Abstract
Porcine astrovirus type 3 (PoAstV3) is an emerging virus in the family Astroviridae that has been recently associated with polioencephalomyelitis/encephalitis. Herein, we describe the experimental oral and intravenous inoculation of an infectious central nervous system (CNS) tissue homogenate containing PoAstV3 to cesarean-derived, colostrum-deprived pigs, and the subsequent development of clinical signs, histologic lesions, specific humoral immune response, and detection of viral particles by electron microscopy (EM) and viral RNA by RT-qPCR (reverse transcriptase quantitative polymerase chain reaction) and in situ hybridization (ISH). IgG against a portion of the PoAstV3 ORF2 capsid was first detected at 7 days post-inoculation (DPI) in 2 of 4 inoculated animals and in all inoculated animals by 14 DPI. At 21 and 28 DPI, 2 of 4 inoculated animals developed ataxia, tetraparesis, and/or lateral recumbency. All inoculated animals had histologic lesions in the CNS including perivascular lymphoplasmacytic cuffs, multifocal areas of gliosis with neuronal necrosis, satellitosis, and radiculoneuritis, and PoAstV3 RNA as detected by RT-qPCR within multiple anatomic regions of the CNS. Consistent viral structures were within the soma of a spinal cord neuron in the single pig examined by EM. Of note, PoAstV3 was not only detected by ISH in neurons of the cerebrum and spinal cord but also neurons of the dorsal root ganglion and nerve roots consistent with viral dissemination via axonal transport. This is the first study reproducing CNS disease with a porcine astrovirus strain consistent with natural infection, suggesting that pigs may serve as an animal model to study the pathogenesis of neurotropic astroviruses.
Porcine astrovirus type 3 (PoAstV3, tentatively Mamastrovirus 22) is an emerging virus of the genus Mamastrovirus, family Astroviridae. 4,35 This single-stranded, positive sense RNA virus was initially detected in 2011 in a swine fecal sample collected in Canada in 2006. 47 Since then, additional strains have been identified in the United States, Japan, Croatia, Hungary, Kenya, Slovakia, Spain, and China. 2,14,20,24,40,57,62,73
Over the last several years, astroviruses have been identified and associated with central nervous system (CNS) disease in multiple mammalian species. 16 Frequently, these neurotropic strains cluster within the VA/HMO clade originating from humans, mink, sheep, cattle, and pigs including PoAstV3, suggesting the presence of common genetic determinants and the potential for cross-species transmission. 17,25,26,60,71
The pathogenesis and histologic lesions of neurotropic astroviruses appear similar across species. Cases of disease are frequently characterized by lymphoplasmacytic encephalomyelitis; in cattle and sheep, ganglioneuritis and ganglionitis have also been described, which may be a result of viral dissemination via axonal transport. 45,56 In humans, there is extensive information assessing the specific immune response mounted against human astroviruses suffering of enteric disease by human astroviruses, which has made possible to elucidate multiple aspects of their epidemiology and immunogenicity. 3,9,34,37,38,43,53 In contrast, there is no information available for swine regarding the development of diagnostic techniques used to evaluate the specific humoral response for porcine astrovirus type 3.
The isolation of neurotropic strains of astroviruses from clinical samples is an arduous process and only 2 successful isolations involving human strains have been described. 41,72 To date, and in contrast to PoAstV1, no isolation methods or permissive cell lines have been identified for PoAstV3. 31,39,64 To overcome this impediment and investigate the pathogenesis and immunopathogenesis of PoAstV3 infection, herein we describe the experimental inoculation of colostrum-deprived, cesarean-derived (CDCD) pigs with an infectious PoAstV3 CNS tissue homogenate and subsequent development of neurologic disease consistent with those observed in naturally occurring cases and a specific immune response using a newly developed ELISA.
Materials and Methods
Animals and Experimental Infections
Six 5-week-old, crossbred CDCD pigs were purchased from a commercial source and held in a BSL-2 large animal facility. Individual fecal swabs were collected prior to inoculation and tested PoAstV3 negative by RT-qPCR (reverse transcriptase quantitative polymerase chain reaction). All procedures were approved by Veterinary Resources Inc Institutional Animal Care and Use Committee (Log Number: ISU-S-18-1229). CNS tissues from 2 cases (US/IA/48142/2018 and US/IA/53214/2018) sent for routine diagnostic investigation to the Iowa State University Veterinary Diagnostic Laboratory (ISU-VDL) and diagnosed with PoAstV3-associated polioencephalomyelitis were identified. Both cases were submitted in 2018 and each case included multiple animals from a single farm with intermittent outbreaks of PoAstV3-associated disease. Sections of cerebrum, cerebellum, brainstem, and spinal cord were collected and stored at −80 °C until further processing. For each case, CNS tissues were combined, minced, and diluted with Eagle’s minimum essential medium (EMEM, Life Technologies) at a 1:2 ratio, and processed using a Geno/Grinder 2010 (SPEX SamplePrep LLC). Tissue homogenates (inocula) were filtered with 0.22 µm filters (Milipore) and stored at −80 °C. Both oral (US/IA/48142/2018) and intravenous (US/IA/53214/2018) inocula were tested via RT-qPCR and were negative for teschovirus A (PTV), sapelovirus A (PSV), enterovirus G (PEV), pseudorabies virus (PRV), porcine reproductive and respiratory virus (PRRSV), and porcine circovirus type 2 (PCV2). PoAstV3 quantification cycles (Cq) values from the oral (Cq = 20.3) and intravenous (Cq = 21.1) inoculum corresponded to 6.1 × 107 viral copies/ml and 3.5 × 107 viral copies/ml, respectively. The equation for PoAstV3 genome copy number/ml of sample was 10((Cq-46.332)/-3.3434). Cq values ≥40 were considered negative. Additionally, a sample of the intravenous inoculum was taken via a sterile culture swab (Puritan sterile swab) and cultured on blood agar at 35 °C, blood agar with Staphylococcus hyicus nurse at 35 °C with 7.5% CO2, and MacConkey agar at 35 °C. No bacterial growth was evident after 72 hours.
