Abstract
In the past decade, different members of the genus Mamastrovirus have been associated with outbreaks of neurologic disease in humans, cattle, sheep, mink, and, most recently, porcine astrovirus 3 (PoAstV3) in swine. We performed a retrospective analysis of 50 cases of porcine neurologic disease of undetermined cause but with microscopic lesions compatible with a viral encephalomyelitis to better understand the role and pathogenesis of PoAstV3 infection. Nucleic acid was extracted from formalin-fixed paraffin-embedded (FFPE) tissue for reverse transcription quantitative polymerase chain reaction (RT-qPCR) testing for PoAstV3. In addition, 3 cases with confirmed PoAstV3-associated disease were assayed by RT-qPCR to investigate PoAstV3 tissue distribution. PoAstV3 was detected in central nervous system (CNS) tissue via RT-qPCR and in situ hybridization in 13 of 50 (26%) FFPE cases assayed. PoAstV3 was rarely detected in any tissues outside the CNS. Positive cases from the retrospective study included pigs in various production categories beginning in 2010, the earliest year samples were available. Based on these results, PoAstV3 appears to be a recurring putative cause of viral encephalomyelitis in swine that is rarely detected outside of the CNS at the time of clinical neurologic disease, unlike other common viral causes of neurologic disease in swine.
Astroviruses (AstVs) are viral agents in the family Astroviridae that are able to infect and cause disease in a wide variety of host species. 1 There are significant gaps in knowledge concerning the epidemiology, ecology, and pathophysiology of most AstVs. 9 AstVs are 28–35 nm, icosahedral, and nonenveloped virions, characterized by a starlike surface structure when visualized using electron microscopy. 12 The current International Committee for Taxonomy of Viruses classification contains 19 recognized species of Mamastrovirus (MAstV1-19), with an additional 14 strains awaiting classification considered as tentative new species including 4 porcine lineages. 3
AstV infections are considered to be one of the most common causes of gastroenteritis in children, but in animals, their association with enteric disease is not well characterized, with the exception of turkey and mink AstV infections. 6,17,20,29 AstVs can also be found in tissues outside the enteric system of poultry. More recently, AstVs have been identified in central nervous system (CNS) tissue of humans, cattle, sheep, mink, and pigs with neurologic disease. *
Porcine AstVs (PoAstVs) are distributed worldwide. 15 Five PoAstV lineages (PoAstV1–PoAstV5) have been identified, possibly reflecting different origins, interspecies transmission, and recombination events, some presumably with human strains. 22,24,30,33 The pathogenic significance of PoAstVs remains unsettled with the likely exception of PoAstV3, which has recently been associated with outbreaks of encephalitis-polioencephalomyelitis in swine in the United States and Hungary. 5,8
PoAstV3 was first described in 2011 in the feces of healthy suckling pigs that were collected in Canada in 2006. 24 Subsequent reports have identified other strains from this lineage in swine from the United States, Canada, Japan, Kenya, Croatia, and Hungary. 2,8,13,19,33 In one of these studies, the overall frequency of detection of PoAstV3 by polymerase chain reaction (PCR) in the feces of pigs from the United States was reported to be 1.2% (6/509), based on samples from suckling to adult pigs from 255 farms located in 19 states (53.4% were from Iowa). 33
To better understand the importance and pathogenesis of PoAstV3 infection, a retrospective analysis of swine neurologic cases of unknown etiology and lesions compatible with a viral cause was undertaken.
Materials and Methods
Case Selection of FFPE Samples
Cases were identified with 1 or more of 5 diagnostic codes including encephalomyelitis, viral (unidentified) encephalitis, idiopathic encephalitis, viral miscellaneous encephalitis, and nonsuppurative encephalitis. The cases were compiled after searching the Iowa State University Veterinary Diagnostic Laboratory (ISU-VDL) information management system database. The search included cases with formalin-fixed paraffin-embedded (FFPE) tissue available from diagnostic cases between March 26, 2010, and August 1, 2017. Corresponding hematoxylin and eosin–stained sections were reviewed by 1 of 9 veterinary diagnostic pathologists who selected which block(s) to be tested by reverse transcription quantitative polymerase chain reaction (RT-qPCR) based on lesions consistent with a neurotropic viral infection. Lesions used as inclusion criteria were gliosis, lymphohistiocytic perivascular cuffs, and neuronal necrosis with lymphocyte satellitosis. The distribution of histologic lesions in the CNS was recorded for each case. Lesions outside the CNS, neurologic clinical signs, and concurrent detection of other porcine pathogens were also recorded.
Case Selection of Fresh Samples
Three cases (cases 14–16), submitted to the ISU-VDL between August 1, 2017, and February 1, 2018, and diagnosed with viral polioencephalomyelitis due to PoAstV3 were identified, and fresh tissues were retrieved for further testing. Animals were between 20 days and 8 weeks of age with compatible histologic lesions of a neurotropic virus, in which PoAstV3 was detected in the CNS via RT-qPCR. All animals were from a nursery endemically infected with PoAstV3 reporting 2% to 10% mortality attributed to PoAstV3 (Suppl. File 1) over the course of at least 5 years. This site was originally identified in the retrospective FFPE samples (case 9).
