Abstract
Periostin is a matricellular protein important in regulating bone, tooth, and cardiac development. In pathologic conditions, periostin drives allergic and fibrotic inflammatory diseases and is also overexpressed in certain cancers. Periostin signaling in tumors has been shown to promote angiogenesis, metastasis, and cancer stem cell survival in rodent models, and its overexpression is associated with poor prognosis in human glioblastoma. However, the role of periostin in regulating tumorigenesis of canine cancers has not been evaluated. Given its role in bone development, we sought to evaluate mRNA and protein expression of periostin in canine osteosarcoma (OS) and assess its association with patient outcome. We validated an anti-human periostin antibody cross-reactive to canine periostin via western blot and immunohistochemistry and evaluated periostin expression in microarray data from 49 primary canine OS tumors and 8 normal bone samples. Periostin mRNA was upregulated greater than 40-fold in canine OS tumors compared to normal bone and was significantly correlated with periostin protein expression based on quantitative image analysis. However, neither periostin mRNA nor protein expression were associated with time to metastasis in this cohort. Gene Set Enrichment Analysis demonstrated significant enhancement of pro-tumorigenic pathways including canonical
Periostin, or osteoblast-specific factor 2 (OSF-2), is a multifunctional matricellular protein with an important role in regulating normal bone, tooth, lung, and cardiac tissue development. 12,26,77 The protein has been shown to be important in cardiac mesenchymal stem cell differentiation to support early valvulogenesis. 70 In development, periostin has been shown to induce collagen fibrillogenesis, increasing the integrity of the ventricular wall. 13 Periostin has also been described to be important in neonatal lung remodeling and alveolar simplification, with periostin expression to be increased in mice and infants with hyperoxic lung injury. 5 Pathologically, aberrant expression of periostin is associated with a variety of inflammatory processes, such as pulmonary fibrosis, cancer, and asthmatic and allergic airway inflammation. 36,47,55,78
On initial characterization of periostin, it was noted that the protein has distinct homology to the insect protein fasciclin I, an adhesion protein found to play an important role in neuronal development. 18,76,83,87 This protein contains 4 fasciclin-like domains (FAS1 domains) that are responsible for interacting with various integrin receptors. Indeed, similar to its insect homologue, periostin functions as more than just a structural protein and also binds various integrin receptors (namely, αvβ3 and αvβ5) to modulate cell behavior. 25,69,76 Periostin is known to activate numerous signal transduction pathways including focal adhesion kinase (FAK) and phosphatidylinositol 3-kinase (PI3-K)/Akt, pathways which have all been implicated in cancer development and are known to be important for angiogenesis and cellular survival, proliferation, and migration. 1,8,12,25,32,45,49,59,69,75,84
The role of periostin in cancer has been evaluated in a variety of human cancers. Elevated periostin levels in tumor tissue or blood serum compared to healthy tissue has been demonstrated in breast, 1,45 ovarian, 25,75,85 glioblastoma, 49,59,84 colon, 1,2 and many other types of cancers. 42,53 Periostin overexpression in these cancers is associated with increased biological aggressiveness and worse clinical outcome. Pro-tumorigenic functions demonstrated in human in vitro cell models and rodent preclinical models include periostin enhancement of invasion, metastasis, cancer stem cell survival, angiogenesis, and tumor growth. 1,2,25,42,45,49,53,59,69,75,84,85 Yet very few studies have evaluated the role of periostin in osteosarcoma (OS), 6,32,33 despite first being described in this cancer. 76
Osteosarcoma is the most common primary bone cancer found in humans 50,56 and dogs. 15,60 Roughly 20% of human OS patients will have detectable pulmonary metastases at the time of initial diagnosis, 37,50,56 with the likelihood of subsequent development of micrometastases in those patients initially free of distant disease remaining very high. Micrometastatic tumor cells can be present in the bone marrow and blood of 70% of individuals with no signs of metastatic disease at the time of diagnosis, 7 and approximately 25% of all patients will eventually develop pulmonary metastases despite treatment with surgery and standard-of-care chemotherapy. 7 Patients with lung metastases have a 30% 5-year survival, 50,56 which has not improved in over 30 years. Thus, investigations into new model systems and treatments, specifically to stop development and progression of pulmonary metastasis, are desperately needed to improve the outcome of OS patients. 21,39
Dogs with OS provide a unique opportunity to study this disease due to the high rate of natural occurrence in this species and similarity of the disease between humans and dogs. 15,16,19,50,52,56 Canine OS occurs at greater than a 10-times higher rate than humans, with an incidence ranging from 7.9 to 13.3 per 100 000 dogs 16,64 compared to adolescent human OS at 4.5 per 1 000 000 children. 50,56 Canine OS shares many characteristics with human OS such as clinical behavior including primary tumor location, predilection for pulmonary metastasis and shared prognostic factors, as well as biological similarities including histological appearance and overlapping genomic and transcriptomic landscape. 19,40,58,80 The use of canine OS as a model for human OS can provide a better understanding of the disease, and improve treatment and survival for both species. 19,39,80 Already canine OS has been part of the development of novel surgical techniques and new therapies for their human counterpart. 35,43,48,72,79,80
Given periostin’s role in normal bone development and correlation of its aberrant expression with many biological hallmarks of cancer progression across multiple tumor types, we hypothesized that periostin expression would be elevated in canine OS and that increased periostin would be associated with worse clinical outcome in canine OS patients. To investigate this, we validated an anti-human periostin antibody cross-reactive to canine periostin via western blot, immunocytochemistry, and immunohistochemistry. Utilizing formalin-fixed paraffin-embedded (FFPE) archived cases of canine OS and a previously generated microarray expression dataset associated with these cases, we evaluated mRNA and protein expression levels of periostin in a subset of 49 primary appendicular canine OS tumors and determined its association with patient progression-free survival. Furthermore, utilizing Gene Set Enrichment Analysis, we sought to determine whether periostin overexpression in canine OS was associated with in vivo enrichment of tumor-promoting biological functions similar to those previously reported for periostin in human and rodent cancer models.