Experimental Design
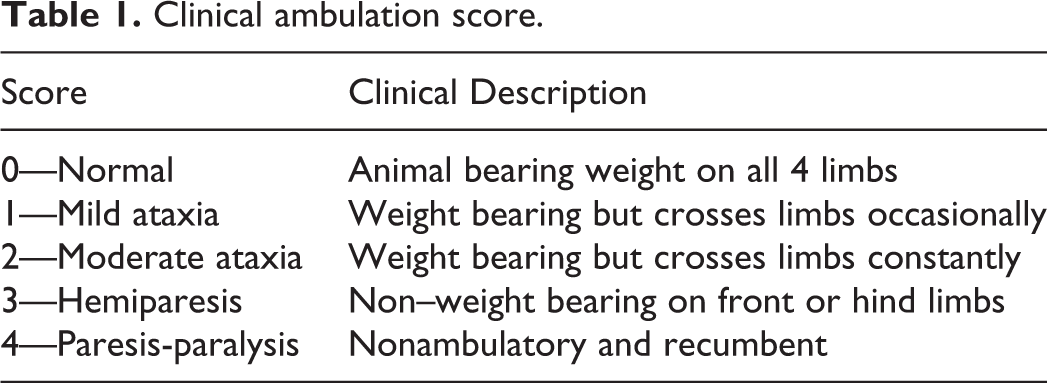
Pigs were randomly assigned to either the negative control group (pigs 1 and 2) or the PoAstV3 inoculated group (pigs 3–6) and housed separately. Animals were fed ad libitum with commercial feed, and drinking water was provided by automatic nipple drinkers. For inoculation, pigs were sedated with an intramuscular injection of Telazol 500 mg (Zoetis), ketamine 250 mg (VET One), and xylazine 250 mg (VET One) at 4.4 mg/kg. The inoculum was administrated orally using a nasogastric feeding tube (Covidien, IRL) and intravenously using a 25-gauge butterfly catheter (Terumo Surflo, Terumo Corporation). Negative control pigs were inoculated orally (10 ml) and intravenously (5 ml) with EMEM (Life Technologies). Pigs 3 to 6 were given an equivalent volume of PoAstV3-positive CNS tissue homogenate using the same administration routes as control pigs. Animals were evaluated daily for the presence of ambulatory/neurologic clinical signs including ataxia, hemiparesis, paresis, and paralysis (Table 1). Serum and fecal samples were collected on days post-inoculation (DPI) 0, 3, 7, 10, 14, 21, and 28. Animals were necropsied at DPI 29, with the exemption of pigs 5 and 3 that were euthanized at DPI 21 and 28, respectively, according to humane endpoint criteria preestablished for this study. The rectal temperature of all pigs, and respiratory rates (breaths/min), with the exception of pig 5, were recorded prior to euthanasia.
Clinical ambulation score.
Pathologic Evaluation and Sample Collection
At necropsy, fresh and fixed (10% neutral buffered formalin) sections of cerebrum, cerebellum, brainstem, spinal cord with adjacent spinal ganglion (cervical, thoracic, and lumbar), cerebrospinal fluid, lung, heart, liver, spleen, kidney, tonsil, jejunum, ileum, cecum, colon, lymph node (tracheobronchial, mesenteric, and inguinal), feces, and serum from all animals were collected. Fresh samples were stored at −80 °C until further processing. Sections of cerebrum, cerebellum, spinal cord with adjacent spinal ganglion (cervical, thoracic, and lumbar) from all animals were fixed in 2.5% glutaraldehyde in 0.1 M cacodylate buffer solution. Gross lesions were independently recorded by 2 veterinary diagnosticians.
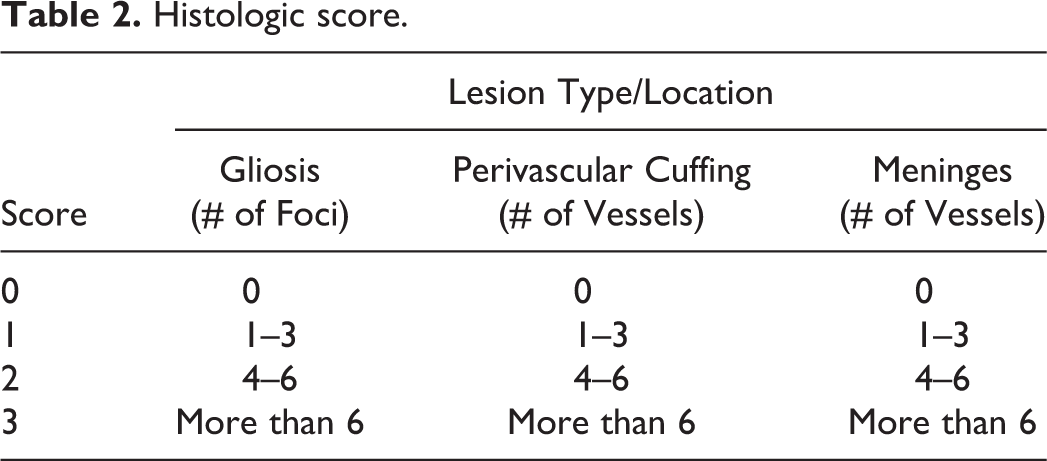
Briefly, after 48-hour fixation in 10% buffered neutral formalin, CNS samples were embedded in paraffin blocks, cut on 4 μM sections, and stained with hematoxylin and eosin. Sections of cerebrum (at the level of the thalamus), cerebellum, brainstem, and cervical, thoracic, and lumbar spinal cord were scored by a blinded veterinary diagnostician. Lesions were scored based on the number of glial nodules and number of perivascular mononuclear inflammatory infiltrates observed in the leptomeninges and Virchow–Robin spaces (Table 2). After recording lesions in each of 6 anatomic regions, a sum of all scores was compiled for a cumulative score. The cumulative score was assigned a level of severity (normal, mild, moderate, and severe). Animals with a cumulative score of 0 to 2 were considered normal, and animals with 2 to 18, 19 to 36, and 37 to 54 cumulative points were scored as mild, moderate, and severe, respectively.
Histologic score.
In Situ Hybridization
For each inoculated animal, the hematoxylin and eosin–stained slide with the most severe histologic lesions was selected and a sequential slide from the formalin-fixed paraffin-embedded (FFPE) block was used. Location-specific matching slides from control animals were used as negative controls. RNA in situ hybridization (ISH) was performed with RNAscope 2.5 HD Reagent Kit-Red (Advanced Cell Diagnostics) as previously described. 49
Electron Microscopy (EM)
Sections of cerebrum, cerebellum, brainstem, spinal ganglion and sections of cervical, thoracic, and lumbar spinal cord from pig 5 were selected for EM based on the severity and distribution of histologic lesions. Samples were post-fixed with 2% osmium and then processed through graded alcohols, propylene oxide, and Eponate 12 resin followed by 48-hour polymerization. Thick sections (1 µm) were performed on select samples and toluidine blue and basic fucsin stains were applied. Polaroid photos were taken of these images, and the area of interest for thin sections were selected. A uranyl acetate and Reynold’s lead stain were performed on the thin section before being examined with a ThermoFisher FEI Tecnai G 2 BioTWIN electron microscope (FEI Co). 19 Images were taken with Nanosprint12 camera (AMT Corp).
PoAstV3 RT-qPCR
Aliquots of frozen CNS samples (cerebrum, cerebellum, brainstem, cerebrospinal fluid, cervical, thoracic, and lumbar spinal cord) and non-CNS tissues (lung, heart, spleen, kidney, liver, tonsil, jejunum, ileum, colon, and mesenteric, inguinal, and tracheobronchial lymph nodes) from each animal were individually minced with sterile forceps and scissors and diluted with EMEM (Life Technologies) at a 1:10 ratio and processed at 1000 rpm for 180 seconds using a Geno/Grinder 2010 (SPEX SamplePrep LLC). At time of collection, fecal swabs were diluted in 1 ml of phosphate-buffered saline (PBS) solution while CSF samples were undiluted. All samples were stored at −80 °C until further processing. Viral RNA extraction was performed using the MagMAX Pathogen RNA/DNA Kit isolation kit (Thermo Fisher Scientific), a Kingfisher 96 Flex Purification System (Thermo-Fisher Scientific), and a high-volume procedure as previously described. 59 Primers and probe targeting the polymerase gene were as follows: forward primer ATGACYCTCTATGGGAAACTCCTT, reverse primer GTGCCTRGCAACAACCTCCAA, minor groove binding probe FAM-TTGGCCAYAACCTCCCTGA-MGB. Cq values ≥40 were considered negative.
