Abstract
We describe molecular testing for felid alphaherpesvirus 1 (FHV-1), carnivore protoparvovirus 1 (CPPV-1), feline calicivirus (FCV), alphacoronavirus 1 (feline coronavirus [FCoV]), feline leukemia virus (FeLV), feline immunodeficiency virus (FIV), and canine distemper virus (CDV) in whole blood samples of 109 free-ranging and 68 captive neotropical felids from Brazil. Samples from 2 jaguars (
Introduction
Wild felids are believed to be susceptible to the same pathogens that affect domestic cats. However, physiologic and ecologic patterns are different among felids and may be responsible for different patterns observed in the evolution of infectious diseases in felid species. Declines in wild felid populations caused by infectious diseases have been reported,30,40 and although well studied in domestic cats, little is known about the impact of pathogens on wild species.
The most important viruses of domestic cat are carnivore protoparvovirus 1 (feline parvovirus [CPPV-1]), felid alphaherpesvirus 1 (FHV-1), feline calicivirus (FCV), alphacoronavirus 1 (feline coronavirus [FCoV]), feline leukemia virus (FeLV), and feline immunodeficiency virus (FIV). Domestic cats are thought to be resistant to infection by canine morbillivirus (canine distemper virus [CDV]), but CDV is an important threat to the conservation of large wild felids. 45
Feline panleukopenia caused by CPPV-1 (previously called feline parvovirus or feline panleukopenia virus) is characterized by fever, depression, dehydration, and diarrhea in domestic cats.
2
FHV-1 can cause rhinotracheitis, and FCV is responsible for stomatitis and gingivitis.
19
Some FCoV isolates induce feline infectious peritonitis and others cause mild-to-severe enteritis.
19
Exposure to these viruses has been described in wild felids, including lion (
FeLV infection is rare in wild felids.
25
However, Iberian lynx, European wild cat (
Eight species of neotropical felids are in decline and under some degree of threat in Brazil according to the International Union for Conservation of Nature (http://www.iucnredlist.org), and a considerable number of individuals are kept in captivity. 31 Wild felids, belonging to the order Carnivora, play important roles in the ecosystems in which they live. 43 Several surveys of pathogens have been conducted in free-ranging and captive neotropical felids, and epidemiologic surveys are currently being addressed in conservation programs in Brazil.3,10,14,22,35,41 We investigated the occurrence of FHV-1, CPPV-1, FCV, FCoV, FeLV, FIV, and CDV in free-ranging and captive neotropical felids in Brazil, to better understand the virus circulation in these populations.
Materials and methods
Between January 1999 and March 2011, samples of 177 neotropical felids from Brazil were analyzed, including 109 free-ranging from 6 different states and 68 kept in captivity in São Paulo Zoological Park in São Paulo State, Brazil. The origin of the captive felids (wild-caught or zoo-born) was also recorded (Table 1). Felids were captured using leg-hold and metal cage traps, and for large felids, trained hounds were also used. After chemical immobilization, whole blood samples were collected by puncture of the femoral, cephalic, or jugular vein in vacuum tubes with EDTA anticoagulant. Whole blood was aliquoted and first stored at −20°C and then stored at −80°C, or stored immediately at −80°C, when available.
