Abstract
Amyloid fibrils are characterized by a linear morphology and a cross-β structure. Polymorphic and multiple fibril morphologies can be found when amyloid fibrils are extracted from amyloid-laden tissue. In this study, we report on the purification and transmission electron microscopic analysis of amyloid fibrils from 5 different animal species (mouse, cow, goat, dog, and camel) with AA amyloidosis. The results show that amyloid fibrils had a linear morphology with a cross-structure and irregular length in vivo. Although the fibrils from these different species showed highly similar conformations, there were significant differences in fibril width and crossover distance. We analyzed the sequences of homologous amyloid proteins and serum amyloid A, an evolutionarily conserved protein and a major amyloid precursor. We found 78.23% homology between the most distant amyloid proteins. The findings suggested similar fibril width and crossover distance in different animal species that displayed high homology of amyloid protein sequences. Dog and camel, as well as goat and cow, showed high genetic homology and similar fibril morphology. These data indicate that the fibrils from different animal species have similar genetic homology and morphology, which may provide a better understanding of the pathogenesis of amyloidosis.
Amyloidosis is a group of diseases characterized by the deposition of fibrils that are comparatively resistant to degradation, because they have a β-sheet structure that leads to hydrogen bonding between misfolded monomers in tissues, resulting in organ dysfunction. 14,18,35 There are currently more than 50 different peptides or proteins associated with amyloidosis in humans and domestic animals, and amyloid fibrils contribute to the pathogenesis of many disorders including rheumatoid arthritis, tuberculosis, type 1 diabetes, and others. 2,6,25 Serum amyloid A (SAA) is a highly evolutionarily conserved protein in vertebrates and invertebrates that is encoded by a family of closely related genes. 22 SAA is a multifunctional acute phase protein, a biomarker of inflammation in plasma, and an amyloid precursor, and it is mainly synthesized in the liver. 12,30 Synthesis of SAA is stimulated by multiple agonists such as interleukins (IL)-1 and -6, TNF-α, lipopolysaccharide (LPS), and NF-κB signaling, which induce an abnormal increase in plasma SAA concentrations. 1,3,29 In AA amyloidosis, SAA is cleaved to form AA, which is deposited as amyloid fibrils in kidneys, spleen, and other organs, resulting in organ damage. 17,21,37 The transmission of SAA, which is similar to that of prions, is possible via oral uptake and across different animal species. AA amyloidosis was experimentally induced by inoculating AA from different animal species, namely, cow, cats, and mice. 24,28,34,36
The same amyloid protein, which from different species or between in vivo and in vitro conditions, can assemble into amyloid fibrils with different arrangements of subunits or width and crossover distance. 6,7,9 Moreover, cross-species infection may occur between different animals like bovine, feline, and mouse, although the relationship between the structure and sequence of SAA and cross-species infection remains unclear. 11 Although the various amyloid proteins have unrelated sequences, they can all form fibrils with a similar ultrastructural appearance. 39 In this study, we purified and extracted the amyloid fibrils from 5 animal species (mouse, cow, goat, dog, and camel) and examined them by transmission electron microscopy (TEM). We measured their width and crossover distance, which impact the helical twisting of amyloid fibrils extension width and periodicity twisting. Furthermore, we analyzed the sequences of amyloid fibrils and SAA of these 5 animal species and humans and established a phylogenetic tree to assess the relationship between gene homology and fibril forms. Our analyses provide insight into the mechanism of misfolding and fibril cross-seeding in SAA-mediated AA amyloid formation.
Materials and Methods
Animals
The study used frozen samples and formalin-fixed paraffin-embedded samples of liver or kidney from one case from each species: mouse (Mus musculus), cow (Bos taurus), goat (Capra aegagrus hircus), dog (Canis lupus familiaris), and camel (Camelus bactrianus). All cases had been diagnosed with AA amyloidosis by histopathology (Congo red stain) and immunolabeling of the amyloid deposits for SAA1.1. Additional information on the cases is provided in Supplemental Table S1. Research was approved by the Research Center of Global Agromedicine, Obihiro University of Agriculture and Veterinary Medicine, Obihiro, Japan. Mouse AA amyloidosis was experimentally induced as previously described. 5,34 Briefly, interleukin-I receptor antagonist knockout (IL-I raKO) mice were intraperitoneally (ip) injected with 500 µl of amyloid enhancing factor (AEF) and subcutaneously (sc) injected with 500 µl of 2% AgNO3 to induce amyloidosis. Mice were euthanized by whole blood collection under deep anesthesia, then samples were collected. Other samples were collected from natural cases of AA amyloidosis.