Sequencing and Phylogenetic Analysis
Briefly, nucleic acids were extracted from 100 µl aliquots of oral and IV inocula using the MagMAX Pathogen RNA/DNA Kit (Thermo Fisher Scientific) and a KingFisher Flex Purification System (Thermo Fisher Scientific) following the instructions of the manufacturer. Primer sets targeting approximately 1000 base pair overlapping sections of the whole genome were designed based on 2 previously published sequences (GenBank accession nos. JX556691 and KY940545). The viral genomic RNA was amplified using a 25 µl reaction utilizing custom qScript One-Step RT-PCR Kit (Quanta Biosciences) following the manufacturer’s recommendations. Each primer was present in the final reaction at 320 nM, and 5 µl of RNA template was used per reaction. PoAstV3 RNA amplification was performed on an Applied Biosystems SimpliAmp thermal cycler (Thermo Fisher Scientific) under the following conditions: initial reverse transcription at 48 °C for 20 minutes, followed by initial denaturation at 94 °C for 3 minutes and 45 cycles of denaturation at 94 °C for 30 seconds; annealing at 55 °C for 50 seconds; extension at 68 °C for 90 seconds; and a final elongation at 68 °C for 7 minutes. Primer sequences are presented in Supplemental Table S1. PBS was extracted as a negative extraction control and nuclease free water was used as a negative amplification control. The RT-qPCR products were visualized using QIAxcel Advanced System (Qiagen) and purified using the ExoSAP-IT PCR Product Cleanup Reagent (Thermo Fisher Scientific) according to the manufacturer’s instructions. Sequencing was completed via Sanger sequencing at the Iowa State University DNA Facility. Sequences were assembled using Geneious Prime 2019.2.1 software.
Full genome sequence of USA/IA/48142/2018 (GenBank accession MT394895) and USA/IA/53214/2018 (GenBank accession MT394896) were aligned with 27 full or near-full genome sequences (+6000 nt in length) of neurotropic astroviruses and the original full-genome PoAstV3 US-MO123 strain. All sequences were obtained from GenBank. Multiple sequence alignment and sequence comparisons were carried out using the Clustal Omega algorithm by Jalview V2.11.1.0 software. 69 Phylogenetic trees were reconstructed with the Neighbor Joining (NJ) method using the Tamura–Nei model by MEGA X. 44,61,66 The confidence of the internal branches was evaluated performing 1000 bootstrap replications. 32
Protein Expression, Purification, and Indirect ELISA
The coding region of the vp27 gene, part of the PoAstV3 ORF2 capsid, was synthesized (Shanghai Genery Biotech Co), amplified, and cloned into pCold II expression plasmid, and confirmed by sequencing (Genewiz Inc). Plasmid was transformed into Escherichia coli BL21(DE3) (Invitrogen), grown in Luria-Bertani medium (Invitrogen) containing 50 µg per ml ampicillin (Amresco) at 37 °C with shaking at 250 rpm. When an A600 of 0.6 to 0.8 was reached, 0.3 mM isopropyl-β-thio-D-galactopyranoside (IPTG; Amresco) was added, and cultures were grown for an additional 16 hours at 20 °C with shaking at 250 rpm. Cells were chilled and harvested by centrifugation at 3500 × g for 15 minutes, resuspended in Lysis Buffer A (20 mM Tris, 300 mM NaCl, pH 8.0), sonicated (Φ3, 15%, 3 seconds on/6 seconds off, 10 minutes) on ice, and analyzed by polyacrylamide gel electrophoresis (SDS-PAGE). Analysis indicated that the protein was insoluble under native buffer conditions, and it was therefore extracted from the cell pellet under denaturing conditions. The pellet was resuspended with Lysis Buffer B (20 mM Tris, 300 mM NaCl, 8 M urea, pH 8.0), and sonicated (Φ15, 15%, 3 seconds on/6 seconds off, 20 minutes; Suppl. Fig. S1a). Supernatant was collected and subjected to affinity chromatography purification using Ni NTA Beads 6FF (Shanghai YuanYe Bio-Technology Co) according to the manufacturer’s instructions. In brief, the protein supernatant and beads were mixed and incubated for 1 hour at 4 °C. Beads were then washed with Wash Buffer C (20 mM Tris, 300 mM NaCl, 8 M urea, 20 mM imidazole, pH 8.0), followed by Wash Buffer D (20 mM Tris, 300 mM NaCl, 8 M urea, 50 mM imidazole, pH 8.0), and eluted with Elution Buffer E (20 mM Tris, 300 mM NaCl, 8 M urea, 500 mM imidazole, pH 8.0; Suppl. Fig. S1b). Purified VP27 protein (0.3 mg/ml) was dialyzed into Final Buffer F (PBS, 0.5% SKL, pH 7.4), analyzed by Western Blot (Suppl. Fig. S1c), aliquoted, and stored at −80 °C until use. The nucleotide and amino acid sequence of the VP27 protein is presented in Supplemental Materials 1.
Recombinant PoAstV3 VP27 protein was used to develop antibody isotype indirect ELISAs. Ninety-six well polystyrene ELISA plates (Maxisorp, Thermo Fisher Scientific) were coated with 100 µl of recombinant VP27 (1.2 µg/ml) in carbonate buffer, pH 9.6, and incubated at 4 °C for 16 hours. The plates were washed 5 times with PBS-T (0.1% Tween 20), blocked (300 µl/well) with 1% (w/v) bovine serum albumin blocking solution (BSA, Jackson ImmunoResearch Inc), incubated at 20 to 22 °C for 2 hours, and dried at 37 °C for 4 hours. Serum samples were tested at 1:100 dilution (100 µl reaction volume), incubated at 37 °C for 1 hour, and washed 5 times with PBS-T. Then, 100 µl of peroxidase (HRP)-conjugated goat anti-pig IgG (Fc) antibody (1:30 000), goat anti-pig IgA (1:3000) or goat anti-pig IgM (1:5000) (Bethyl Laboratories Inc) were added to each well and the plates were incubated at 37 °C for 1 hour. After a washing step, the reaction was visualized by adding 100 µl of tetramethylbenzidine-hydrogen peroxide (TMB; Surmodics IVD Inc) substrate solution to each well, and incubated for 10 minutes (IgG) or 5 minutes (IgA, IgM) at 20 to 22 °C. The reaction was stopped by adding 100 µl of stop solution (Surmodics IVD Inc) to each well, and the absorbance was read (450 nm) using an ELISA reader (Molecular Devices). Antibody responses were expressed as sample-to-positive (S/P) ratios. For statistical analysis, a generalized linear model with mixed effects was used to establish a quantitative relationship between S/P ratios, treatment (control vs inoculated), and DPI. In the model, treatment and DPI were used as fixed effects, and individual pig as random effect. Since the S/P data were right skewed, we adopted the log link with Gaussian distribution. As some S/P IgA and IgM observations had negative values, data were transformed by adding a specified value (+0.05) to all observations, so all values were positive and log link was feasible. A significance level of P ≤ .05 was used. All statistical analyses were performed with R software with the addition of “lme4” and “ggplot2” packages. 10,70
Metagenomic Sequencing
Oral and intravenous inocula and pooled CNS tissues (cerebrum, cerebellum, cervical, thoracic, and lumbar spinal cord) from pig 5 were sent to a company for metagenomic sequencing analysis (Cambridge Technologies). In brief, samples were clarified by filtration through a 0.2 µm filter followed by digestion of unprotected nucleic acids with a cocktail of DNases and RNases as previously described. 36 Following RNA isolation, samples were reverse transcribed followed by second strand synthesis using barcoded random hexamers. Viral DNA was next amplified using barcode-specific primers. Following cleanup of the amplicons, sequencing libraries were constructed using a Nextera XT kit and sequenced on a MiniSeq instrument using paired 151 bp sequencing. Sequencing reads were trimmed of barcode and adaptor sequences and low-quality sequences were removed by Illumina onboard software. Sequencing reads were next assembled de novo using CLC genomics software. Assembled contigs were analyzed by BLASTX to identify homology to sequences in the GenBank nonredundant protein sequence database. Further characterization of multiple viruses reads initially detected with Miseq Illumina platform were further corroborated at ISU-VDL by RT-qPCR.
