Abstract
Keywords
CPPV-1 is a nonenveloped single-stranded DNA virus with an approximately 5 kb genome size containing 2 open reading frames encoding 2 nonstructural proteins (NS1 and NS2) and 2 capsid proteins (VP1 and VP2). Several mutations of amino acid residues in the VP2 protein play a central role in the susceptible host selection in domestic animals and expand infectivity to other wild carnivores. 1,17,18,25,27,28 Although mutations in the VP2 gene have been a basis for variant classification and understanding the potential host-dynamic range, mutations in the NS1 region of the FPV isolated from domestic cats have been recently considered as potential points for pathological insights. 6 Moreover, the NS1 of CPPV-1 is now important for molecular epidemiological investigations. 15 However, the information regarding both NS1 and VP2 mutations of CPPV-1 isolated from wild carnivores is limited.
The FPV, a member of CPPV-1, was reported in wild felids, such as American tiger cat (
Focusing on CPPV-1 infection in the
Materials and Methods
Clinical History and Sample Collection
In September 2013, 2 out of 55 captive small Indian civets (
Pathological Examination
Gross lesions of the 7 necropsied civets (cases 18–24) were evaluated by a veterinary pathologist (WB). Various organs, including the heart, lung, liver, kidney, small intestine, large intestine, cecum, rectum, mesenteric and peripheral lymph nodes, thymus, genital organs, and brain, were collected and preserved in 10% (v/v) neutral buffered formalin and routinely processed for histopathological examination.
Polymerase Chain Reaction (PCR) and RFLP Analysis
Eighteen fecal swabs (cases 1–17, 24) were subjected to viral genomic extraction using a viral DNA/RNA extraction kit II (GeneAid, Taipei, Taiwan). The VP1/VP2 gene encoding for the capsid proteins of CPPV-1 was amplified using primers VPF and VPR (Supplemental Table S1) as previously described
16
with minor modifications. Briefly, the PCR thermal cycling was performed as 30 cycles of 94 °C for 30 seconds, 55 °C for 2 minutes, and 72 °C for 2 minutes. The 2246 bp PCR products were resolved by 1.5% (w/v) agarose-gel electrophoresis and visualized under UV transillumination after ethidium bromide staining, and then subjected to RFLP analysis to identify the CPPV-1 as FPV or CPV. Amplicons were digested with
To further differentiate between the virus in these civets and vaccine strains given to the nearby client-owned cats, separate PCR reactions and RFLP analyses were performed. Civet viral samples and the FPV-commercial vaccine strains Felocell (Zoetis) and Purevax (Merial) were PCR-amplified by primers specific to the 1755 bp of VP2 gene (P5 and P6, Supplemental Table S1) as reported,
20
with thermal cycling performed as above. The PCR products were digested with
The complete nonstructural NS1 and NS2 genes were amplified from the dead civet cat that had shown neurological signs of circling and seizures (case 24). The PCR amplification was performed in separate reactions using previously described primer pairs for NS gene (Supplemental Table S1). 22 Briefly, PCR thermal cycling was performed as 30 cycles of 94 °C for 30 seconds, 55 °C for 1 minute, and 72 °C for 1 minute.
Other common canine/feline viral pathogens, including canine distemper virus (CDV), canine parainfluenza (CPIV), canine herpesvirus-1 (CaHV-1), canine adenovirus-1 and -2 (CAdV-1/-2), and feline and canine enteric coronaviruses (FCoV/CCoV) were also tested using civet fecal samples by specific PCRs as described previously. 13,21,23
Genomic Sequencing, Phylogenetic and Recombination Analysis
Eight VP1/VP2 PCR-positive samples derived from fecal swabs of cases 2, 4, 8, 10, 11, 13, 14, and 24 were further sequenced for genome analysis. Multiple PCR reactions with different primer pairs (Supplemental Table S1) were used for the complete genetic characterization of the civet CPPV-1 VP gene as described. 16 Moreover, the civet cat that showed neurological signs (case 24) was additionally sequenced for the complete NS gene (as mentioned above). All PCR products were purified using NucleoSpin Extract II (Macherey-Nagel, Düren, Germany) and submitted for commercial genetic sequencing (SolGent Ltd, Daegeon, South Korea). Nucleotide alignments and deduced amino acids were analyzed and compared with published data deposited in GenBank using Bioedit Sequence Alignment Editor Version 7.2.5.
