Abstract
Five chimney swift fledglings died following a progressive loss of appetite and condition while being cared for by an experienced wildlife rehabilitator. All animals had severe necrotizing and heterophilic ventriculitis, with myriad epithelial cells characterized by karyomegaly with intranuclear inclusion bodies. Transmission electron microscopy showed distention of epithelial cell nuclei and chromatin peripheralization by nonenveloped, icosahedral, 75- to 85-nm-diameter virions. Degenerate nested PCR for a highly conserved region of the adenovirus DNA polymerase gene was positive. BLAST analysis of the amplicon sequence indicated the presence of a novel adenovirus, with 74% homology to Antarctic penguin adenoviruses and 72% homology to a bat adenovirus, at low query coverages of only 65% and 63%, respectively. BLAST analysis of the predicted amino acid sequence generated the highest scores for squamate adenoviruses at 100% query coverage. Based on phylogenetic analysis of the partial amino acid sequence of the DNA polymerase, the chimney swift virus was a novel adenovirus most closely related to the Atadenovirus genus. Using a probe based on the novel viral sequence, DNA in situ hybridization identified viral nucleic acid in the nucleus. While the tentatively named chimney swift adenovirus-1 (CsAdV-1) is so far classified with the Atadenoviruses, it is relatively divergent from other members of that genus and may represent the first identified member of a new genus of Adenoviruses.
Keywords
Chimney swifts are members of the Apodidae family that breed in the eastern United States and Canada and winter in South America. As with other members of this family, chimney swifts are unable to perch and must cling from vertically oriented surfaces, such as chimneys; they also lack ingluvia and ceca. 33 According to mitochondrial genome analysis, chimney swifts are most closely related to the common swift (Apus apus). 40 Swifts are insectivores that hunt on the wing and are adapted to feeding at high elevations, all of which lead to difficulty in captive husbandry and thus limited information on diseases. 33 Trauma and malnourishment are common causes of mortality in swifts. There is a report of disseminated Erysipelothrix rhusiopathiae in little swifts (Apus affinis), and chimney swifts have been reported to be sensitive to toxic fumes. 33,36 As with many wild animals, parasitism is common in chimney swifts, including infections with members of the genera Hippoboscidae (louse flies), Ixodidae (ticks), and Hepatozoon (hematozoa). 4,35,37 To our knowledge, there have been no prior descriptions of any viral infections in any members of the family Apodidae.
Adenoviruses are ubiquitous, nonenveloped, double-stranded DNA viruses with icosahedral capsids. The viruses are distributed among 5 genera that affect all vertebrate classes. Birds are infected by members of the genera Aviadenovirus, Atadenovirus, and Siadenovirus. 11,23 The most widely investigated among the disease-causing adenoviruses in birds are fowl adenoviruses, which cause inclusion body hepatitis, respiratory disease, and ventriculitis, with the latter occurring in broiler chickens infected with fowl adenovirus-1. 13 –15,20,26 While there are recent reports of novel adenoviruses in numerous species of wildlife including birds, such as chin-strap penguins (Pygoscelis antarcticus), skua (Catharacta maccormicki), and gulls (Larus argentatus and Larus fuscus), there have been no prior reports of viral infections, including adenovirus infections, in chimney swifts. 6,23,28 In this report, we describe a novel adenovirus associated with intranuclear inclusions and lesions of necrotizing and heterophilic ventriculitis in 5 wild chimney swifts (Chaetura pelagica) from Maine, USA.
Materials and Methods
Animals and Histopathology
Animals were nestlings and were submitted to the same wildlife rehabilitation facility between 2014 and 2016. These animals died naturally or were humanely euthanized. Rehabilitation facility staff made incisions through the coelomic wall and observed and partially dissected organs in situ, and the entire bodies were then immersed in 10% neutral buffered formalin.
Formalin-fixed whole birds were submitted to the New Hampshire Veterinary Diagnostic Laboratory (NHVDL) for histopathologic processing and evaluation. Tissues were routinely processed, embedded in paraffin, sectioned at 5-µm thickness, mounted on charged slides, and stained with hematoxylin and eosin.
Transmission Electron Microscopy
Ventriculus was removed from paraffin-embedded blocks and deparaffinized. Samples were fixed in a mixture of 2.5% glutaraldehyde/2.5% paraformaldehyde in 0.1 M cacodylate buffer at 4°C for 24 hours, then fixed in 1% osmium tetroxide, and subsequently dehydrated in a graded acetone series. Samples were infiltrated and embedded in Poly/Bed 812 resin (Polysciences, Inc. Warrington, PA, USA). Thin sections (70-nm thickness) were cut with a PTXL ultramicrotome (RMC, Boeckeler Instruments, Tucson, AZ, USA), placed on 200 mesh copper grids, stained with uranyl acetate and lead citrate, and imaged using a JEOL 100CX Transmission Electron Microscope (JEOL, Tokyo, Japan) at a 100 kV accelerating voltage, as previously described. 41
PCR and Sequencing
DNA was extracted from formalin-fixed paraffin-embedded (FFPE) ventriculus tissue sections (QIAamp DNA FFPE Tissue Kit, Qiagen, Valencia, CA). A previously described consensus nested PCR method for the detection of a partial fragment, ∼320 base pairs (bp), of the DNA polymerase gene of adenoviruses was performed, with a slight modification of the reagent mix. 38 Briefly, for the first round amplification, the 50 µl PCR reaction mixture contained Qiagen Taq PCR Master Mix reagent (Qiagen), 2.5 µl dimethyl sulfoxide, forward outer primer 5’- TNMGNGGNGGNMGNTGYTAYCC-3’ and reverse outer primer 5’-GTDGCRAANSHNCCRTABARNGMRTT-3’ (each at 1 µM final primer concentration), and 5 µl of extracted template DNA. For the second round of PCR, the 50-µl reaction mixture contained similar buffer and enzyme composition, with the forward inner primer 5’- GTNTWYGAYATHTGYGGHATGTAYGC-3’ and reverse inner primer 5’-CCANCCBCDRTTRTGNARNGTRA-3’, each at 1 µM final primer concentration; 5 µl of the PCR product from the first round PCR was used as template. The following cycling parameters were used for each round of PCR: denaturation at 95°C for 5 minutes, then 45 cycles of 94°C for 30 seconds, 46°C for 60 seconds, and 72°C for 60 seconds, ending with a final extension step of 72°C for 5 minutes.