PTV, PSV, and Pestivirus K RT-qPCR
PTV and PSV RT-qPCR were performed on tissue homogenate from brainstem samples. Pestivirus K RT-qPCR was performed on tissue homogenates from cerebrum and cerebellum. Briefly, tissue homogenates of all animals, and previously used for PoAstV3 RT-qPCR, were subjected to testing. Viral RNA extraction was performed using the MagMAX Pathogen RNA/DNA Kit isolation kit (Thermo Fisher Scientific), a Kingfisher 96 Flex Purification System (Thermo-Fisher Scientific), and a high-volume procedure as previously described. 59 Primers and probes and RT-qPCR conditions for PTV, PSV, and pestivirus K were previously described. 7,49 Cq values ≥36 were considered negative.
Results
Clinical Evaluation
Ambulatory scores are summarized in Table 3 and reported by DPI in Supplemental Table S2. On DPI 20, pig 5 developed posterior paresis, and the following day (DPI 21) was laterally recumbent and unable to ambulate or support weight (tetraparesis; Suppl. Video 1). On DPI 25, all inoculated animals were lethargic and reluctant to move. At DPI 28, pig 3 had intermittent posterior ataxia. Rectal temperatures were similar across groups except for pig 5, which was hypothermic (Suppl. Table S3). Prior to euthanasia, the respiratory rate of all inoculated animals was increased (mean = 114) compared to control animals (mean = 60; Table 3). No ambulatory deficits were observed in negative control animals throughout the study (Suppl. Table S2).
Clinical and histopathologic findings, and identification of viral RNA by in situ hybridization (ISH) and RT-qPCR in pigs after inoculation of cerebral homogenate of porcine astrovirus type 3.
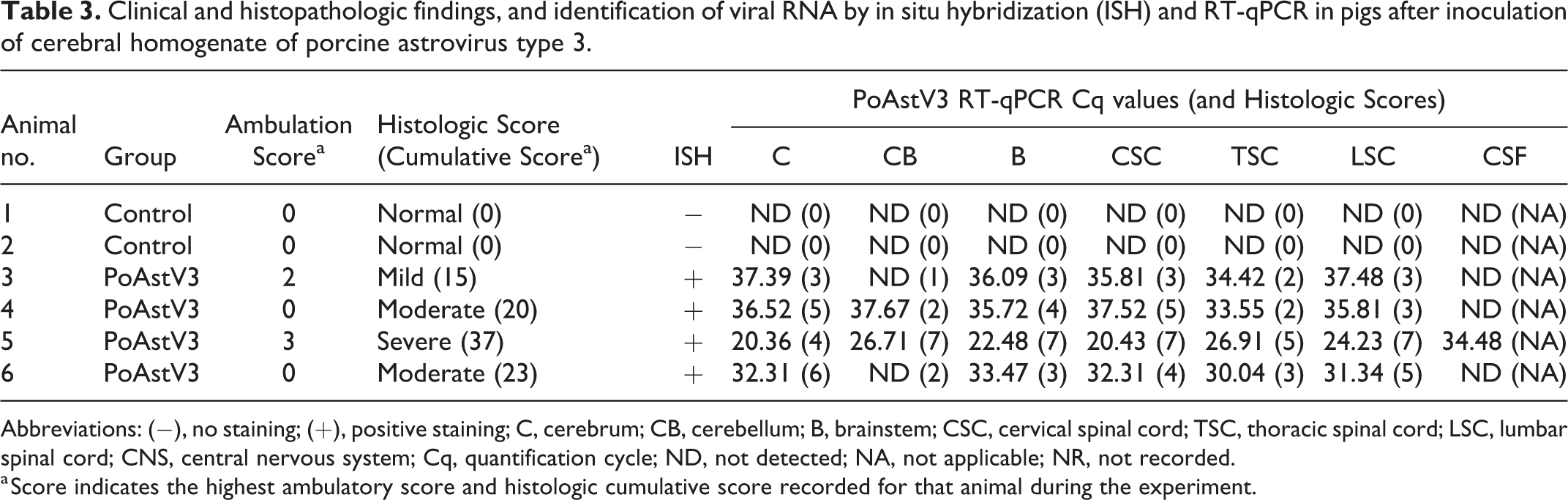
Abbreviations: (−), no staining; (+), positive staining; C, cerebrum; CB, cerebellum; B, brainstem; CSC, cervical spinal cord; TSC, thoracic spinal cord; LSC, lumbar spinal cord; CNS, central nervous system; Cq, quantification cycle; ND, not detected; NA, not applicable; NR, not recorded.
a Score indicates the highest ambulatory score and histologic cumulative score recorded for that animal during the experiment.
Pathologic Evaluation
Gross lesions were not observed in pigs 1 to 5. Mild bilateral exophthalmos was noted in pig 6. All 4 inoculated animals had histologic lesions in the CNS that varied from mild to severe lymphoplasmacytic polioencephalomyelitis with multifocal areas of gliosis and perivascular cuffing (Fig. 1). Other lesions included neuronal necrosis (angular eosinophilic cytoplasm, N = 4), satellitosis (perineuronal clustering of glial cells and lymphocytes, N = 4; Fig. 2), and dilation of myelin sheaths with rare spheroids in sections of cerebrum, cerebellum, and spinal cord (N = 4). Overall, lesions predominantly affected the gray matter. Additionally, pigs 3 and 5 had mild to severe lymphoplasmacytic ganglioradiculoneuritis, respectively (Fig. 3). The most severe histologic lesions were observed in pig 5, followed by pigs 6 and 4, and pig 3 (Table 3). No CNS histologic lesions were observed in control animals. The distribution of CNS histologic lesions are summarized in Supplemental Table S4.
