencing
Partial sequencing of the 16S ribosomal and OmpA genes was performed using the high-fidelity AmpliTaq Gold™ DNA Polymerase with Buffer I (ThermoFisher Scientific). A 50 μl of reaction mixture was composed of 5 μl of 10× Buffer I, 1 μl of dNTP (2.5 mM), 1.25 μl of 16S or OmpA-specific forward and reverse primers (Table 2, 20 μM, final concentration 0.5 μM7,8), 0.5 μl of AmpliTaq Gold™ DNA Polymerase, 36 μl of nuclease-free water, and 5 μl of template. The cycling conditions included a PCR activation step at 95°C for 5 minutes, 40 cycles of denaturation (95°C for 30 seconds), annealing (58°C for 30 seconds), and extension (72°C for 1 minute); and a final extension at 72°C for 7 minutes. Polymerase chain reaction products were gel-purified using the QIAquick® Gel Extraction kit (Qiagen) according to the manufacturer’s recommendations. DNA was submitted for Sanger sequencing to GeneLab, Louisiana State University School of Veterinary Medicine. Both DNA strands of partial 16S and OmpA amplicons were sequenced using the primers specified in Table 2.
Sequence and Phylogenetic Analysis
Sequence analysis was performed using Geneious R7 (Biomatters Inc., Newark, N.J., USA). Multiple sequence alignment of nucleotide sequences was performed using MUSCLE 9 and phylogenetic analysis was performed using MEGAX. 17 The phylogenetic tree was constructed using the Neighbor-joining method with 1000 bootstrap replicates. Waddlia chondrophila WSU 86-1044 OmpA gene was used as an outgroup (GenBank accession number CP001928.1). The sequences generated in this study were deposited in GenBank under accession numbers OL880546 and OL956539.
Statistical Analysis
Correlation analysis between antigen scores and lesion severity scores was performed using the non-parametric Spearman’s rank correlation coefficient (ρ). A P-value <.05 was considered significant.
Results
Postmortem and Histological Findings