Description and origin of neotropical felids samples analyzed.
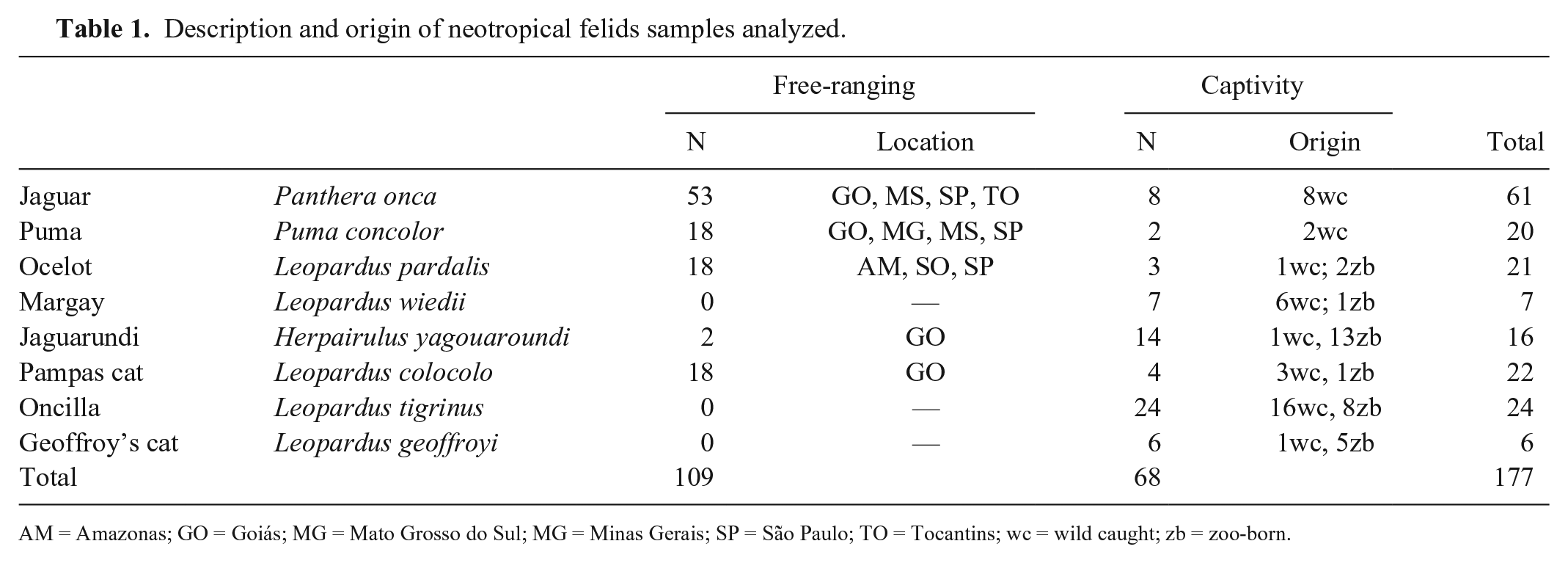
AM = Amazonas; GO = Goiás; MG = Mato Grosso do Sul; MG = Minas Gerais; SP = São Paulo; TO = Tocantins; wc = wild caught; zb = zoo-born.
Animal handling procedures were approved by the Bioethics Commission of the School of Veterinary Medicine and Animal Science of University of São Paulo (FMVZ-USP; permit 2017/2010). The Chico Mendes Institute for Biodiversity Conservation/ICMBio granted field work (permits 11214, 11628, 13249, 14637, and 239114).
DNA extraction was performed for 63 blood samples of free-ranging felids as described previously, 5 with some modifications using guanidine thiocyanate, and for the other 114 blood samples using a DNA extraction kit (NucleoSpin blood QuickPure kit, Macherey-Nagel, Düren, Germany). Extracted DNA was stored at −20°C in cryotubes. RNA was extracted from 177 blood samples (Ambion PureLink RNA mini kit, Life Technologies, Carlsbad, CA) and stored at −80°C.
Synthesis of complementary DNA (cDNA) for reverse transcription PCR (RT-PCR) was performed for detection of FCV, FCoV, and CDV. Reaction was prepared with 3.5 µL of RNA samples denatured at 94°C for 5 min and added to the RT mix containing 1× first strand buffer (Invitrogen, Carlsbad, CA), 1 mM of dNTPs, 10 mM of DTT, 25 ng of random primer (Invitrogen), 100 U of M-MLV reverse transcriptase (Invitrogen), and ultrapure water to a final volume of 10 µL. RT-PCR was performed at 37°C for 60 min.
PCR reactions were carried out with 1× PCR buffer (Invitrogen), 0.2 mM of dNTPs, 1.5 mM of MgCl2, 0.5 µM of each specific primer, 0.625 U of platinum Taq DNA polymerase (Invitrogen), 2.5 µL of cDNA or DNA sample, and ultrapure water to obtain a final volume of 25 µL. For CDV, we used 1 µL of each cDNA sample and 0.3 µM of each primer.
Amplification conditions for RNA viruses were: 95°C for 5 min (FCV) or 2 min (FCoV and CDV), 40 cycles of 95°C for 30 s, 57.6°C (FCV), 56°C (FCoV), or 60°C (CDV) for 30 s and 72°C for 30 s, and 72°C for 10 min for final extension using FCV24 and FCV414 primers for FCV, FCoV Tor2 11 and FCoV Tor2 13 primers for FCoV, and CDV-NP-F1 and CDV-NP-R1 primers for CDV (Table 2). Nested PCR was performed for CDV using 1 µL of PCR product, 0.5 µM of CDV inner primers (CDV-NP-F1 and CDV-NP-R2; Table 2), and the same reagents and amplification conditions described above. As positive controls, we used a commercial vaccine against FCV (Fel-O-Vax LVK IV + CaliciVax, Fort Dodge, Overland Park, KS), virus suspensions of canine coronavirus cultivated in canine fibrosarcoma cells (cells A72; Biovet Laboratory, Vargem Grande Paulista, São Paulo, Brazil), and a commercial vaccine against CDV (Nobivac puppy DP, MSD Animal Health, Upper Hutt, Wellington, NZ) for FCV, FCoV, and CDV, respectively.
Primer sequences used for detection of felid viruses, expected fragment size, and reference used.