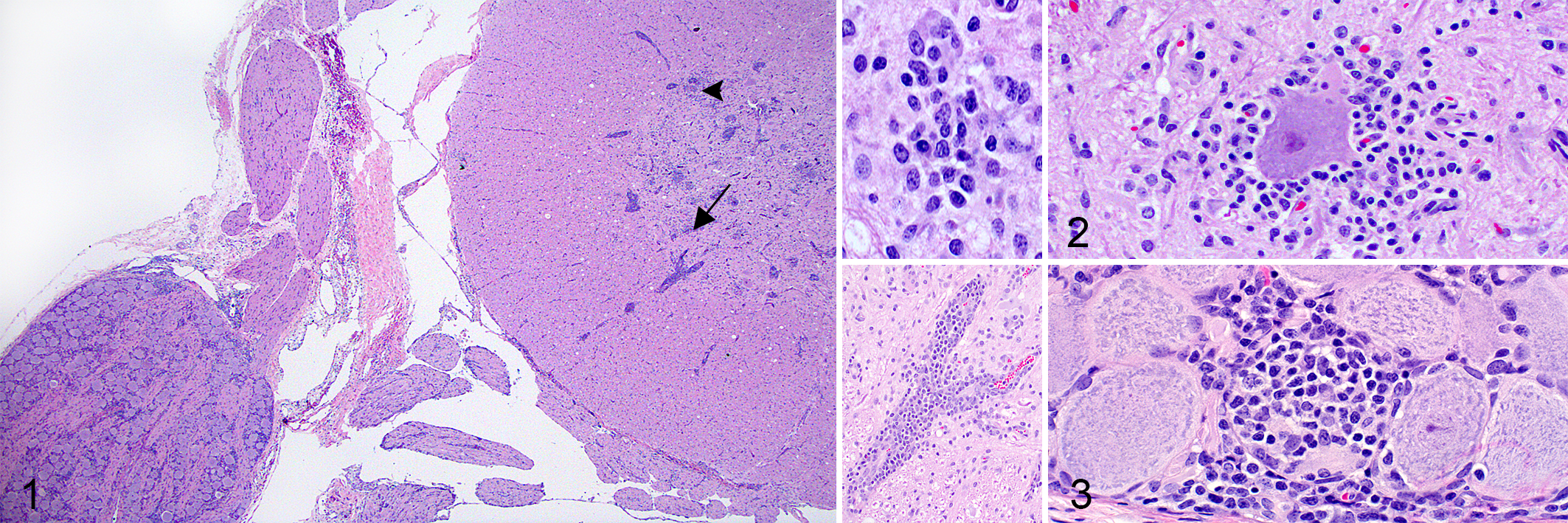
Porcine astrovirus type 3 (PoAstV3), pig 5, 21 days post-infection.
In Situ Hybridization
Neurons (soma, dendrites, and/or axonal processes) and neuroglia of selected sections from all inoculated animals had discrete to marked, punctuate, intracytoplasmic labeling (Figs. 4, 5). PoAstV3-specific labeling was also observed within neurons (soma) of the dorsal root ganglion and axonal processes of dorsal and ventral root fibers in pig 5 (Fig. 6). No specific labeling was observed in sections from control animals.
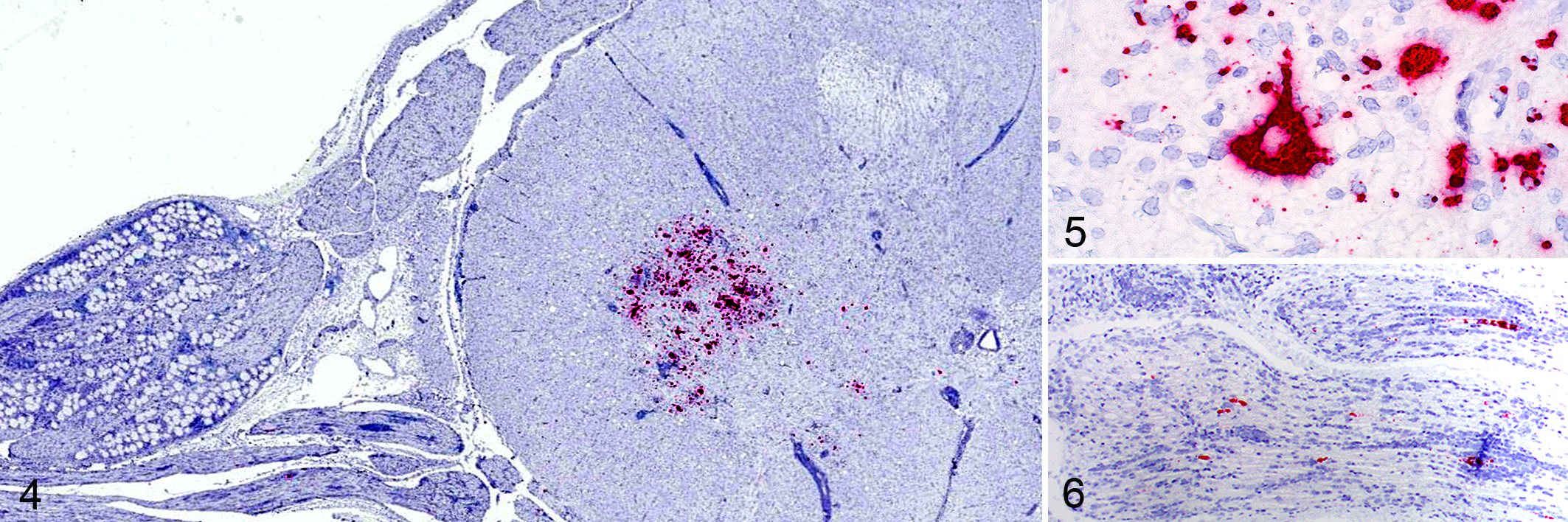
Porcine astrovirus type 3 (PoAstV3), pig 5, 21 days post-infection. In situ hybridization for PoAstV3 (ISH).
Electron Microscopy
In the single animal assayed, pig 5, electron-dense round particles between 20.2 and 28.6 nm in diameter arranged in paracrystalline array—consistent with viral structures—were observed within the soma of a lumbar spinal neuron (Figs. 7, 8).
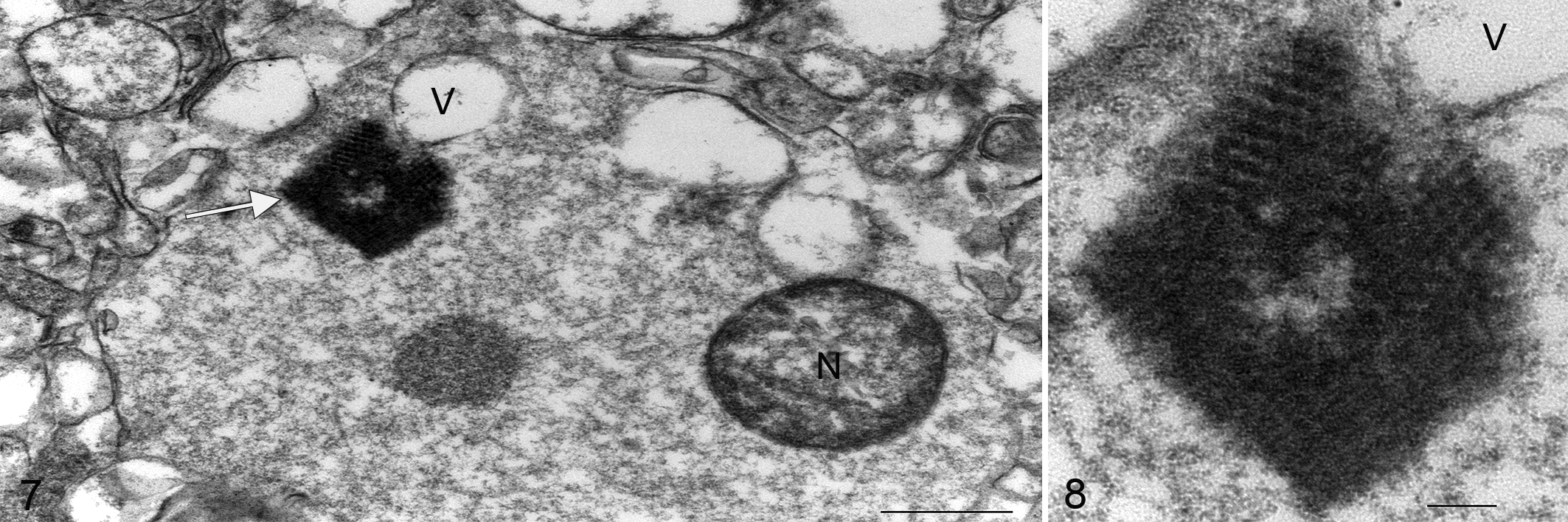
Porcine astrovirus type 3 (PoAstV3), lumbar spinal cord (ventral horn), pig 5, 21 days post-infection.