RT-qPCR of FFPE Samples
Selected paraffin blocks were retrieved, and two 40-μm-thick scrolls of FFPE tissue were prepared using a Thermo Scientific HM 355 S automatic microtome (Thermo Fisher Scientific, Kalamazoo, MI) and stored at room temperature in 1.5-ml Fisherbrand Premium Microcentrifuge Tubes (Catalog No. 05408130, Thermo Fisher Scientific) until further processing. FFPE scrolls were obtained after removal of the 3 most superficial layers of the block. For each individual block, Surgipath DB80 LX microtome blades (Catalog No. 14035843497, Leica Biosystems, Wetzlar, Germany) were replaced, and the microtome’s movable parts were disinfected using a mixture of sodium hypochlorite (10% v/v), ethanol (50% v/v), and purified water (40% v/v). Cases in which PoAstV3 was detected by RT-qPCR from extracts of CNS tissue were also tested for PoAstV3 by RT-qPCR on FFPE blocks having 1 or more of the following tissues: spinal peripheral nerve, heart, liver, kidney, spleen, lung, lymph node, thymus, tonsil, ileum, jejunum, duodenum, colon, and skeletal muscle. Briefly, for the extraction of viral RNA, FFPE scrolls were processed using Mag-Bind FFPE RNA Kit (Catalog No. M2551-01, OMEGA Bio-tek, Norcross, GA) and a Kingfisher flex instrument (Thermo Fisher Scientific) according to the manufacturer’s recommendations. PoAstV3 amplification was carried out in FAST mode using an Applied Biosystems 7500 thermal cycler (Applied Biosystems, Foster City, CA) with TaqMan Fast Virus 1-Step Master Mix (Catalog No. 4444434. Life Technologies, Carlsbad, CA) according to the manufacturer’s instructions. The final protocol consisted of the addition of 5 μL of isolated RNA to 15 μL of RT-qPCR mix containing a final concentration of 0.4 μM primers and 0.2 μM fluorogenic probe and 0.2 μM primers and 0.075 μM fluorogenic probe targeting xeno internal positive control and RNase-free water. The reaction was first subjected to a reverse transcription step at 50°C for 5 minutes, followed by 20 seconds at 95°C and 40 cycles of 3 seconds at 95°C and 30 seconds at 60°C.
RT-qPCR of Fresh Samples
PoAstV3, Teschovirus A (PTV), and Sapelovirus A (PSV) RT-qPCR was performed on tissue homogenate from samples stored individually at –80°C prior to processing. Tissue aliquots were homogenized with 15 ml of Earl’s balanced salt solution (Catalog No. 24010-043, Thermo Fisher Scientific). The following samples, when available, were assayed: spleen, thymus, tonsil, adrenal gland, kidney, liver, heart, lung, stomach mucosa, jejunum, ileum, colon, feces, cerebrum, cerebellum, brainstem, pooled or individual sections of spinal cord (cervical, thoracic, lumbar), and lymph node (mesenteric, tracheobronchial, and inguinal). RT-qPCR for PoAstV3 and PTV was done as previously described 4,5,25,34 ; RT-qPCR for PSV used the same methods. Primers and probes are listed in Table 1.
Reverse Transcription Quantitative Polymerase Chain Reaction Primers and Probes.
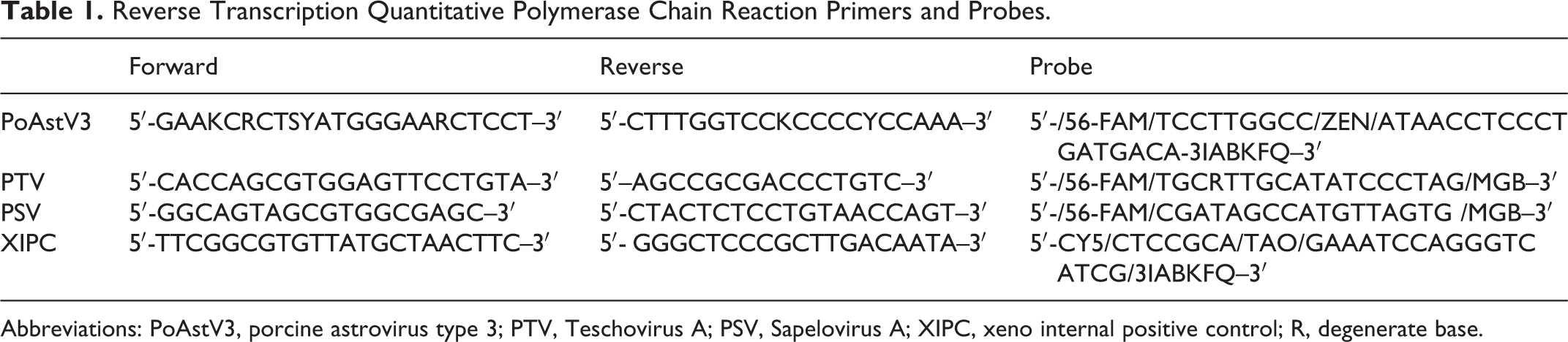
Abbreviations: PoAstV3, porcine astrovirus type 3; PTV, Teschovirus A; PSV, Sapelovirus A; XIPC, xeno internal positive control; R, degenerate base.