Extraction and Purification of Amyloid Fibrils From Fresh Tissues
Amyloid fibrils were extracted according to Pras’ method. 20 Briefly, 1 to 1.5 g of tissue was homogenized with 15 ml of saline on ice for 30 seconds (5 kHz) in a sonicator (TAITEC). The supernatant was transferred to an ultracentrifuge tube and ultracentrifuged (Optima L-100 XP Ultracentrifuge, Beckman Coulter) at 35 000 × g for 50 minutes at 4 °C. After centrifugation, the supernatant was discarded, and the pellet was suspended in 15 ml of saline. This washing step was repeated 10 times. After washing, the pellet was suspended in 15 ml of distilled water (DW), and centrifuged at 35 000 × g for 45 minutes. The first supernatant was discarded, and the pellet was suspended in DW and stored at 4 °C overnight. After quick sonication (5 kHz), the solution was centrifuged at 35 000 × g for 45 minutes. The second supernatant was collected, and the pellet was resuspended in DW, centrifuged similarly to the first supernatant, and the third supernatant was collected. The second and third supernatants were transferred to a new tube and centrifuged at 125 000 × g for 60 minutes. The supernatant was discarded, and the pellet was collected. After centrifugation at 125 000 × g for 60 minutes, pellets were suspended in 1 ml of DW and kept in autoclaved centrifuge tubes at −80 °C until use.
Histopathological Analysis
Histopathological sections were prepared from formalin-fixed paraffin-embedded tissues. Sections were stained with Puchtler’s alkaline Congo red. Immunohistochemistry (IHC) was performed using goat anti-mouse SAA1.1 polyclonal antibody (1:400, R&D Systems) and mouse anti-human AA monoclonal antibody (Kyowa Medex) as previously described. 33 Peroxidase-conjugated donkey anti-goat IgG (1:400, Bethyl) and an EnVision Polymer Detection System (Agilent Technologies) were used for signal detection.
Immunoblot and SDS-PAGE Analysis of Amyloid Extracts
Amyloid fibril extracts were mixed with Laemmli sample buffer (TAKARA) and boiled at 100 °C for 5 minutes. Samples were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE; 15% polyacrylamide gel; Bio-Rad). 8 Gels were stained with Coomassie brilliant blue (CBB; Bio-Rad) for 15 minutes and then decolored with 7.5% acetic acid. To confirm whether the prepared amyloid extracts were SAA proteins, we performed dot blot and Western blot analyses. Western blot analysis was performed according to our previous method. 34 Dot blot analysis was performed according to the following method: 2 µl of amyloid fibril extracts were transferred onto a polyvinylidene fluoride membrane after drying at room temperature, and blocked with 1% skimmed milk in phosphate-buffered saline with Tween-20 for 1 hour at room temperature. In these immunoblot analyses, we used rabbit anti-mouse SAA polyclonal antibody (R&D Systems) as the primary antibody and horseradish peroxidase–conjugated goat anti-rabbit IgG (Santa Cruz Biotechnology) as the secondary antibody. After Clarity Western CEL substrate (Bio-Rad) was stained, membranes were detected by ImageQuant LAS 500 (GE Healthcare Life Sciences).
Homology Analysis
A total of 29 sequences of SAA protein from 5 vertebrate animal species (cow, goat, mouse, dog, and camel) and humans were searched on the NCBI website (http://www.ncbi.nlm.nih.gov). DNAMAN software (Lynnon Biosoft) was used to analyze SAA sequence homology and to establish a phylogenetic tree by protein multiple sequence alignment.
Observation of Amyloid Fibrils by Transmission Electron Microscopy (TEM)
A total of 10 µl of extracted and purified amyloid fibrils were prepared. Each sample was transferred onto 200-mesh copper grids (Nisshin), incubated for 10 minutes, then negatively stained with 1% phosphotungstic acid for 5 minutes. This procedure was repeated twice at room temperature. Specimens were examined under an H-7600 transmission electron microscope (Hitachi) at 80 kV. The width and crossover distance of amyloid fibrils were measured and analyzed by ImageJ software. The data are based on 5 technical replicates of one case per animal species.