The phylograms were analyzed using the MEGA 7.0 software (http://www.megasoftware.net/). Phylogenetic trees were constructed by the maximum likelihood (ML) method and bootstrap analysis with 1000 replicates was used to assess the confidence level of the branch pattern. Bootstrap values of >70% were considered to be significant. To detect possible genetic recombination event(s) in these CPPV-1 strains, recombination analysis was conducted with CPPV-1 sequence derived from the infected civet (case 24), and CPPV-1 sequences available in GenBank served as sequence queries. A cocktail of statistical methods, including RDP, GENECONV, BootScan, MaxChi, Chimaera, SiScan, and 3Seq were used with default settings in the Recombination Detection Program (RDP) package, version 4.0, to check for potential recombination events. Any potential breakpoint signals revealed by at least 4 methods with
Immunohistochemistry (IHC)
To localize CPPV-1 antigen in formalin-fixed paraffin embedded tissue, 3 necropsied civets (cases 18, 19, and 24) were used for IHC investigation using the EnViSion system horseradish peroxidase labeled with polymer anti-mouse (EnVision polymer, Dako, Glostrup, Denmark). Sections were cut at 4-µm thickness, deparaffinized, and hydrated. They were then pretreated with trypsin for 30 minutes. Endogenous peroxidase and nonspecific reactions were blocked by incubation with 3% (w/w) hydrogen peroxide and 5% (w/v) skim milk in 1% PBS, respectively. Subsequently, sections were incubated with mouse monoclonal anti-canine parvovirus antibody (1:250 dilution in 5% skim milk) to detect the CPVs, mink enteritis virus (MEV), and FPV (ab59832, Abcam, Cambridge, UK), at 4 °C overnight, washed 3 times in 1% PBS, and then sections were incubated with the EnViSion system as per the manufacturer’s instructions before being immersed in 3,3′-diaminobenzidine and counterstained with Mayer’s hematoxylin before mounting. Sections of the intestinal tissue of confirmed FPV-infected cats were used as a positive control, while sections from the same source but omitting primary monoclonal antibody (replaced with distilled water) was applied as a negative control. Immunopositive cells were noted for interpretation of cellular tropism and viral distribution in organs. Immunoreactivity was scored, semiquantitatively analyzed, and averaged in 5 different areas under high-power field (400×) as follows: − (no immunopositive cells); + (≤25% immunopositive cells); ++ (26% to 50% immunopositive cells); +++ (51% to 75% immunopositive cells); ++++ (≥76% immunopositive cells).
Results
Clinical Findings
All captive civets in this study were small Indian civets,
Pathological Findings
The 7 necropsied civets (cases 18–24) were markedly emaciated with pale mucous membranes and severe dehydration. All 7 civets had remarkable macroscopic lesions of segmental hemorrhagic enteritis containing a watery dark-brown content in the lumen (Fig. 1; Supplemental Table S3). Hematopoietic organs including spleen and mesenteric lymph nodes were enlarged and congested, and there were foci of thymic hemorrhage in 2 civets. Marked congestion was observed in the brain, lung, and liver. One dead civet (case 24) was promptly tested with rapid test kits for CDV/CPV/CCoV antigen (Bionote, Gyeonggi-do, Republic of Korea) using a fecal sample, and showed a strong positive result only for CPV (data not shown).