Materials and Methods
Cell Lines and Culture
Flint Animal Cancer Center (FACC) canine tumor cell lines were obtained from other institutions, purchased from the American Type Culture Collection (ATCC), or established from tumor biopsy samples obtained at the FACC, as previously described. 14,22 Tumor cells were cultured in RPMI 1640 culture medium containing 10% fetal bovine serum (FBS), penicillin (100 units mL−1), and streptomycin (100 µg mL−1) and incubated under standard conditions at 37 °C in a humidified atmosphere of 5% CO2:95% air. Prior to expression profiling, all cell lines were confirmed to be mycoplasma-negative and the species of origin and cell line identity validated by polymerase chain region for short tandem repeat (STR) analysis.
Canine Osteosarcoma Primary Tumor Samples and Clinical Data
Treatment-naïve primary tumor samples were obtained from the Colorado State University (CSU) Flint Animal Cancer Center’s tissue archive. These patient samples are archived in compliance with the CSU Institutional Animal Care and Use Committee following obtainment of owner consent. Forty-nine tumors were selected based on clinical characteristics as described previously. 57 Case selection was based on those primary appendicular OS tumors obtained from dogs treated only with surgical amputation followed by adjuvant chemotherapy with doxorubicin and/or a platinum-based drug (choice of adjuvant drug has been shown to not result in significantly different survival times between treatment groups in previous studies). 67 All dogs were free of lung metastases by thoracic radiographic analysis at diagnosis, and follow-up consisted of evaluation by clinical examinations including repeat thoracic radiographs every 2 to 3 months after initial treatment. For 8 of the appendicular OS tumor samples collected, samples of matched normal metaphyseal bone were also harvested from the same limb (at minimum separated by at least one joint space from the tumor) following amputation. Tumor and normal bone tissues collected at amputation were flash-frozen in liquid nitrogen and stored at −80 °C prior to downstream analysis.
Western Blot
The canine Moresco osteosarcoma cell line was used as a positive control for western blot validation of the manufacturer-reported canine cross-reactivity of the periostin antibody. The Moresco cell line was provided by Dawn Duval (Flint Animal Cancer Center) and was originally obtained from the University of Wisconsin–Madison. Cells were cultured as described above and then supernatants collected as well as cell lysates generated by incubation of flasks with protein extraction buffer: 7 ml M-PER (ThermoFisher), 140 mg sodium dodecyl sulfate (ThermoFisher), 70 µl of 100 mM phenylmethylsulfonyl fluoride (ThermoFisher), 35 µl of 200 mM sodium orthovanadate (Sigma-Aldrich), and ½ tablet Complete Mini protease inhibitor (Roche) for 5 minutes on ice. Lysates were removed from the flasks, triturated on ice using a pipette, centrifuged at 13 000 rpm for 5 minutes, and supernatant removed for protein concentration determination using a BCA protein assay (Pierce).
Proteins were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Protein was then wet transferred to nitrocellulose membranes, and membranes were blocked for 1 hour at room temperature (RT) in a 5% non-fat dry milk in Tris-buffered saline Tween 20 solution (TBST). Membranes were then incubated with the primary antibody (rabbit anti-canine periostin, LifeSpan Bio, LS-B3986) diluted 1:1000 (1 μg/ml) in 5% non-fat dry milk in TBST, overnight at 4 °C. The following day, membranes were rinsed (×3 with TBST), incubated with the secondary antibody (HRP-linked goat anti-rabbit IgG; Thermo Scientific) diluted 1:3000 in 5% milk-TBST for 1 hour at RT. Membranes were rinsed again (×3 with TBST). Last, membranes were imaged with chemiluminescent substrate (Clarity Western ECL, BioRad) using a Chemi Doc XES + system (BioRad).
Immunohistochemistry and Immunocytochemistry
FFPE tumor tissue blocks were sectioned at 5 µm and mounted on Superfrost Plus slides (Fisher Scientific) for periostin immunolabeling. Briefly, tissue slides were deparaffinized in xylenes and rehydrated using a series of graded alcohols. Antigen retrieval was performed using 10 mM sodium citrate buffer, pH 6.0, for 20 minutes at 125 °C in a DAKO PT link module. For immunocytochemistry, cultured Moresco tumor cells were immobilized on to Superfrost Plus slides using a cytocentrifuge and subsequently fixed in 4% paraformaldehyde prior to commencing with immunolabeling, using the same autostainer and protocol for FFPE tumor tissue, as follows. Immunolabeling was performed using a Dako autostainer link 48. Tissues were blocked for endogenous peroxidase by incubation in 3% H2O2 for 5 minutes. Subsequently, sections were then incubated with polyclonal rabbit anti-canine periostin primary antibody (LifeSpan Biosciences, LS-B3986; 2.5 μg/ml) for 1 hour at RT. Bound primary antibody was detected using the universal labeled streptavidin-biotin2 system (Dako), which consists of incubation with a mixture of biotinylated goat anti-mouse and rabbit IgG secondary antibodies followed by horseradish peroxidase–labeled streptavidin. Positive staining was visualized using DAB chromogen substrate (Dako). Purified rabbit immunoglobulin (Dako) was used in place of the primary antibody as a negative control to ascertain nonspecific labeling. Slides were counterstained with Mayer’s hematoxylin (Sigma Aldrich).
Image Analysis
For quantification of periostin immunoreactivity, five 400× magnification, intratumoral independent fields of each primary tumor were captured using standardized exposure times and a Nikon Eclipse 80i microscope and Olympus DP70 camera. Representative tumor fields digitally captured for quantitative image analysis of periostin immunolabeling were chosen based on regions of highest expression. Five nonoverlapping 400× magnification images were captured from these regions by an individual blinded to clinical outcome or any other study data (ocular field number = 22 mm, total field of view area analyzed = 1.19 mm2). In order to ensure accurate quantitative assessment of periostin-positive immunolabeling, we used the color deconvolution algorithm developed for the NIH open source image analysis software, ImageJ. Using the H DAB (hematoxylin and DAB chromogen) vector for this algorithm, digitized intratumoral images of periostin immunolabeling were separated into 8-bit gray scale images representative of the chromogen color (DAB) only. A lower threshold limit was then set at a value corresponding to the mean of the isotype control, and universally applied to every single image. Any pixel value falling above this lower threshold value was measured as positive for periostin and used to determine % area positive within the field. For quality control, image masks of the “thresholded,” positive counted area were also generated and directly visually compared to the originally captured photomicrographs by a board-certified pathologist, to ensure accuracy in representation of periostin positive immunoreactivity and overall tumor area.