The PCR products were analyzed by agarose gel electrophoresis, stained with ethidium bromide, and visualized by UV-transillumination. Products of the expected size were cut and purified using the QIAquick Gel Extraction Kit (Qiagen). Pooled purified amplicons from duplicate secondary PCR reactions were submitted to the Research Technology Support Facility Genomics Core at Michigan State University for bidirectional Sanger sequencing.
Sequence assembly and analyses, including multiple sequence alignment, were performed using the Lasergene 12 Core Suite (DNASTAR, Inc., Madison, WI, USA). Nucleotide and protein BLAST searches against the GenBank database were performed (https://blast.ncbi.nlm.nih.gov/Blast.cgi). 1
Phylogenetic analysis of the aligned amino acid sequences was performed with the software MEGA6, using the neighbor-joining (NJ) and the maximum likelihood methods, both with bootstrapping at 500 replicates, and computation of the evolutionary distances by the Poisson correction method for the NJ tree. 10,32,34,39,42
In Situ Hybridization
DNA in situ hybridization (ISH) was performed using routine protocols for detection of psittacine beak and feather disease virus and avian polyomavirus nucleic acids, which were modified to detect DNA from a psittacine or budgerigar variant adenovirus, using PCR-generated dsDNA probes (GeneArt Strings DNA Fragment, Life Technologies Corporation, Carlsbad, CA, USA: PsittAdV probe homologous to GenBank EF442329.1; BudgieAdV probe homologous to GenBank AB485763) labeled with multiple digoxigenin moieties (DIG-PCR) per manufacturer’s instructions (Roche Diagnostics Corporation, Indianapolis, IN, USA). 22,30 Similarly, a dsDNA probe (SwiftAdV probe) was later designed and manufactured (GeneArt) to target the novel chimney swift adenovirus sequence, resulting in a 226 bp oligonucleotide encompassing bases 3 to 228, as was a primer pair (Sigma-Aldrich, St. Louis, MO, USA). The primers corresponded to each end of the manufactured oligonucleotide: Swiftadenoforward, 5’-CAGCTTTGACTCACCCAATG-3’ and Swiftadenoreverse, 5’- GTCCAACATAAACGCCCTCC-3’.
The ISH protocol utilized the dsDNA DIG-PCR probes, as previously described, using a manual capillary-action workstation (Fisher Scientific Co, Suwanee, GA, USA), and 3-aminopropyltrieth-oxylane-coated slides (ProbeOn Plus, Fisher Scientific Co). 16 For slides tested with the SwiftAdV probe, a 105°C incubation was followed by incubation overnight at 37°C only. To further increase nonspecific binding to adenoviruses with base differences, slides incubated with PsittAdV or BudgieAdV probes at both temperatures were washed 3 times in a low-stringency solution of 2.0X SSC, 4% Tween 20, and 2.5% Brij 35 to remove nonhybridized probe. Nonhybridized SwiftAdV probe was removed from slides by 2 graded concentrations of SSC solution to minimize nonspecific hybridization: (1) 3 washes in 2.0X SSC, 4% Tween 20, 2.5% Brij 35 and (2) 3 high-stringency washes in 0.5X SSC, 4% Tween 20, 2.5% Brij 35. Slides were labeled with anti-digoxigenin antibody conjugated to alkaline phosphatase enzyme (Roche Diagnostics Corporation) and counterstained with 0.5% fast green, dehydrated, coverslipped, and examined by light microscopy for deposition of blue-black chromagen, indicating probe/viral DNA hybridization.
Results
History, Clinical Signs, and Gross Necropsy
Animals were part of annually received clutches of chimney swifts submitted for rehabilitation. Broods of swifts are routinely received. Between July 20 and August 15, 2015, 36 chimney swifts were received at the clinic. Of these, 10 were released (27.8%), 12 died naturally (33.3%), and 14 were euthanized (38.9%). According to the wildlife rehabilitator, this rate of mortality and euthanasia was uncommonly high. Most of the birds that survived and were released came from clutches received earlier in the season. Five of the 26 dead birds were selected by the rehabilitator and submitted for histopathologic examination.