Results
Histopathology
In the mouse, cow, and goat liver samples, diffuse amyloid deposits were observed in the space of Disse. In the dog and camel kidney samples, multifocal amyloid deposits were observed in the interstitium of the medulla and glomeruli. Amyloid deposits were immunopositive for anti-SAA or anti-AA antibody (Suppl. Figs. S1–S5). All amyloid deposits stained by Congo red displayed apple green birefringence under polarized light (Suppl. Figs. S6–S10).
SDS-PAGE and Immunoblot Analyses of Amyloid Extracts From Different Animal Species
To analyze amyloid extracts from different animal species, we performed SDS-PAGE and immunoblot analyses. The full amyloid extracts of the 5 animal species consisted of the amyloid fraction and many other unidentified minor molecules (Fig. 1 and Suppl. Fig. S11). The amyloid extracts contained a large amount of water-soluble amyloid fibrils, 34 which are approximately 10 to 15 kDa in size, as observed by SDS-PAGE. 31 The result of dot blot analysis (Fig. 2 and Suppl. Fig. S12) showed that the mouse SAA antibody reacted with the amyloid extracts from all 5 animal species, suggesting that the prepared amyloid extracts indeed contained AA proteins. In addition, Western blot analysis of mouse amyloid extract detected strong bands at around 10 to 15 kDa (Fig. 1 lane 6, Suppl. Fig. S13 lane 1), similar to our recent report. 34 Western blot analysis also detected oligomer and high-order aggregates. On the other hand, Western blots could not detect amyloid of the remaining 4 animal species (Suppl. Fig. S13 lanes 2–7). This may reflect limited or no cross-reaction of the polyclonal antibody with amyloid from other species denatured by SDS-PAGE.
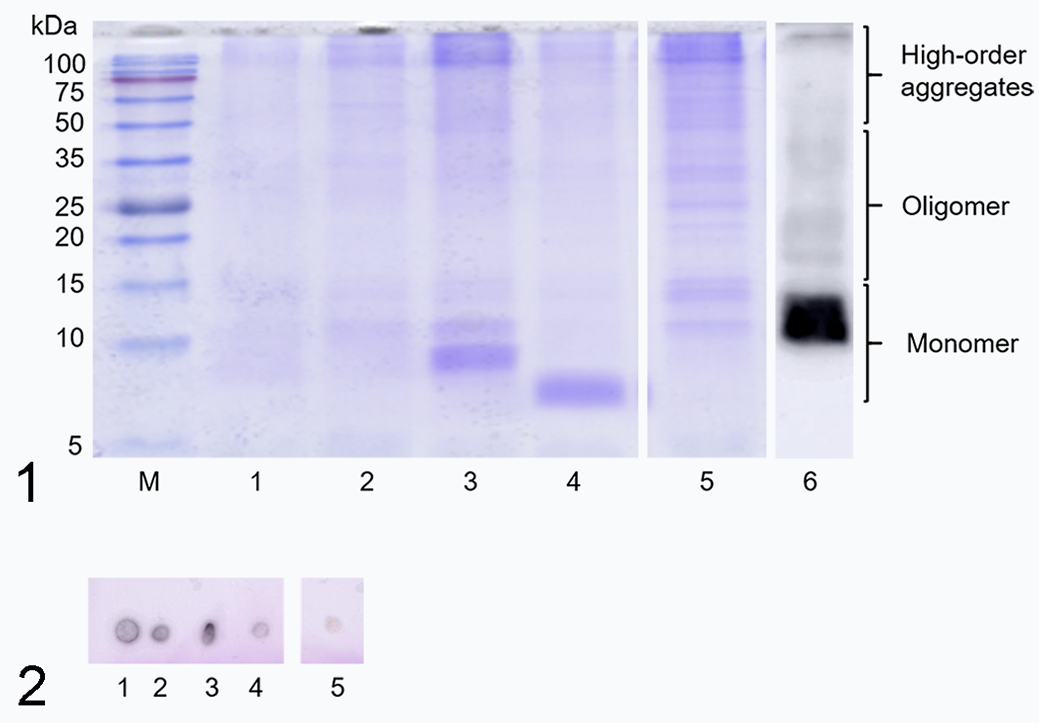
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) of full amyloid extracts from 5 animal species (liver or kidney). M, molecular weight marker; lane 1, mouse; lane 2, dog; lane 3, camel; lane 4, goat; lane 5, cow; lane 6, Western blot analysis of the amyloid extract from mouse, probed with a polyclonal antibody to mouse serum amyloid A protein.