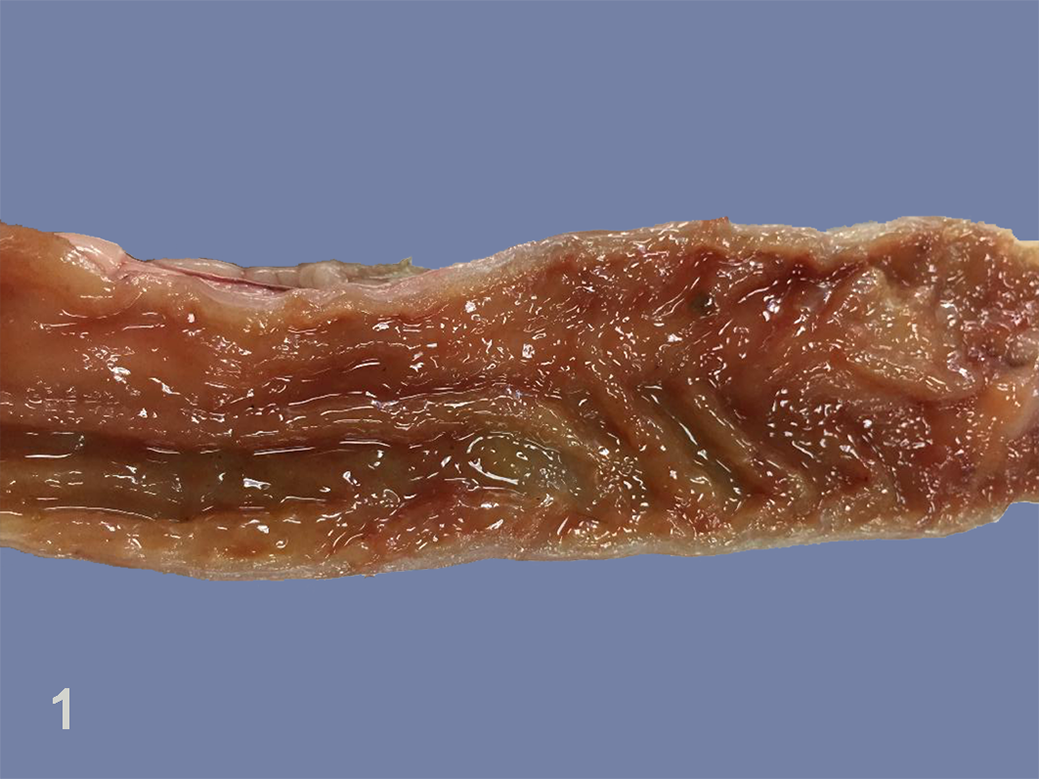
Carnivore protoparvovirus infection, jejunum, small Indian civet, case 24. There is reddening of the mucosal surface.
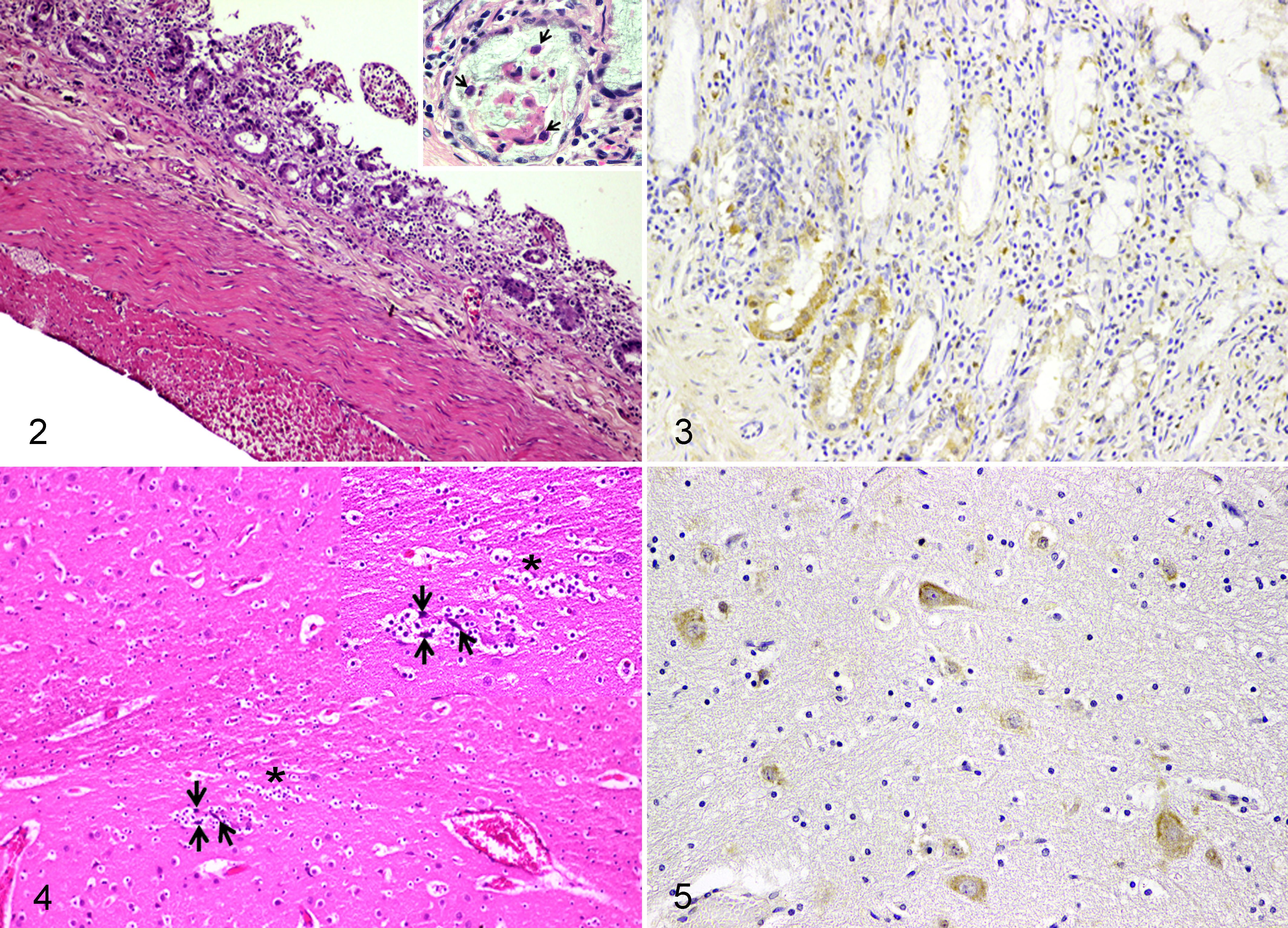
Histopathological findings in the small intestine revealed severe diffuse desquamative hemorrhagic enteritis, characterized by atrophied and fused villi, crypt necrosis, and lymphoplasmacytic infiltration in the lamina propria (Fig. 2). All 7 examined civet cats showed numerous amphophilic intranuclear inclusion bodies in small intestinal crypt epithelial cells and villous enterocytes (Fig. 2, inset; Supplemental Table S4). There was severe lymphoid depletion in the Peyer’s patches of the ileum, colonic lymphoid follicles, mesenteric lymph nodes, and spleen in all 7 examined cases. For the central nervous system, the meninges and choroid plexus were moderately congested. There was neuronal degeneration with satellitosis in the cerebral cortex while there were no lesions in white and gray matters of cerebellum (Fig. 4).
PCR and RFLP Results