Phylogenic Analysis
Because of the fragmented nature of formalin-fixed RNA, 4 primer sets targeting the PoAstV3 ORF2 gene were designed to amplify smaller overlapping sequences that, when combined, would generate a longer final sequence. Sequencing from fresh tissue was performed in 1 reaction. The target sequences were amplified from RT-qPCR products obtained from CNS tissues, used as templates in 25-μL reactions using a custom qScript One-Step RT-PCR Kit (Quanta Biosciences, Beverly, MA) following the manufacturer’s recommendations. Each primer was present in the final reaction at 320 nM, and 5 μL of template was used per reaction. PoAstV3 RNA amplification from FFPE scrolls was performed on an Applied Biosystems 2720 thermal cycler (Applied Biosystems) under the following conditions: initial reverse transcription at 48°C for 20 minutes, followed by initial denaturation at 94°C for 3 minutes and 45 cycles of denaturation at 94°C for 30 seconds; annealing at 50°C for 30 seconds; extension at 68°C for 30 seconds; and a final elongation at 68°C for 7 minutes. RNA amplification of the fresh tissues was performed on an Applied Biosystems 2720 thermal cycler (Applied Biosystems) under the following conditions: initial reverse transcription at 48°C for 20 minutes, followed by initial denaturation at 94°C for 3 minutes, and 45 cycles of denaturation at 94°C for 30 seconds; annealing at 55°C for 30 seconds and extension at 68°C for 30 seconds with a final elongation at 68°C for 7 minutes. Primer sequences for PoAstV3 sequencing are presented in Table 2. The PCR products were visualized using QIAxcel Advanced System (Qiagen, Venlo, the Netherlands) and purified using the ExoSAP-IT PCR Product Cleanup Reagent (Catalog No. 78200.200.UL, Thermo Fisher Scientific) according to the manufacturer’s instructions. Sequencing was completed via Sanger sequencing at the Iowa State University DNA Facility. For phylogenic analysis, 12 other PoAstV3 sequences and PoAstV1, PoAstV2, PoAstV4, and PoAstV5 reference strains and 30 neurotropic AstV sequences from human, cattle, sheep, mink, and swine sequences were retrieved from GenBank and aligned using MUSCLE v3.8.31. 16 Neighbor-joining trees were built using ClustalW 2.1 made available through the European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI). 18,21 Resultant trees were rendered using FigTree v1.4. 28
PoAstV3 Sequencing Primers.
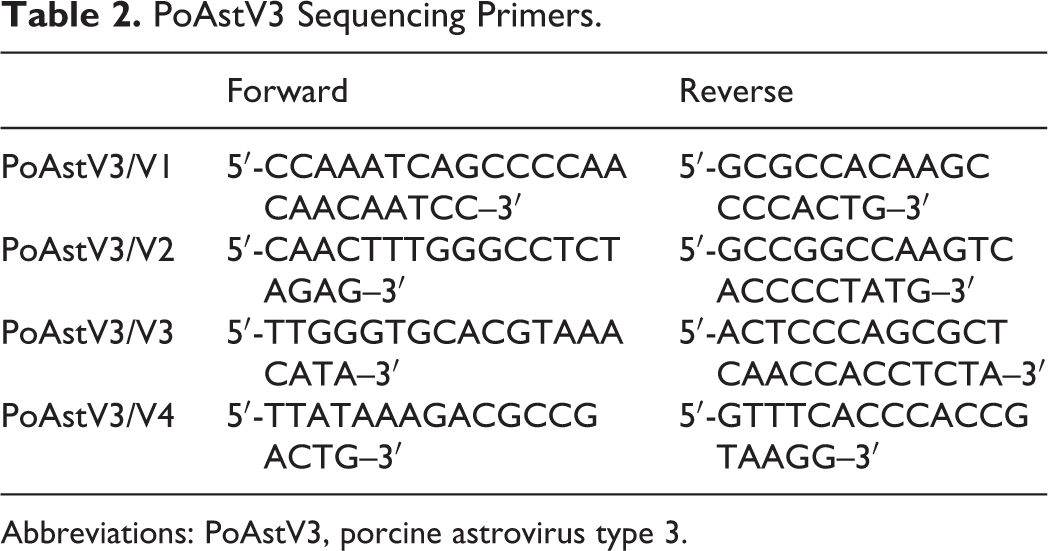

Abbreviations: PoAstV3, porcine astrovirus type 3.
In Situ Hybridization
RNA in situ hybridization (ISH) was performed with RNAScope 2.5 HD Reagent Kit-Red (Catalog No. 322350. Advanced Cell Diagnostics, Newark, CA) according to the manufacturer’s instructions. Paraffin blocks stored at room temperature from each selected case were retrieved, and 5-µm sections were trimmed and mounted on Superfrost Plus slides (Catalog No. 4951PLUS4, Thermo Fisher Scientific). Slides were then heated for 1 hour at 60°C followed by a deparaffinization treatment consisting of 2 consecutive washes of 100% xylene and 2 washes of 100% ethanol. Slides were treated with Pretreat 1 solution (hydrogen peroxide) at room temperature for 10 minutes and rinsed with distilled water. Slides were then immersed in target retrieval solution for 30 minutes at 100°C, with subsequent protease treatment for 30 minutes at 40°C. Probes were then hybridized for 2 hours at 40°C. RNAScope probes targeting PoAstV3 capsid RNA (Catalog No. 516231), RNAScope positive control probe Sc-PPIB targeting swine PPIB gene (Catalog No. 428591), and RNAScope negative control probe DapB (Catalog No. 310043) were designed and synthesized by Advanced Cell Diagnostics. After this step, 6 rounds of amplification were performed (solution kit Amp1-6) followed by the incubation with red chromogenic detection solution for 10 minutes at room temperature. Slides were counterstained with 50% hematoxylin solution for 2 minutes, rinsed in a 0.02% ammonia solution for 10 seconds, and then washed 5 times in distilled water. Slides were then dried at 60°C for 15 minutes and cooled down at room temperature for 5 minutes. Finally, slides were submerged into a 100% xylene solution and immediately coverslipped with EcoMount solution (Catalog No. EM897 L, Biocare Medical, Pacheco, CA). Slides were visualized using an Olympus BX43 bright-field microscope (Olympus Corporation, Tokyo, Japan).
