Abstract
Regulatory T cells (Tregs) can be targeted in cancer immunotherapy. A previous study has shown that the chemokine CCL17 and the receptor CCR4 play a role in Treg recruitment in canine urothelial carcinoma. Here, we describe the association of tumor-infiltrating Tregs with CCL17/CCR4 expression in dogs with other carcinomas. In this study, we investigated 23 dogs with mammary carcinoma, 14 dogs with oral squamous cell carcinoma, 16 dogs with pulmonary adenocarcinoma, and 8 healthy control dogs. Immunohistochemistry showed that Foxp3+ Tregs and CCR4+ cells were increased in the tumor tissues of mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma, when compared with the healthy tissues. The number of CCR4+ cells was associated with that of Foxp3+ Tregs. Double immunofluorescence labeling confirmed that most tumor-infiltrating Foxp3+ Tregs expressed CCR4. In vitro, canine carcinoma cell lines expressed CCL17 mRNA. Quantitative RT-PCR (reverse transcriptase-polymerase chain reaction) showed that CCL17 mRNA expression in canine carcinomas was increased approximately 10- to 25-fold relative to that of healthy tissues. These results suggest that the CCL17/CCR4 axis may drive Treg recruitment in a variety of canine carcinomas. CCR4 blockade may be a potential therapeutic option for tumor eradication through Treg depletion.
Regulatory T cells (Tregs) are a subset of CD4+ helper T lymphocytes expressing the IL-2 receptor alpha chain (CD25) and transcription factor forkhead box P3 (Foxp3). The function of Tregs is to suppress activity of effector immune cells. Tregs play a role not only in suppression of immune response and inflammation but also in tumor progression, presumably by inhibiting antitumor immunity. In humans, Treg infiltration has been reported in certain tumors and has been related to progression and prognosis. 5,24 High tumor-infiltrating Tregs are associated with poor prognosis in patients with ovarian, 3,29 breast, 1,9 lung, 36 gastric, 25 squamous cell, 10 and hepatocellular carcinomas. 6,8 There is a growing interest in Treg infiltration in canine tumors; numerous studies have reported on the association of tumor-infiltrating Tregs with poor prognosis in various canine tumors, such as melanoma, 27,28 intestinal lymphoma, 21 mammary carcinoma, 2 squamous cell carcinoma, 28 and pulmonary adenocarcinoma. 28
Evidence suggests that tumor-infiltrating Tregs are a potential target for cancer immunotherapy. In mice and humans, several molecules are candidates for Treg depletion or attenuation of their function, including CD25, CD15s, CD134, cytotoxic T-lymphocyte associated protein 4 (CTLA-4), glucocorticoid-induced TNF receptor (GITR), lymphocyte-activation gene 3 (LAG3), programmed cell death protein-1 (PD-1), and C-C chemokine receptor 4 (CCR4). 24,35 Anti-Treg therapy is under investigation for human malignant tumors, such as melanoma, adult T-cell leukemia/lymphoma, lung cancer, and esophageal cancer. 14,16,34,38 The mechanisms of Treg infiltration and the therapeutic efficacy of its depletion in canine tumors remain unclear.
CCR4 is a receptor for C-C chemokines CCL17 and CCL22, both of which are produced by mature dendritic cells. 12,13 It has been shown that CCR4 is preferentially expressed on type 2 helper T (Th2) cells or Tregs in both mice and humans. 43 A recent study by our group has demonstrated that Tregs infiltrate tumor tissues through the interaction between tumor-producing CCL17 and CCR4 on Tregs, and CCR4 blockade exerts an antitumor effect in dogs with urothelial carcinoma. 19 However, little is known about the role of CCR4 in Treg infiltration in other canine tumors.
In the present study, we investigated the number of Foxp3+ Tregs and CCR4+ cells, and the gene expression of ligands for CCR4, in dogs with mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma, to elucidate the therapeutic potential of anti-CCR4 treatment for these canine carcinomas.
Materials and Methods
Canine Tumor Samples
For immunohistochemistry and immunofluorescence analyses, we obtained archival formalin-fixed, paraffin-embedded tumor tissues that were surgically removed from 53 dogs at the Veterinary Medical Center of the University of Tokyo. The following diagnoses were confirmed by histopathology: mammary simple carcinoma (n = 23), oral squamous cell carcinoma (n = 14), and pulmonary adenocarcinoma (n = 16). Lymph node involvement and lung metastasis were assessed by palpation, ultrasound, and radiographic examinations, respectively. In this study, lymph node involvement was observed in 4/23 dogs with mammary carcinoma, 4/14 dogs with squamous cell carcinoma, and 2/16 dogs with pulmonary adenocarcinoma. There were no cases with identified lung metastasis. No dogs received chemotherapy or radiation therapy before surgery. Normal tissues (mammary gland, oral mucosa, and lung) were obtained from 8 healthy beagles via necropsy for other experiments and used as controls.
For gene expression analysis, fresh tumor tissues from 15 dogs with mammary carcinoma, 10 dogs with oral squamous cell carcinoma, and 12 dogs with pulmonary adenocarcinoma were immediately submerged in RNAlater (Thermo Fisher Scientific, Waltham, MA) and stored at −20°C. Archival frozen normal tissues (mammary gland, oral mucosa, and lung) collected from 5 healthy beagles euthanized for another experimental purpose were used as a control. All procedures in this study were conducted in accordance with the guidelines of the Animal Care Committee of the Graduate School of Agricultural and Life Sciences, the University of Tokyo.
Immunohistochemistry
The expression of Foxp3 and CCR4 was examined by immunohistochemistry. 19 Heat-induced antigen retrieval was performed by autoclaving the sections for 10 minutes at 121°C in 10 mM sodium citrate buffer (pH 6.0). Endogenous peroxidase activity was blocked by incubation with REAL Peroxidase-Blocking Solution (DAKO, Glostrup, Denmark) at room temperature for 10 minutes. Sections were blocked with 5% skim milk in Tris-buffered saline with 0.1% Tween 20 (TBST) at room temperature for 60 minutes, and then incubated with primary antibodies, rat anti-Foxp3 (1:400 dilution, clone FJK-16s, eBioscience, San Diego, CA), or mouse anti-CCR4 (1:100 dilution, clone 1G1, BD Biosciences, Franklin Lakes, NJ) at 4°C overnight. The cross-reactivity of these primary antibodies with canine Foxp3 and CCR4 was confirmed in previous studies. 20,23 Secondary antibodies were applied as follows: samples were incubated with EnVision polymer reagent anti-mouse (Dako) at room temperature for 45 minutes, or biotin-labeled anti-rat IgG (Vector Laboratories, Burlingame, CA) at 37°C for 30 minutes followed by HRP-labeled streptavidin (Dako) at room temperature for 30 minutes. For Foxp3 and CCR4, mesenteric lymph nodes were used as positive controls. Incubation of sections with TBST only served as negative controls. The reaction products were visualized with 3,3′-diaminobenzidine. Cells with clear lymphocyte morphology, distinct nuclear immunolabeling for Foxp3, or membranous immunolabeling for CCR4 were evaluated as positive. Foxp3+ and CCR4+ cells were manually counted in 10 representative fields (400× magnification: 0.238 mm2 per field) with the most intense positivity and >50% of tumor cells using ImageJ software. 28,30
For double immunofluorescence labeling, antigen retrieval, endogenous peroxidase blocking, and milk blocking were performed as described above. Primary antibodies, rat anti-Foxp3 (1:100 dilution, clone FJK-16s, eBioscience) and mouse anti-CCR4 (1:100 dilution, clone 1G1, BD Biosciences), were applied at 4°C overnight. Immunofluorescence was performed using secondary antibodies, namely, Alexa Fluor 594 goat anti-rat IgG (1:500, Invitrogen, Carlsbad, CA) and Alexa Fluor 488 donkey anti-mouse IgG (1:500, Invitrogen). Images were captured using a fluorescence microscope (BZ-X800; Keyence, Osaka, Japan).