The birds included in this report were submitted to the rehabilitator as apparently abandoned nestlings; no information regarding the health of the adults or reason for abandonment is known. Clinical signs began 1 to 3 weeks after admission to the facility and often began as difficulty swallowing that progressed to thick mucoid saliva and sometimes vomit. Clinical signs were generally progressive and lasted 1 to 4 days. No animals were noted to have clinical signs on admission. Two of the birds were from the same clutch, while the rest were all from different clutches. The birds from the same clutch (birds Nos. 1 and 2) were observed to be vomiting or regurgitating before death. Vomiting was also noted in 2 other birds that were not among the subset submitted for histopathology. Bird No. 3 had difficulty swallowing. Bird No. 4 was older than chimney swifts typically treated by the rehabilitator; this bird was thriving until it showed sudden onset of mucoid saliva and hyporexia followed by weight loss and unexpected death. Finally, Bird No. 5 was the smallest of a clutch and failed to thrive over a period of 2 weeks; this bird was initially a vigorous feeder but did not gain weight and eventually became lethargic and hyporectic.
Of the 26 animals that died or were euthanized, the wildlife rehabilitator noted few lesions in the opened coelom and organs examined in situ. The most common finding was luminal blood in the gastrointestinal tract and mucus in the sinuses, though none of these lesions were noted specifically in the subset of cases submitted for histopathology. A clutch-mate of bird No. 3 perished, and the necropsy performed by the rehabilitator demonstrated blood in the intestinal lumen and pale lungs. This bird’s tissues were not submitted for histopathology. Additionally, 4 fledglings from the previous year (2014) and 3 from the subsequent year (2016) from the same clinic were submitted for histopathology.
Histopathology
The 5 animals from 2015 had segmentally severe, erosive to ulcerative heterophilic ventriculitis that was most notable at the aboral portion of the organ (Fig. 1). The koilin layer overlying and adjacent to this lesion was disrupted and frayed. There was partial to full thickness necrosis of the ventricular epithelium and underlying lamina propria, with individualized, fragmented, often hypereosinophilic cells mixed with amorphous granular eosinophilic necrotic debris (Fig. 2). This necrotic pseudomembrane and lamina propria contained a moderate to marked infiltrate of variably degenerated heterophils and some free erythrocytes (Figs. 2, 3). There was often aggregated necrotic debris and degenerated heterophils in the lumen overlying severe mucosal lesions (Figs. 1, 2). In numerous consecutive epithelial cells, the nuclei were expanded 2 to 3 times the normal diameter by a large (15–30 µm), slightly refractile eosinophilic to basophilic inclusion body, causing peripheralization of the chromatin (Fig. 3).
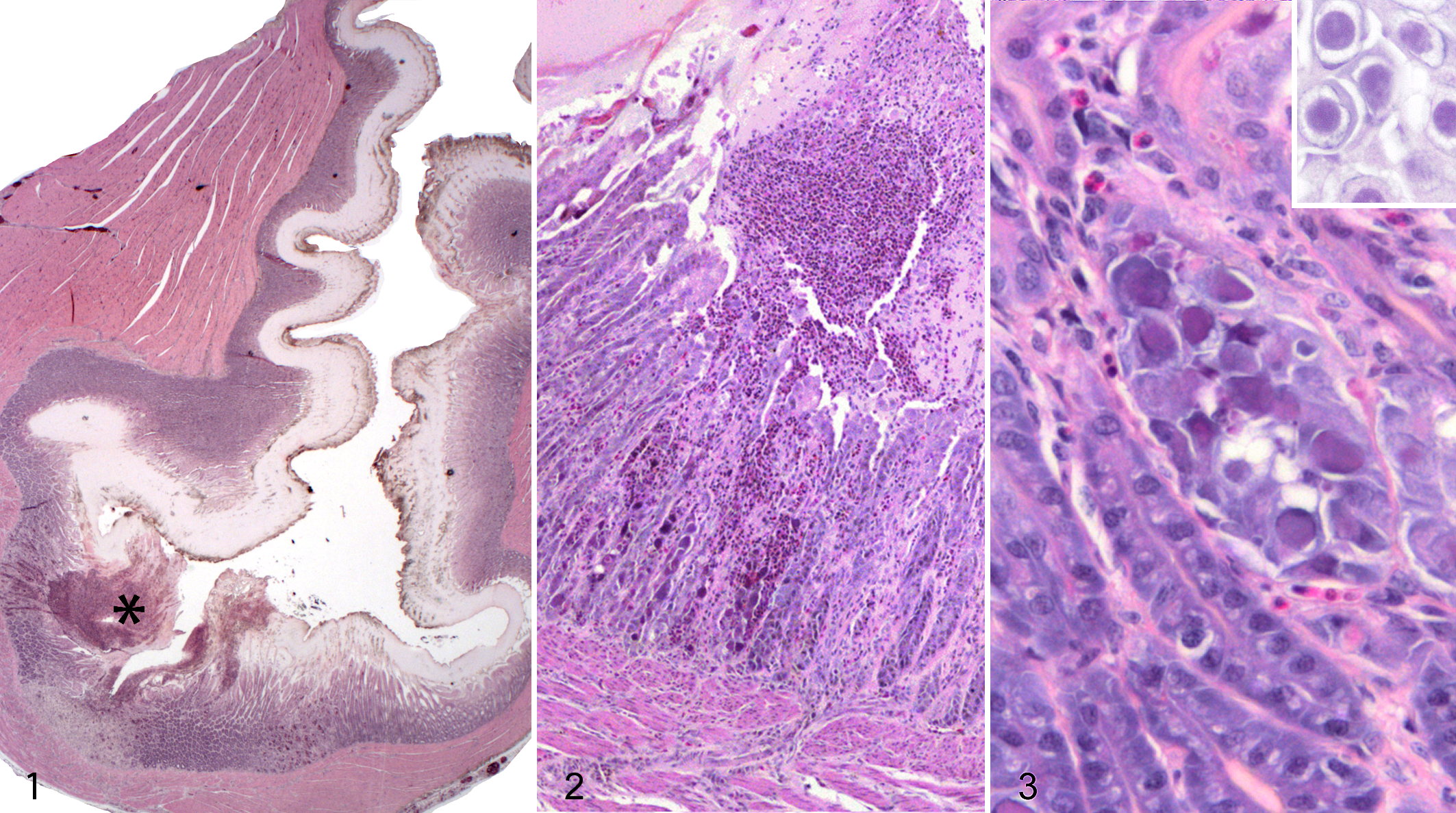
The only other lesion was necrotizing, heterophilic dermatitis with intralesional bacterial bacilli affecting the skin folds at the commissures of the mouths of 2 animals (bird Nos. 2 and 4).