Results
Case Selection of FFPE Samples
A total of 153 cases were found matching 1 or more of the 5 diagnostic codes described in the search criteria; 50 had lesions meeting inclusion criteria and were selected for further testing (Fig. 1A). Of these 50 cases, 29 (58%) were originally from Iowa, 13 (26%) from undefined locations, 2 (4%) from Illinois, and 1 (2%) from each of the following states: Nebraska, Virginia, Minnesota, Indiana, Kentucky, and North Carolina.
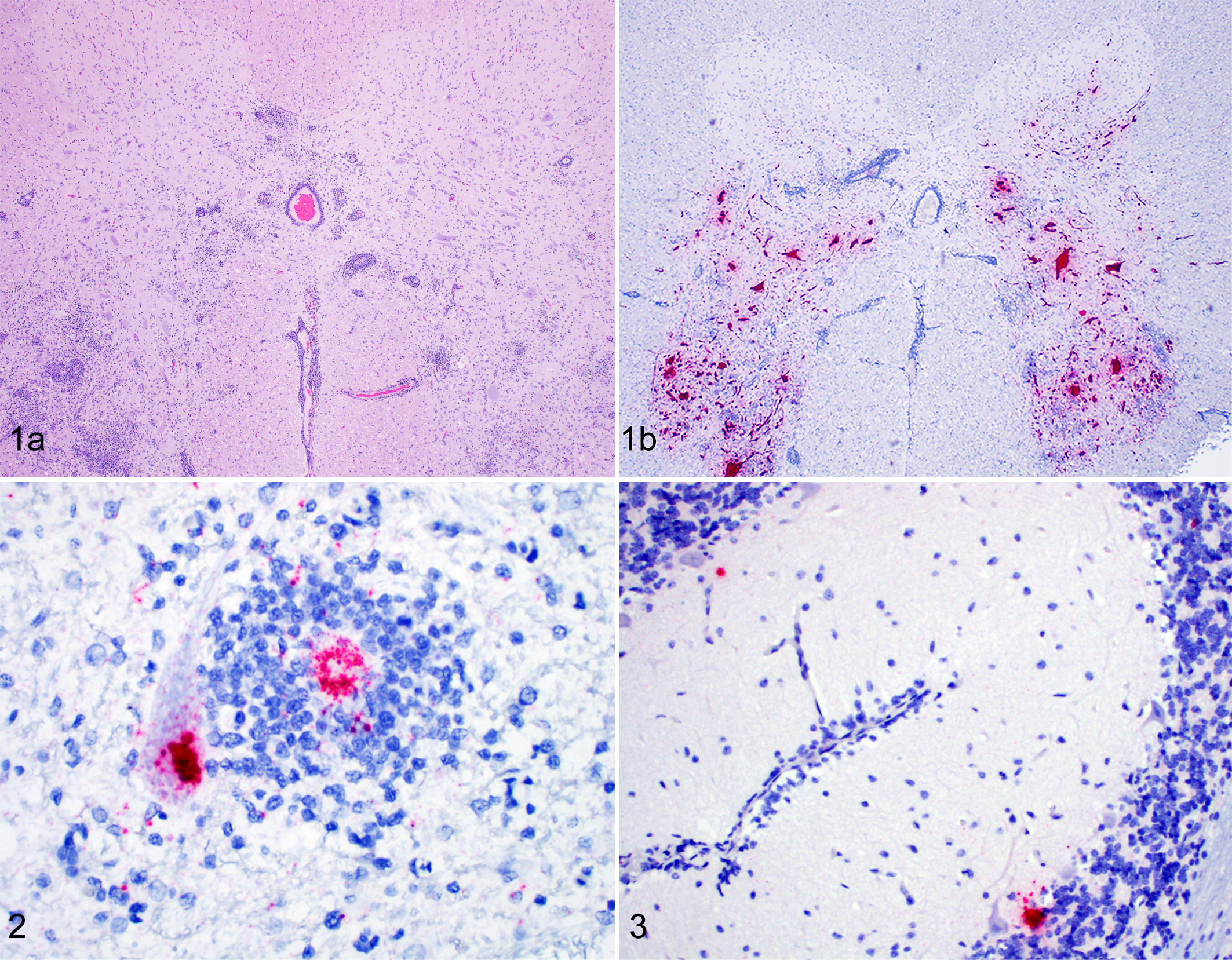
Porcine astrovirus type 3 (PoAstV3)–associated polioencephalomyelitis, pig.
RT-qPCR on FFPE Samples
PoAstV3 was detected in FFPE CNS tissue by RT-qPCR in 13 of the 50 (26%) cases (cases 1–13; Table 3). However, PoAstV3 was not detected in any FFPE non-CNS tissue from these 13 cases (Suppl. Table S1). Of the 13 RT-qPCR–positive cases, 3 occurred in 2016 and 2 cases per year occurred in 2015, 2014, 2012, 2011, and 2010 (Table 3). The most commonly affected age category was adult females (1 year old or more, 6/13), followed by finishing pigs (10 to 24 weeks old, 4/13), and nursery pigs (5 to 6 weeks old, 3/13). Each case originated from a different site with no apparent connection. Clinical signs frequently reported on the diagnostic submission form included lateral recumbency and paresis (Suppl. Table S2).