All 4 alligators in Group 1 were in good body condition at the time of postmortem examination. Postmortem findings are summarized in Figs. 1–4. Macroscopically, all livers had an overall accentuated, orange-tinged lobular pattern. In one alligator, extensive areas of the hepatic parenchyma were gray and firm while the remaining parenchyma was brown with a soft texture and containing multiple yellow pinpoint foci. The capsular surface of the liver was discolored yellow in all Group 1 animals (Fig. 1). The lungs in all Group 1 animals oozed a clear, yellow mucoid exudate. The myocardium of 2 animals had scattered white pinpoint foci on the epicardial surface (Fig. 4). Moderate to severe pericardial effusion was noted in 3 Group 1 animals, which also had multifocally reddened intestinal mucosa and white to yellow fibrinous exudate in the intestinal lumen. No gross changes were noted in the eyes of these animals.
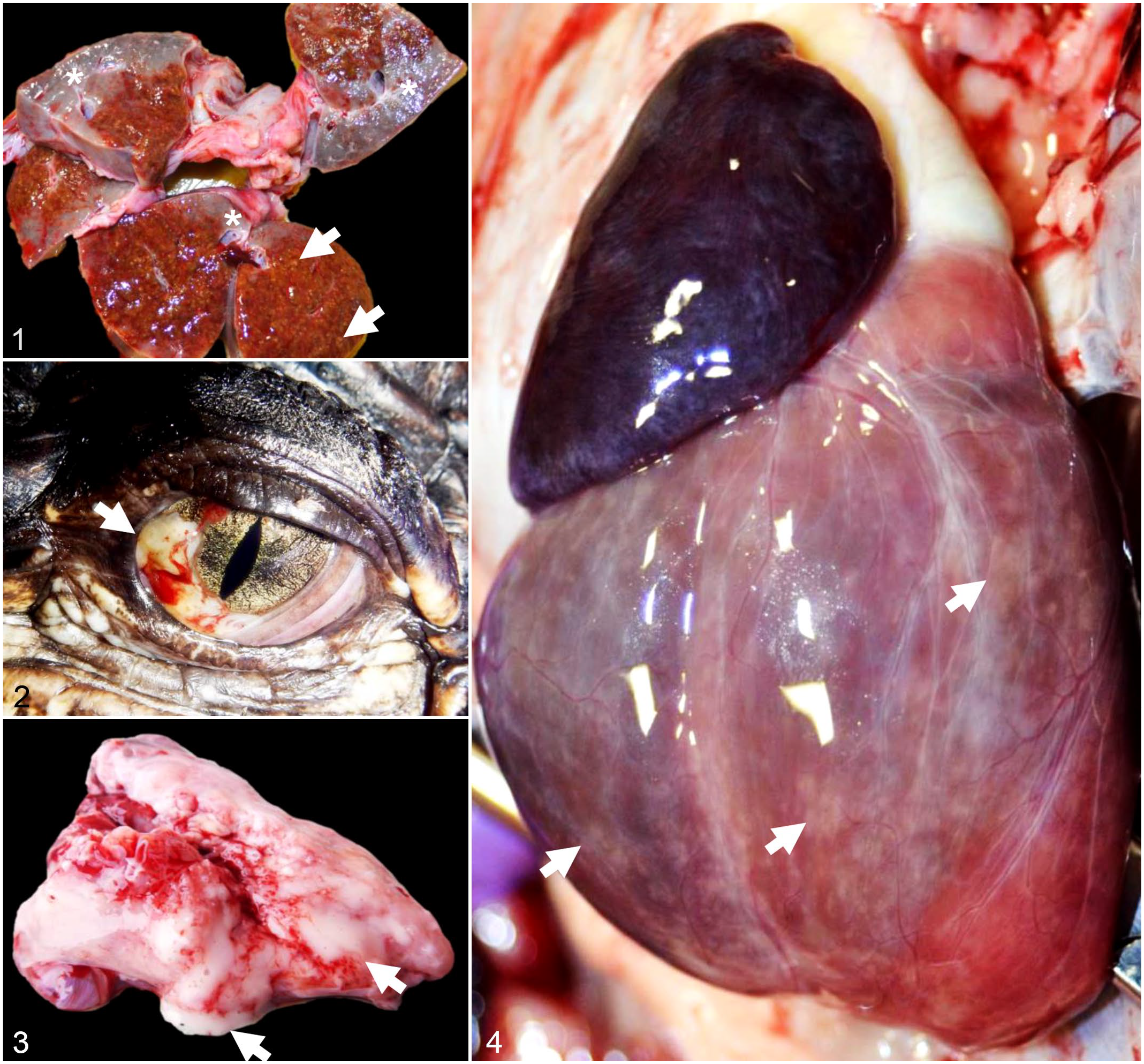
Chlamydial infection, farmed American alligators.
Of the 4 alligators in Group 2 (with clinical signs of conjunctivitis), full postmortem examinations were performed in 2 animals while tissues from field necropsies were submitted for the other two. The examined animals were in good body condition. There was bilaterally reddened conjunctiva with mucoid exudate (Fig. 2) in 4 of 4 cases. There were multiple pinpoint white foci throughout the myocardium, spleen, and liver, and moderate to severe coelomic and pericardial effusions. In 2 of 4 animals, the lungs oozed abundant frothy white fluid on cut surface (Fig. 3).
The histologic changes identified in the liver, heart, lungs, and spleen were similar in all cases from both groups with variation in the lesion severity and distribution. The alterations in the intestinal tract mostly affected alligators in Group 1 while examination of the eyes was restricted to those alligators in Group 2. Histologic findings are summarized in Table 3.
Histological findings associated with chlamydiosis in farmed American alligators (Group 1, n = 4; Group 2, n = 4).