Gene Expression Microarray Analysis
Periostin expression across the FACC tumor cell line panel was evaluated using a microarray expression data set previously generated by the Duval Laboratory at the Flint Animal Cancer Center. 22 For microarray analysis of the FACC tumor cell line panel, RNA was extracted from cells using the RNeasy kit (Qiagen) according to manufacturer’s protocol, as described previously. 22 Quantity and quality of RNA was evaluated via a NanoDrop 1000 spectrophotometer (Thermo Scientific) and a Bioanalyzer (Agilent). RNA samples were then sent to the Genomics and Microarray Core at the University of Colorado. Briefly, samples were hybridized onto Affymetrix GeneChip Canine Genome 2.0 arrays, Canine Gene 1.0 ST arrays, and GeneChip miRNA 4.0 arrays. Resulting CEL files of expression data were then imported into Bioconductor, 24 and intensity values were preprocessed with the Robust Multi-Array Average (RMA) algorithm. 34
Evaluation of periostin expression in canine primary OS tumor samples and normal bone was done using microarray expression data also previously generated by the Duval Laboratory at the Flint Animal Cancer Center.
57
For microarray analysis of fresh frozen primary OS tumor samples, total RNA was extracted as described previously.
57
Briefly, samples were freeze-fractured, homogenized, extracted with Trizol reagent (Invitrogen) and further cleaned up and purified with RNeasy extraction kit (Qiagen) following manufacturer’s protocols. Normal bone RNA was also extracted following this same protocol, with the exception of an additional spin of 800 ×
Pathway (Gene Set Enrichment) Analysis
Gene Set Enrichment Analysis (GSEA; https://www.gsea-msigdb.org/gsea/downloads.jsp) was performed according to the user guide (https://www.gsea-msigdb.org/gsea/doc/GSEAUserGuideFrame.html) to determine biological pathways differentially enriched between periostin “high” and “low” expressing canine primary OS tumors. GSEA is a statistical tool developed for analyzing microarray or other biological expression data based on applying a metric for ranking all genes in an expression dataset and then determining the degree to which gene sets in specific molecularly defined gene signatures are overrepresented at the top (positive enrichment score) or bottom (negative enrichment score) of the ranked list of genes.