Detection of Porcine Astrovirus 3 in Cases 1–13.
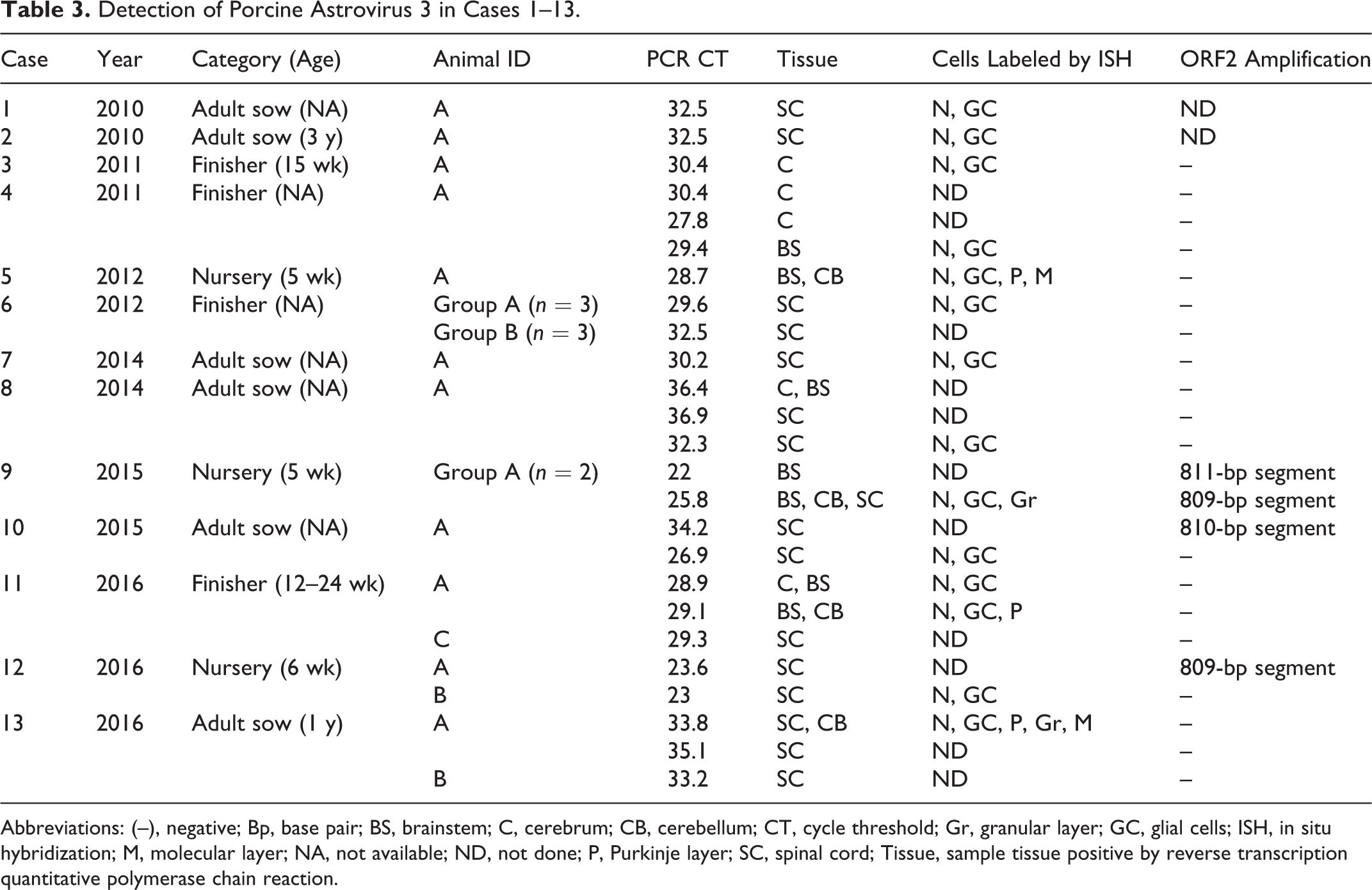
Abbreviations: (–), negative; Bp, base pair; BS, brainstem; C, cerebrum; CB, cerebellum; CT, cycle threshold; Gr, granular layer; GC, glial cells; ISH, in situ hybridization; M, molecular layer; NA, not available; ND, not done; P, Purkinje layer; SC, spinal cord; Tissue, sample tissue positive by reverse transcription quantitative polymerase chain reaction.
Distribution of Histologic Lesions
Histologic lesions were most commonly observed in sections of brainstem (12/12 cases), spinal cord (11/11 cases) and less commonly in the cerebrum (9/10 cases), cerebellum (5/9 cases), and leptomeninges (6/16 cases; Suppl. Table S2). When the spinal cord was available for evaluation, inflammation was commonly noted in the ventral horns. In cases in which lesions were severe, inflammation frequently extended into the white matter and leptomeninges. In cases 14 to 16, the most severe lesions were observed in the brainstem and spinal cord, while the cerebellum and cerebrum had more mild lesions. Often, non-CNS tissues did not contain significant histologic lesions. However, pneumonia was observed in 3 of 10 cases in which lung was available for evaluation. Animal A from case 7 had purulent bronchopneumonia attributed to a bacterial cause, and a single animal in case 9 and 11 had concurrent lymphohistiocytic interstitial pneumonia attributed to porcine reproductive and respiratory syndrome virus (PRRSV).