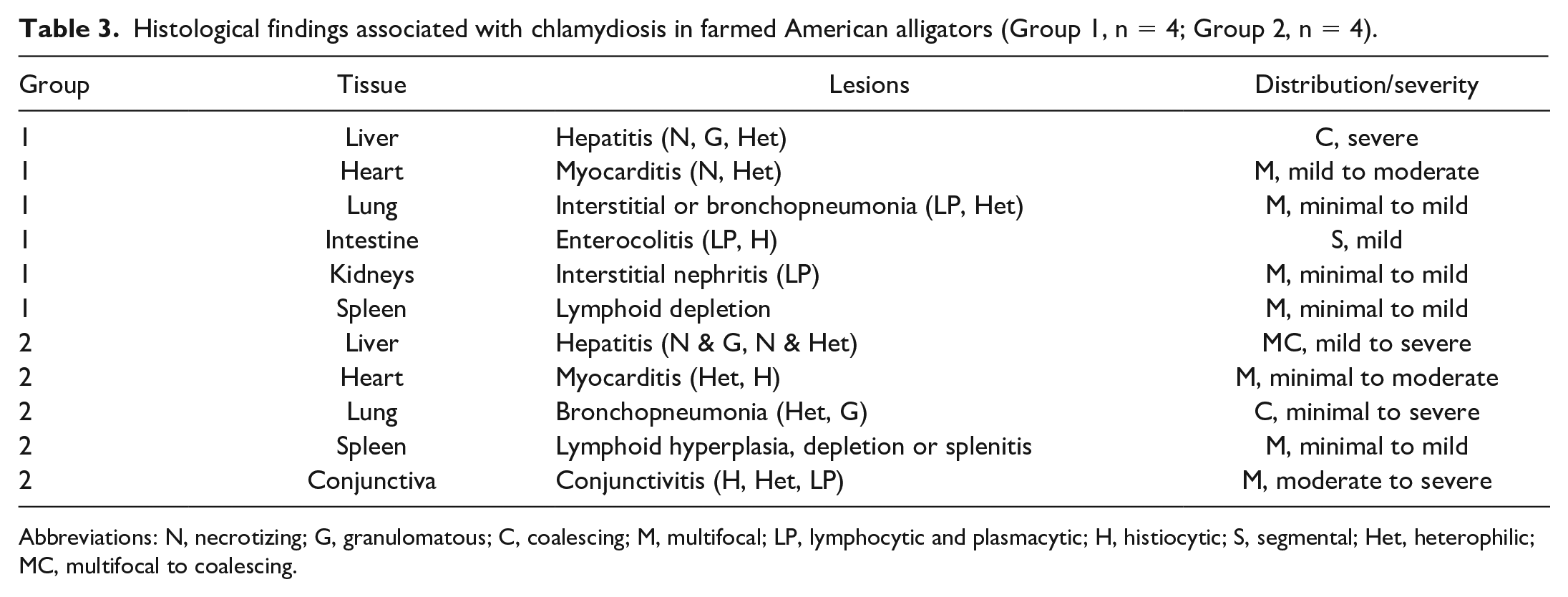

Abbreviations: N, necrotizing; G, granulomatous; C, coalescing; M, multifocal; LP, lymphocytic and plasmacytic; H, histiocytic; S, segmental; Het, heterophilic; MC, multifocal to coalescing.
Group 1
Severe coalescing necrotizing, heterophilic, and granulomatous hepatitis (Figs. 5–7) and mild to moderate multifocal necrotizing and heterophilic myocarditis (Fig. 8) were the prevailing lesions in Group 1 (4/4). Other lesions included moderate multifocal heterophilic bronchopneumonia (1/4; Fig. 9), interstitial lymphoplasmacytic pneumonia (3/4), moderate splenic lymphoid depletion (4/4), minimal to mild lymphoplasmacytic interstitial nephritis (3/4), and mild segmental lymphoplasmacytic and histiocytic enterocolitis (4/4) with rare (acute) vascular thrombosis (1/4). Intrahepatocellular granular chlamydial inclusions ranged from sporadic to frequent in sections stained with Machiavello, Gimenez, or hematoxylin and eosin (Fig. 7), but were not microscopically identifiable in other tissues examined (lung, spleen, kidneys, heart, stomach, intestines, cloaca, brain, eye, and conjunctiva). No acid-fast or argentaffin-positive organisms were detected.