The 4 animals from 2014 had degeneration of the epithelial cells of the ventriculus, with no intranuclear viral inclusions and minimal associated inflammation. The 3 animals from 2016 did not have lesions in the ventriculus. Animals from both of these years had variable degrees of dermatitis of the commissures of the mouth, as described previously.
Transmission Electron Microscopy
The nuclei of the epithelial cells were markedly expanded by 75- to 85-nm-diameter, nonenveloped, icosahedral virions, often arranged in a paracrystalline array (Fig. 4). The virions had variably electron-dense cores of genetic material, often surrounded by a halo of electron-lucent material and then a peripheral capsid (Fig. 5). The chromatin was peripheralized and compressed against the nuclear envelope. Nuclei contained large, irregular, and occasionally multiple, moderately electron-dense nucleoli. In some cells, which were often degenerated or necrotic, there were fewer virions that were scattered within and displaced by a large amount of granular, moderately electron-dense amorphous material.
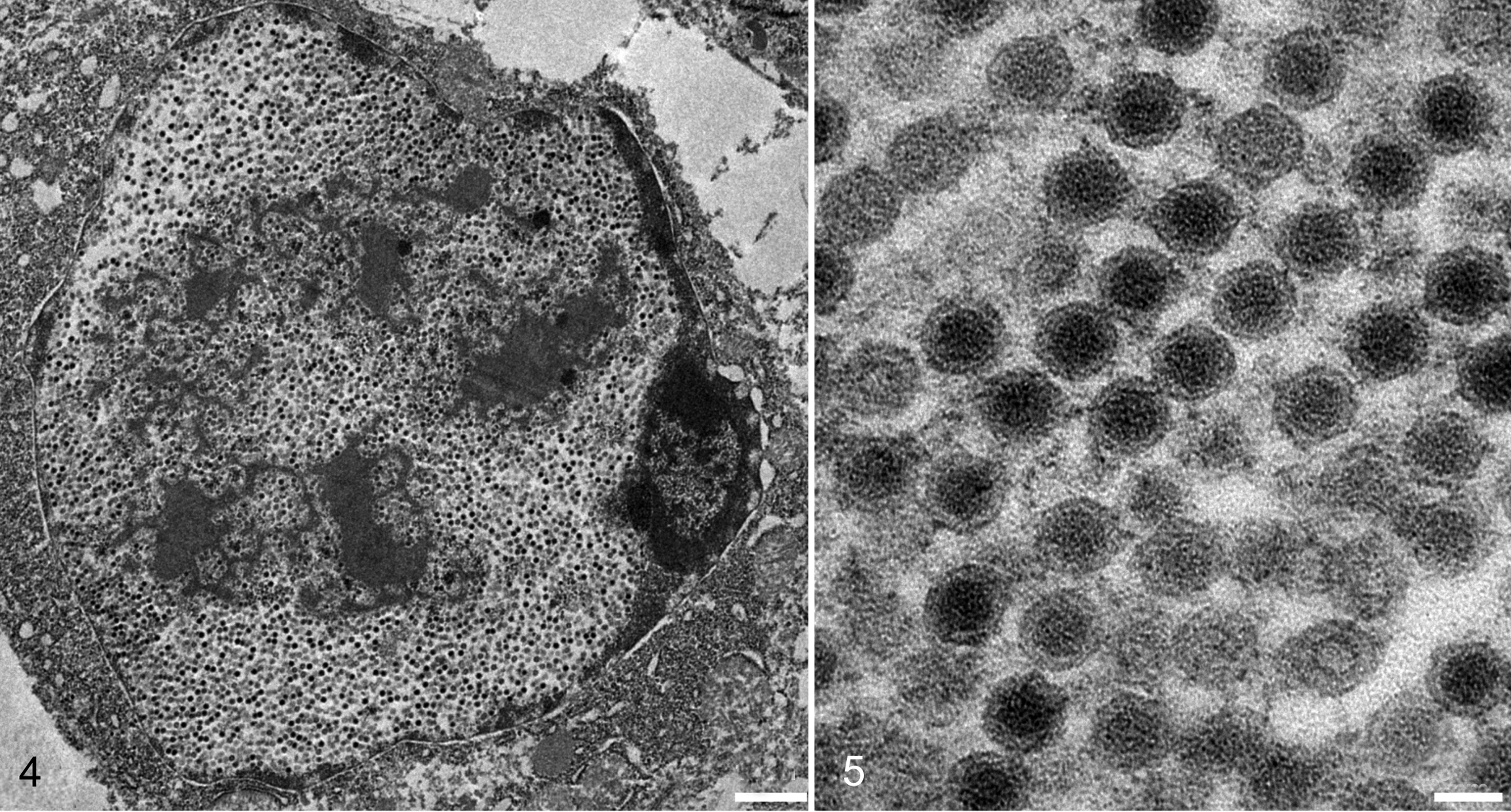
Adenoviral ventriculitis, ventriculus, chimney swift, case 2. Transmission electron micrograph, bar = 1 µm. The nucleus is expanded by a dense paracrystalline array of virions that cause margination of chromatin. There is a moderately electron-dense material within and disrupting the array.