When the results of these amyloid-related bands in SDS-PAGE (Fig. 1 lanes 1–5) were compared, differences between the 5 animal species were revealed. On the other hand, 3 groups of bands at 10 to 15 kDa (monomer), 15 to 50 kDa (oligomer), and >50 kDa (higher-order aggregates) were crudely detected in all animal species. Interestingly, camel and goat showed larger bands at about 9 kDa and 7 kDa, respectively (Fig. 1 lanes 3 and 4, Suppl. Fig. S11 lanes 3, 4, and 5).
Transmission Electron Microscopy
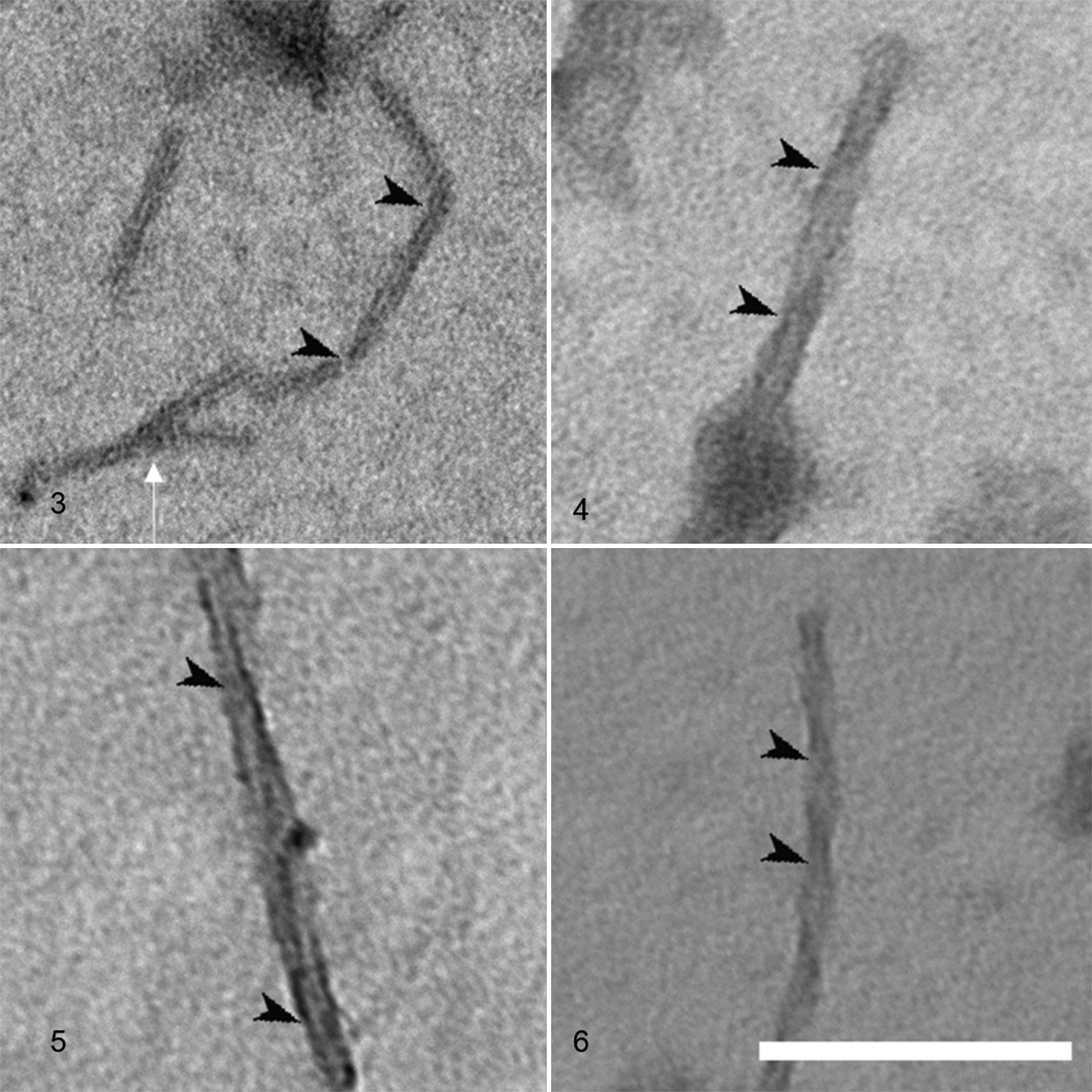
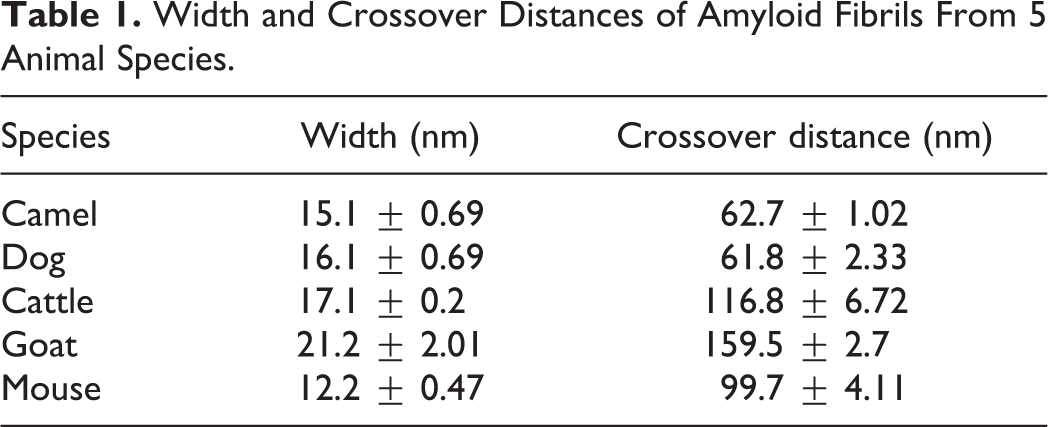
We successfully observed amyloid fibrils that had been extracted and purified from the 5 animal species (Figs. 3–7). All fibrils had a linear morphology with a cross-structure and irregular length. Although it is possible that fibrils can break or cleave during the preparation steps, we selected fibrils with a complete