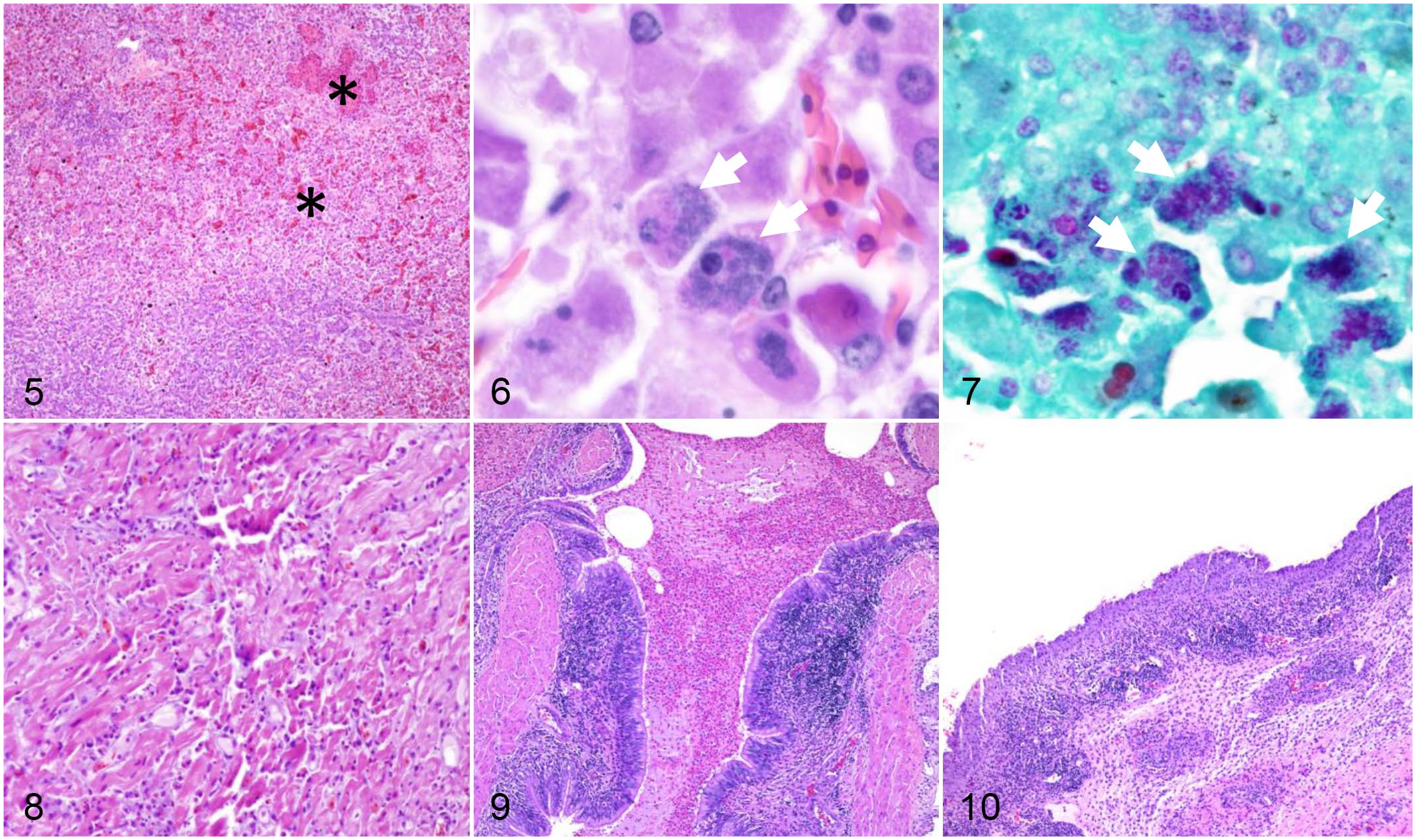
Chlamydial infection, alligators.
Group 2
Overall, histologic alterations in this group were mostly subacute rather than acute as noted for alligators in Group 1. The eyes were the commonly affected organ among animals within this group. Specifically, lesions included moderate to severe multifocal lymphoplasmacytic, histiocytic, or heterophilic and erosive conjunctivitis and keratoconjunctivitis (Fig. 10), and mild diffuse heterophilic, histiocytic, and lymphoplasmacytic anterior uveitis or panuveitis (4/4). Alterations in the liver were less severe compared to alligators in group 1, ranging from mild to moderate, and characterized by multifocal necrotizing and heterophilic to granulomatous hepatitis (4/4). Similarly, while myocarditis was a feature in alligators from group 2, it ranged from minimal to moderate and was mostly heterophilic to histiocytic (4/4). There was also evidence of bronchopneumonia (heterophilic to granulomatous) of variable severity (2/4). Changes in the spleen were variable (4/4) and included prominent reactive lymphoid hyperplasia (2/4), mild lymphoid depletion (1/4), or moderate multifocal heterophilic splenitis (1/4).