In Situ Hybridization
PoAstV3 was detected in situ most commonly in the cytoplasm of neurons and dendritic processes and less commonly in glial cells (cases 1–13; Table 3; Figs. 1 and 2). In the 3 cases with cerebellum available, PoAstV3 was rarely detected in Purkinje cells or cells of the molecular layer and internal granular cell layer (Fig. 3).
RT-qPCR of Fresh Samples
PoAstV3 was detected throughout the CNS of cases 14 to 16 (Table 4). Case 14 comprised 4 animals having histologic lesions consistent with a neurotropic virus; PoAstV3 was detected in all 4 animals throughout the CNS (cerebrum, cerebellum, brainstem, and cervical, thoracic, and lumbar spinal cord). Case 15 consisted of 4 animals, and PoAstV3 was detected in all CNS samples from the 2 animals (animals C and D) that had histologic lesions of polioencephalomyelitis. In case 16, which included 3 animals, PoAstV3 was detected in the cerebrum, cerebellum, brainstem, and different anatomical regions of the spinal cord of animals B and C and detected in only the brainstem of animal A at a high Ct value (Ct = 37.6). However, polioencephalomyelitis was observed only in animals B and C. PoAstV3 was not detected in any of the non-CNS tissues examined from cases 14 and 15. In case 16, PoAstV3 was detected in gastrointestinal samples in 2 animals: in the jejunum (Ct = 23.8), ileum (Ct = 25.7), colon (Ct = 28.5), and mesenteric lymph node (Ct = 34.7) of animal A and in the colon (Ct = 33.0) of animal C. PTV and PSV were not detected in CNS tissue from any case and were commonly detected in the gastrointestinal tract, mesenteric lymph node, and tonsil (Suppl. Table S3).
Detection of Porcine Astrovirus 3 by RT-qPCR in Different Tissues From Cases 14–16.a
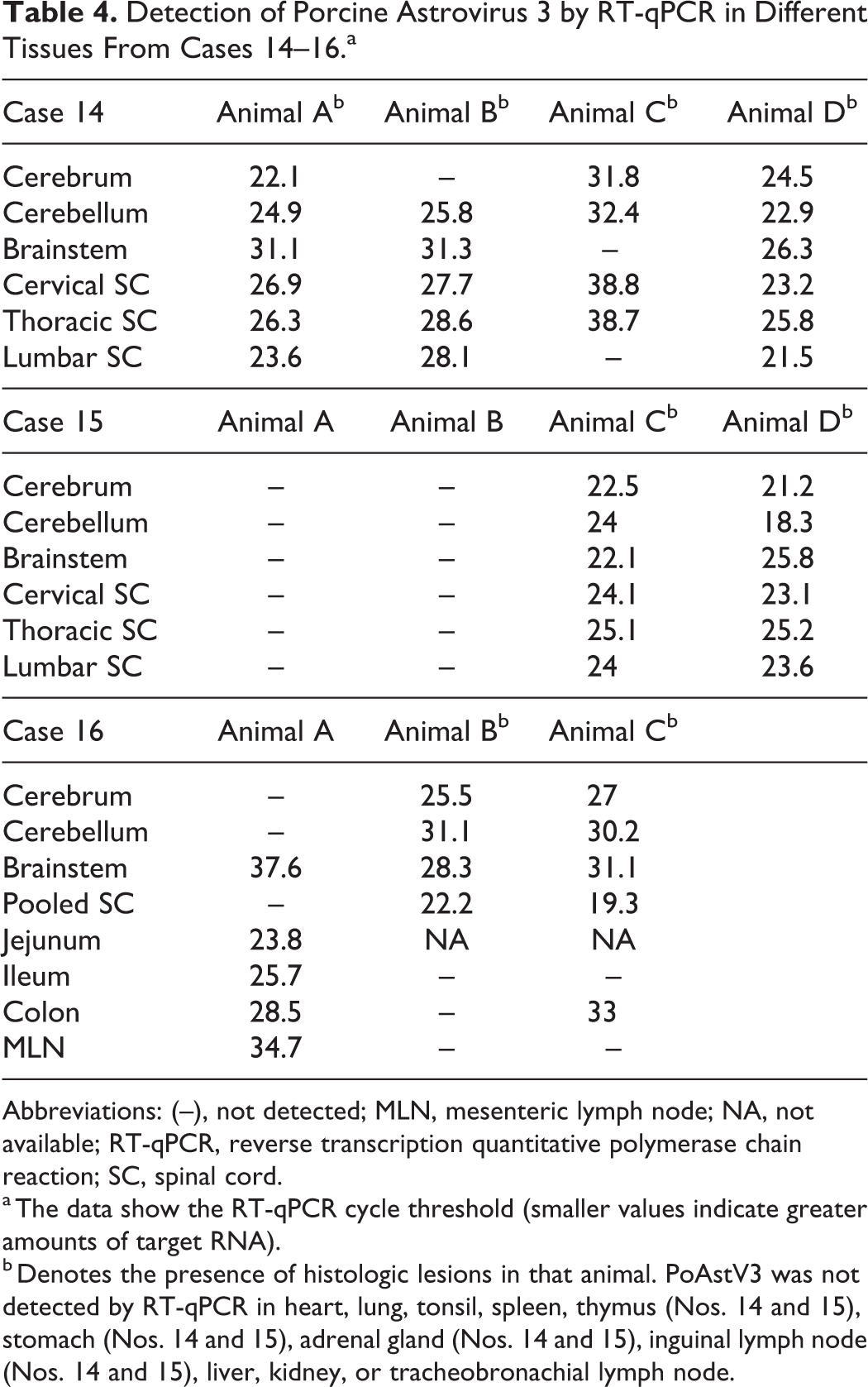
Abbreviations: (–), not detected; MLN, mesenteric lymph node; NA, not available; RT-qPCR, reverse transcription quantitative polymerase chain reaction; SC, spinal cord.