73
These gene sets are publicly available and housed in the Molecular Signatures Database (MSigDB; https://www.gsea-msigdb.org/gsea/msigdb/index.jsp) maintained by the Broad Institute and UC San Diego. The following parameters were used for GSEA analysis: Pathway enrichment between periostin “high” and “low” tumors was done using the Hallmarks gene set (h.all.v6.2.symbols.gmt), with Signal2Noise ranking metric, permutation type set to “gene set,” and 1000 permutations used to calculate statistical significance of enrichment scores. Statistically significant pathways were defined as those with
Immune Cell Type–Specific Gene Signatures
The relative abundance of immune cell types was determined using a previously described gene expression signature.
63
Heatmaps were generated with
Statistical Analyses
Continuous data were expressed as means ± standard deviation and normality tested using the D’Agostino and Pearson method. For comparison of means between 2 groups with normally distributed data a 2-tailed, unpaired Student’s
Results
Study Population
Dogs (
Dogs (
Periostin Expression Is Elevated in Canine Osteosarcoma Cell Lines and Tumors
To first investigate a potential role for periostin overexpression in canine OS, we evaluated periostin mRNA expression across a panel of 37 canine tumor cell lines encompassing the following histo-types: osteosarcoma (
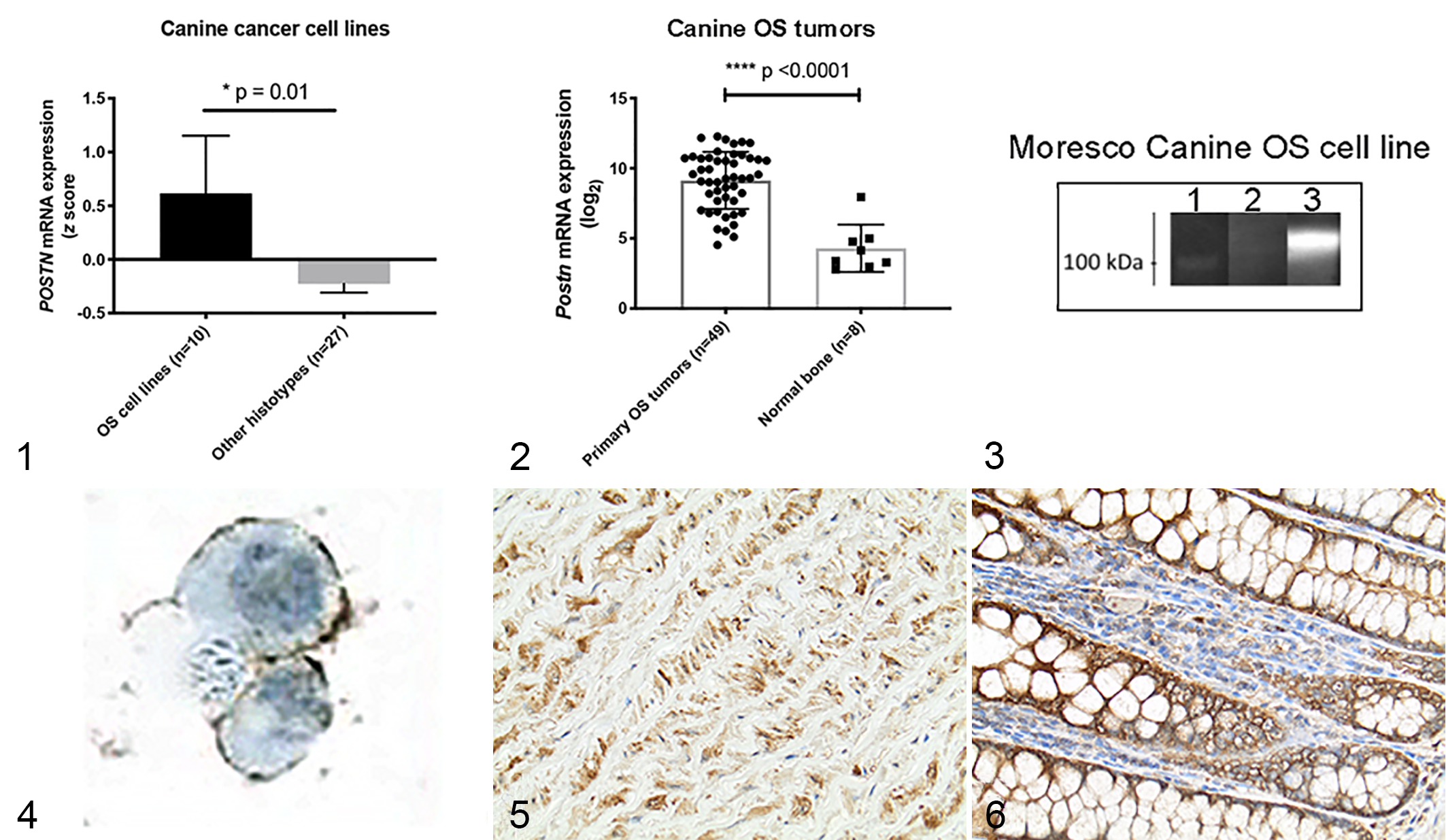
Next, to determine whether these canine cancer cell lines were representative of in vivo periostin expression in OS tumors, we compared periostin mRNA expression between primary appendicular OS tumors (
Immunohistochemical Expression of Periostin in Canine Osteosarcoma
Based on this mRNA expression data, we sought to characterize periostin protein expression and localization in canine OS tumors. To do this, we first validated an anti-human periostin antibody reported by the manufacturer to be reactive to canine periostin. Western bot analysis of the Moresco OS cell line (the cell line with the second highest periostin mRNA expression value) performed with this antibody detected a band of ∼100 kDa, near the predicted molecular weight (∼93 kDa) of canine periostin, in Moresco cell culture supernatant (Fig. 3, lane 3). The observance of periostin in cell culture supernatant is expected given the presence of a signal peptide in its amino acid sequence 76 and is consistent with the secreted nature of this matricellular protein. 