Further Testing
Chlamydiaceae DNA was successfully amplified from the liver of Group 1 and Group 2 alligators as well as from conjunctival specimens of Group 2 alligators. In Group 1 alligators, aerobic bacterial cultures from the liver were negative, but Salmonella sp (serotype III_48:z52,z53) was cultured from the intestinal content of 2 of 4 alligators which also showed gross intestinal lesions as described above. Among Group 2 alligators, moderate growth of hemolytic Escherichia coli and Salmonella sp (not serotyped) were obtained from the liver, heart, and spleen in 1 of 4 alligators. Chlamydia-specific cultures from the liver and RT-qPCR for WNV (performed on a pool of tissues) were negative in all animals.
Immunohistochemical Findings
Sections from multiple tissues were evaluated by IHC using a cross-reactive genus-specific Chlamydia LPS monoclonal antibody (Figs. 11–16 and Supplemental Figs. S1–S3). Distribution of chlamydial antigen is shown in Fig. 17. In Group 1, Chlamydia antigen was most abundant in the liver (within the cytoplasm of hepatocytes and histiocytes; Figs. 11 and 12), heart (within histiocytes in areas of cardiomyocyte necrosis; Fig. 13), spleen (within histiocytes; Fig. 14), and intestine (within histiocytes and enterocytes; Fig. 15 and Supplemental Fig. S1). In Group 2, Chlamydia antigen was less frequent in scattered histiocytes in the lungs (Supplemental Fig. S2), lamina propria of the stomach, renal glomeruli and interstitium, and sporadic renal tubular epithelial cells (Supplemental Fig. S3). Chlamydia sp antigen was detected in low to moderate numbers of conjunctival epithelial cells in most alligators in Group 2 (3/4; Fig. 16). Antigen was otherwise rare in other tissues, except for the liver of one of the animals that had moderate numbers of immunopositive cells. Abundance of chlamydial antigen positively correlated with the severity of the histologic lesions (Fig. 17; ρ = 0.63; P-value <.0001).
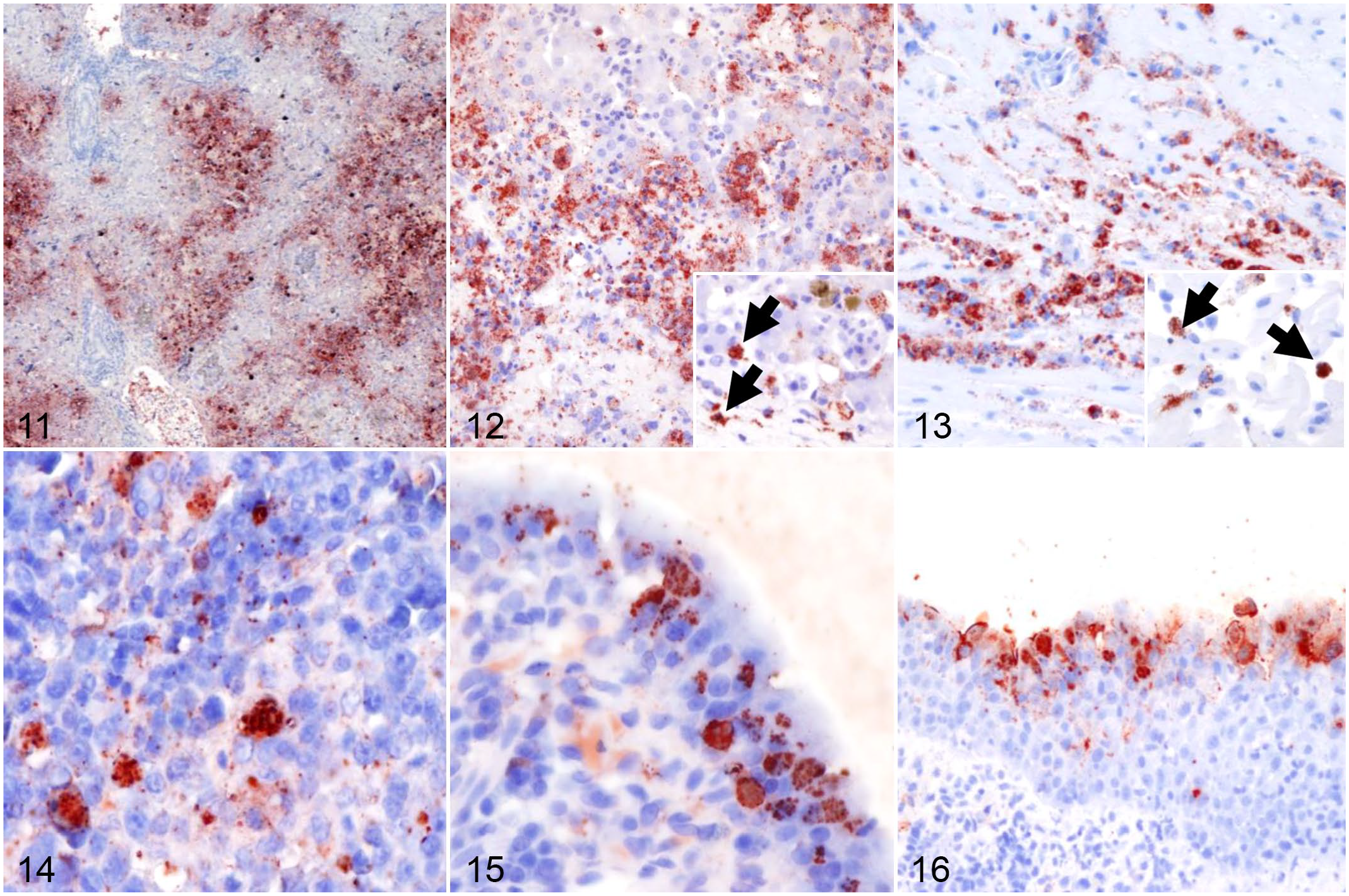
Chlamydial infection, alligators. Immunohistochemistry for chlamydial antigen.
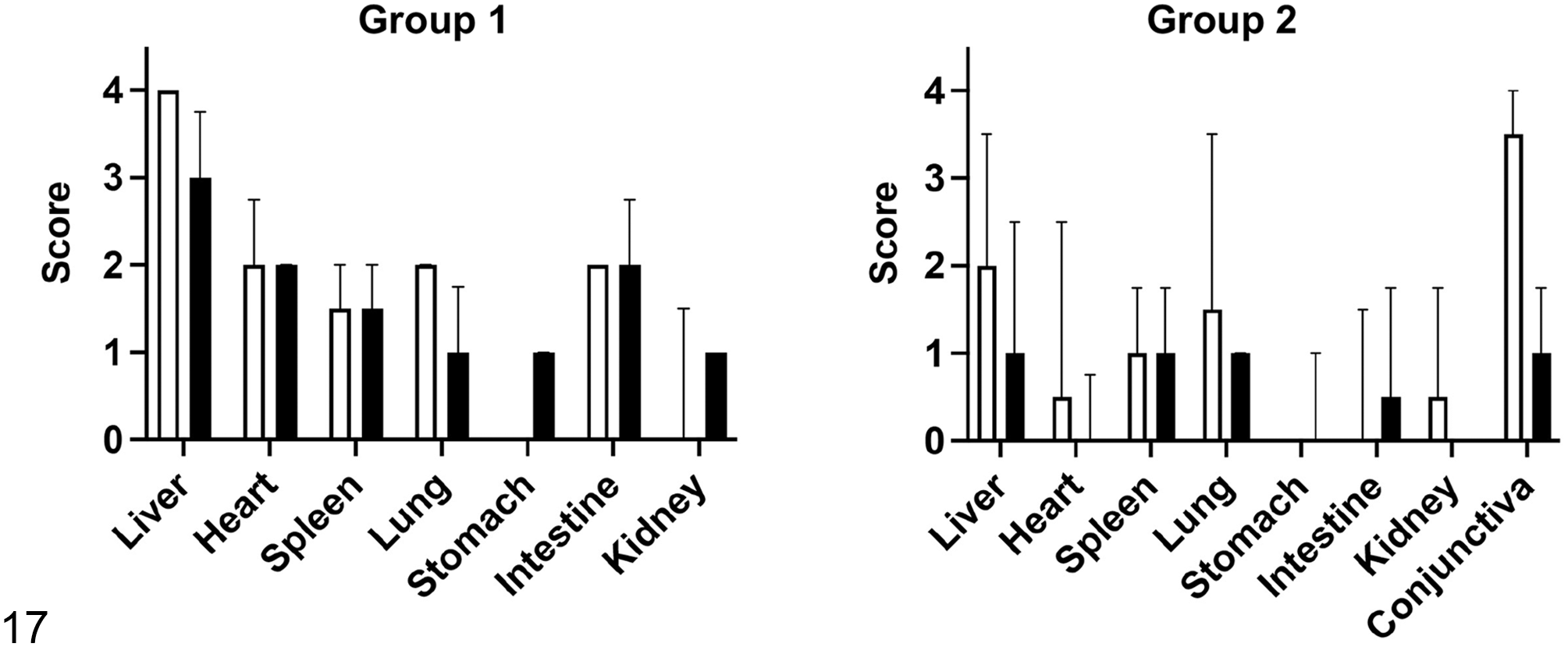
Severity scores of histologic lesions (white bars) and chlamydial antigen abundance (black bars) based on immunohistochemistry in tissues from Group 1 and Group 2 alligators. Median and interquartile range are depicted.