Cell Culture
A total of 7 canine carcinoma cell lines were used. 22,39,44 The cell lines were derived from mammary adenocarcinoma (CIPp, CIPm, and CTBp), squamous cell carcinoma (POCO, SYRUP, and MILK), and pulmonary adenocarcinoma (GEN). CIPp, CIPm, CTBp, and GEN cells were grown in RPMI-1640 (Sigma, St. Louis, MO), supplemented with 10% FBS (Thermo Fisher Scientific) and 100 μg/ml penicillin and streptomycin (Sigma) at 37°C under a humidified atmosphere with 5% CO2. POCO, SYRUP, and MILK cells were grown in DMEM (Thermo Fisher Scientific, Waltham, MA), supplemented with 10% FBS (Thermo Fisher Scientific) and 100 μg/ml penicillin and streptomycin (Sigma) at 37°C under a humidified atmosphere with 5% CO2. Two nontumor cell lines, established from normal lip epithelium COPK 41 and keratinocytes CPEK, 33,41 were purchased from CELLnTEC Advanced Cell Systems (Bern, Switzerland). COPK and CPEK cells were maintained in CnT-09 medium (CELLnTEC Advanced Cell Systems) at 37°C under 5% CO2.
Gene Expression Analysis
We extracted total RNA from each canine cell line using the RNAspin Mini RNA Isolation Kit (GE Healthcare, Buckinghamshire, England). Reverse transcription (RT) was accomplished using a ReverTra Ace (Toyobo, Osaka, Japan). PCR (polymerase chain reaction) was performed over 35 cycles using Taq polymerase (AmpliTaq Gold; Thermo Fisher Scientific) under a 3-step thermal cycling process of 94°C for 30 seconds, 60°C for 30 seconds, and 72°C for 30 seconds. Table 1 lists the primer pair sequences. Amplicons were separated by electrophoresis and stained by ethidium bromide for visualization and confirmation of the amplicon size.
Primer Pair Sequences Used for RT-PCR.
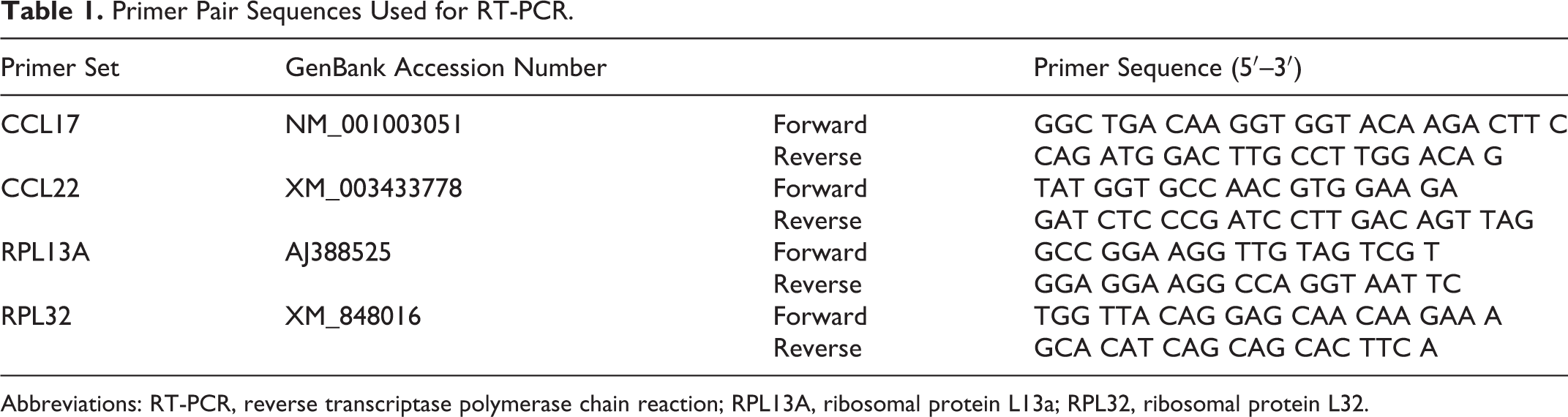

Abbreviations: RT-PCR, reverse transcriptase polymerase chain reaction; RPL13A, ribosomal protein L13a; RPL32, ribosomal protein L32.
For quantitative RT-PCR, we extracted the total RNA from tumor and normal tissues using the RNAspin Mini RNA Isolation Kit (GE Healthcare). Two-step real-time RT-qPCR (StepOnePlus; Applied Biosystems, Waltham, MA) was performed using THUNDERBIRD SYBR qPCR Mix (Toyobo) under the amplification conditions of 95°C for 10 seconds, followed by 40 cycles of PCR (95°C for 5 seconds and 60°C for 30 seconds). RPL13A and RPL32 were used as references. 26 Table 1 lists the primer pair sequences. All primer sets exhibited >95% efficiency, which was determined by using a 10-fold dilution series from representative cDNA samples. The nuclease-free water was used as a negative control. Cycle threshold (Ct) values that indicate points of threshold intersects with the amplification curve of PCR were determined using StepOne Plus v2.3 software (Applied Biosystems). All samples were examined in duplicate, and the mean ΔCt was calculated. Each mRNA expression level (relative quantity) was calculated using 2-ΔCt, resulting in the evaluation of carcinoma tissues as the n-fold difference from the mean value of normal tissues after normalizing based on the reference genes.
Statistical Analysis
We used JMP Pro version 14.2.0 (SAS Institute, Cary, NC) for statistical analyses. Intergroup comparisons were assessed using the Mann-Whitney U test. The Kruskal-Wallis test, followed by the Dunn test, was used for multiple comparisons. Correlation between 2 variables was assessed using Spearman’s rank correlation coefficient. Statistical significance was defined as P < .05.