PCR and Sequencing
A nested PCR for a highly conserved region of the adenovirus DNA polymerase gene was performed on the DNA extracted from the pooled 2015 chimney swift tissue samples, yielding an amplicon of the expected target size of ∼320 bp.
BLAST analysis of the nucleotide sequence (272 bp, excluding primer sequences) obtained from the generic adenovirus PCR product revealed a 74% sequence similarity (at 65% query coverage) to the DNA polymerase gene of adenoviruses from gentoo and chinstrap penguins in Antarctica. 24 The second highest scoring match was to a bat adenovirus, with 72% similarity at 62% query coverage. BLAST analysis of the predicted amino acid sequence (90 residues) of the partial DNA polymerase sequence generated the highest scores (at 100% query coverage) with Gekkonid adenovirus 1 and Amphisbaenian adenovirus 1, at 63% and 61% sequence identities, respectively. We designated this novel adenovirus chimney swift adenovirus-1 (CsAdV-1). The partial DNA polymerase gene sequence of CsAdV-1 was submitted to GenBank and assigned the accession number, MG736957.
Consensus PCR analysis of ventriculus DNA extracted from each of the four 2014 and the three 2016 CS samples did not identify CsAdV-1 DNA.
Phylogenetic Analysis
The predicted partial DNA polymerase amino acid sequence of CsAdV-1 was aligned with corresponding sequences of representative mastadenoviruses, siadenoviruses, aviadenoviruses, atadenoviruses, and box turtle adenovirus (a member of a newly proposed genus, Testadenovirus). 