Sequencing and Phylogenetic Analysis of Uncultured Chlamydia sp
Following positive amplification for the Chlamydiaceae 23S rRNA gene by qPCR, pooled samples from Group 1 alligators were subjected to 16S rRNA gene sequencing for genus confirmation. The sequences generated in this study were deposited in GenBank under accession numbers OL880546 and OL956539. Following analysis using the nucleotide basic local alignment search tool (nBLAST, https://blast.ncbi.nlm.nih.gov/Blast.cgi), the partial sequence obtained was determined to have 99% homology with an uncultured Chlamydia sp identified in a Siamese crocodile and C. psittaci strain CPX0308, for which high homology to Chlamydia sp sequences derived from crocodiles was also identified in the past. 24 Subsequently, OmpA gene sequencing for Chlamydia sp and nBLAST along with phylogenetic analysis were performed. nBLAST analysis revealed 86% sequence homology with C. psittaci strain CPX0308. Phylogenetic analysis of the OmpA sequence determined that the uncultured Chlamydia sp sequence derived from these American alligators (2559) formed a separate cluster in close relationship with Chlamydia sp S15-834C isolated from the snake species Pantherophis guttatus and designated as C. poikilothermis 28 and C. psittaci strain CPX0308 detected in feces of an oriental white stork (Ciconia boyciana) in Japan 5 (Supplemental Fig. S4). The sequence obtained from these American alligators was not closely related to Chlamydia sp detected in crocodiles (Supplemental Fig. S4).
Discussion
Chlamydial infections in crocodilian species of the genus Crocodylus have been frequently described with variable clinical presentation ranging from inapparent to fulminant disease.13,14,24,26,32 Occurrences in this group of reptiles have been reported particularly in hatchling and juvenile Indo-Pacific crocodiles (Crocodylus porosus) in Australia and Papua New Guinea and Nile crocodiles (Crocodylus niloticus) from South Africa and Zimbabwe. 13 The reported lesions mostly involved the conjunctiva and upper respiratory tract. Interestingly, several studies have identified co-infections with Aeromonas sobria 32 and an uncharacterized herpesvirus.23,26 While C. pneumoniae has been reported as the most frequent species affecting reptiles, the chlamydial species affecting reptiles remain largely unknown.3,30 Other potentially novel, poorly characterized Chlamydia species have also been identified in clinically normal snakes and turtles.16,20 Most recent studies have characterized novel chlamydial species in snakes (C. serpentis and C. poikilothermis) and Siamese crocodiles (named C. crocodili). 6 For the latter, recovery of this pathogen from cloacal, pharyngeal, and conjunctival swabs, as well as various tissues (liver, spleen, lung, and brain), was reported, with clinical signs ranging from inapparent infections to occurrence of kyphoscoliosis in juveniles, conjunctivitis, pharyngitis, ascites, depression, anorexia, and death. 6 Subclinically infected animals are suggested to serve as carriers, and it has been proposed that crocodilian Chlamydia may act either as an opportunist or in concert with other pathogens. The most recent epizootics described in Thailand reported severe necrotizing lymphoplasmacytic hepatitis in hatchlings and juvenile Nile and Siamese crocodiles, respectively, but with no evidence of conjunctivitis. 32 Outbreaks as such can have a devastating economic impact in farmed animals and the poor characterization of the chlamydial organisms affecting these reptilian species raises concerns for potential zoonotic transmission. The aim of this study was to characterize the pathological, immunohistochemical, and molecular aspects of an outbreak associated with systemic chlamydial infection in farmed American alligators.