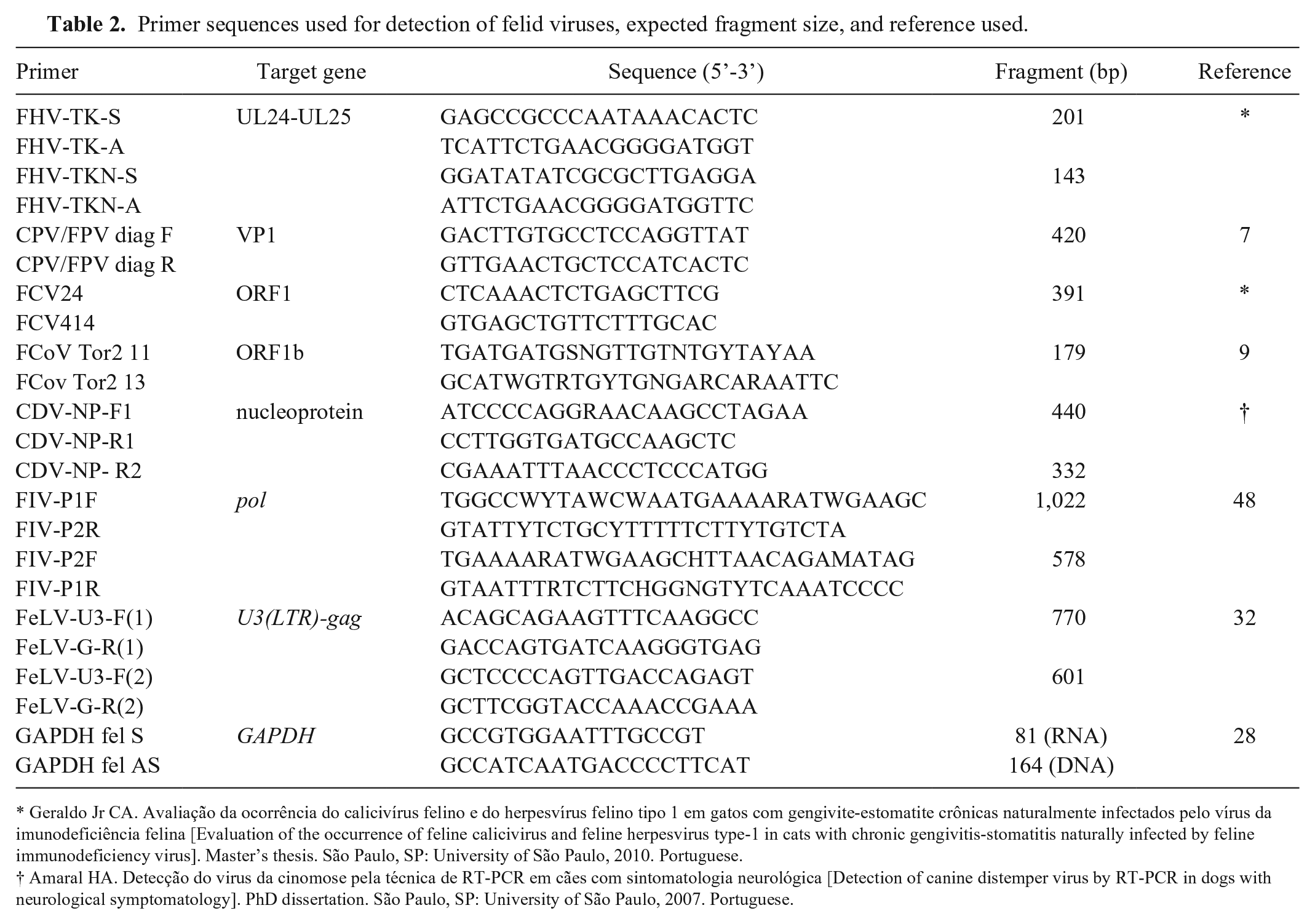
Geraldo Jr CA. Avaliação da ocorrência do calicivírus felino e do herpesvírus felino tipo 1 em gatos com gengivite-estomatite crônicas naturalmente infectados pelo vírus da imunodeficiência felina [Evaluation of the occurrence of feline calicivirus and feline herpesvirus type-1 in cats with chronic gengivitis-stomatitis naturally infected by feline immunodeficiency virus]. Master’s thesis. São Paulo, SP: University of São Paulo, 2010. Portuguese.