a The data show the RT-qPCR cycle threshold (smaller values indicate greater amounts of target RNA).
b Denotes the presence of histologic lesions in that animal. PoAstV3 was not detected by RT-qPCR in heart, lung, tonsil, spleen, thymus (Nos. 14 and 15), stomach (Nos. 14 and 15), adrenal gland (Nos. 14 and 15), inguinal lymph node (Nos. 14 and 15), liver, kidney, or tracheobronachial lymph node.
Phylogenic Analysis
Four partial ORF2 sequences of 809 to 811 bp were obtained from 3 of the 13 PoAstV3 FFPE cases (cases 9, 10, and 12; Table 3). Sequences shared a percentage identity ranging from 93.14% to 99.75% nucleotide (nt) homology. Each FFPE case originated from a different site (farm) with no apparent connection. Eight sequences were obtained from cases with fresh tissue (cases 14–16), ranging from 99.59% to 100% nucleotide homology. Three sequences were obtained from the spinal cord of animals A, B, and D in case 14 (99.6% to 99.73% nt), 2 sequences from the brainstem of animals C and D in case 15 (99.79% nt), and 3 sequences from the colon of animal A and spinal cord of animals B and D in case 16 (99.86% to 100% nt; Suppl. Table S4). Altogether, these 12 sequences shared between 92.8% and 100% nucleotide identity, and with the exception of the sequence obtained from the colon of animal A in case 16, all were obtained from CNS tissues. Tree topology, distances, and relation with other PoAstV3 strains are shown in Supplemental Figs. 1 and 2. For phylogenetic analyses, 2 trees were constructed, the first reflecting the relation of newly identified strains with 12 other PoAstV3 sequences and PAstV1, PAstV2, PAstV4, and PAstV5 reference strains (Suppl. Fig 1), and the second tree reflecting the relation with 30 neurotropic AstV sequences from human, bovine, ovine, mink, and porcine (Suppl. Fig S2). Homology matrices are provided (Suppl. Tables S5 and S6). All sequences were deposited in GenBank under accession numbers MH686390–MH686400.
Discussion
Recently, PoAstV3 has been associated with neurologic disease and lesions of polioencephalomyelitis in swine in the United States and Hungary. 5,8 To improve our understanding of PoAstV3, a retrospective analysis of swine neurologic cases of unknown etiology and lesions compatible with a viral agent was undertaken. PoAstV3 was detected by RT-qPCR and ISH in FFPE CNS tissue in 13 of 50 (26%) cases assayed. These cases included pigs in varying production categories beginning in 2010, the earliest year samples were available. Three cases were detected in 2016. The most commonly affected age category was adult females, followed by finisher pigs and nursery pigs. Clinical signs reported on diagnostic submission forms frequently included lateral recumbency, paresis, and ataxia without mention of paddling or nystagmus. Each case originated from a different site located in Iowa with the exception of cases 14–16, which originated from the same site as case 9. A vast majority of cases submitted to the ISU-VDL originate from sites located in Iowa, which likely accounts for the geographical distribution of cases in this report.
Although not all sections of the CNS were available for histologic evaluation, lesions were most commonly observed in the brainstem and spinal cord and less commonly in sections of cerebrum, cerebellum, and leptomeninges. Inflammation in the spinal cord was consistently observed in the ventral horns. When lesions were noted to be severe, inflammation also frequently extended into the white matter and leptomeninges. The distribution of histologic lesions in the CNS reported in this study is similar to and likely indistinguishable from those described for other neurologic viral causes of swine including PTV and PSV. 4,5,8,25
PoAstV3 was detected by ISH most consistently in the cytoplasm of neuronal cell bodies and dendrites, less commonly in adjacent glial cells, rarely in Purkinje cells, and in the granular layer as well as the molecular layer of the cerebellum. The localization of viral RNA in histologic lesions was similar to that found via ISH in pigs affected by PoAstV3 in Hungary and in cattle affected by neurotropic strains of bovine AstV (BoAstV) by ISH and IHC. 8,10,11 In those studies, viral RNA was detected in neuronal bodies, dendritic processes, and glial cells. In contrast, in a human case of encephalitis from a patient with X-linked agammaglobulinemia, viral antigen was found in astrocytes but not in neurons, oligodendrocytes, and macrophages using IHC. 27 Based on these data, it would appear that the neuron is the primary permissive cell type in the CNS to PoAstV infection in swine and cattle.