Two distinct groups on the same farm differed in their lesions and clinical presentation. Group 1 was characterized by high mortality with widespread necrotizing hepatitis and myocarditis and usually abundant chlamydial antigen, and Group 2 was mostly characterized by subacute conjunctivitis and lesions in other organs indicating systemic disease of lesser severity. Group 1 did not have gross ocular changes although the eyes were not examined histologically. The lesions in Group 2 were predominantly subacute with lower abundance of chlamydial antigen.
Based on immunohistochemistry, the chlamydial organism identified in this outbreak appeared to have a cellular tropism mostly restricted to macrophages, hepatocytes, and specific subsets of epithelial cells including conjunctiva and enterocytes. Based on these results, it is hypothesized that ocular and intestinal shedding might play a major role in environmental spread of the organism.
The OmpA gene is commonly utilized for genotyping Chlamydia sp and assessing its evolutionary dynamics.4,21 Therefore, we performed OmpA sequencing to gain further insight into the relationship with Chlamydia species reported in other species. Phylogenetic analysis revealed that this strain clustered separately from other reported crocodilian Chlamydia species and was closely related to C. poikilothermis from snakes and C. psittaci isolated from healthy wading birds. The partial sequencing performed is insufficient to draw definitive conclusions on the genetic relationship with other species, and the separate clustering from other crocodilian Chlamydia may reflect spatiotemporal differences. Based on the close relationship with the newly detected C. poikilothermis from snakes, 28 it is hypothesized that the Chlamydia species in these farmed alligators could belong to this species. The Chlamydia species could not be cultured using the methods of this study, and while all samples collected were positive for Chlamydia sp by qPCR, only one pooled sample yielded good quality sequences for analysis. Further studies are required to identity this organism and to identify the risk factors linked to disease outbreaks. Two of the affected farmed alligators in Group 1 had concurrent intestinal lesions and intestinal infection with Salmonella sp, and E. coli and Salmonella sp were also cultured from multiple organs of a single alligator in Group 2. The latter case supports concurrent sepsis, while the role of Salmonella sp as a commensal or possible opportunistic pathogen affecting the former case remains unclear. No other concurrent infections with other pathogens including Aeromonas sp or WNV were identified during this outbreak. Samples were not tested for herpesvirus, which has been identified as a concurrent infection in Siamese crocodiles, 23 but no herpesviral inclusion bodies were identified. While these findings suggest that this chlamydial organism may be a primary pathogen in this outbreak, the possibility of another unidentified microorganism serving as the primary pathogen in this mortality event cannot be completely excluded.
In conclusion, this study describes the occurrence of a previously unreported Chlamydia sp associated with a major mortality event in farmed American alligators. Enhanced epidemiological monitoring is needed to gain further insight into the specific cause, disease dynamics, risk factors, and role and prevalence of carrier animals within wild and farmed alligator populations.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858221095269 – Supplemental material for An outbreak of systemic chlamydiosis in farmed American alligators (Alligator mississippiensis)
Supplemental material, sj-pdf-1-vet-10.1177_03009858221095269 for An outbreak of systemic chlamydiosis in farmed American alligators (Alligator mississippiensis) by Mariano Carossino, Javier G. Nevarez, Kanako Sakaguchi, Daniel B. Paulsen, Ingeborg M. Langohr, Keith Strother, Jacqueline Ferracone, Alma Roy, Nicholas A. Crossland and Fabio Del Piero in Veterinary Pathology
Footnotes
Acknowledgements
We would like to acknowledge the members of the Histology and Immunohistochemistry sections at the Louisiana Animal Disease Diagnostic Laboratory (LADDL), School of Veterinary Medicine, Louisiana State University for their assistance. We would also like to acknowledge Mr Robert Poston for attempting isolation of the Chlamydia sp in embryonated chicken eggs and Ms Thaya Stoufflet at GeneLab, Division of Biotechnology and Molecular Medicine, School of Veterinary Medicine, Louisiana State University for performing Sanger sequencing.
Supplemental material for this article is available online.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was partially supported by the Louisiana Animal Disease Diagnostic Laboratory, Louisiana State University School of Veterinary Medicine, and start-up funds provided to M. Carossino (Louisiana State University School of Veterinary Medicine; PG009641).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