For FHV-1 and CPPV-1 detection, amplifications were performed using FHV-TK-S and FHV-TK-A outer primers and CPV/FPV diag F and CPV/FPV diag R primers, respectively (Table 2). Touchdown PCR technique was performed for both with the following conditions: 95°C for 3 min, 12 cycles of 94°C for 30 s, 72°C for 30 s (decreasing 1°C/cycle) and 72°C for 30 s, 30 cycles of 94°C for 30 s, 60°C for 30 s and 72°C for 30 s, followed by a final extension at 72°C for 10 min. For FHV-1, nested PCR was performed using 1 µL of PCR products and the same amplification cycle, using the primers FHV-TKN-S and FHV-TKN-A (Table 2). A commercial vaccine against feline rhinotracheitis and panleukopenia (Fel-O-Vax LVK IV + CaliciVax, Fort Dodge) was used as a positive control.
For FIV, proviral DNA was amplified with P1F and P2R outer primers (Table 2) under the following conditions: 94°C for 3 min, 45 cycles of 94°C for 15 s, 58°C for 30 s and 72°C for 45 s, and a final extension of 72°C for 10 min. Nested PCR was performed using P2F and P1R inner primers (Table 2), with 1 µL of PCR product and the same amplification cycle. DNA from a whole blood sample from a domestic cat naturally infected with FIV was used as a positive control. For FeLV, proviral DNA was amplified with U3-F(1) and G-R(1) outer primers (Table 2) using a touchdown PCR technique with the following cycle: 94°C for 7 min, 14 cycles of 94°C for 45 s, 72°C for 30 s (decreasing 1°C/cycle) and 72°C for 1 min, 26 cycles of 94°C for 45 s, 58°C for 30 s and 72°C for 1 min, and final extension of 72°C for 7 min. Nested PCR was performed under the same amplification conditions, using inner primers U3-F(2) and G-R(2) (Table 2), with 1 µL of PCR product. A FeLV-positive DNA sample of a domestic cat was used as a positive control. In all reactions, ultrapure water was used as a negative control.
All samples (DNA and RNA) were tested for feline glyceraldehyde-3-phosphate dehydrogenase (