31,76 Supporting the results of the western blot analysis, immunocytochemistry for periostin performed with this antibody on the Moresco cell line demonstrated strong membranous localization of periostin (Fig. 4). This membranous localization may be reflective of autocrine binding of secreted periostin to its cognate αvβ3/5 receptors, 25,69 which we have observed to be overexpressed in these canine OS cell lines (unpublished data, Regan laboratory).
Immunohistochemistry (IHC) performed with this antibody on FFPE canine aorta and colon (positive control tissues) demonstrated strong immunoreactivity and expected localization of periostin expression within myocytes and the extracellular space of the aortic tunica media as well as peri-cryptal immune and stromal cells and mucosal epithelial cells of the colon (Figs. 5, 6; Supplemental Figures S1, S2)
41,82
(and https://www.proteinatlas.org/ENSG00000133110-POSTN/tissue). Next, we performed immunohistochemistry for periostin expression on a subset of 18 of the 49 primary OS tumors evaluated for mRNA expression. These 18 cases were chosen based on FFPE block availability as well as equal representation of significant differences in clinical outcome based on disease-free interval (DFI) >250 days (good responders,
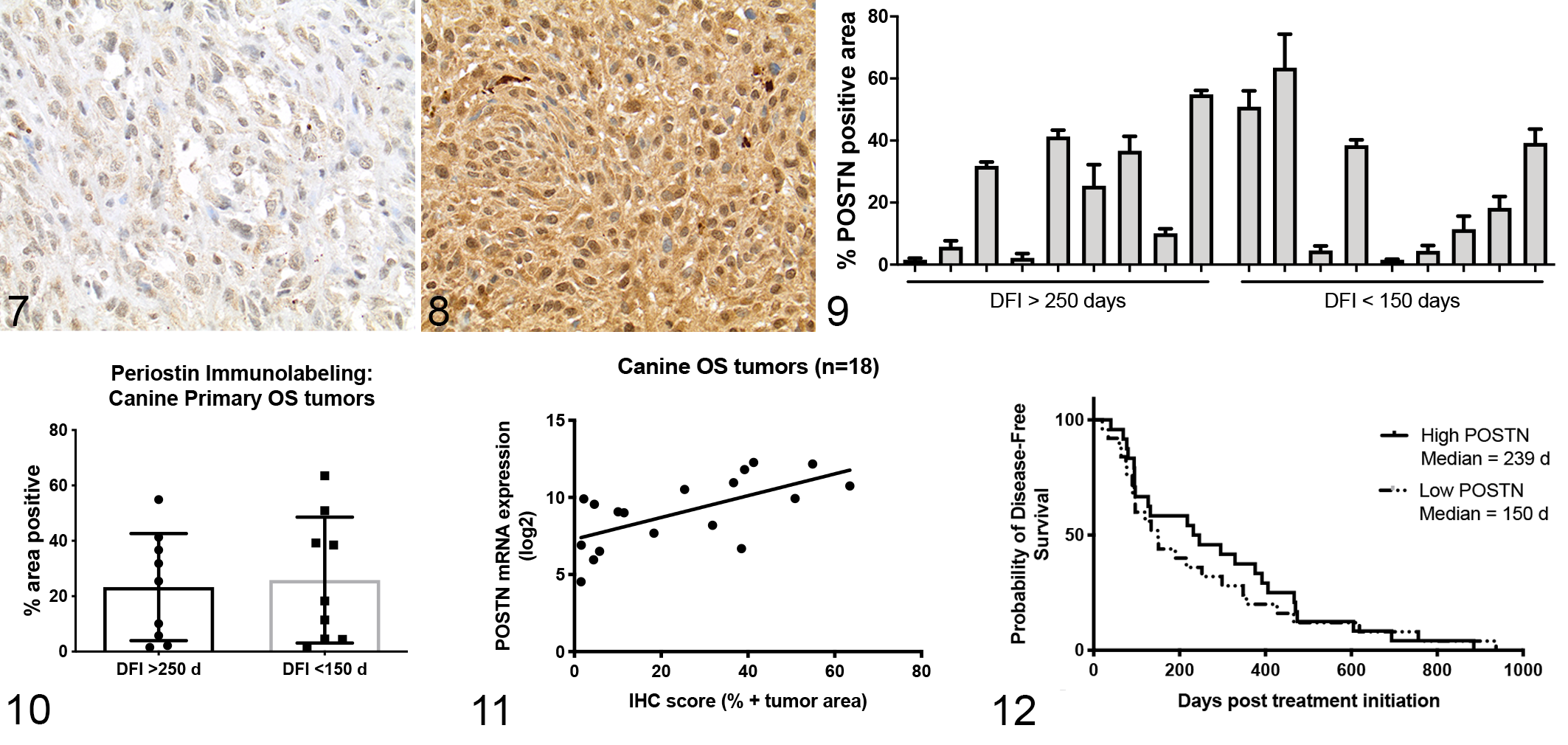
Association of Periostin Protein and mRNA Expression With Clinical Outcome in Canine Osteosarcoma
Data from the quantitative analysis of periostin immunolabeling was then used to determine whether periostin protein expression was different between those subsets of dogs with favorable clinical outcome (DFI > 250 days) versus those dogs with poor clinical outcome (DFI < 150 days). Immunohistochemical expression of periostin was not significantly different between those dogs with DFI <150 days (mean % periostin positive = 25.8) or >250 days (mean % periostin positive = 23.3; Fig. 10). Similarly, periostin protein expression was not correlated to DFI (
Given the low power and limitations associated with performing survival analysis in only 18 dogs, we sought to determine whether periostin mRNA expression was correlated with disease-free interval in our larger 49 dog subset. This approach was validated by the strong positive linear correlation we observed between periostin mRNA and protein expression in our initial 18 dog cohort (Fig. 11, Spearman
