Abstract
There is a growing need to quantitate or “score” lesions in mouse models of human disease, for correlation with human disease and to establish their clinical relevance. Several standard semiquantitative scoring schemes have been adapted for nonneoplastic lesions; similarly, the pathologist must carefully select an approach to score mouse models of cancer. Genetically engineered mouse models with a continuum of precancerous and cancerous lesions and xenogeneic models of various derivations present unique challenges for the pathologist. Important considerations include experimental design, understanding of the human disease being modeled, standardized classification of lesions, and approaches for semiquantitative and/or quantitative scoring in the model being evaluated. Quantification should be considered for measuring the extent of neoplasia and expression of tumor biomarkers. Semiquantitative scoring schemes have been devised that include severity, frequency, and distribution of lesions. Although labor-intensive, scoring mouse models of cancer provides numerical data that enable statistical analysis and greater translational impact.
Keywords
Scoring schemes exist for dozens of histologic lesions in mouse model systems. 25 Histopathologic assessment is a crucial part of the evaluation and validation of mouse models of cancer. Genetically engineered mouse (GEM) models of human cancer are widely used to evaluate the initiation and progression of cancer with the ultimate goals of understanding the human disease and early detection, treatment, and prevention. Scoring mass lesions or metastases as an incidence may be employed for large mass lesions or few foci of metastases; however, mouse models that include many foci of preneoplastic and precancerous progressive lesions and malignant lesions pose a significant challenge. Furthermore, little is documented regarding best practices in scoring tumors with a xenogeneic natural history. When presented with a mouse cancer model, scoring approaches depend heavily upon the research question, the pathologist’s role in the study, what materials are available, and natural limitations of the model and its observer. There are many considerations involved when scoring mouse models of cancer (Table 1).
Important Considerations When Scoring Mouse Models of Cancer.
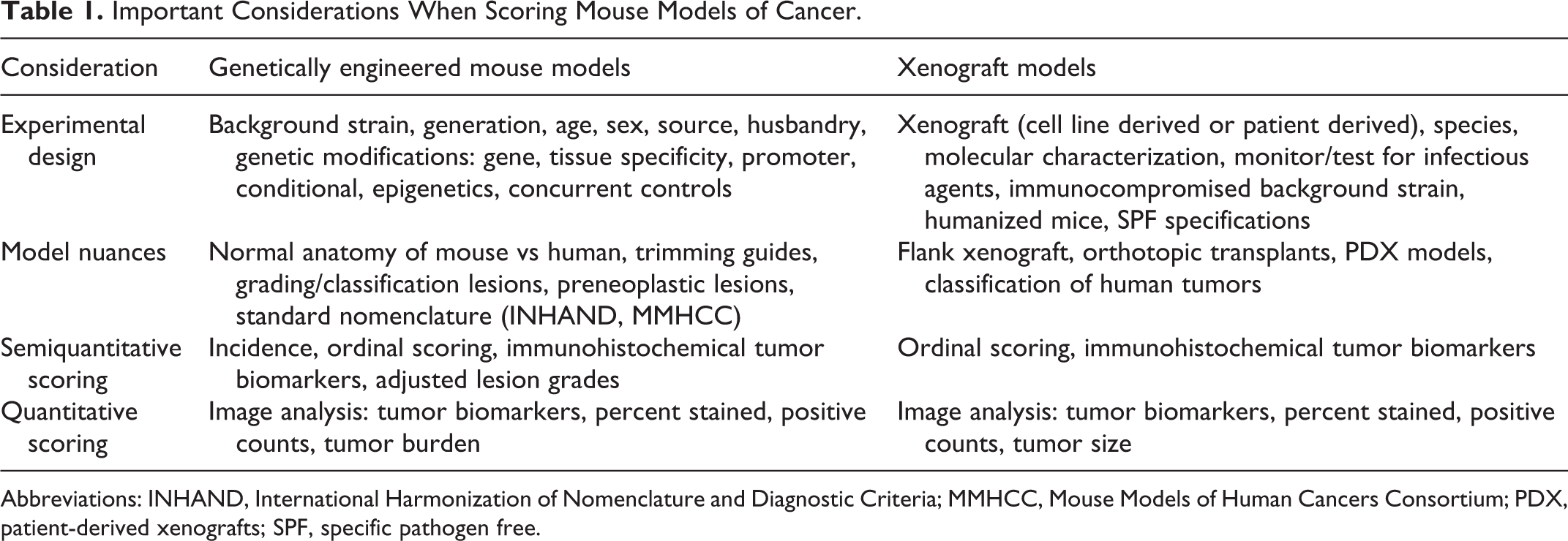

Abbreviations: INHAND, International Harmonization of Nomenclature and Diagnostic Criteria; MMHCC, Mouse Models of Human Cancers Consortium; PDX, patient-derived xenografts; SPF, specific pathogen free.