All PCR products were visualized by electrophoresis on 1.5% (for CPPV-1, FIV, and FeLV) or 2% (for FHV-1, FCV, FCoV, CDV) agarose gel stained with GelRed (0.1 µL/mL; Biotium, Uniscience, São Paulo, SP, Brazil). Reactions with fragments of expected size (Table 2) were considered positive.
Products considered positives were purified (GFXTM PCR DNA and Gel Band purification kit, GE Healthcare, Little Chalfont, Buckinghamshire, UK) and quantified in comparison to a low-mass DNA ladder (Invitrogen). Bidirectional sequencing was performed to verify the specificity of PCR and confirm the agent (BigDye Terminator v.3.1 cycle sequencing kit, Applied Biosystems, Foster City, CA; ABI 3500 sequencer, Applied Biosystems). The quality of the sequences was determined using Phred Electropherogram Quality Analysis software (http://asparagin.cenargen.embrapa.br/phph). Final sequences were obtained (ABI 3500 sequencer, Applied Biosystems), and compared with sequences in GenBank (http://www.ncbi.nlm.nih.gov/BLAST). Values of nucleotide and amino acid identity were calculated using BioEdit Sequence Alignment Editor v.7.2.5. 17
FIV nucleotide sequences generated from our study were aligned with homologous sequences of FIV retrieved from GenBank using CLUSTAL W method with BioEdit Sequence Alignment Editor v.7.2.5.
17
The FIV nucleotide sequence of this dataset was deposited in GenBank as accession KT445903. A phylogenetic tree for FIV nucleotide sequences was constructed with MEGA 6.06 program
44
using the maximum likelihood method and Hasegawa–Kishino–Yano (HKY) evolutive model with 1,000 bootstrap replicates and
Results
All of the 177 samples analyzed were found to be positive for the endogenous control
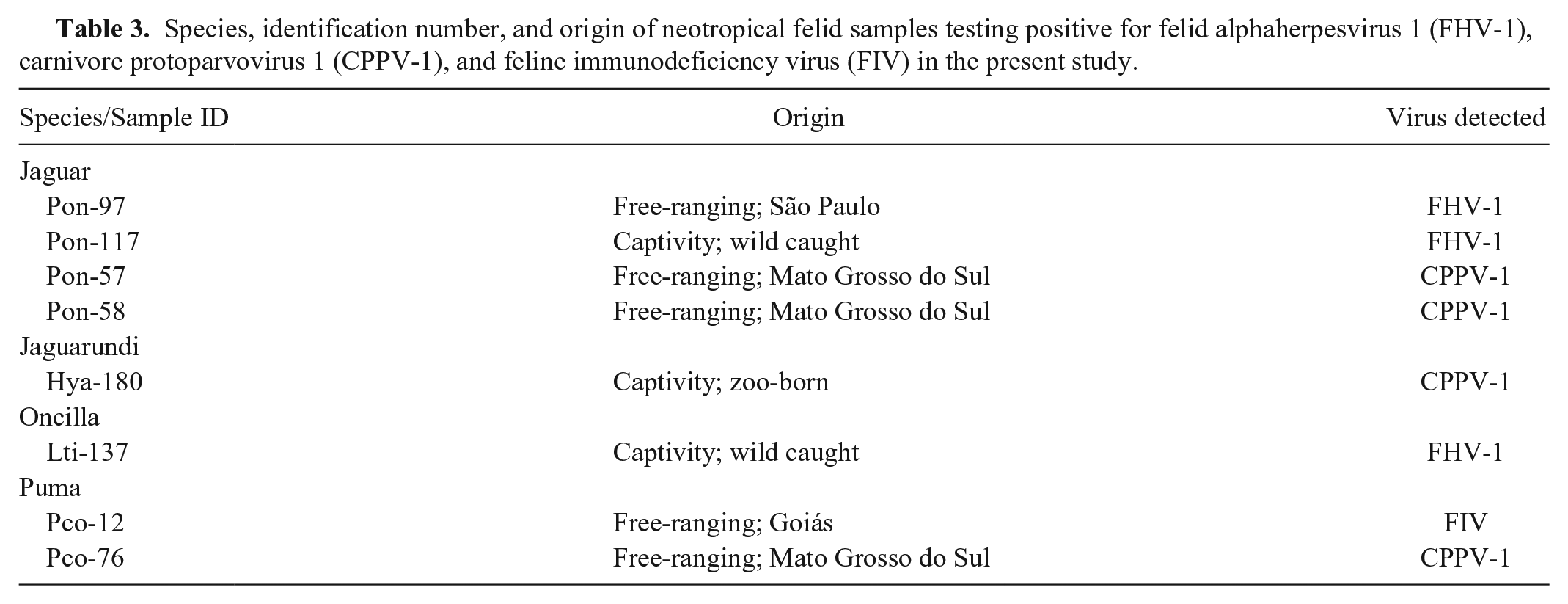
Species, identification number, and origin of neotropical felid samples testing positive for felid alphaherpesvirus 1 (FHV-1), carnivore protoparvovirus 1 (CPPV-1), and feline immunodeficiency virus (FIV) in the present study.

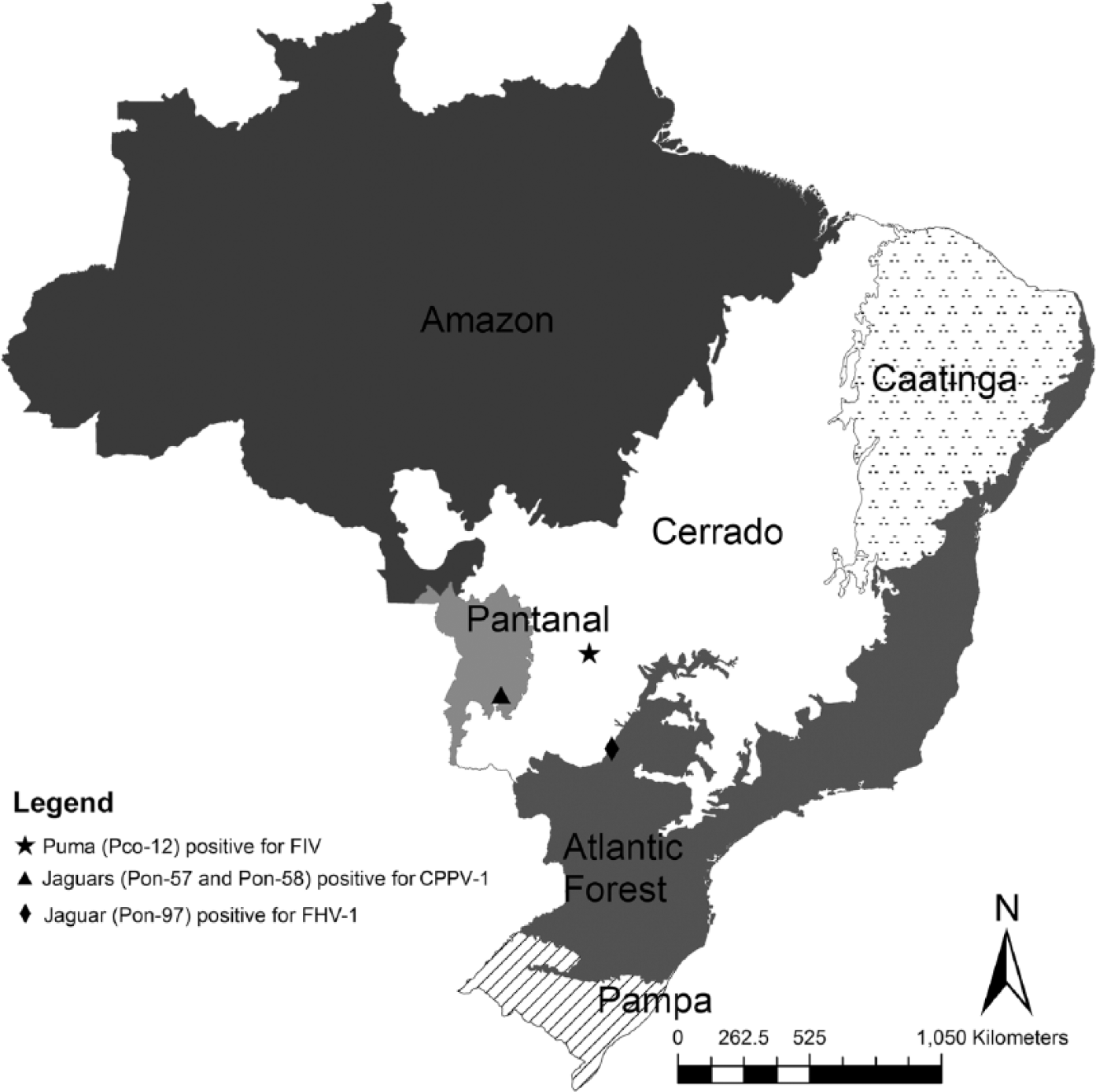