Results
Foxp3+ Tregs and CCR4+ Cells Infiltrate Into Canine Carcinoma Tissues
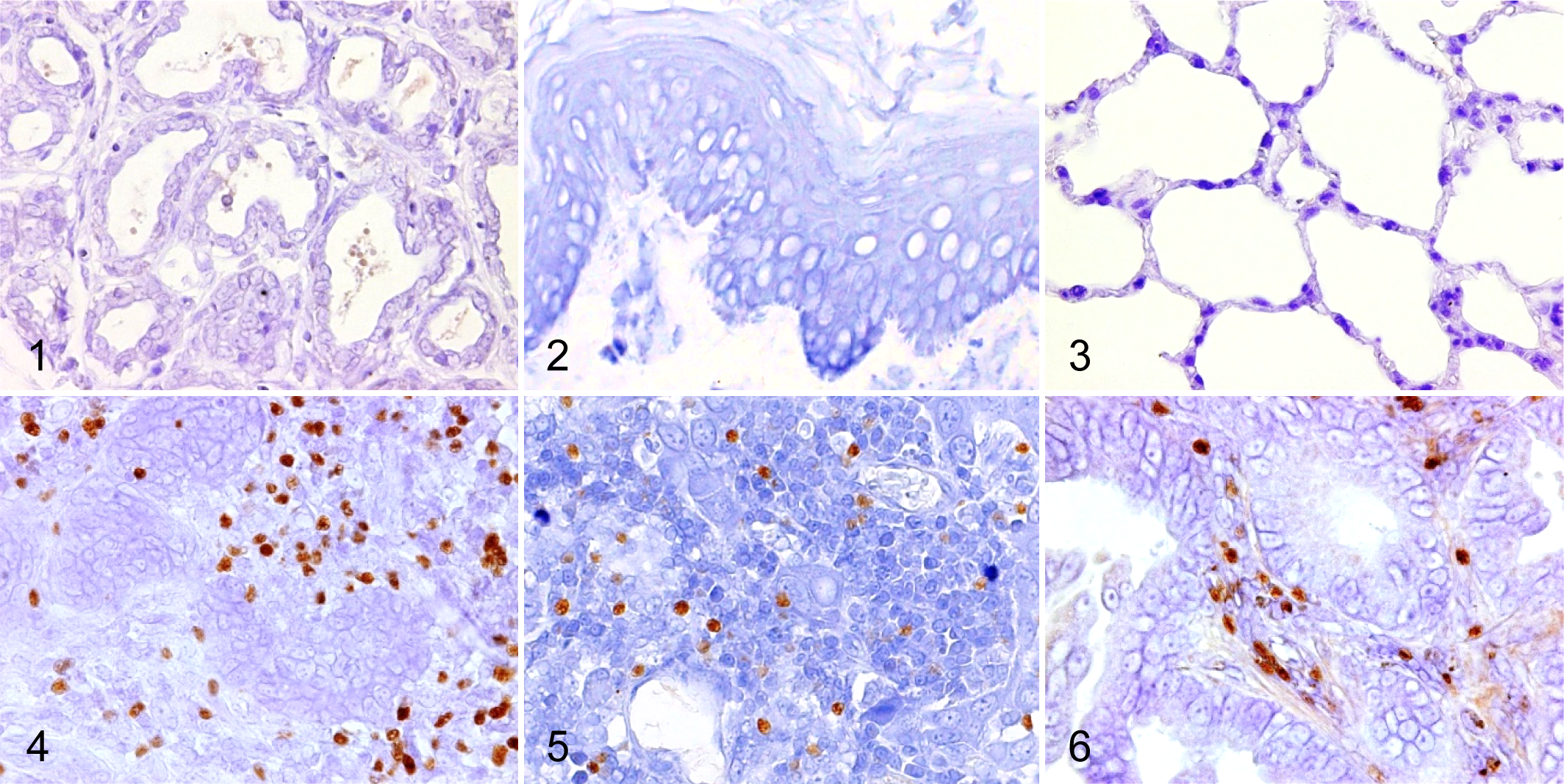
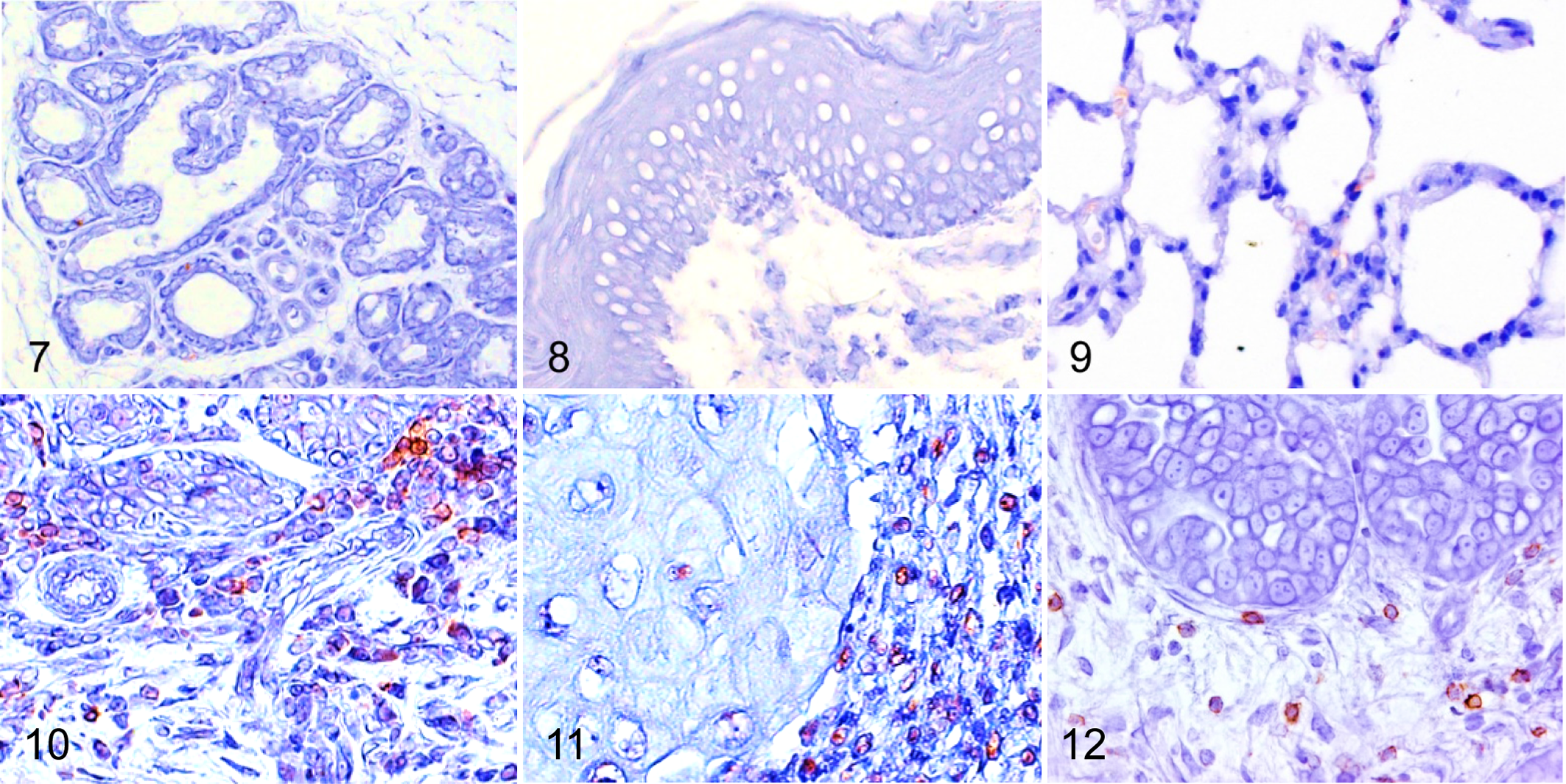
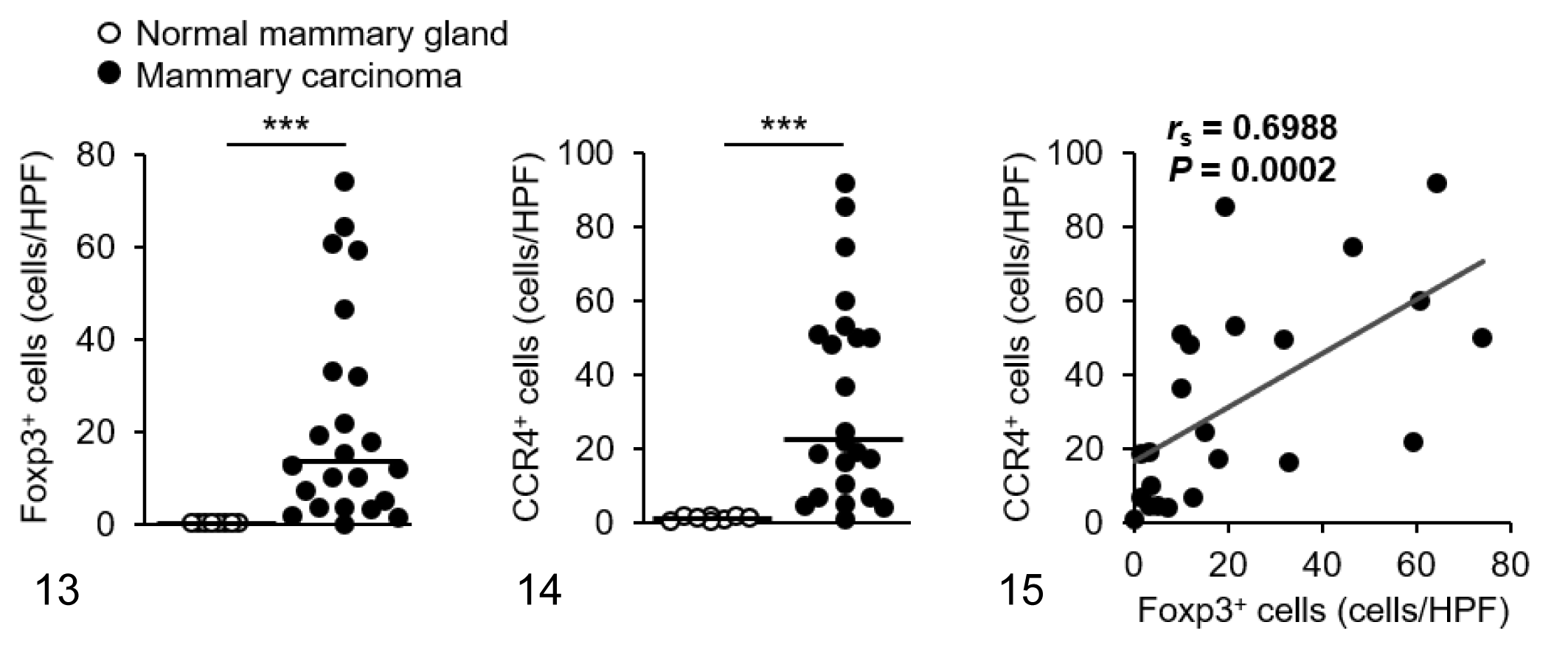
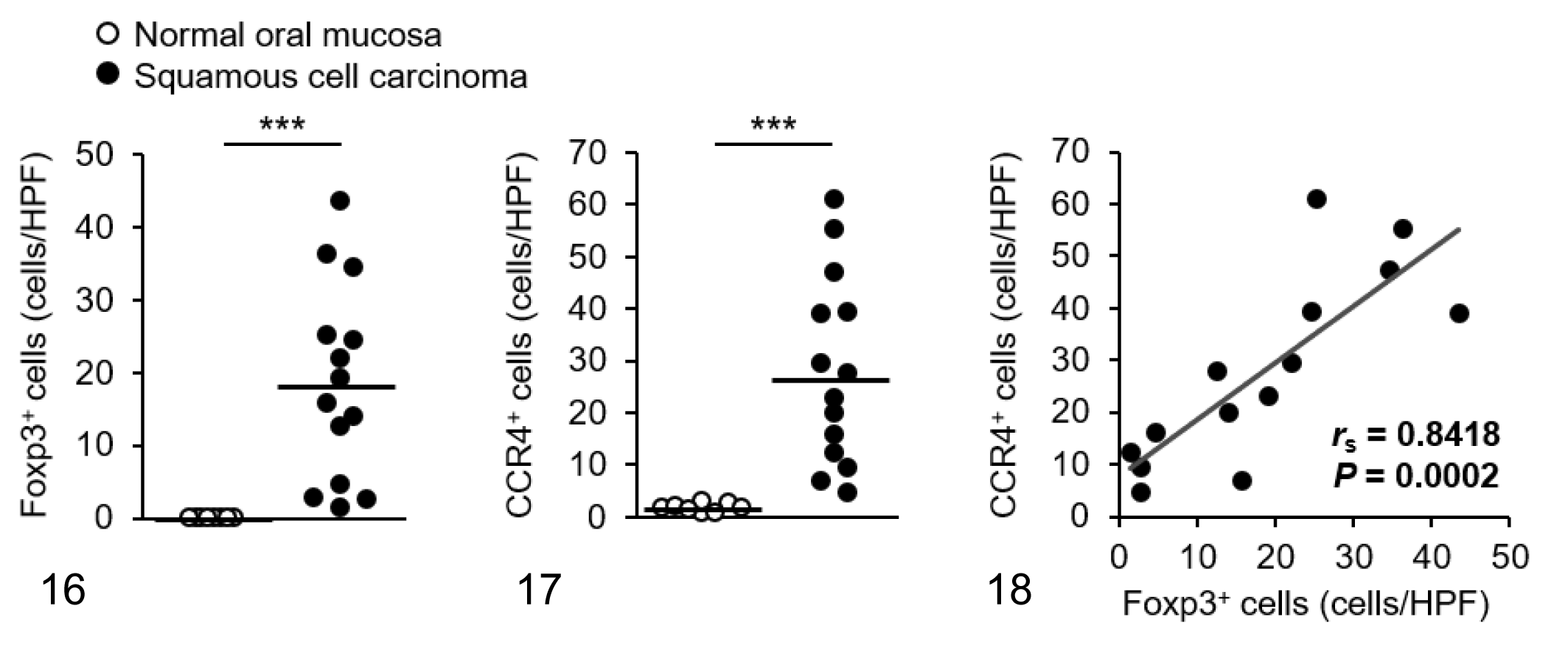
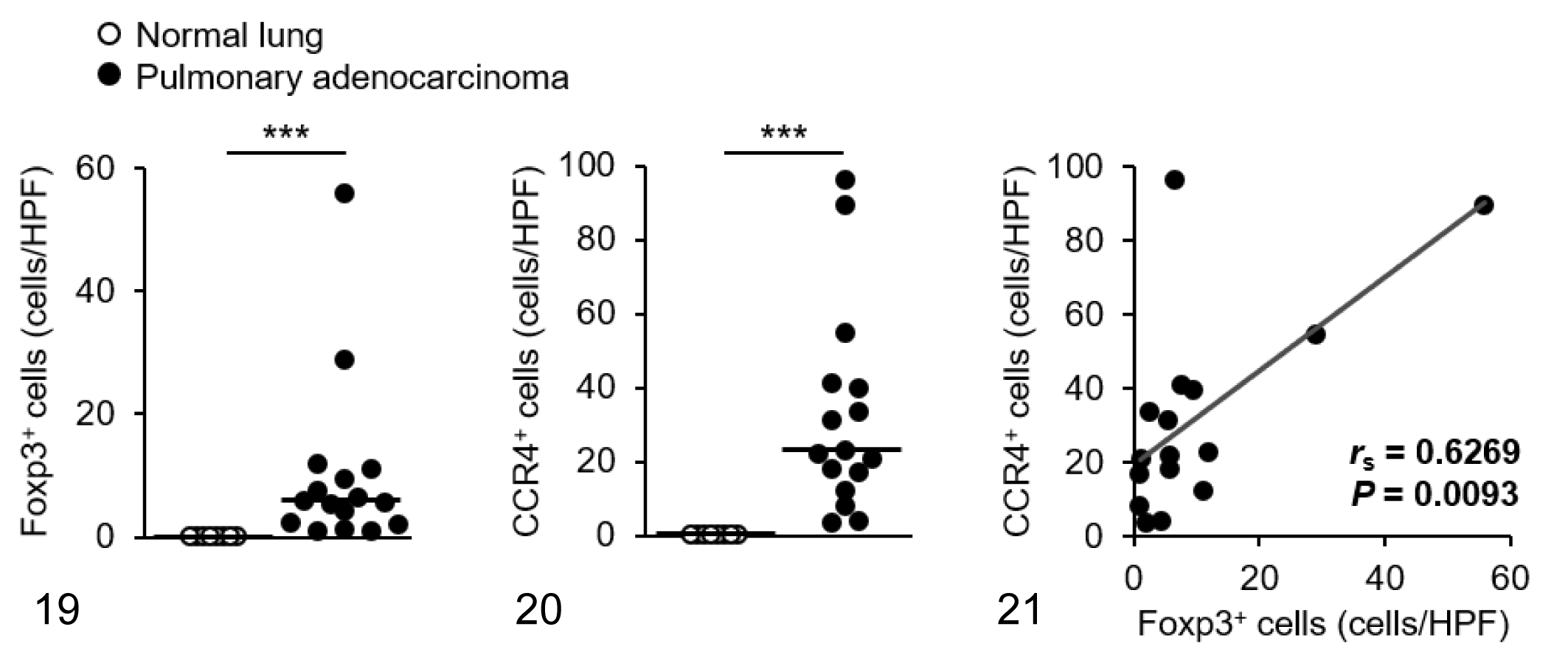
We visualized Foxp3 and CCR4 with immunohistochemistry and examined the localization and number of tumor-infiltrating Tregs and CCR4+ cells. We observed only a few Foxp3+ Tregs in the normal tissues (Figs. 1–3), whereas Foxp3+ Tregs were densely present in mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma (Figs. 4–6). Tumor-infiltrating CCR4+ cells were also increased in these carcinomas but not in the normal tissues (Figs. 7–12). Both Foxp3+ Tregs and CCR4+ cells were observed mainly in the peritumoral stroma, and some of them were present in the intratumoral area. Compared to normal tissues, Foxp3+ Tregs and CCR4+ cells were more frequent in dogs with mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma (Figs. 13 –21). The density of Foxp3+ Tregs was positively correlated to the number of CCR4+ cells (Figs. 13 –21).