structure for measurements. The results show that dog and mouse fibrils were shorter, while the camel and dog fibrils were straighter than the fibrils of the other animals. The width and crossover distance of the fibrils are summarized in Table 1, which shows that there are similarities in the amyloid fibrils of camel and dog, and of cow and goat. The data show that camel and dog amyloid fibril widths were 15.1 ± 0.69 nm and 16.1 ± 0.69 nm, and crossover distances were 62.7 ± 1.02 nm and 61.8 ± 2.33 nm, respectively. On the other hand, in cow and goat, fibril widths were 17.1 ± 0.2 nm and 21.2 ± 2.01 nm, and crossover distances were 116.8 ± 6.72 nm and 159.5 ± 2.7 nm, respectively. The mouse amyloid fibrils were 12.2 ± 0.47 nm in width (similar to dog and camel), and had a crossover distance of 99.7 ± 4.11 nm (different from the other animals). In addition, the amyloid fibrils of mouse and camel had a branched pattern (Figs. 7 and 11), whereas those of cow, goat, and dog were linear without branched pattern.
Width and Crossover Distances of Amyloid Fibrils From 5 Animal Species.
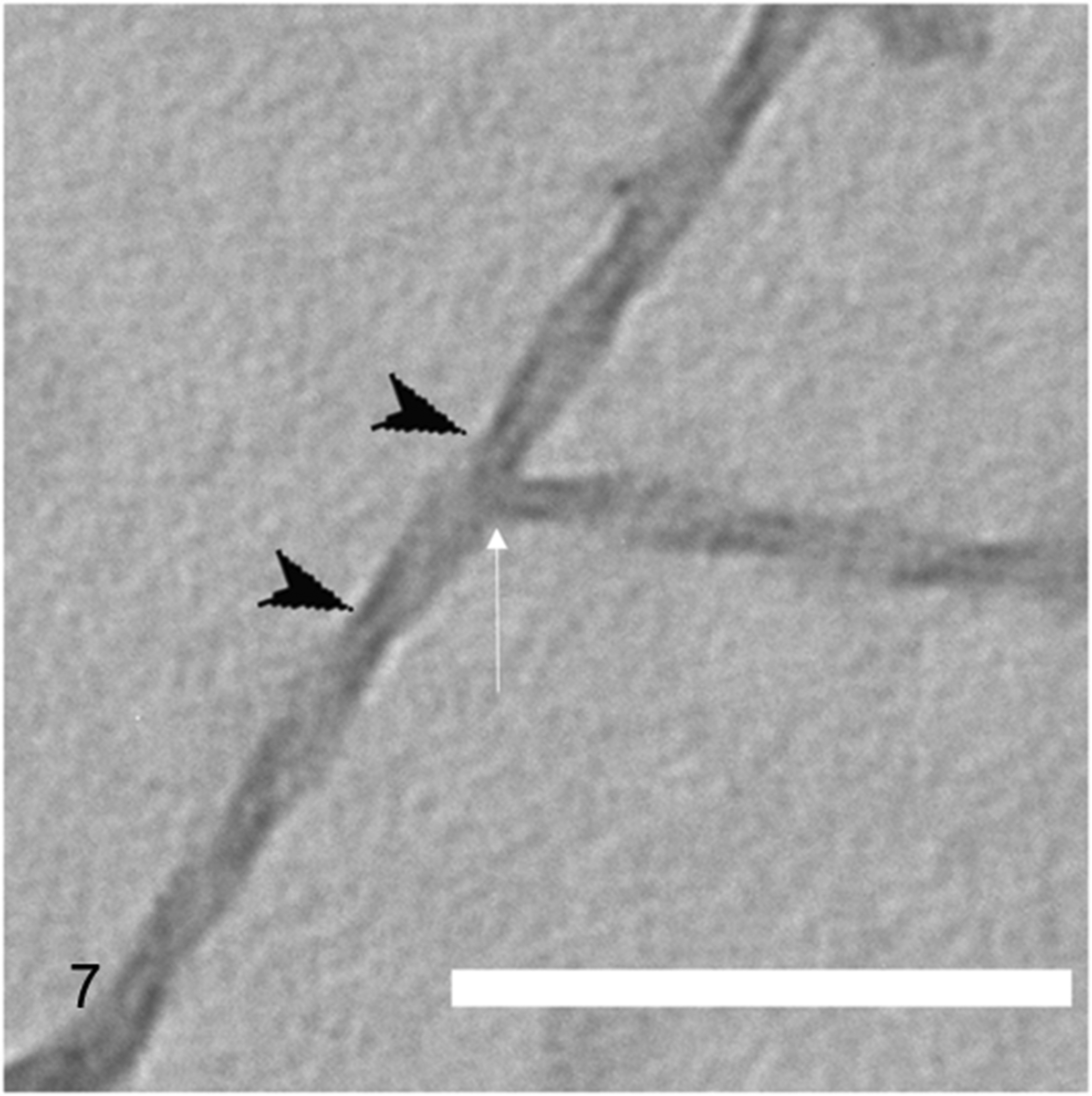
A camel amyloid fibril with branching pattern (white arrow). Black arrowheads: crossover positions of the twisting fibrils; this interval is the crossover distance. Transmission electron microscopy, negatively stained with 1% phosphotungstic acid. Scale bar: 200 nm.
Homology Analysis
SAA protein had high homology of at least 78.23% across the 6 vertebrate species studied (Fig. 8 and Suppl. Fig. 14). A phylogenetic tree showed 3 groups, the camel and dog group (green box), the cow and goat group (red box), and the human and mouse group (blue box). The highest similarities were between camel and dog (92.22%), cow and goat (88.48%), and human and mouse (80.85%).