Map of Brazil showing the sites of capture of free-ranging felids testing positive for felid alphaherpesvirus 1 (FHV-1), carnivore protoparvovirus 1 (CPPV-1), and feline immunodeficiency virus (FIV).
For FHV-1, despite the small length of the amplified products (143 bp), agent confirmation was possible by DNA sequencing. Nucleotide sequences (103 bp) of the captive jaguar (Pon-117) had 99% identity to free-ranging jaguar (Pon-97) and captive oncilla (Lti-137), whereas the latter two had 98% identity to each other. The captive jaguar (Pon-117) showed 100% identity to FHV-1 samples described in domestic cats (M26660, FJ478159) and free-ranging jaguar (Pon-97), and the captive oncilla (Lti-137) showed 99% identity with these same samples. Both captive felids (Pon-117 and Lti-137) were wild caught and introduced to the captive settings.
For CPPV-1, 294-bp nucleotide sequences of the free-ranging jaguars (Pon-57 and Pon-58) showed 100% identity between them and 99.6% and 99.3% with the sequences identified in samples of the free-ranging puma (Pco-76) and the captive zoo-born jaguarundi (Hya-180), respectively. Nucleotide sequences of the puma (Pco-76) were identical to parvovirus of a domestic dog (
For FIV, nucleotide sequences (444 bp) had 82.4% identity with the lentivirus sequence identified in a free-ranging puma from Yellowstone National Park in the United States (DQ107148) and 79.5% of identity with the sequence from a puma kept in captivity in Brazil (AY878236). Comparing the amino acid sequence (148aa), the sample of the free-ranging puma (Pco-12) had high identity with puma samples: 90.5% identity with an amino acid sequence identified in a free-ranging puma from Canada (AY307116), 88.5% from a free-ranging puma from Yellowstone National Park (DQ107148), and 85.8% from a puma kept in captivity in Brazil (AY878236). The phylogenetic tree of FIV nucleotides sequences showed that sample Pco-12 segregated with puma samples retrieved from GenBank, diverging from samples identified in other feline species (Fig. 2).