Experimental Design
Planning a study ahead of time is significant to ensure proper model selection and scoring. It is important to understand the biologic activity of the model for adequate, accurate evaluation. Pathology end points may be determined based on previous findings, published reports, and natural history. Essential model parameters include age, source, background strain and substrain, sex, any genetic manipulations, and genetic quality control measures. Details regarding genetic modifications include the specific gene of interest, if the mouse is transgenic, tissue specificity of the model, type of promoter, and if the model is conditional. 9,46 Husbandry and environmental conditions are significant as changes in environment may alter flora, predispose to opportunistic infections, and ultimately alter tumor kinetics. 9,46 Thus, without detailed methodology regarding study design, models may be misinterpreted. With complete background information, the pathologist can identify potential pathology end points that will ultimately characterize the biologic activity of the model and aid in histologic analysis and scoring.
The number of mice evaluated is a critical consideration when planning for pathology end points. Consulting with a statistician and running a sample size calculation will help determine the number of mice needed for a particular study. In xenograft studies, the number of mice needed can be calculated based on the published or predicted take rate of a given cell line or transplanted tumor. In both xenograft and GEM models, the experimental plan should account for any unexpected deaths secondary to congenital disease, background strain related lesions, tumor progression, and any other unplanned causes of death. 42 For GEM, evaluation of age- and sex-matched concurrent controls of the same genetic background, ideally littermates, is essential to determine if the observed lesion, such as a neoplasm, results from a particular mutation or if it is the result of background pathology of the strain used. 9,46 For example, primary lung tumors are common in certain background strains and must not be mistaken for metastasis. Background strain may have an impact on phenotype and the development of neoplasia. Some strains are more resistant to tumor development, and some strains are predisposed to tumor development. 9,34 Background strain has affected the spectrum of tumor phenotypes in established models. 23 Age is a relevant consideration as the incidence of background neoplasia and other background lesions increases with time. 10 Specific GEM models of cancer may progress and vary with age, where the progression to malignancy may only occur in older animals. 10 In the case of prostate cancer models, time points that mark reproductive maturity should be considered. Thus, an increased number of mice at various ages (serial evaluation by age) may need to be evaluated to fully understand a model, its biologic activity, which is crucial for accurate scoring. Pathologists might consider running a “time course” to assist with phenotype evaluation and/or therapeutic evaluations.