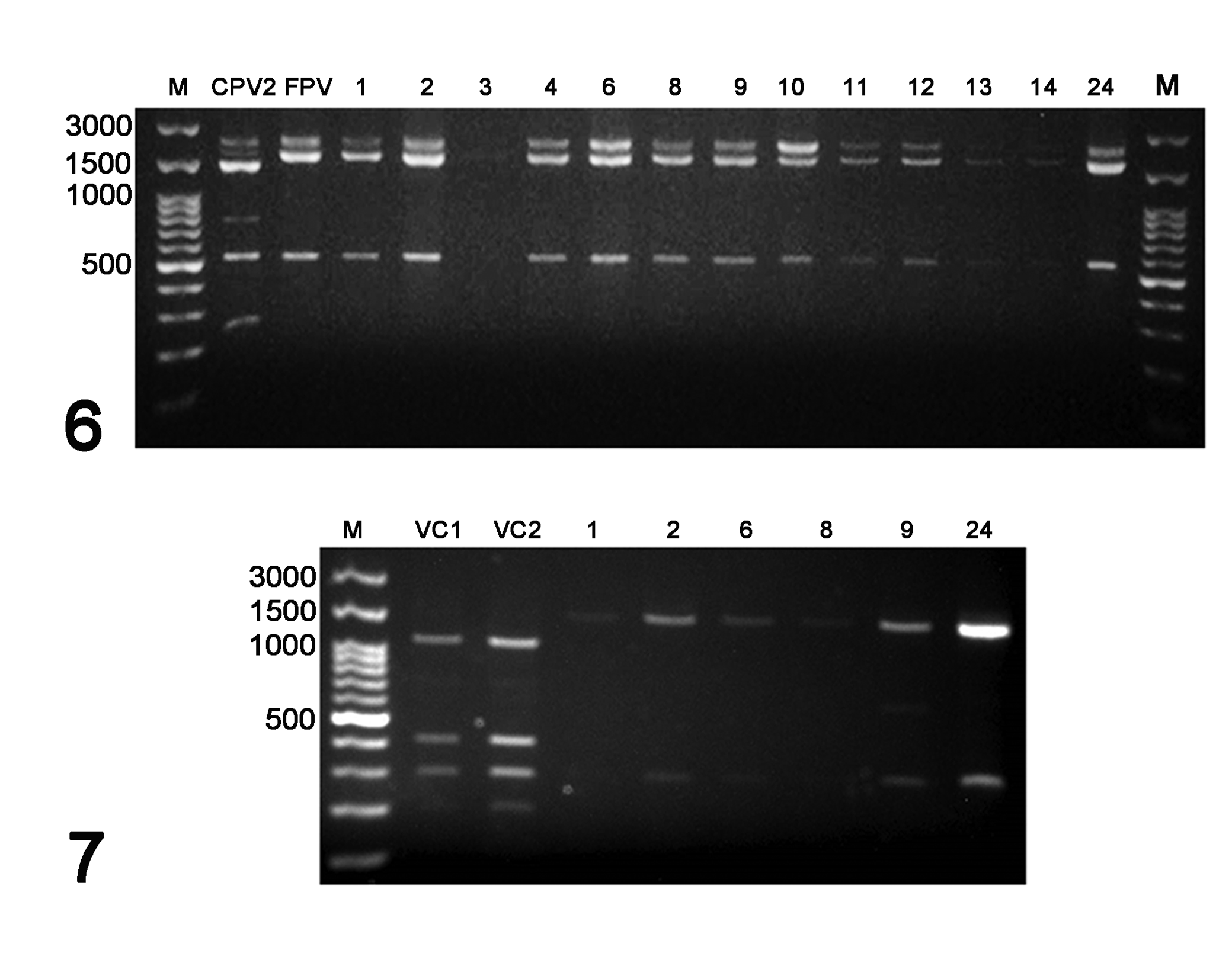
Out of 18 fecal swab samples (cases 1–17 and 24), 13 (72%) had positive 2246 bp PCR products for the VP1/VP2 gene of CPPV-1. All were negative using PCR assays specific for CDV, CPIV, CaHV-1, CAdVs, FCoV, and CCoV (Supplemental Table S2). The 2246 bp VP1/VP2 amplicons from each of the 13 CPPV-1 positive civets were subjected to RFLP analysis using the
Viral Genome Analysis
The VP2 gene was further sequenced for 7 cases that had positive VP1/VP2 amplicons (cases 2, 4, 8, 10, 11, 13, 14), while one other (case 24) was further sequenced for the full-length CPPV-1 genome. The obtained sequences have been submitted to the GenBank database with accession nos. KP019617 to KP019620 for cases 2, 8, 10, and 13, respectively; KT357491 to KT357493 for cases 4, 11, and 14, respectively; plus KP019621 for the full-length genome from case 24 (Supplemental Table S2). The civet-derived nucleotide sequences and deduced amino acids were compared to other CPPV-1-annotated sequences retrieved from the GenBank database.
The full-length genome of the civet CPPV-1 (case 24) shared 99.4% nucleotide similarity with the FPV HH-1/86 strain from a jaguar (
For the in silico deduced amino acid analysis, the civet CPPV-1 (case 24) revealed 99.7% and 99.4% amino acid similarity to NS1 of the jaguar FPV HH-1/86 and the FPV GX01 strain isolated from domestic cat (accession no. MG924893), respectively. For the amino acid similarity to NS2, this civet CPPV-1 demonstrated 98.8% similarity to both jaguar FPV HH-1/86 and the cat FPV GX01 (data not shown), while the VP2 showed 100% amino acid similarity to the FPV lineage at specific amino acid residues (Table 1). Interestingly, the civet CPPV-1 had a distinct amino acid mutation, G573S, in the NS1 gene, that was not observed in other FPV isolates, but this mutation was present in some CPV-2a (accession no. AY742936) and CPV-2c (accession no. MH476584) isolates (Table 1).
Host-Specific Amino Acids of the Viral NS1 and VP2 Proteins for Carnivore Protoparvovirus-1 Viruses Analyzed in This Study.