PoAstV3 RT-qPCR
PoAstV3 was detected in all CNS tissues from inoculated pigs (22 of 24 samples, mean Cq = 31.75, SD: 5.73) except for cerebellum of pigs 3 and 6. The highest viral RNA load was detected in the cerebrum (Cq = 20.36; 5.86 × 107 copies/ml) and cervical spinal cord (Cq = 20.48, 5.39 × 107 copies/ml) of pig 5. The lowest CNS viral RNA load was detected in the cerebellum of pig 4 (Cq = 37.67; 389 copies/ml). PoAstV3 was only detected in the CSF of pig 5 (Cq = 34.48; 223 copies/ml; Table 3). Feces were positive at DPI 3 from pigs 3 (Cq = 37.42, 461 copies/ml) and 5 (Cq = 37.7, 381 copies/ml), and at DPI 14 from pig 6 (Cq = 38.53, 215 copies/ml) (mean Cq = 37.88, 337 copies/ml, SD = 0.33, range = 1.1). PoAstV3 was not detected in any other samples of feces (Suppl. Table S5), sera (Suppl. Table S6), or non-CNS tissues including lung, heart, spleen, kidney, tonsil, jejunum, ileum, colon, inguinal lymph node, mesenteric lymph node, and tracheobronchial lymph node (Suppl. Table S7) across groups.
Genome Features and Phylogenic Analysis
The genomes of USA/IA/48142/2018 and USA/IA/53214/2018 have highly homologous sequences (99.9% nt identity), and excluding the poly A tail, are both 6434 nt in length, with a GC content of 47.1% and 47.4%, respectively. Both strains cluster in proximity and have 2 deletion foci within the 3′ UTR when compared to PoAstV3 US-MO123 strain (feces) and USA/IA/7023/2017 (CNS tissue; Fig. 9, Suppl. Table S8).
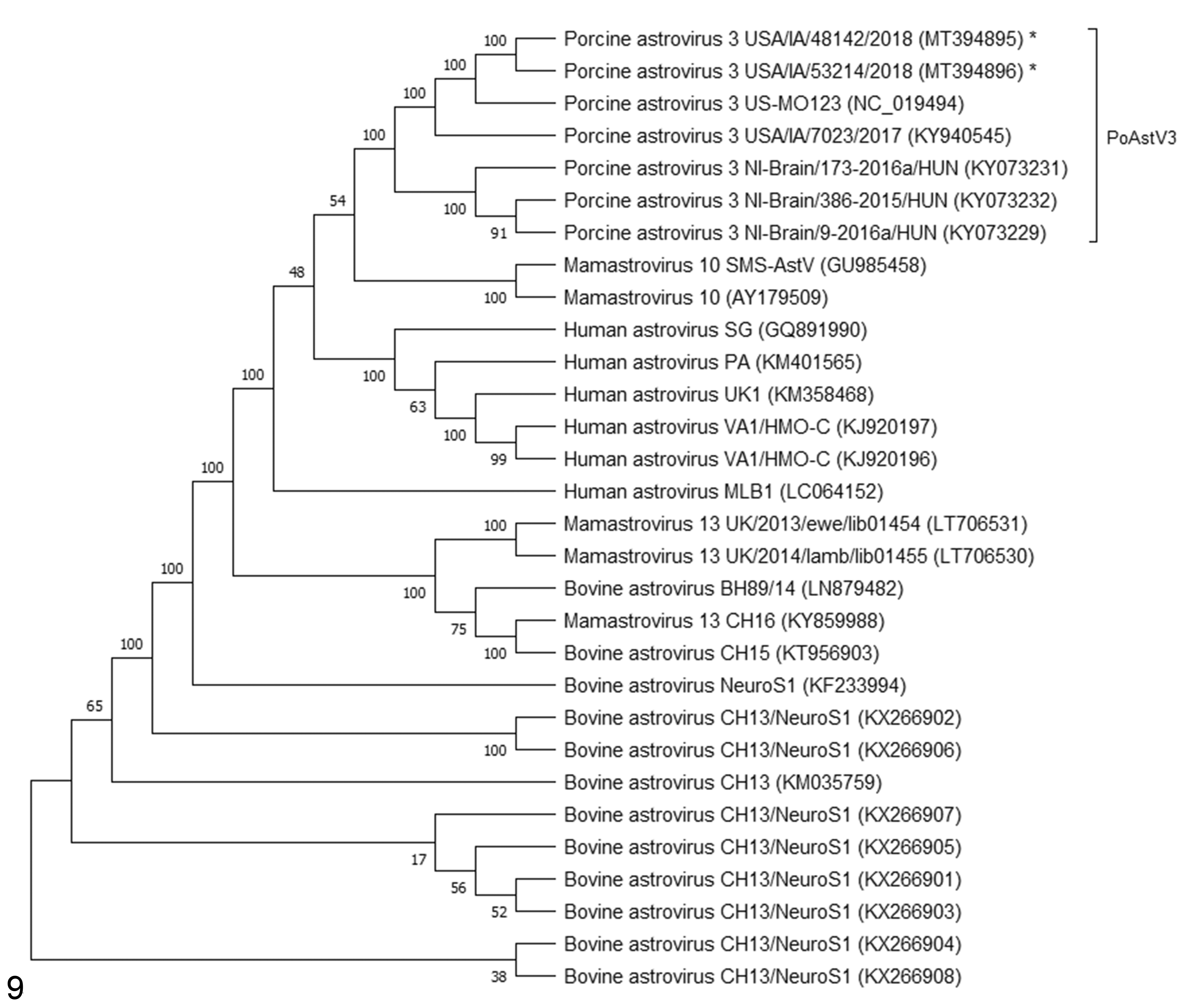

Neighbor joining phylogenetic tree involving 29 full or near-full genomes of neurotropic astrovirus sequences and original porcine astrovirus type 3 (PoAstV3) US-MO123 strain. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test are shown next to the branches. The asterisk indicates strains identified in this study. GenBank accessions are shown in parentheses.