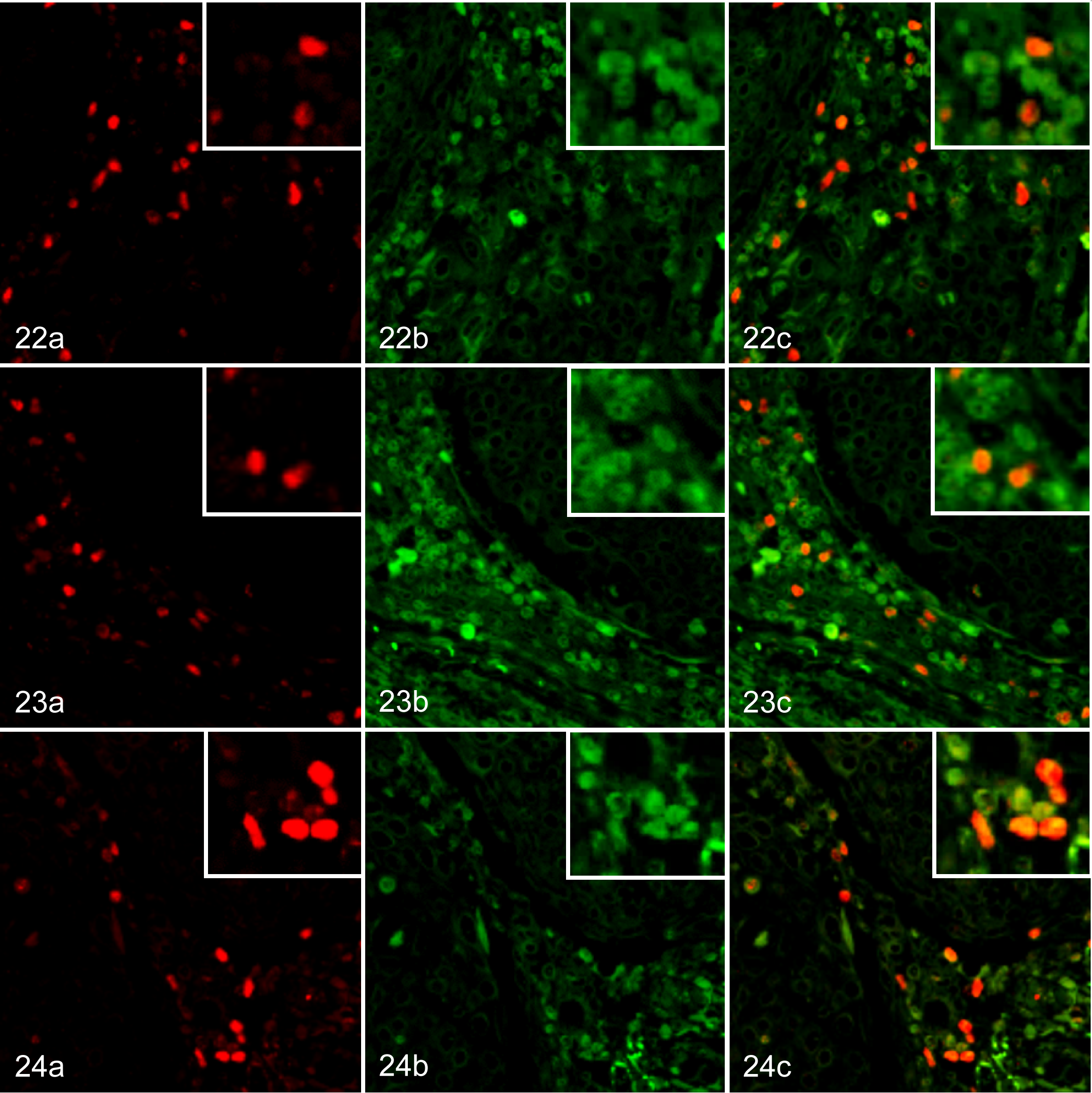
Tumor-Infiltrating Tregs Express CCR4 in Various Canine Carcinomas
To confirm the expression of CCR4 in tumor-infiltrating Tregs, we conducted double immunofluorescence labeling. Tumor-infiltrating Foxp3+ Tregs expressed CCR4 in mammary carcinoma (Fig. 22), squamous cell carcinoma (Fig. 23), and pulmonary adenocarcinoma (Fig. 24). In addition, Foxp3− CCR4+ non-Treg cells were also present in the carcinoma tissues (Figs. 22–24). However, Foxp3+CCR4− Tregs were only rarely observed in the carcinoma tissues (Figs. 22–24). The median proportions of Foxp3+CCR4+ cells, Foxp3-CCR4+ cells, and Foxp3+CCR4− cells were 28.4% (range, 20.1% to 44.3%), 69.5% (58.1% to 79.4%), and 1.2% (0% to 2.8%) in dogs with mammary carcinoma; 27.7% (20.6% to 39.1%), 72.4% (47.3% to 81.6%), and 0% (0% to 2.2%) in dogs with squamous cell carcinoma; and 34.1% (25.8% to 48.2%), 63.1% (43.1% to 74.2%), and 0% (0% to 3.5%) in dogs with pulmonary adenocarcinoma, respectively.
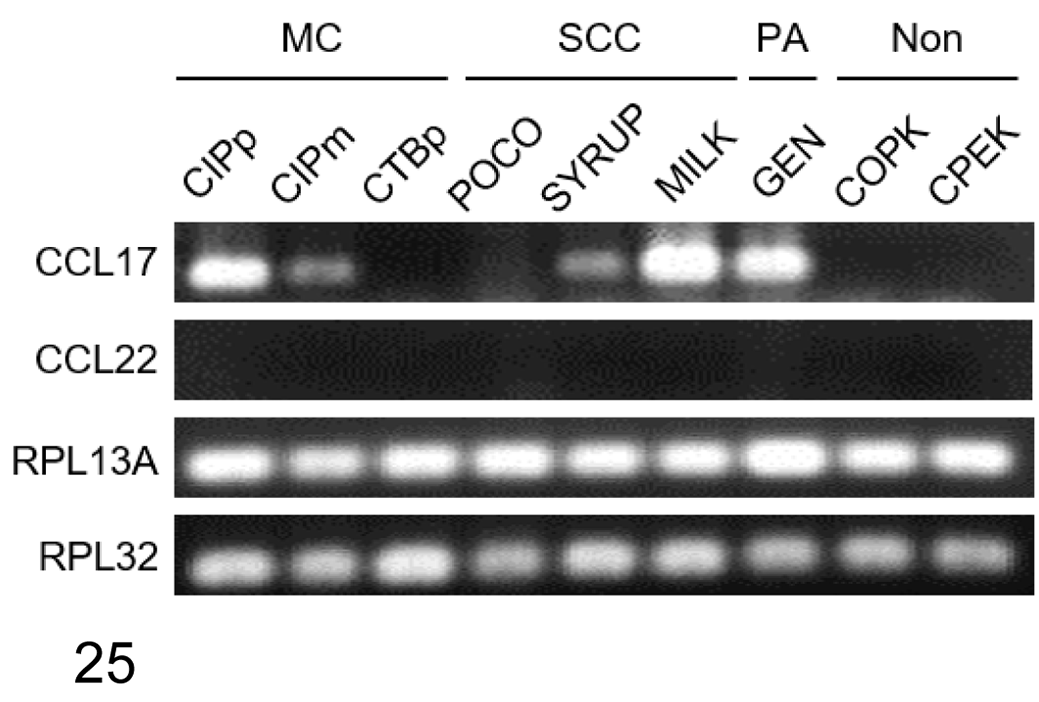
CCL17 mRNA Expression Is Elevated in Various Canine Carcinomas
To examine the mRNA expression of CCL17 and CCL22 in tumor cells, RT-qPCR was performed using canine carcinoma cell lines. We detected mRNA expression of CCL17, but not CCL22, in mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma cell lines (Fig. 25). Interestingly, CCL17 mRNA expression varied among cell lines of the same tumor type. CCL17 mRNA was detected in 2/3 mammary carcinoma cell lines, 2/3 squamous cell carcinoma cell lines, and 1/1 pulmonary adenocarcinoma cell line (Fig. 25). The normal canine squamous epithelial cell line COPK and the epidermal keratinocyte cell line CPEK did not express CCL17 or CCL22 (Fig. 25).