PoAstV3 was most commonly detected in neurologic tissue, with the exception of 2 animals from case 16. In these animals, PoAstV3 was detected in the colon of both animals as well as the jejunum, ileum, and mesenteric lymph node in a single animal. These findings are somewhat similar to those reported in the initial description of PoAstV3 in the United States and Hungary, in which PoAstV3 was not detected in any tissue outside the CNS and was less commonly detected in serum, feces, ileum, myocardium, tonsil, lung, liver, and lymph nodes, respectively. 5,8 AstVs are most commonly reported to have a fecal-oral transmission. 1 These results suggest that fecal shedding may be a route of viral excretion. As seen in case 16, it is evident that virions with 100% ORF2 nucleotide homology can reside within the gastrointestinal tract of an animal without histologic lesions in the CNS (animal A) and in CNS tissues of animals with histologic lesions (animals B and C), suggesting that the gastrointestinal tract may be a likely portal of entry. However, detection of PoAstV3 in the gastrointestinal tract of animals with PoAstV3 neurologic disease is uncommon at the time of diagnosis, whereas both PTV and PSV are commonly found in the feces of pigs with PTV- or PSV-associated neurologic disease at the time of diagnosis. 4,25 This could suggest that there are important differences in the pathogenesis of PoAstV3 compared with PTV and PSV infections. This is also supported by the finding that PoAstV3 is able to cause disease in sows, whereas this does not appear to be the case with the current strains of PTV or PSV circulating in the United States swine herd.
Cases 14 to 16 originated from a farm first identified in the retrospective study, which continues to experience PoAstV3-associated neurologic disease. These findings suggest that affected farms could experience ongoing disease that spans months to years with a clinical prevalence varying from 2% to 3% up to 10%. Interestingly, this farm reports no neurologic disease on the sow farm or finisher. This may be a result of numerous factors, including herd immunity, environmental stressors, infection dynamics, comorbidities, and viral load.
Impaired immunity is commonly reported in human cases of AstV-associated neurologic disease. However, PRRSV was detected via PCR only in pooled lung samples from animals in cases 9 and 11, and influenza A virus (IAV) was not detected via PCR in the cases in which testing was warranted. Lesions consistent with porcine circovirus 2 systemic disease (PCV2-SD) were not observed in any case. These findings suggest that at the time of neurologic disease development, concurrent disease due to PRRSV, IAV, and/or PCV2-SD is not a common finding. However, risk factors associated with PoAstV3-associated neurologic disease including immunosuppression warrant additional investigation.
The phylogenetic analysis based on ORF2 sequences described in this article are clustered closer to neurotropic strain USA/IA/7023/2017 (KY940545), fecal strain US-MO123NC (019494.1), and Japanese fecal isolates (LC201595-LC201599) compared with Hungarian neurotropic strains (KY073229-KY073232). Because strain US-MO123 was identified from feces collected in Canada in 2006, it seems plausible that this strain is a predecessor or shares common ancestors with the neurotropic strains identified in this study and strain PoAstV3 USA/IA/7023/2017. 24
Lastly, all PoAstV3 strains described in this study are closely related to strains belonging to the HMO cluster containing a mink strain (GU985458) and 5 human strains (KJ920196, KJ920197, KM358468, KM401565, GQ891990) identified in cases of neurologic disease, suggesting that they share a common ancestor. This cluster consistently includes strains known to cause neurologic disease and could suggest the presence of a common viral determinant that confers neurotropism. 9,14,23,24,31
Conclusions
Based on these data, PoAstV3 appears to be a recurring putative cause of viral polioencephalomyelitis in swine at various production stages. Significant gaps in knowledge concerning the epidemiology, ecology, and pathophysiology of PAstV3 remain. In vivo studies, field investigations, virus isolation, and serologic assay development will provide the necessary tools and foundational knowledge to assist in the prevention, control, and/or elimination of PoAstV3.
Supplemental Material
Supplemental Material, DS1_VET_10.1177_0300985819875741 - Polioencephalomyelitis in Domestic Swine Associated With Porcine Astrovirus Type 3
Supplemental Material, DS1_VET_10.1177_0300985819875741 for Polioencephalomyelitis in Domestic Swine Associated With Porcine Astrovirus Type 3 by Franco S. Matias Ferreyra, Laura K. Bradner, Eric R. Burrough, Vickie L. Cooper, Rachel J. Derscheid, Phillip C. Gauger, Karen M. Harmon, Darin Madson, Pablo E. Piñeyro, Kent J. Schwartz, Gregory W. Stevenson, Michel A. Zeller and Bailey L. Arruda in Veterinary Pathology
Supplemental Material
Supplemental Material, DS2_VET_10.1177_0300985819875741 - Polioencephalomyelitis in Domestic Swine Associated With Porcine Astrovirus Type 3
Supplemental Material, DS2_VET_10.1177_0300985819875741 for Polioencephalomyelitis in Domestic Swine Associated With Porcine Astrovirus Type 3 by Franco S. Matias Ferreyra, Laura K. Bradner, Eric R. Burrough, Vickie L. Cooper, Rachel J. Derscheid, Phillip C. Gauger, Karen M. Harmon, Darin Madson, Pablo E. Piñeyro, Kent J. Schwartz, Gregory W. Stevenson, Michel A. Zeller and Bailey L. Arruda in Veterinary Pathology
Footnotes
Acknowledgements
We would like to express our gratitude to the staff and faculty of the ISU-VDL for helping with this study. B.L.A., E.R.B., V.L.C., R.J.D., P.C.G., D.M., P.E.P., G.W.S., and K.J.S. examined slides. B.L.A., E.R.B., V.L.C., R.J.D., P.C.G., D.M., F.S.M.F., P.E.P., G.W.S., K.J.S., L.K.B., K.M.H., and M.A.Z. analyzed the data. F.S.M.F. and B.L.A. wrote the article. All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