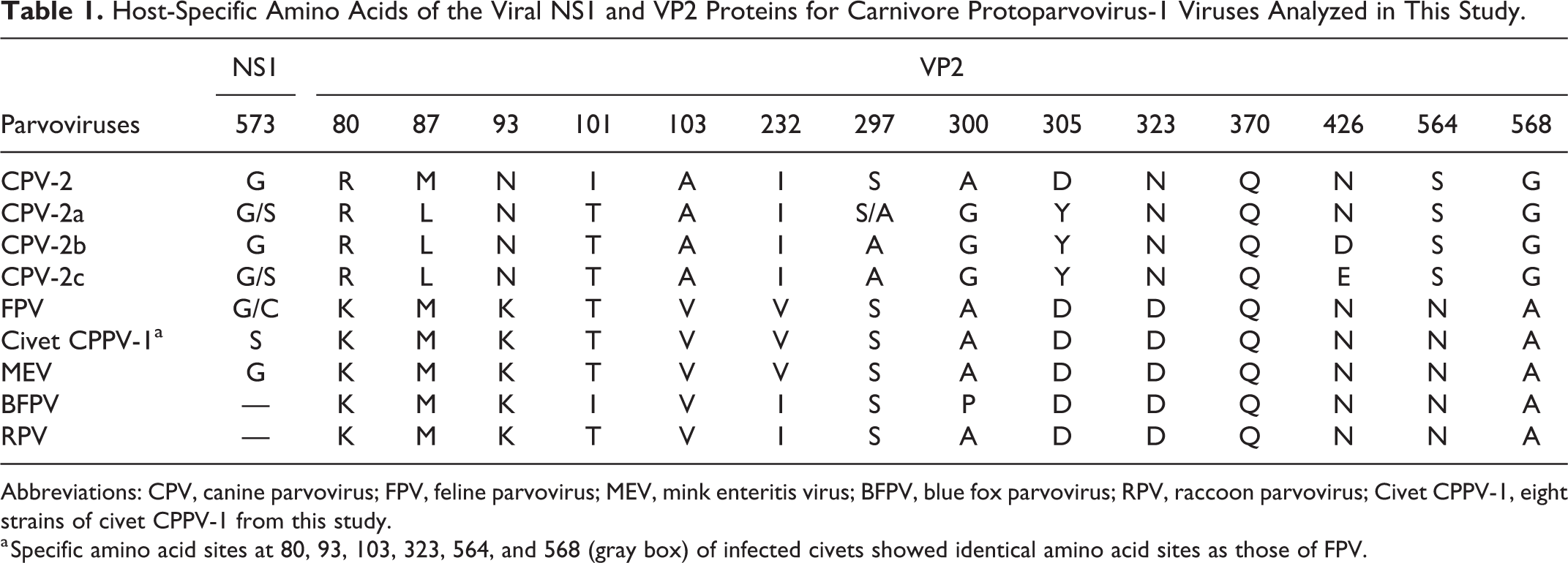
Abbreviations: CPV, canine parvovirus; FPV, feline parvovirus; MEV, mink enteritis virus; BFPV, blue fox parvovirus; RPV, raccoon parvovirus; Civet CPPV-1, eight strains of civet CPPV-1 from this study.
a Specific amino acid sites at 80, 93, 103, 323, 564, and 568 (gray box) of infected civets showed identical amino acid sites as those of FPV.
Phylogenetic analysis of NS1, VP2, and whole genome sequences of civet CPPV-1 depicted a distinctively monophyletic branch in the FPV lineage, which was separated from the other CPV (and MEV) lineages (Supplemental Figs. S1-S3). The phylogram of reported CPPV-1 infections in animals of the family
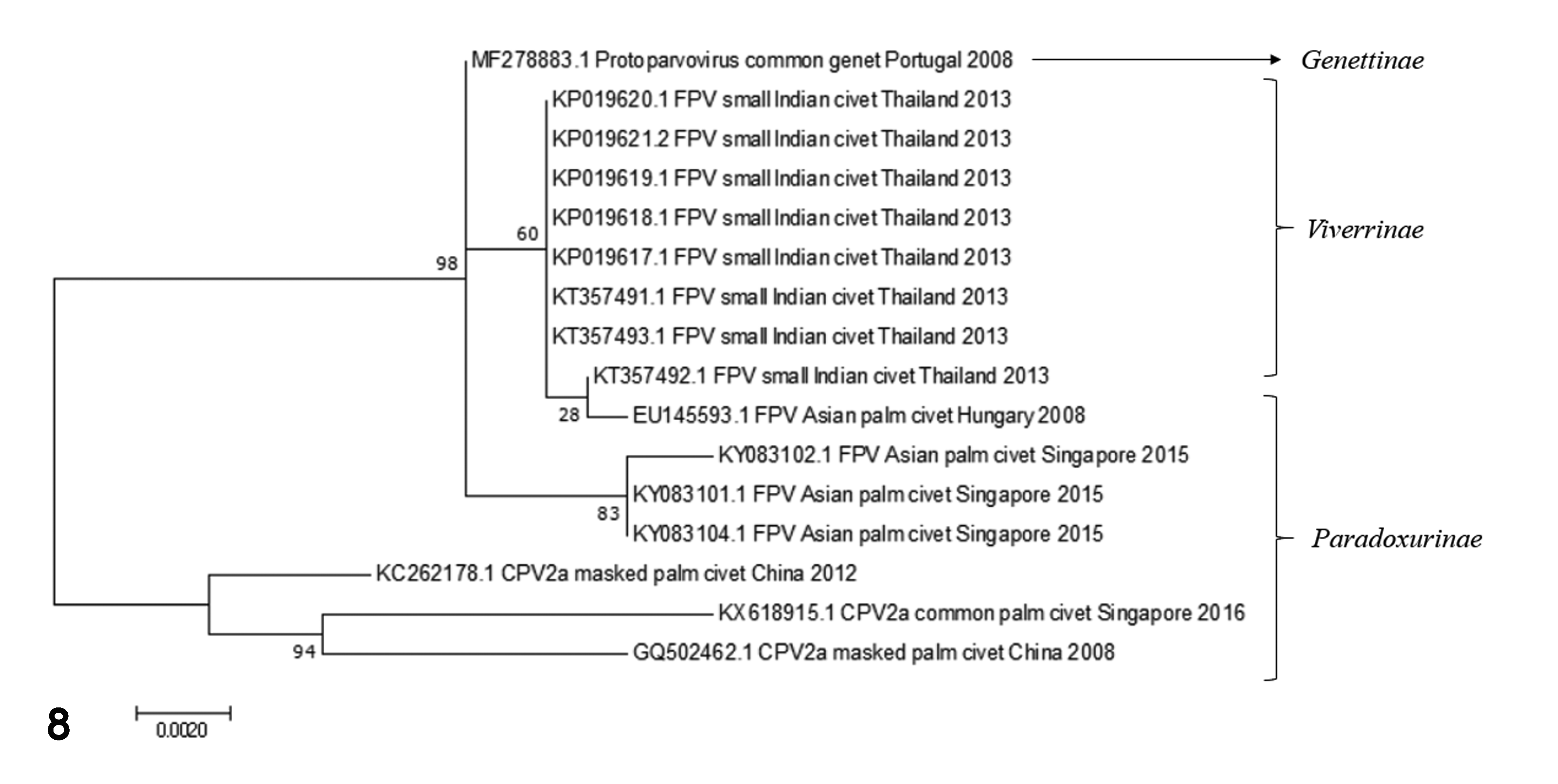
Phylogram of CPPV-1 (FPV and CPV) infection in family
To investigate the possibility of genomic recombination in these CPPV-1-infected civets, the potential recombination event(s) in the CPPV-1 VP2 gene of the 8 sequences obtained from the different infected civets in this study and the other CPPV-1 sequences used for the phylogenetic analysis were analyzed using the RDP software. However, no recombination breakpoints were found in any of the civet CPPV-1 sequences (data not shown).
Cellular Tropism and Tissue Localization
Three necropsied civets (cases 18, 19, and 24) were analyzed using IHC to determine the localization of CPPV-1 antigen. Immunolabeling was observed in both the cytoplasm and nucleus of the villous enterocytes, crypt epithelial cells, and myenteric ganglionic neurons, which was compatible with the observed abundant infiltration of mononuclear inflammatory cells in the mucosa of the small intestine (Fig. 3). The infiltrated histiocytes in the spleen and mesenteric lymph node also showed intense immunolabeling. In the central nervous system, neurons in the cerebral cortex and cerebellar Purkinje cells showed strong immunoreactivity in their cytoplasm (Fig. 5). The IHC scores are shown in Supplemental Table S5.
Discussion
The fatal gastroenteritis disease caused by protoparvovirus infection in dogs and cats is common, and it is sporadically reported in wild carnivores. Unvaccinated free-roaming dogs and cats may serve as potential hosts for the outbreak. Unfortunately, samples of the free-roaming domestic and farmed animals that had access to this Indian civet cat farm were not available for evaluation of their infection status.