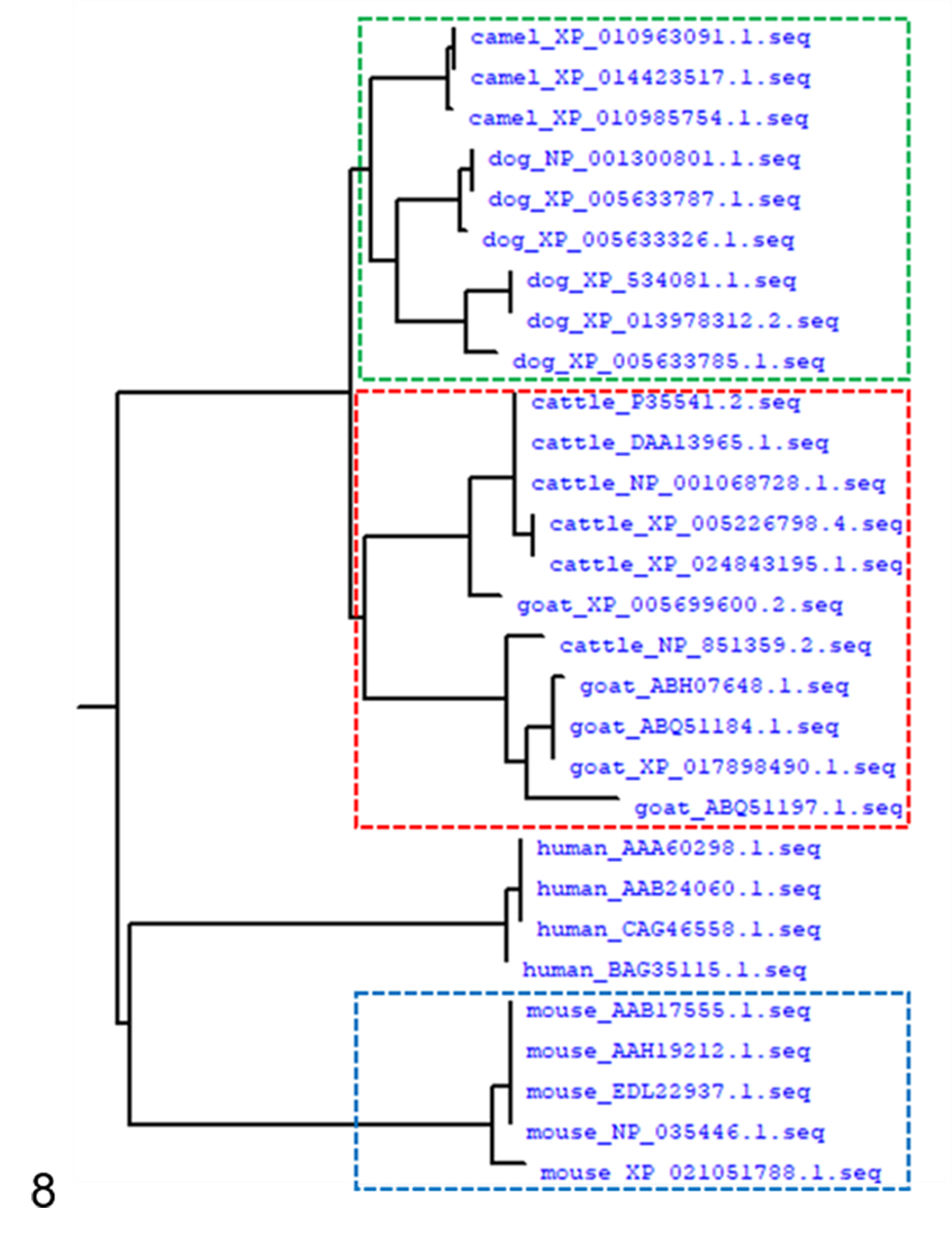
Phylogenetic tree of SAA gene sequences from camel, dog, cow, goat, mouse, and human. The numbers following the species names indicate the NCBI accession numbers. Dotted boxes in green, red, and blue indicate clustering of sequences from camel and dog, cow and goat, and mouse isoforms with similar fibril morphologies.
To clarify the differences in SAA sequences in these 5 animal species, we compared their amino acid sequences. Although the SAA protein sequences in the same species had many subtypes, we selected the same sequence length and different subtype sequences from these 5 animal species to compare their homology. The sequences of different subtypes’ high homology among different samples from the same individual animal exceeded 96.72%. Consequently, the sequence we selected showed no differences between the different subtypes within a given animal species.
Discussion
AA amyloidosis is complication of chronic inflammatory, infectious, and neoplastic diseases in humans and domestic animals. 33,38 Amyloid fibrils with a cross-β structure are deposited as insoluble fibrils, mainly in the extracellular spaces of organs or tissues, as a result of sequential changes in protein folding. Their accumulation results in organ damage and dysfunction. 14,26 Most reports to date have focused on the occurrence, development, and structure of AA amyloid derived from SAA protein. SAA increases in plasma in response to some inflammatory stimuli. Under normal conditions in plasma, the level of SAA is 1 to 5 μg/ml, but SAA is sensitive to inflammation or other damage-related injuries such as trauma, infection, or tumors, and in a short period of time, the concentration can increase by nearly 1000 to 2000 times more than the normal in vivo concentration. 4,32 For this reason, the concentration of SAA in body can serve as a biomarker of AA amyloidosis. 16 Subsequently, proteolysis of SAA protein may lead to aggregation and deposition of SAA fibrils in organs. The current therapy for AA amyloidosis is anti-inflammatory drugs or colchicine to reduce SAA levels in the body. 19 However, adequate suppression of SAA production is not effective.