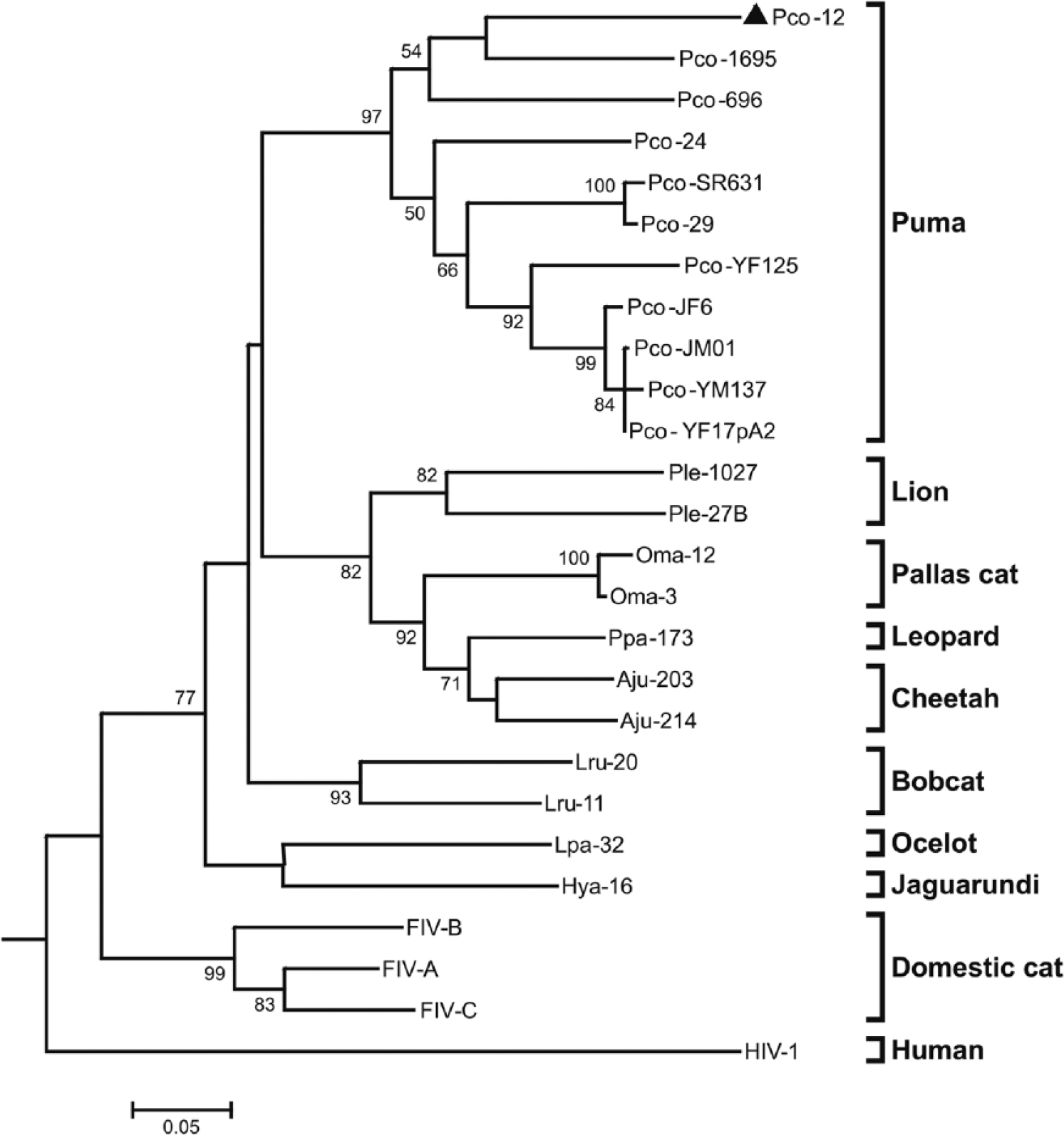
Phylogenetic tree of 444-bp nucleotide sequences of feline immunodeficiency virus FIV
Discussion
Neotropical felid samples tested in our study were found to be positive for 3 of the 7 viral agents surveyed: FHV-1, CPPV-1, and FIV. Presence of FHV-1 in free-ranging and captive jaguars and in a captive oncilla is consistent with other serologic studies,3,10,11 but our study describes molecular detection in neotropical felids. Exposure to FHV-1 is reported, but at low frequency, showing that the virus does not circulate widely in these species, except in the oncilla. 3 PCR positive for FHV-1 can confirm the circulation of the virus in neotropical felids but should be interpreted with caution because the technique is able to detect small amounts of viral nucleic acid, which may or may not be associated with disease. 47 Virus remains latent after recovery, and cats become asymptomatic carriers, but during reactivation, which can be caused by stress of capture or captivity, animals release the virus intermittently again. 47 So, these PCR-positive wild cats are epidemiologically important as sources of infection for susceptible animals. 15 Because the FHV-1 primer sequence chosen in our study detected a 143-bp sequence from a conserved region, it was not possible to perform phylogenetic analysis. Further studies should be performed with other primers that target other regions of the viral genome in order to examine genetic diversity in FHV-1 present in domestic and wild populations.
CPPV-1 was detected infecting 2 free-ranging jaguars and 1 puma, and 1 captive zoo-born jaguarundi. This low frequency is consistent with previous serologic studies of exposure to the agent in wild felids in Brazil, which detected higher numbers of animals with antibodies against the virus.10,11 High CPPV-1 resistance in the environment and the fecal–oral route of transmission favor exposure to the agent. Infected felids are viremic for many days and excrete the virus in feces, urine, saliva, or vomitus.
49
Thus, when feces are not available for testing, a blood sample is indicated for detection,
49
as used in our study. Identity of CPPV-1 nucleotide sequence of positive samples was >98%, which is consistent with other parvoviruses described in the literature.