ELISA
The PoAstV3 antibody isotype (IgM, IgA, and IgG) responses are shown in Figures 10 to 12 and Supplemental Table S9. A statistically significant difference in the levels of IgG (P = .0002) and IgM (P = .008) isotypes was observed between inoculated and control animals. A statistically significant increase in the IgG isotype of inoculated animals was observed at DPI 7, 10, 14, and 21 (P = .02, .005, .0001, .002, respectively). The mean S/P value of IgG and IgM in the inoculated group was 2.52 and 4.41 times higher than the control group, respectively. For IgA, a statistically significant difference in the level of specific circulating antibodies was only found at DPI 14 (P = .0003). The maximum IgA S/P value was detected in pig 3 at DPI 14, while the maximum IgG and IgM S/P values were detected in pig 4, both at DPI 14. Overall, the highest levels of IgG in the inoculated group were detected at DPI 14 (pigs 3, 4, and 6) and DPI 21 (pig 5). The highest IgM levels detected in inoculated pigs were detected at DPI 14 (pigs 3, 4, and 5) and DPI 28 (pig 6; Figs. 10–12). Overall, regardless of the antibody isotype, S/P values from control pigs were below the mean S/P value obtained for each isotype, with the exception of DPI 7 IgA S/P value from pig 1.
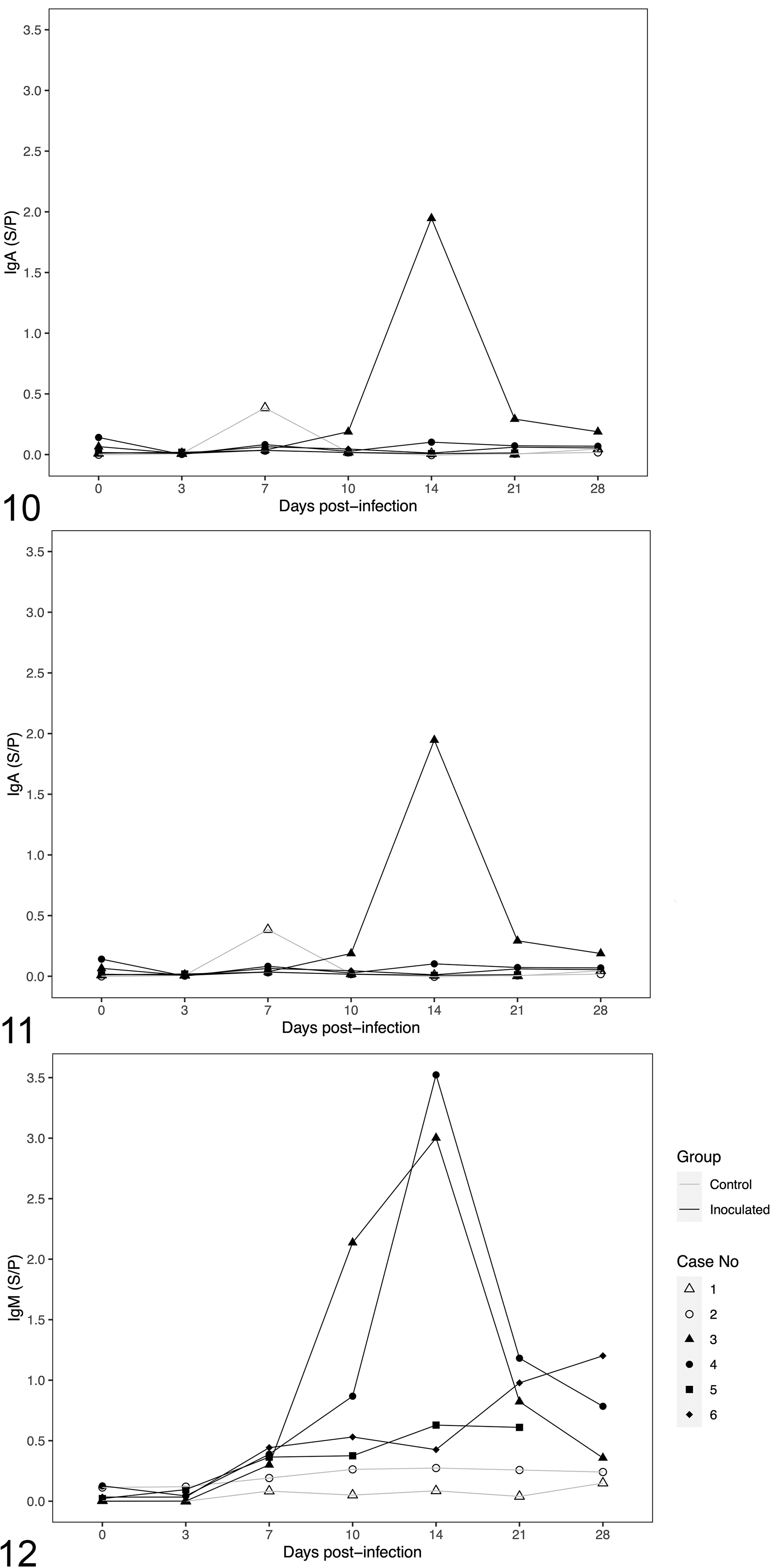
Concentrations of immunoglobulin (Ig) in serum of pigs at different days post-inoculation of porcine astrovirus type 3.
Metagenomic Sequencing
No viral or bacterial reads were identified in the CNS tissues from pig 5.
Exogenous mammalian, viral, and bacterial reads were detected within the intravenous inoculum (Suppl. Table S10). RT-qPCR testing of the intravenous inoculum for pestivirus K (positive, Cq = 32.36) and rotavirus C (positive, Cq = 33.59) was conducted at the ISU-VDL (Suppl. Table S11). PoAstV3 and host reads were detected in the oral inoculum (Suppl. Table S12).
PTV, PSV, and Pestivirus K RT-qPCR
All brainstem samples were negative for PTV and PSV (Suppl. Table S6) and pestivirus K was not detected in the cerebellum or cerebrum of any animal. Pestivirus K was detected at high Cq values in the serum from all PoAstV3-inoculated animals at least twice (Suppl. Table S13).
Discussion
In the last decade, the number of viruses in the genus Mamastrovirus associated with encephalitis/polioencephalomyelitis in different animal hosts has notably increased. 6,12,15,18,26,28,45,56,58,60,68 These neurotropic strains commonly cluster within the VA/HMO clade. 16,21,25,35 Mink intracranially inoculated with a tissue homogenate containing neurotropic strain SMS-AstV (GenBank accession GU985458, tentatively Mamastrovirus 21) developed clinical disease and histologic lesions consistent with shaking mink syndrome (SMS). 12 Interestingly, based on phylogenetic analysis, strain SMS-AstV is closely related to the strain used in this study and other PoAstV3 strains previously described. 49 Additionally, sequence analysis of both USA/IA/48142/2018 and USA/IA/53214/2018 showed multiple deletion foci all located within the 3′UTR when compared to PoAstV3 US-MO123. Previous studies with human astroviruses and other single-stranded RNA viruses indicated that sequence changes in the 3′UTR region may lead to conformational changes related to viral replication and translation. 1,30 Therefore, additional studies are needed to further elucidate if these conformational changes in PoAstV3 are associated with changes in virulence or cellular tropism. 11,27,29
In swine, studies have associated the detection of PoAstV3 with consistent histologic lesions in the CNS and clinical neurologic disease. 6,14,49 However, there are no previous reports of experimental reproduction of CNS disease in swine. In this study, all PoAstV3 inoculated animals had histologic lesions in the CNS consistent with a viral etiology, with PoAstV3 RNA detected by RT-qPCR and ISH in multiple anatomic sections of the CNS.