The 3D structure of human SAA1 protein is a hexamer, which is characterized by subunits forming a unique 4-helix bundle fold and stabilized by a long C-terminal tail. 10 Moreover, the N-terminal of amyloid fibril proteins are the most hydrophobic and amyloidogenic segment of the protein sequence. They are a driver of AA amyloidosis, are crucial for forming the disease-associated AA amyloid fibrils, and provide stability to the analyzed amyloid fibrils. 9,13 In this research, we successfully extracted and purified amyloid fibrils from 5 animal species. Results of SDS-PAGE analysis indicated that the SAA monomers have different sizes in the different animal species. Since camel and goat samples showed smaller bands (at about 7 and 9 kDa) than the other animal species, this may reflect a different progression of amyloidosis.
TEM was used to observe human amyloid fibrils that accumulate in extracellular plaques and intracellular inclusions associated with disease and that are especially prevalent during ageing. 27 However, it is difficult to distinguish the features of amyloid fibrils because almost all of them show a linear morphology with a cross-structure and irregular length. In this study, we used TEM to observe amyloid fibrils extracted and purified from 5 animal species. The species formed 3 groups based on measurements of the amyloid fibril width and crossover distances: camel and dog; cow and goat; and mouse. However, we did not confirm whether the morphology of fibrils within the same species differs depending on sex, age, and the degree of amyloidosis. Some reports have shown that the same amyloid fibrils from different species have differences in their spiral morphology in vivo and in vitro, while fibrils in liver and in feces also have differing morphology in width and crossover distance. 7,39 In this study, amyloid fibrils extracted from different animals by the same method had different morphology, and morphology was related to clustering of SAA gene sequences in the phylogenetic tree. Further investigation is required to assess differences in the morphology of fibrils within the same species. Amyloid fibrils share a common underlying architecture, in which the β-strands within each protofilament align perpendicularly to the long axis of the fibril. 6 Using cryo-EM, Liberta et al revealed that human and murine AA amyloid fibrils have different fundamental properties, such as the presence of right-handed or left-handed twisted cross-β sheets and overall folding. 9 Furthermore, amyloid fibrils were defined as rigid, nonbranching features. 26 Interestingly, our data showed that the amyloid fibrils of mouse and camel had a branching pattern.
SAA protein is known as an evolutionarily conserved protein in vertebrates with high homology in different animal species. Since AA amyloid can be transmissible between different animal species, it is possible that the processes underlying amyloidosis may be similar in different species. 15,34 In our study, SAA gene sequences of human and 5 animal species (dog, cow, mouse, camel, and goat) had only 78.23% homology but there was high homology between cow and goat (88.48%), and between dog and camel (92.22%). Of interest, it was shown that species with similar homology also had fibrils with similar width and crossover distance, but these differed from those in mice. These findings may suggest that differences in amino acid sequences are related to differences in the width and crossover distance of fibrils. In mice with induction of amyloidosis, treatment with the C-terminal peptide can inhibit further extension of amyloid fibrils in AApoA2 amyloidosis. 23 Moreover, the N-terminus contributes to the stability of amyloid fibril morphology. 9 These approaches may provide a novel strategy to combat this disease by interfering with specific fibril polymorphs. In conclusion, we provide novel insight into amyloid fibrils from different animal species displaying high genetic homology and morphological similarity. We hope that this information will assist to better understand amyloid disease and the development of novel therapies.
Supplemental Material
Supplemental Material, Combined_supplemental_materials-Lin_et_al - Comparison of AA Amyloid Fibril Morphology and Serum Amyloid A Gene Sequence in 5 Animal Species
Supplemental Material, Combined_supplemental_materials-Lin_et_al for Comparison of AA Amyloid Fibril Morphology and Serum Amyloid A Gene Sequence in 5 Animal Species by Xuguang Lin, Masahiro Kuragano, Kenichi Watanabe and Kiyotaka Tokuraku in Veterinary Pathology
Footnotes
Author Contributions
Conception and design: KT and KW. Experiments performed: XL. Data analysis and acquisition: XL, MK, and KW. Drafting the manuscript: XL. Supervision: KT and KW. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Japan Society for the Promotion of Science (KAKENHI, JP16H03288; grant awarded to KT).
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