The CPV and FPV are members of the contagious CPPV-1 virus group in which VP2, the main structural protein, plays a major role in infection in various animals. Therefore, recent studies of CPPV-1 genome have focused on amino acid mutations in the VP2 gene. In this study, 6 mutated amino acid residues of VP2 of civet CCPV-1 (sites 80, 93, 103, 323, 564, and 568) were observed that can differentiate FPV from CPV isolates. 28 Furthermore, a deduced amino acid mutation (G573S) in the NS1 gene of civet CPPV-1 was observed, that was distinct from the amino acid sequence of FPV. Therefore, civet CPPV-1 was tentatively named as a potentially novel virus detected in this outbreak. Nevertheless, a small number of the observed sequences did not have this mutation. Thus, the relationship of CPPV-1 to FPV needs further clarification.
The obtained sequences demonstrated a high nucleotide similarity to other FPV isolates and the phylogenetic analysis revealed that civet CPPV-1 grouped with FPV in a monophyletic lineage that was separate from CPV isolates. This supports the conclusion that CPPV-1 detected in this civet cat outbreak was of FPV origin, which is also supported by the RFLP analysis. The use of RFLP is useful for distinguishing origins of CPPV-1. However, one sample (case 3) that was PCR positive for the VP2 gene revealed no digestion in the RFLP assay. Although not sequenced to confirm the presence or absence of the
The origin of the viral carrier in this outbreak remained unclarified. It might have occurred from civet-spread parvovirus itself or been from other animal reservoirs living in the area surrounding this farm. To date CPPV-1 detection in
From the pathological studies, CPPV-infected civets showed lesions similar to parvovirus infection, including severe acute segmental muco-hemorrhagic enteritis with collapsed, blunted villi, and necrosis and inflammation of intestinal crypts. Moreover, viral amphophilic intranuclear and intracytoplasmic inclusion bodies were abundant in enterocytes and cryptal epithelial cells, in accordance with previous findings in parvovirus-infected dogs and cats. 9,14,19,26
The IHC analysis also supported the parvovirus infection and indicated its localization. Interestingly, immunopositive cells were present not only in intestinal crypts but also in cerebral neuronal cells and cerebellar Purkinje cells. There are some previous reports of CPPV-1 localization in the brain tissue in companion animals. 12,29 The mechanism of neuron invasion of the FPV-infected cat was related to the L582S mutation of NS1, changing residue 582 from leucine to serine. 6 However, this specific mutation was absent in CPPV-1, even though the neuronal cells were immunolabeled for parvoviral antigen. These discrepant findings warrant further investigation, due to the different affected hosts, using a large-cohort study.
In conclusion, we report here the first identification of CPPV-1 infection in small Indian civet cats (
Supplemental Material
Supplemental Material, Combined_supplemental_materials-Chaiyasak_et_al - Carnivore Protoparvovirus-1 Associated With an Outbreak of Hemorrhagic Gastroenteritis in Small Indian Civets
Supplemental Material, Combined_supplemental_materials-Chaiyasak_et_al for Carnivore Protoparvovirus-1 Associated With an Outbreak of Hemorrhagic Gastroenteritis in Small Indian Civets by Surangkanang Chaiyasak, Chutchai Piewbang, Wijit Banlunara and Somporn Techangamsuwan in Veterinary Pathology
Footnotes
Acknowledgements
We are thankful to Dr Piyaporn Kongmakee for excellent coordination with the Monitoring and Surveillance Center for Zoonotic Disease in Wildlife and Exotic Animals (MoZWE) of Mahidol University, Thailand, and the Zoological Park Organization under The Royal Patronage of H.M. The King, Thailand.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: S. Chaiyasak was granted by Chulalongkorn University tuition fee and Mahasarakham University domestic tuition fee. C. Piewbang was supported by Ratchadapisek Somphot Fund for Postdoctoral Fellowship, Chulalongkorn University. This study was financially supported by the 90th Anniversary of Chulalongkorn University Fund (Ratchadaphiseksomphot Endowment Fund) and partially funded by the Veterinary Pathogen Bank, Animal Virome and Diagnostic Development Research Group, and Diagnosis and Monitoring of Animal Pathogens Research Unit, Faculty of Veterinary Science, Chulalongkorn University.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