25 Phylogenetic analyses based on multiple amino acid sequence alignment showed that CsAdV-1 appeared to be most closely related to members of the Atadenovirus genus by both neighbor-joining and maximum likelihood methods (Figs. 6, 7). A notable difference observed by using the maximum likelihood method was that CsAdV-1 grouped with Gekkonid AdV-1 (Fig. 7). The trees also showed the grouping of the other adenoviruses into each of their respective genus as previously classified. A slight topology change was observed within the maximum likelihood tree for a subset of the siadenoviruses, forming a separate clade with the aviadenoviruses, but was supported only by a low bootstrap value.
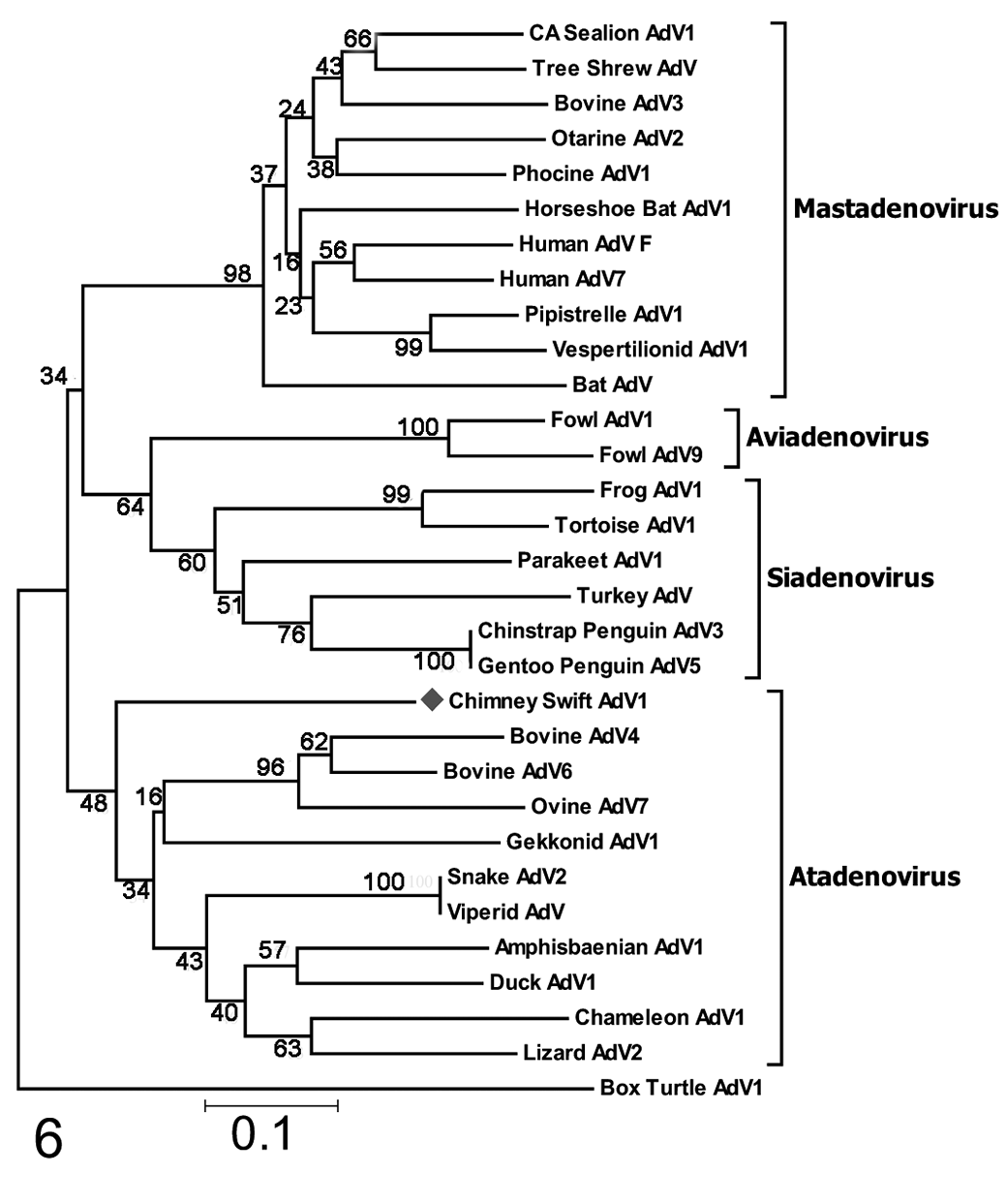
Neighbor-joining tree, based on deduced partial amino acid sequences of the DNA-dependent DNA polymerase proteins of chimney swift adenovirus-1 (CsAdv-1, this study). The CsAdV-1 sequence clusters closest to those of the atadenoviruses but has low relative homology to the most closely related atadenovirus (bovine AdV4); therefore, its placement in the genus is tentative. Bootstrap values are indicated at the nodes. GenBank accession numbers are in the table available in the online Supplemental Material.
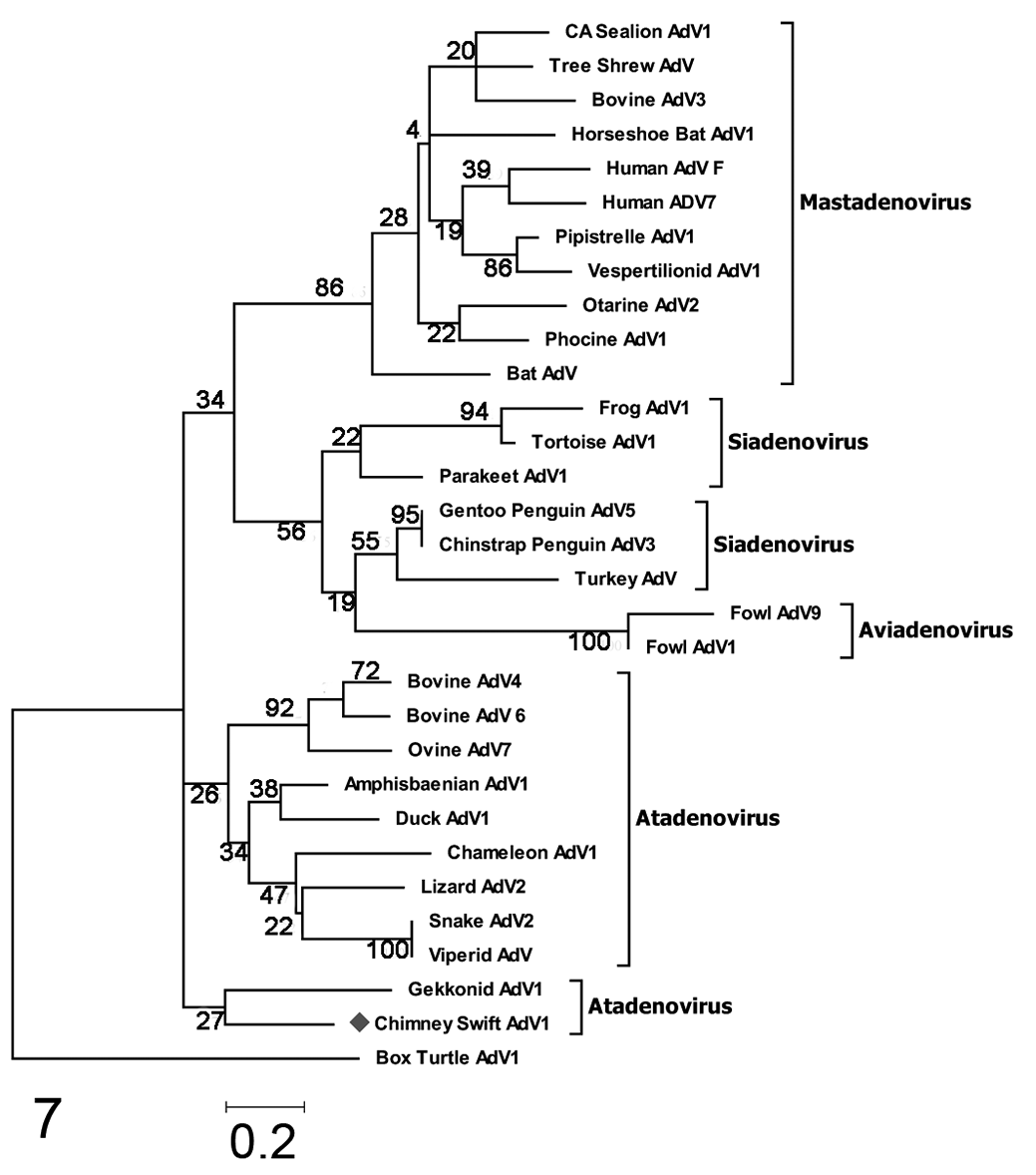
Maximum likelihood tree based on deduced partial amino acid sequences of the DNA-dependent DNA polymerase proteins of chimney swift adenovirus-1 (CSAdV-1, this study). As in the neighbor-joining tree, the closest homology is to an atadenovirus (Gekkonid AdV-1), and there is relatively low homology between CsAdV-1 and gekkonid AdV-1. Bootstrap values are indicated at the nodes. GenBank accession numbers are in table available in the online Supplemental Material.