42
For future studies, we suggest using the
FIV has not been reported previously in free-ranging pumas from Emas National Park in Goiás State. In Brazil, FIV or a similar lentivirus has been detected in captive pumas, jaguar, jaguarundi, oncilla, margay, and ocelot.4,26 Antibodies to the agent were also detected in free-ranging pumas from the Pantanal (wetlands located primarily in Mato Grosso do Sul). 10 FIV is transmitted by direct contact, through inoculation of virus from saliva of an infected cat via biting. 21 Thus, the solitary habit of neotropical cats does not favor transmission of this agent.
Nucleotide and amino acid sequences from the free-ranging puma (Pco-12) grouped with other lentiviruses described in pumas, confirming the presence of a species-specific strain of FIV in free-ranging pumas in this region of Brazil. Nucleotide sequences from Pco-12, a free-ranging puma from Vancouver Island (Pco-1695; AY307116), and a captive puma from Brazil (Pco-696; AY878236) are closely related and quite distant phylogenetically from the other pumas in our study. As well as Pco-12, the captive puma (Pco-696; AY878236) from Brazil is also from the State of Goiás, but we do not know if the animal was wild-caught or zoo-born. Absence of clinical signs of disease in puma may reflect a longer period of coevolution between virus and host in this species.
36
However, further disease monitoring of FIV-infected pumas would be important to evaluate the pathogenicity of the virus for this species. For future studies, in order to perform molecular characterization of FIV detected in Pco-12, we recommend the use of the
Absence of detection of FeLV shows that neotropical felids in our study probably were not in acute, regressive, or progressive stages of disease, characterized by the presence of proviral DNA. 29 Corroborating this result, in a previous study we performed serologic detection in some of these same samples of free-ranging wild felids and they had no FeLV antigen, 14 but 2 captive jaguarundis were FeLV antibody positive by western blot technique and anti-FeLV gp 70 in ELISA. 11 Although proviral DNA of FeLV has been detected also in a captive zoo-born ocelot and oncilla in Brazil, 16 when it is present, it normally occurs at low frequency. Secretions and excretions are the routes of elimination of the agent, but because of their fragility in the environment, transmission occurs primarily through direct contact with saliva and nasal secretions.20,25 Thus, the solitary habit of free-ranging neotropical felids studied here does not favor transmission of the virus, although it can be a problem in captive settings.
CDV, responsible for important outbreaks in big cats,1,40 was not detected in wild felids in our study. Antibody detection to CDV in free-ranging jaguars was reported previously,14,35 but fatal cases were not reported. Antibody detection and PCR-negative results may indicate prior exposure to the agent but absence of infection at the time of capture or collection of samples from these individuals. Because the virus is not always present in the bloodstream, the moment of capture or collection must coincide with the acute phase of infection and elimination of the virus in exudates. 6 At this stage, ocular and nasal secretions, saliva, and urine could be harboring the agent, but were not investigated in our study.
Although serologic detection of FCV has been reported in various species of wild cats,27,37,46 we did not detect the FCV genome. FCV is widely distributed in felid populations, and virus circulation in Brazil was detected by serologic tests in wild and domestic felids.10,18 Sensitivity of molecular testing depends on primers and strain used, given the great variability of the viral genome. 39 The recently deposited complete genome of FCV isolate from a tiger (GenBank KJ9944377) could be used to design specific primers for wild felids. Oral and nasal secretion swabs collected during the acute phase of disease should preferably be used for FCV detection.
Absence of FCoV was not expected for the captive felids, given that 55 individuals tested in our study were antibody-positive to the agent in a previous study. 11 However, negative results may have occurred because RT-PCR detects the agent but does not detect previous exposure to the agent. In addition, negative results may be the result of low amounts of virus present in the sample, or high FCoV genetic variability.16,24 Given that FCoV is shed in feces, these could have been better samples for agent detection. But, FCoV resistance in the environment is low. In captive neotropical felids from Brazil, the agent was not reported in feces, 16 but previous studies show evidence of FCoV infection by direct immunofluorescence assay. 41
Our results indicate the circulation of FHV-1, CPPV-1, and FIV in wild cats in Brazil, but no inference can be made as to whether these agents are historically associated with neotropical felid populations or were introduced by interactions with domestic carnivores. Neither is it possible to infer about the impact of these agents in the studied populations or the health of positive individuals. Our results expand our knowledge about virus circulation in felid species in Brazil, and emphasize that they may represent a threat. For free-ranging animals, this information can also serve as the basis for future projects involving translocation and reintroduction of wild cats.
Footnotes
Acknowledgements
We thank the Centro Nacional de Pesquisa e Conservação de Mamíferos Carnívoros–CENAP/ICMBio for providing neotropical felids samples. We thank Gisele Oliveira de Souza, Cassia Yumi Ikuta, and Sheila Oliveira de Souza Silva for laboratory assistance.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received financial support from FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo, process 11/09970-7). MF received a scholarship from FAPESP (process 11/07944-9).