Periostin Overexpressing Osteosarcoma Tumors Are Enriched for Gene Pathways Associated With Hallmarks of Cancer Progression
Despite a lack of association between periostin expression and clinical outcome, we still sought to better understand the functional impact of periostin expression in canine OS, given the wide range in expression values across our cohort of tumors. To do this, we performed pathway analysis with the gene microarray expression data from the 49 canine primary OS tumors using Gene Set Enrichment Analysis (GSEA) software (https://www.gsea-msigdb.org/gsea/index.jsp). GSEA analysis evaluates changes in genes between 2 user-defined phenotypes, in this case “periostin mRNA-High” and “periostin mRNA-Low,” using a ranking metric for each gene in order to more globally evaluate gene expression changes and detect differential enrichment in gene sets associated with specific biological processes between these phenotypes. 51,73
GSEA analysis using the hallmarks gene sets
44
comparing “periostin-High” versus “periostin-Low” tumors (again based on median dichotomization) identified statistically significant enrichment (
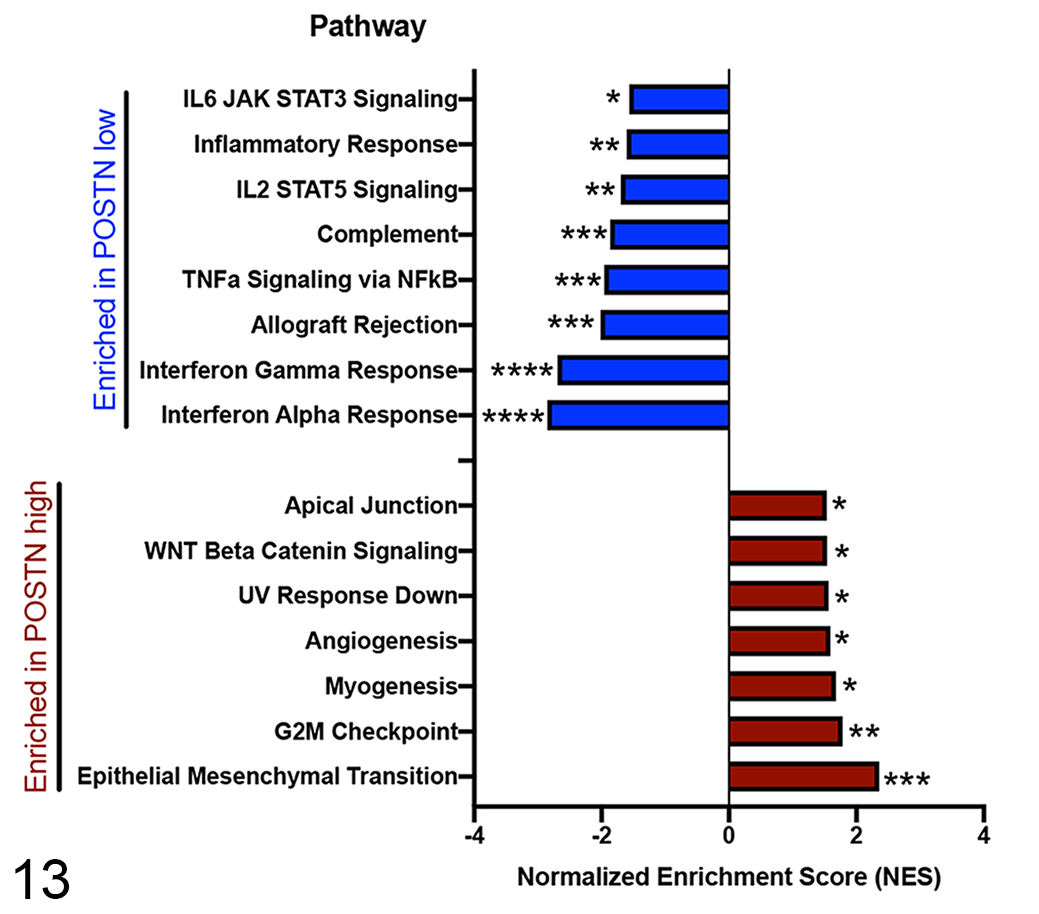
Gene Set Enrichment Analysis (GSEA; Hallmark gene sets of Molecular Signatures Database) of differentially enriched pathways in canine osteosarcoma primary tumors with high (
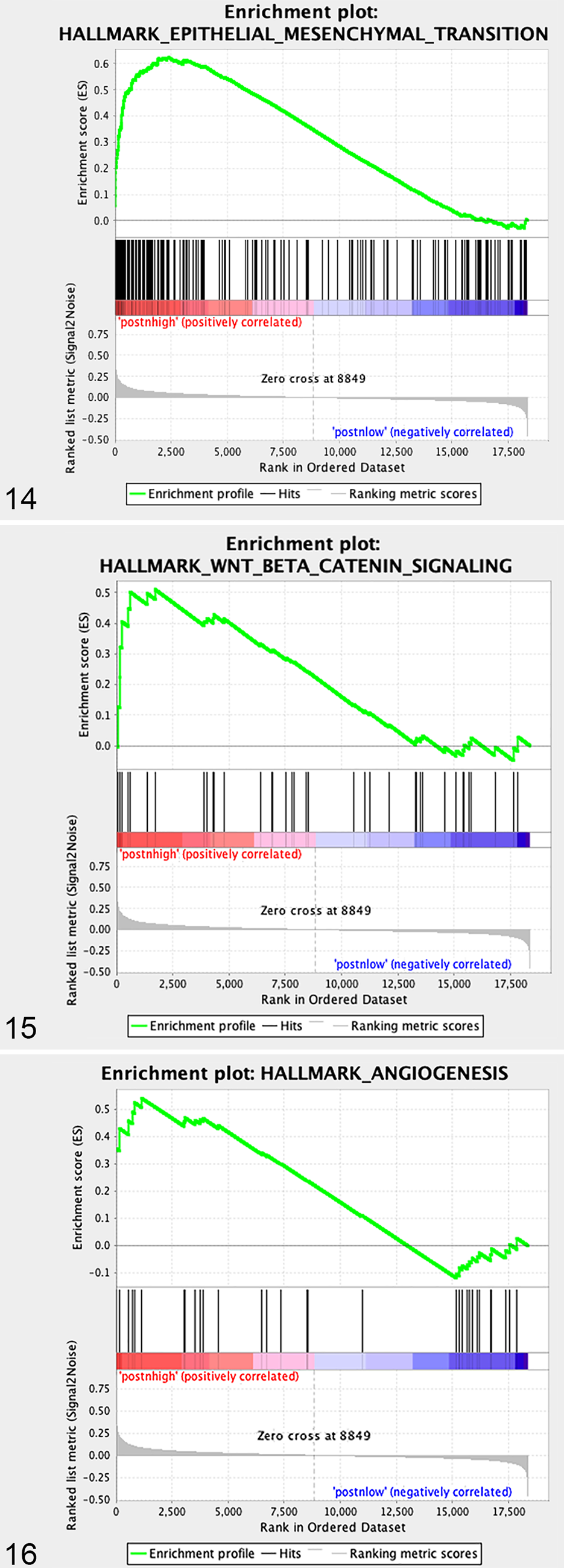
GSEA enrichment plots of specific gene sets upregulated in periostin “high” versus periostin “low” canine OS primary tumors.
“Periostin-Low” Osteosarcoma Tumors Are Associated With Heightened Immune Activity
GSEA analysis of “periostin-low” expressing tumors demonstrated statistically significant enrichment in 8 pathways (Fig. 13 and Figs. 17-19,
For our study, we utilized previously published cell-type specific gene signatures
63
to transcriptomically compare the immune infiltrate between those tumors in the lower quartile (lowest 25%) of periostin expression (POSTN low—blue,
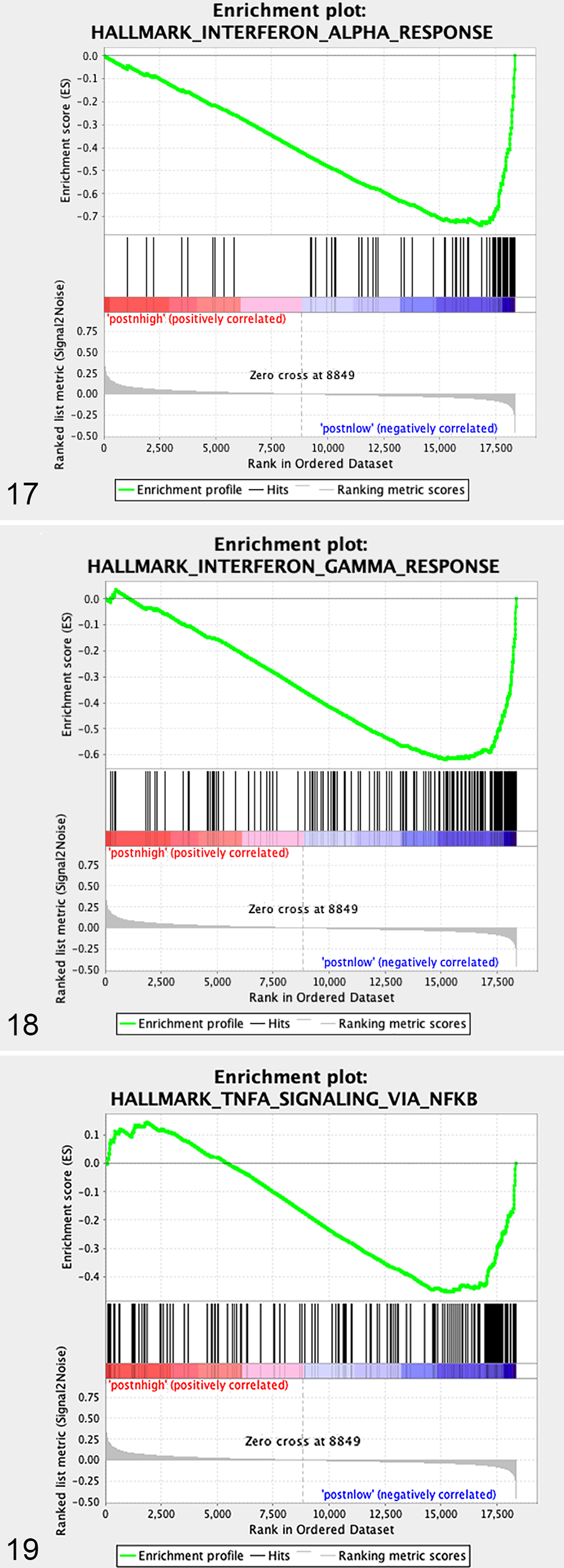
GSEA enrichment plots of specific gene sets upregulated in periostin “low” versus periostin “high” canine OS primary tumors.