In Situ Hybridization
Initial in situ hybridization using the PsittAdV and BudgieAdV probes showed no labeling of chimney swift tissues. ISH using the SwiftAdV probe, derived from the unique CsAdV-1 polymerase gene sequence, resulted in distinct labeling of karyomegalic nuclei containing inclusions (Fig. 8). There was no off-target labeling in the chimney swift tissues, and the probe did not detect viral DNA in control tissues containing either psittacine or budgerigar adenoviruses.
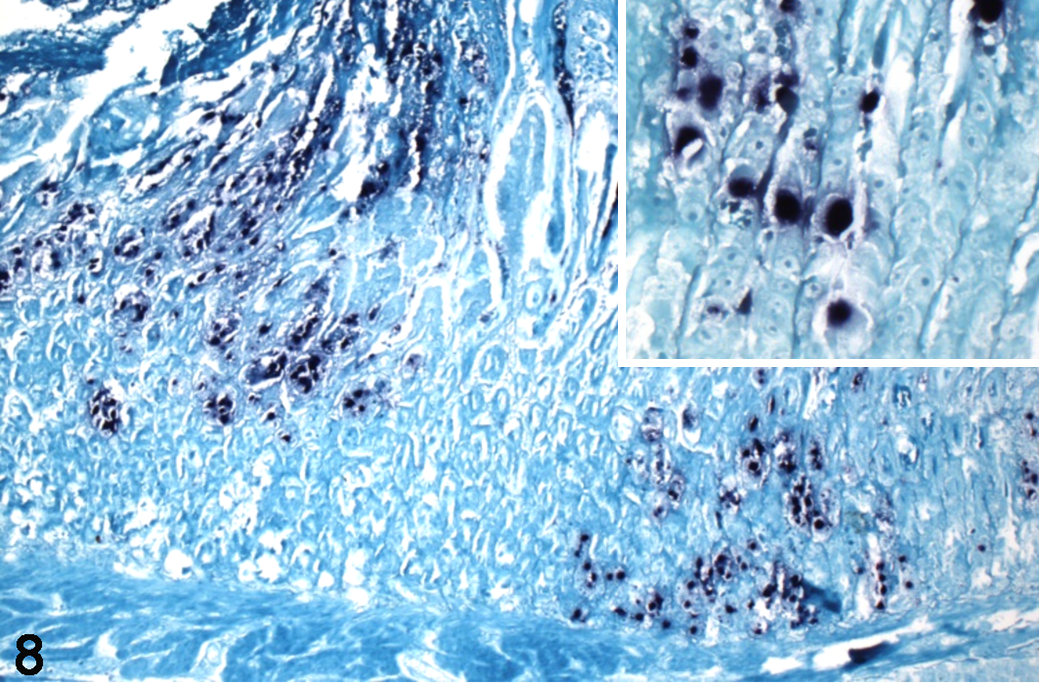
Adenoviral ventriculitis, ventriculus, chimney swift, case 3. In situ hybridization with custom probes for novel chimney swift adenovirus-1 (CsAdV-1) DNA shows positive labeling (dark blue) of adenoviral inclusions in affected nuclei. Inset: Higher magnification of intranuclear labeling. Nitroblue tetrazolium chromagen and fast green counterstain.
Discussion
The lesions in the affected swifts contained typical adenoviral intranuclear inclusion bodies, and the necrotizing and ulcerative lesions in the ventriculus parallel those described in broiler chickens with fowl adenovirus-1 infection. 14 Examination of the karyomegalic nuclei with inclusions via transmission electron microscopy revealed distended nuclei with attenuated chromatin, peripheralized by icosahedral virions arranged in a paracrystalline array. These characteristics and the virion size are typical of adenoviruses. 25 The specific ISH labeling of the viral inclusions in affected tissues with the Swift probe further substantiates that the inclusions belong to the novel adenovirus that was the only detectable intralesional organism. Specificity for CsAdV-1 was supported by a lack of labeling by SwiftAdV probe of control tissues containing either psittacine or budgerigar adenoviruses and lack of labeling of the chimney swift inclusions with the psittacine adenovirus probe.