Histologic lesions were distributed throughout the CNS including the cerebrum, cerebellum, brainstem, and spinal cord. The most consistent histologic feature throughout the CNS was perivascular lymphoplasmacytic infiltrates followed by glial nodules both of which were most frequently observed in the gray matter of cerebrum, brainstem, and ventral horns of the spinal cord. Perivascular aggregates of mononuclear cells were uncommonly observed in the leptomeninges. Other features less frequently observed included lymphohistiocytic radiculoneuritis and ganglioneuritis, scattered neuronal degeneration, and necrosis.
The presence of radiculoneuritis and ganglioneuritis, observed in this study, would suggest axonal transport as one possible mode of viral dissemination similar to other neurotropic viruses. 22,50,54 The likelihood of this route of viral dissemination is further substantiated by the detection of PoAstV3 RNA in the ganglia and nerve roots. Moreover, Rawal et al previously demonstrated that myenteric neurons are permissive to PoAstV3 infection, and combined with the results of this study, could further strengthen the hypothesis that PoAstV3 uses axonal transport as one possible mechanism of viral dissemination. 59
Despite the presence of histologic lesions and detection of PoAstV3 in the CNS by RT-qPCR and ISH in all PoAstV3 inoculated animals, clinical signs varied among infected animals from subclinical to clinical. The variability in clinical signs across inoculated animals is similar to what has been previously reported in another experimental inoculation study using a swine neurotropic virus, PTV. 48 In this study, the onset of CNS signs was first observed at DPI 21 and were similar to those described in naturally infected swine, but also to those reported in humans and across species including paresis, ataxia, paraplegia, and paralysis. 6,14,21,28,33,46,49,65
Three inoculated animals (pigs 3, 4, and 6) were noted to be lethargic and reluctant to move at DPI 25. Only pig 3 developed clinical signs 3 days later. Interestingly, compared to pig 3, pigs 4 and 6 had more severe histologic lesions and lower CNS viral titers as determined by RT-qPCR. These findings suggest that pigs with PoAstV3 infections may have no overt CNS clinical signs or disease. However, as these pigs were euthanized it is unknown whether either would eventually have developed clinical neurologic disease.
In a previous epidemiologic study, PoAstV3 RNA was frequently detected in feces of a large percentage of asymptomatic piglets (86% to 100%) and sows (52% to 90%), suggesting that fecal shedding is a common epidemiologic feature of PoAstV3. 59 Surprisingly, fecal shedding of PoAstV3 was rarely detected in this study, and when detected, Cq values were high consistent with a low viral load. This finding could suggest that PoAstV3 does not need to replicate to high viral titers within the enteric system to cause CNS disease, and/or that intravenous inoculation of PoAstV3 provides access to the CNS through some other mechanism. In reports of neurotropic astrovirus disease, human astrovirus strains clustering within the HAstV-VA1/HMO-C clade were detected inconsistently in feces, and infrequently in serum/plasma and CSF samples. 21,23,33,52,58,63,72 In this study, PoAstV3 RNA was not detected by RT-qPCR in any non-CNS tissue samples or serum from inoculated animals and was detected in a single CSF sample from the most severely affected animal (pig 5). These findings are consistent to those described in diagnostic cases of PoAstV3 disease in which PoAstV3 is rarely detected in samples outside of the CNS at the time of disease by RT-qPCR. 6,49 Overall, these findings suggest that CNS tissue is the most diagnostically sensitive sample type for neurologic astrovirus infection.
As noted before, the inability to adapt, isolate, and propagate the virus in cell culture systems is a major impediment for the study of neurotropic astroviruses. 16,25 To circumvent this, we opted to use a CNS tissue homogenate as inoculum. At the time of inoculation, CNS homogenates were tested by RT-qPCR for the presence of other swine viruses (ie, PTV, PSV, PEV, PRV, PRRSV, and PCV2); however, subsequent metagenomic analysis detected viral reads within the intravenous inoculum that were later confirmed by RT-qPCR at high Cq values. The authors acknowledge that the presence of spurious material within the intravenous inoculum is a limitation of this study. Pestivirus K was detected in sera of inoculated animals but not in CNS tissues (Suppl. Table S12) and is known to cause congenital tremors in piglets as a result of in utero infection but does not have any known clinical or pathologic effect following infection in growing pigs. 7,42 However, we cannot exclude synergic or antagonistic effects of coinfection.
To better understand the immunopathogenesis of PoAstV3, a novel serologic assay based on VP27 of PoAstV3 was developed. VP27 derives from the hypervariable region of the capsid protein, and together with VP25, forms the dimeric spikes of the virus. 5,8,13 Previous studies have shown that spike proteins are highly immunogenic and contain a cell receptor-interaction domain. 5 Our results indicate that inoculated animals had a significant increase in specific circulating antibodies, mainly IgG and IgM isotypes. In humans, extensive epidemiologic studies have relied on ELISA testing to study population parameters, and one study reported dual detection of human and porcine strains and possible recombination events between these strains. 43,51,55,67 The ELISA developed and described herein provides a new tool to study the ecology, epidemiology, and immunopathogenicity of PoAstV3 in both the experimental and field settings, although further evaluation of this assay including a larger sample size is needed to better understand its functionality and sensitivity.
Conclusions
This is the first study reproducing CNS disease with a porcine astrovirus strain consistent with natural infection in swine and across other species with neurologic astrovirus infections. Histologic lesions in the CNS were predominately characterized by perivascular mononuclear inflammatory infiltrates and gliosis. Additionally, based upon ISH results, this study elucidated axonal transport as a probable novel mode of astrovirus dissemination. In the future, swine may serve as a model to study the immunopathogenesis of neurotropic astroviruses.
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211025794 - Experimental porcine astrovirus type 3-associated polioencephalomyelitis in swine
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211025794 for Experimental porcine astrovirus type 3-associated polioencephalomyelitis in swine by Franco Matias Ferreyra, Paulo E. H. Arruda, Laura K. Bradner, Karen M. Harmon, Min Zhang, Luis Giménez-Lirola and Bailey L. Arruda in Veterinary Pathology
Supplemental Material
Supplemental Material, sj-xlsx-1-vet-10.1177_03009858211025794 - Experimental porcine astrovirus type 3-associated polioencephalomyelitis in swine
Supplemental Material, sj-xlsx-1-vet-10.1177_03009858211025794 for Experimental porcine astrovirus type 3-associated polioencephalomyelitis in swine by Franco Matias Ferreyra, Paulo E. H. Arruda, Laura K. Bradner, Karen M. Harmon, Min Zhang, Luis Giménez-Lirola and Bailey L. Arruda in Veterinary Pathology
Footnotes
Acknowledgements
The authors would like to express their gratitude to the staff and faculty from the Iowa State University Veterinary Diagnostic Laboratory; Judith A. Stasko at the National Veterinary Services Laboratories (NVSL) for excellent assistance with electron microscopy; and Dr Alejandro Ramirez at ISU College of Veterinary Medicine, Department of Veterinary Diagnostic and Production Animal Medicine (VDPAM).
Author Contributions
BA, PA, LB, KH, MZ, LGL, and FMF analyzed the data. FMF and BA wrote the paper. All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