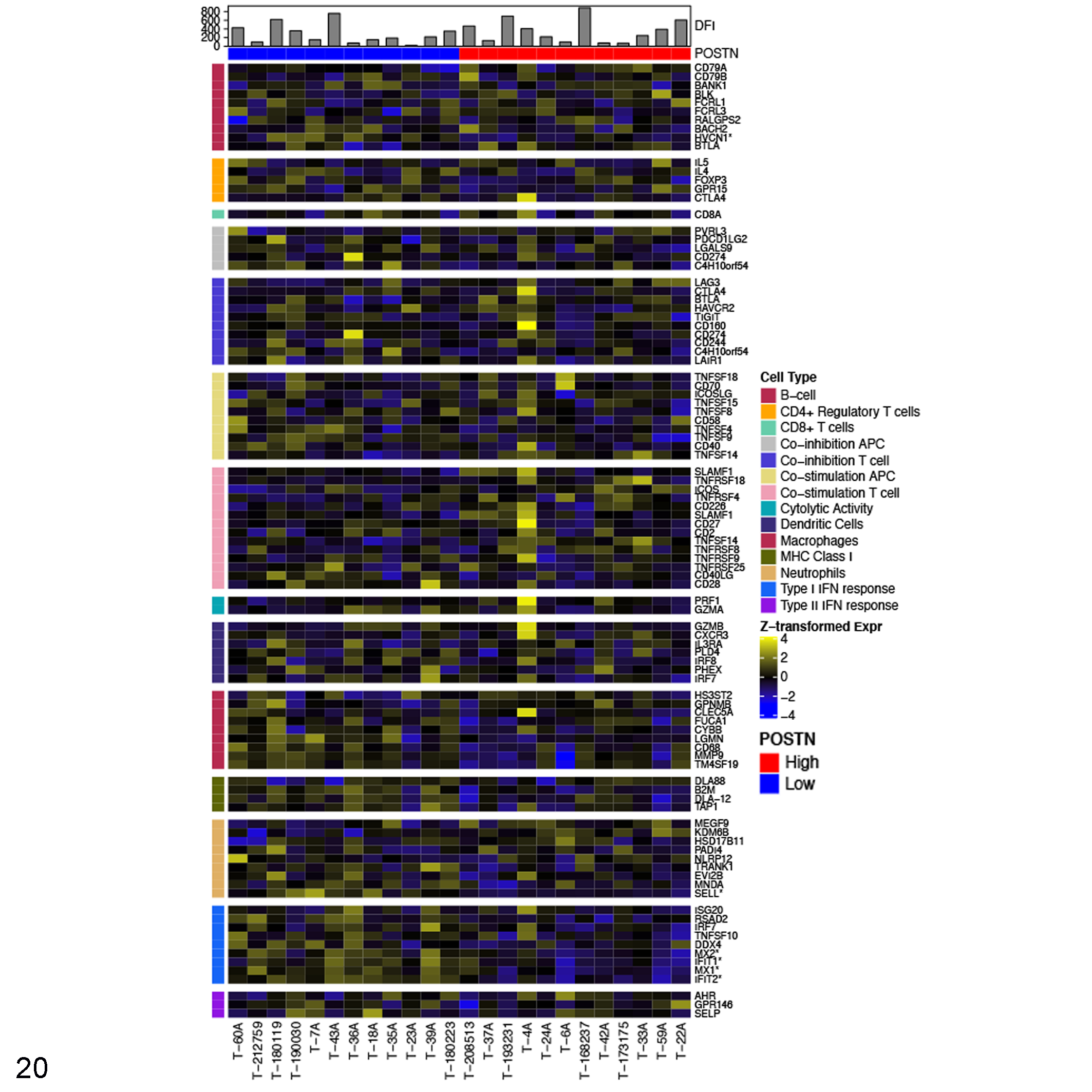
Relative expression of immune cell lineage markers in high (
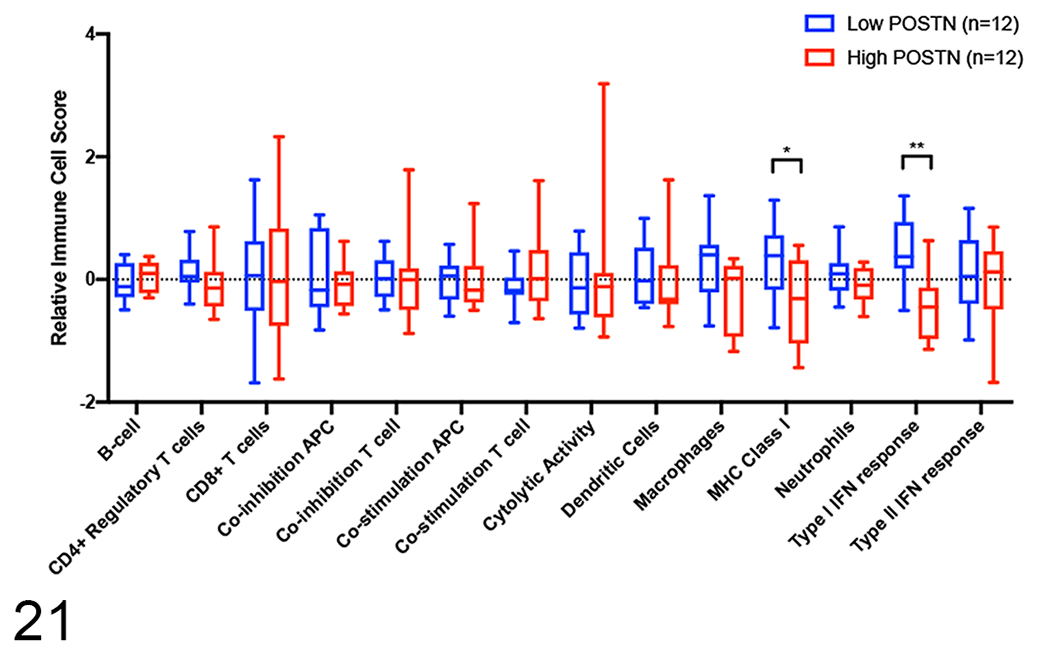
Immune cell scores between high and low periostin-expressing samples. Statistical significance was determined using multiple unpaired
Discussion
Based on periostin’s originally described role in the proliferation and survival of osteoblasts, 86 as well as an increasing body of literature describing its pro-tumorigenic function in human cancers through enhancement of tumor cell migration, metastasis, and angiogenesis, 1,2,32,33,45,49,69 we sought to investigate whether periostin overexpression was associated with worse clinical outcome in canine OS. Our studies validated an anti-human periostin antibody cross-reactive to canine periostin via western blot, immunocytochemistry, and immunohistochemistry. While periostin mRNA or protein expression alone were not associated with clinical outcome in our subset of 49 dogs with OS, we demonstrate statistically significant differential pathway enrichment between periostin high versus low expressing OS tumors. These pathways strongly overlapped with those tumor-promoting functions reported with periostin overexpression in human cancers, and also suggested that variation in periostin expression levels may be associated with differences in the composition of the canine OS immune landscape.