There have been multiple adenoviruses recently described in birds. Some of these, like the adenovirus from chinstrap penguins, are not associated with clinical illness or disease, while others are lesion-associated viruses like CsAdV-1. The virus we identified caused stereotypical intranuclear inclusions, similar to those noted in necrotizing cloacitis in herring gulls (Larus argentatus) and lesser black-backed gulls (Larus fuscus) and the virus that causes enterotyphlocolitis and hepatitis in long-tailed ducks (Clangula hyemalis) in Alaska. 6,8,17,23,24
Phylogenetic analyses based on the partial amino acid sequence of the DNA polymerase of CsAdV-1 was not definitive due to the limited sequence data obtained. Nevertheless, both trees generated showed that this novel adenovirus is most closely related to previously described Atadenoviruses. The DNA polymerase gene region amplified and sequenced in this study has been used previously by investigators to discover novel adenoviruses and phylogenetically determine their genus assignment. 38 Phylogenetic calculations based on distance matrix analysis of the DNA polymerase amino acid sequence is currently 1 of the criteria used for Atadenovirus species demarcation. 18 The Atadenovirus genus was described in 1998 and named due to its high A/T content genome. 5,7 Consistent with this description, the 272 bp partial DNA polymerase sequence obtained for CsAdV-1 has a >60% A/T content. Other genetic characteristics of the genus include a distinct fiber gene sequence and gene product structure, a unique hexon gene sequence, and the presence of a p32 K gene that is potentially of bacterial origin. 9,27,31 With the limited sequence data obtained so far for CsAdV-1, conclusive classification of its membership in the Atadenovirus genus would require further genomic sequencing and characterization. Furthermore, the bootstrap values associated with this clustering are low (<50), precluding any strong conclusions regarding evolutionary events. Our attempts to amplify a region of the hexon gene and another area of the DNA polymerase gene using published degenerate primers were unsuccessful. Based on the marked divergence of the fragment we were able to amplify, it is possible that CsAdV-1 may represent the first identified member of a novel genus of adenovirus.
Adenoviruses, which are nearly ubiquitous, show a high degree of host specificity and are thus fairly genetically divergent from each other—which is likely why the initial psittacine adenovirus in situ hybridization was negative in our cases and why the CsAdV-1-specific probe did not label the control tissues positive for psittacine and budgerigar adenoviruses. This highlights the advantage of the degenerate adenovirus DNA polymerase PCR for detection and sequence analysis for identification over the use of the more specific adenovirus DNA ISH in cases of suspected novel adenoviral infections.
In general, adenoviruses have been thought to be host-specific, so species-jumping events were viewed as outliers and events likely to play a role in evolution of the family. 25 There is evidence that adenoviruses may in fact be less host-specific than prevailing dogma indicates, with 1 study identifying 4 atadenoviruses infecting multiple lizard species. 2 CsAdV-1 may either represent a previously unrecognized, host-adapted strain associated with disease and mortality only in stressed chimney swift nestlings or a species-jumping event of a previously undescribed adenovirus from an unknown host. While Mastadenoviruses, Aviadenoviruses, and the single Ichtadenovirus are specific to mammals, birds, and fish, respectively, the evolution of Atadenoviruses and Siadenoviruses appears to include multiple significant host-jumping evolutionary events that have led to spread of these viruses from the purportedly initial squamate and amphibian host, respectively. 19 This information may indicate that a species-jumping event leading to CsAdV-1 evolution could be more likely; however, this is tenuous based on the limited certainty in CsAdV-1 phyolgeny, as discussed previously.
Regardless of evolutionary origin, infection by non-host-specific adenoviruses tend to have greater pathogenicity than adenoviruses that coevolved in specific hosts. 21 Alternatively, the genetics of the novel host population may confer poor immunologic control or tolerance of the infection. It has been suggested that canine adenoviruses evolved from bat adenovirus-2, and as novel viruses, they possessed varied or novel virulence-associated genes, allowing them to infect multiple species and cause significant disease. 3,12,21,29
The recognition of CsAdV-1, a novel virus with potential pathogenicity, leads to the larger questions of the origin of this virus and how these infections occurred. Although transmission during captivity cannot be completely ruled out, the rehabilitation facility kept each brood in isolation without a shared direct air supply. Cages were disinfected between occupants and implements, and feed and bedding were not shared between cages. Gloves were changed between working with each cage.
The cases we present show that adenovirus infection should be a differential diagnosis for gastrointestinal disease in chimney swifts, especially in young birds, and those with hemorrhagic lesions in the oral segment of the gastrointestinal tract with typical intranuclear inclusion bodies in infected cells. The relative divergence of the sequence we were able to amplify suggests that there may be more genera of Adenovirus then those currently defined.
Supplemental Material
Supplemental Material, DS1_VET_10.1177_0300985819861717 - Necrotizing Ventriculitis in Fledgling Chimney Swifts (Chaetura Pelagica) Associated With a Novel Adenovirus, Chimney Swift Adenovirus-1 (CsAdV-1)
Supplemental Material, DS1_VET_10.1177_0300985819861717 for Necrotizing Ventriculitis in Fledgling Chimney Swifts (Chaetura Pelagica) Associated With a Novel Adenovirus, Chimney Swift Adenovirus-1 (CsAdV-1) by David B. Needle, Annabel G. Wise, Christopher R. Gregory, Roger K. Maes, Inga F. Sidor, Branson W. Ritchie and Dalen Agnew in Veterinary Pathology
Supplemental Material
Supplemental Material, DS2_VET_10.1177_0300985819861717 - Necrotizing Ventriculitis in Fledgling Chimney Swifts (Chaetura Pelagica) Associated With a Novel Adenovirus, Chimney Swift Adenovirus-1 (CsAdV-1)
Supplemental Material, DS2_VET_10.1177_0300985819861717 for Necrotizing Ventriculitis in Fledgling Chimney Swifts (Chaetura Pelagica) Associated With a Novel Adenovirus, Chimney Swift Adenovirus-1 (CsAdV-1) by David B. Needle, Annabel G. Wise, Christopher R. Gregory, Roger K. Maes, Inga F. Sidor, Branson W. Ritchie and Dalen Agnew in Veterinary Pathology
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