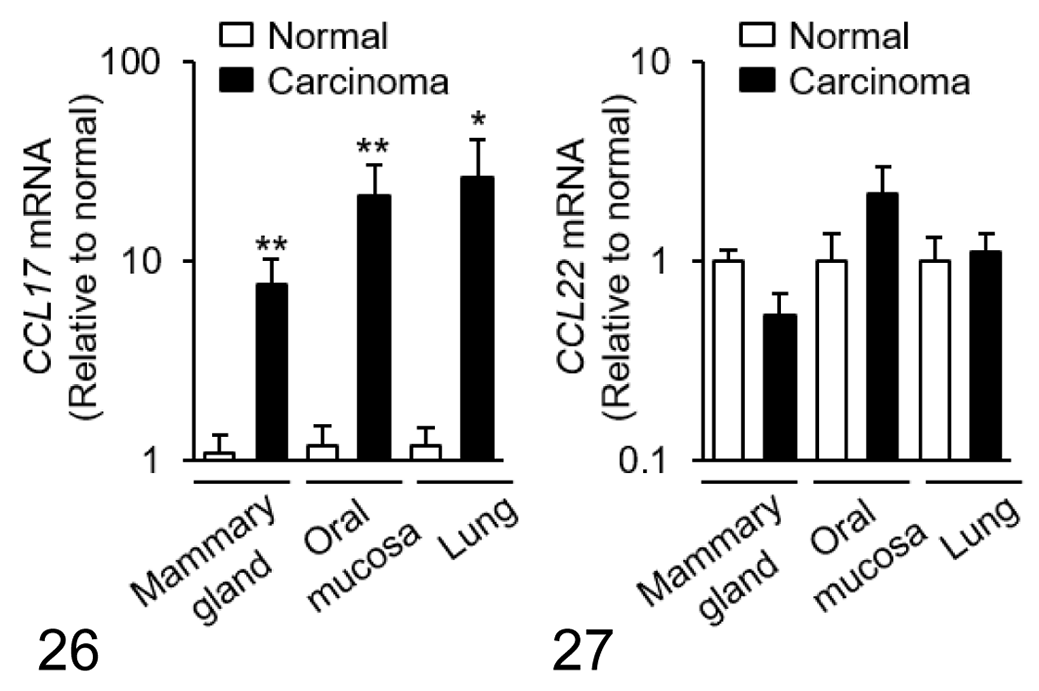
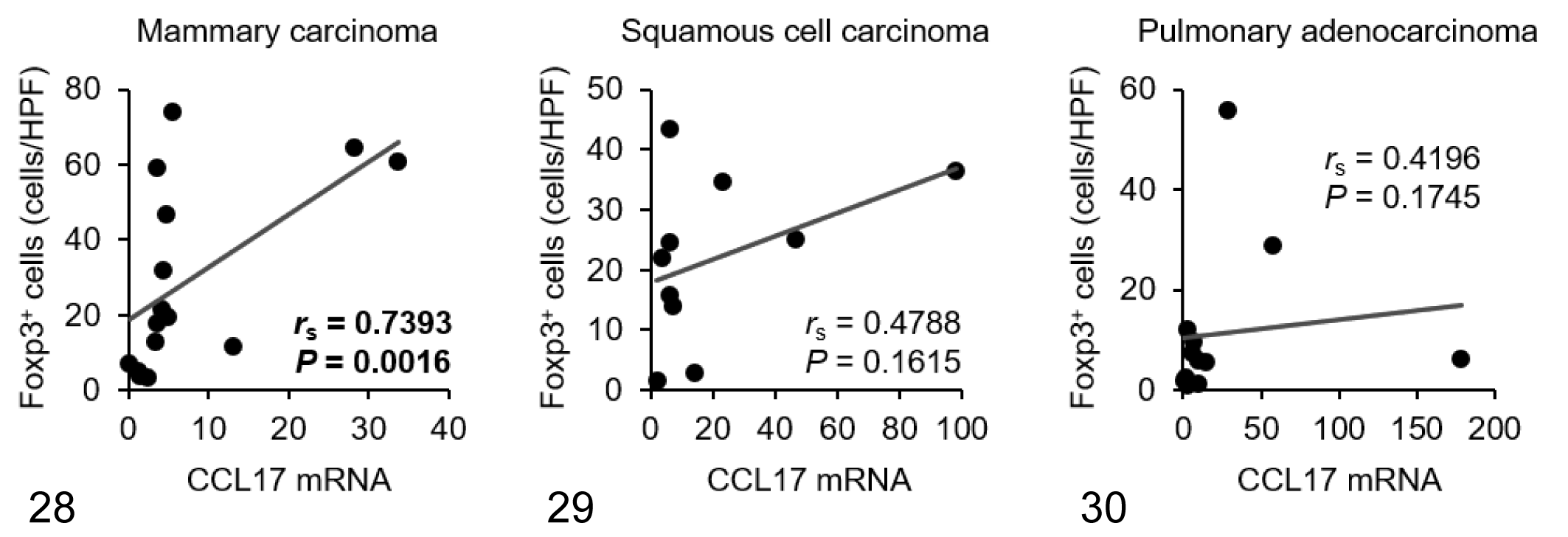
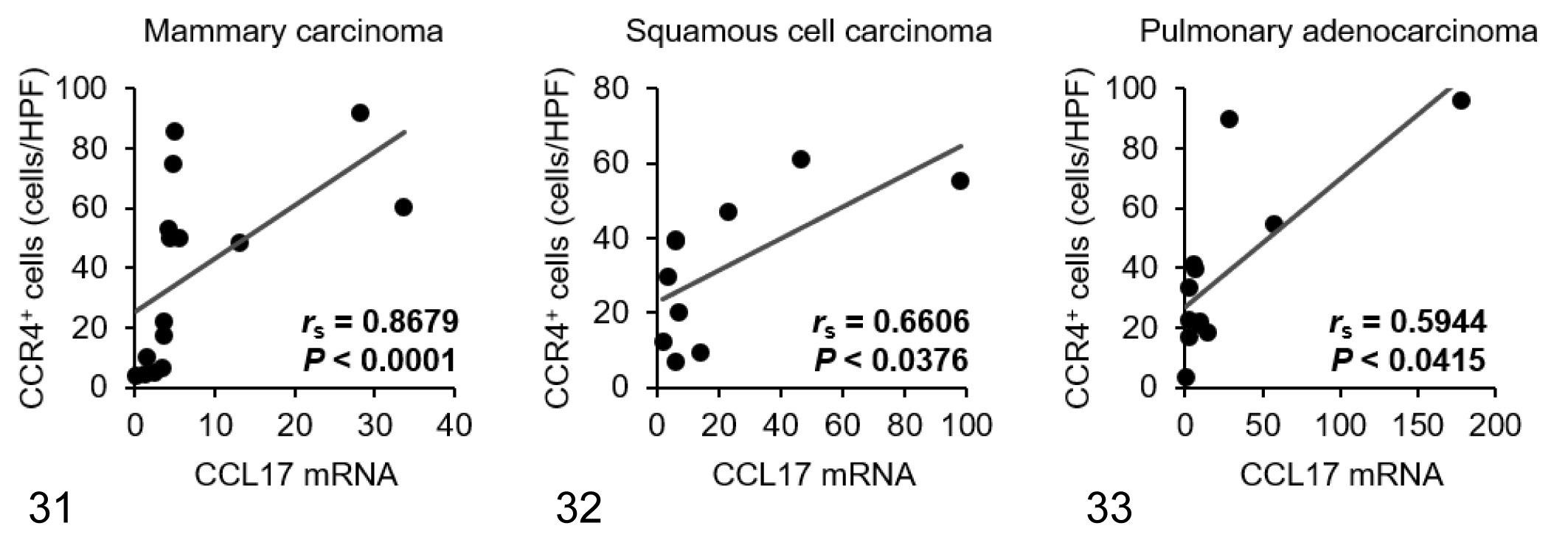
Within samples of tumor tissue, mRNA expression of CCL17 in mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma was increased by approximately 8-, 21-, and 26-fold compared with normal control tissues, respectively (Fig. 26). There was no significant difference in CCL22 mRNA expression between normal and tumor tissues in each carcinoma type (Fig. 27). Tumoral CCL17 mRNA was positively correlated to the number of Foxp3+ Tregs in dogs with mammary carcinoma (Fig. 28), but not in dogs with squamous cell carcinoma (Fig. 29) or pulmonary adenocarcinoma (Fig. 30). Tumoral CCL17 mRNA was positively correlated to the number of CCR4+ cells in dogs with mammary carcinoma (Fig. 31), squamous cell carcinoma (Fig. 32), and pulmonary adenocarcinoma (Fig. 33). However, there was no association between CCL17 and tumor grade (data not shown). Tumoral CCL22 mRNA expression did not correlate to the number of Foxp3+ Tregs and CCR4+ cells (data not shown).