Immunohistochemical analysis demonstrated periostin immunoreactivity in 18/18 canine OS primary tumors evaluated. In these cases, localization of periostin expression was predominately in the cytoplasm of neoplastic cells and extracellular matrix immediately surrounding tumor cells, suggesting that neoplastic OS cells, and not stromal cells such as cancer-associated fibroblasts, are the primary source of periostin expression in these tumors. These observations are further supported by the significant elevation of periostin mRNA expression we demonstrated in our canine OS tumor cell lines. While the degree of periostin mRNA and protein expression varied widely on a case-to-case basis in our cohort, the significant preferential overexpression in canine OS tumor cell lines and OS primary tumors as compared to other tumor histo-types and normal bone, respectively, suggests an important role for this protein in canine OS progression. Importantly, this overexpression demonstrates another similarity between canine and human OS, 32,33,76 and provides a foundation and rationale for future studies aimed at investigating periostin and its associated signal transduction pathways in the development of new treatments for OS.
Despite our demonstration of significant and preferential elevation of periostin in canine OS, our study did not demonstrate an association between periostin mRNA or protein expression and disease-free interval in this cohort of 49 dogs. This is in contrast to observations in human cancers, where periostin expression is associated with clinical outcome in OS, ovarian, colon, and glioma. 2,32,33,49,75,85 A general limitation of our study, which could account for this lack of association with survival, is our small sample size. For example, one study demonstrating significant association of periostin expression with clinical outcome in glioma patients was performed on gene expression data from subsets ranging in size from 77 to 674 patients. 49 Furthermore, these investigators used the statistical power of these large datasets to define a multi-gene “periostin expression signature,” 49 something which we were unable to do, and another limitation of our current study. It is likely that incorporation of multiple genes co-regulated with periostin expression may better identify those tumors which are truly dependent on the pro-tumorigenic functions of periostin, leading to better prognostic utility.
Although worse clinical outcome was not associated with elevated periostin levels in canine OS, we provide evidence that specific gene programs and molecular pathways significantly differ between “high” and “low” expressing periostin tumors. In our pathway analysis of periostin overexpressing OS tumors, we observed epithelial-to-mesenchymal transition (EMT) to be the most significantly enriched pathway in these tumors. This enrichment in EMT may seem like a paradox considering OS is already a mesenchymal-derived tumor; however, recent reports have demonstrated other sarcomas to have enrichment for an EMT program. 11,17,20,59,66 As primarily demonstrated in epithelial-derived tumors, EMT is related to multiple tumor-promoting functions including increased invasion, motility, metastasis, and drug resistance, and enhanced expression of EMT signatures have been previously associated with patient tumor stage and prognosis. 53,66,71 However, these early reports on EMT in sarcomas also suggest similar increased biological aggressiveness associated with this phenotype. Importantly, transforming growth factor β (TGFβ), which plays a key role in initiating EMT transcriptional programs, is one of the principal molecules known to induce periostin secretion and is a potent immune suppressive molecule. 45,61,62,81
GSEA analysis also demonstrated significant enrichment of
Interestingly, our pathway analysis of “low periostin” expressing tumors demonstrated significant enrichment in 8 gene sets, of which all are associated with genes upregulated in immune responses. Gene targets upregulated in response to interferon-α (IFN-α) and interferon-γ (IFN-γ) were the 2 most significantly enriched pathways in our “periostin-low” tumor subset. IFN-α and IFN-γ are both critical mediators of antitumor immunity, with well-defined requirements for both molecules in effective tumor immune surveillance and response to immune-based therapies such as vaccines, oncolytic agents, and checkpoint inhibitors. 23,28,38,88 These data suggest that periostin expression may dictate fundamental differences in the immune landscape of canine OS tumors, which in part is supported by our gene signature–based immune cell profiling performed on these tumors. The association of periostin overexpression with those OS tumors which appear more immunologically “cold” may indirectly reflect enrichment of TGF-β in the tumor microenvironment, given the dependency for this immune suppressive molecule in periostin secretion. 45,81 While both IFN-α and IFN-γ have been reported as independent prognostic factors in human cancers, 46,54 this association was not observed in our case cohort despite significant upregulation of these pathways. Nonetheless, these data warrant future expanded investigations as to whether the level of enrichment for periostin and/or other ECM protein signatures is associated with response to immune-based therapies in canine OS tumors.
In conclusion, our study validated an anti-human periostin antibody cross-reactive to canine periostin via western blot, immunocytochemistry, and immunohistochemistry. Furthermore, we demonstrate an association between increased periostin expression in canine OS and enhancement of pro-tumorigenic transcriptional programs including canonical
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_0300985821996671 - Role of Periostin Expression in Canine Osteosarcoma Biology and Clinical Outcome
Supplemental Material, sj-pdf-1-vet-10.1177_0300985821996671 for Role of Periostin Expression in Canine Osteosarcoma Biology and Clinical Outcome by Lauren N. Alfino, Kai C. Wilczewski-Shirai, Kathryn E. Cronise, Jonathan Coy, Kristina Glapa, E. J. Ehrhart, Joseph B. Charles, Dawn L. Duval and Daniel P. Regan in Veterinary Pathology
Footnotes
Acknowledgements
The authors thank the oncology service and clinical trials staff at the Flint Animal Cancer Center for procurement of canine osteosarcoma biopsies for gene expression analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Research reported in this publication was supported by The National Institutes of Health, Office of the Director, Award Number K01ODO22982 (DPR) and R03OD028265 (DPR); Short-Term National Research Award, Award Number T35OD015130 (LNA); and National Center for Advancing Translational Sciences, Award Number L30 TR002126 (DPR). This work was also supported by Boehringer Ingelheim Veterinary Scholar Fellowship (LNA).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