Discussion
In this study, we observed an increased number of Foxp3+ Tregs expressing CCR4 in canine mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma. Furthermore, high mRNA expression of CCL17, but not CCL22, was evident in these carcinomas, suggesting that Tregs infiltrate the tumor tissue through the CCL17/CCR4 axis in a variety of canine carcinomas, including urothelial carcinoma. 19 Previous studies have shown that high tumor-infiltrating Tregs are associated with poor prognosis in dogs with these carcinomas, 2,28 and our recent investigation has demonstrated that Treg-targeting therapy, using anti-CCR4 antibodies, prolongs survival with low incidence of clinically relevant toxicities in dogs with urothelial carcinoma. 19 Thus, blocking CCR4 or tumor-producing CCL17 may be a rational approach that could be clinically tested as a potential treatment option for dogs with carcinomas.
Although numbers of tumor-infiltrating Tregs were increased in canine mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma, the degree of Treg infiltration varied from case to case. Likewise, the mRNA expression of the Treg-attracting chemokine CCL17 was also different between cell lines and clinical cases regardless of tumor grade. Tumoral CCL17 mRNA expression was positively correlated to tumor-infiltrating CCR4+ cells. Although not all CCR4+ cells were Foxp3+ Tregs, the very low number of Foxp3+CCR4− cells suggested that most Tregs express CCR4 in the tumor tissues. These observations suggest that the degree of Treg infiltration might be influenced by the amount of tumor-producing CCL17.
The underlying mechanisms of differences in CCL17 expression remain uncertain. In human and canine keratinocytes, CCL17 production was induced by tumor necrosis factor-α (TNF-α) and interferon-γ (IFN-γ) through nuclear factor-κB (NF-κB) and p38 mitogen-activated protein kinase (MAPK). 7,15,32,40 Furthermore, serum levels of TNF-α and IFN-γ were increased in dogs with mammary carcinoma compared to healthy female dogs, 18 indicating that TNF-α and/or IFN-γ in the tumor-microenvironment might regulate CCL17 production. Additional studies will be necessary to investigate the mechanisms of upregulated CCL17 expression in canine carcinomas.
Among the types of lymphocytes, CCR4 is preferentially expressed on Tregs and Th2 cells. 43 In this study, immunofluorescence showed that a large number of Foxp3−CCR4+ cells, as well as Foxp3+CCR4+ Tregs, were observed in tumor tissues. Previous studies has shown that peripheral CCR4+ lymphocytes exhibit a Th2 phenotype in dogs. 11,20 Therefore, CCR4+ non-Treg cells observed in tumor tissues might be Th2 cells. Th2 cells produce IL-4, IL-5, and IL-13 and are involved in humoral immunity and allergic diseases. 43 In the tumor-microenvironment, the role of Th2 cells is controversial. 5 Several studies have demonstrated the association of tumor-infiltrating Th2 cells and Th2 cytokines with poor prognosis in human patients with ovarian, 17 gastric, 37 and pancreatic cancers. 4 However, other groups have reported that tumor-infiltrating Th2 cells are associated with favorable outcome in patients with breast cancer 42 and Hodgkin’s lymphoma. 31 Thus, it remains unknown whether Th2 cells exacerbate or protect neoplastic diseases. Further study is needed to characterize the phenotype and role of CCR4+ non-Treg cells in canine tumor tissues.
For precision medicine, biomarkers predicting Treg infiltration before anti-Treg treatment in each case are needed. In dogs with urothelial carcinoma, assessment of urinary CCL17, but not serum CCL17, concentrations can predict the response to CCR4 blockade therapy, 19 which is likely due to the direct contact between tumor and urine. Unlike urothelial carcinoma, CCL17 derived from other tumors including mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma could not be detected in urine (data not shown); therefore, it is necessary to examine other detection methods for predicting Treg infiltration. A possible biomarker predicting anti-Treg therapy is peripheral blood Tregs. Our previous study showed that a reduction in the proportion of circulating CCR4+Foxp3+ Tregs correlates to the therapeutic response to anti-CCR4 treatment. 19
One limitation of this study is that healthy control dogs were not well matched to the clinical cases by breed. As it was impossible to obtain normal tissues from various breeds matched to the carcinoma cases, all healthy control dogs were of the same breed (beagle). This limitation could lead to erroneous results regarding the expression of some genes that might differ from breed to breed.
In conclusion, we detected increased numbers of CCR4-expressing Tregs in various canine carcinoma tissues. Furthermore, we found that expression of CCL17, the ligand of CCR4, was elevated in these carcinomas in vitro and in vivo. These results suggest that the CCL17/CCR4 pathway may drive Treg recruitment, contributing to the immune-suppressive nature of the tumor-microenvironment in canine carcinomas. The present study paves the way for clinical trials of CCR4 blockade therapy for dogs with mammary carcinoma, squamous cell carcinoma, and pulmonary adenocarcinoma.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the JSPS KAKENHI Grant-in-Aid for Young Scientists (A) (Grant No. JP16H06208), the Grant-in-Aid for Scientific Research (A) (Grant No. JP19H00968), and the Anicom Capital Research Grant (EVOLVE).