Similarly, in xenograft cancer models, knowledge of the strain is paramount. Recently, NSG (NOD.Cg-Prkdcscid Il2rgtm1Wjl /SzJ), NOG (NOD.Cg-Prkdcscid Il2rgtm1Sug /JicTac), and NRG (NOD.Cg-Rag1tm1Mom Il2rgtm1Wjl /SzJ) mice have largely replaced the athymic nude and severe combined immunodeficiency (SCID) mice of yesteryear, boasting higher, more uniform engraftment rates due to severely tamped-down immune systems. 5,18,26 NSG and NRG mice have a NOD background, conferring reduced dendritic cells (DC) and natural killer (NK) cell function and defective macrophages. They are also deficient in the interleukin (IL)–2 gamma chain, further affecting the ability of T and B cells, NK cells, DCs, and neutrophils to mature and function appropriately. The difference lies in the genetic manipulation affecting V(D)J recombination—Prkdcscid or Rag1—as this most significantly affects the T- and B-cell subsets. Both strains can be humanized through engraftment with human CD34+ hematopoietic stem cells or peripheral blood mononuclear cells (PBMCs). These models can be further built upon by transgenically conferring production of human stem cell factor/KIT ligand, granulocyte-macrophage colony-pstimulating factor (GM-CSF), IL-3, human leukocyte antigen (HLA) I/II, signal regulatory protein–α, and/or thrombopoietin. An example is the NSGS (NOD.Cg-Prkdcscid Il2rgtm1Wjl Tg(CMV-IL3, CSF2, KITLG)1Eav/MloySzJ) model used in leukemia and immuno-oncology studies. 26 With this must come careful consideration of environmental and biosafety factors, as specific pathogen free (SPF) conditions that are “good enough” for nudes and SCIDs are inadequate for newer, severely immunocompromised strains. Infectious agents of human and murine health concern such as Epstein-Barr virus (EBV) and Corynebacterium bovis can be transmitted through patient-derived tumor samples. 29 Furthermore, an EBV-positive solid tumor can develop B-cell lymphoma. 8,42 The specific strain of immunocompromised mouse used in a study will have implications for tumor kinetics and response to therapeutics.
When anticancer agents are used in GEM and xenograft models, the pathologist is tasked with interpreting what effects, if any, an intervention causes on cancer and possibly by what mechanisms. The pathologist should confront such projects with diligent scientific rigor, particularly in the drug discovery setting. For instance, information about metabolism and bioavailability of a drug in the mouse compared to the human counterpart may not be available. Without a full complement of information pertaining to the study, healthy scientific skepticism regarding model validity is warranted.
Genetically Engineered Mice
Classification of Lesions in GEM Models
The biologic potential of the various organ system models of cancer may be assessed via histological and cytological grading schemes. GEM models of epithelial carcinogenesis often exhibit a multistage progression or continuum through preneoplastic/precancerous lesions to malignancy. 12 This multistage progression may be seen in pancreas, prostate, mammary gland, kidney, gastrointestinal tract, lung, liver, and skin. 12 Proliferative epithelial changes are divided into specific grades for each system. The classification of lesions should follow the accepted nomenclature and grading schemes published for mouse models of cancer. An example is for mouse models of pancreatic ductal adenocarcinoma (PDAC). GEM models of PDAC recapitulate the progression of precursor or preneoplastic lesions known as mouse pancreatic intraepithelial neoplasia (mPanIN) to invasive cancer. 16 –19
The National Cancer Institute (NCI) Mouse Models of Human Cancers Consortium (MMHCC) established pathology committees intended for mouse models of human cancer for specific organ systems. 7,13,20,23,32,37,45 The goal was to address questions of relevance to human cancers and to standardize nomenclature and grading schemes. The grading schemes are based on assessment of biologic behavior and adopted human nomenclature with the goal to facilitate consistency in the literature when reporting existing and new mouse models of cancer. Each consensus paper describes the standardized terminology for each grade within a specific system. Many of these consensus papers have been updated over the past several years; therefore, it is recommended to reference the most current consensus terminology when grading and scoring mouse models of cancer. 23,45
Other references for lesion classification include the International Harmonization of Nomenclature and Diagnostic Criteria for Lesions in Rats and Mice (INHAND) project. The INHAND project devised criteria for the harmonization of nomenclature across documents and databases for both proliferative and nonproliferative lesions in rats and mice in toxicologic pathology. 28 The INHAND criteria and terminology are systems based and are primarily relevant to spontaneous and carcinogen-induced lesions. The goal is to provide recommended nomenclature to increase clarity and consistency across toxicity studies. The INHAND publications also have a complimentary nomenclature online reference site, goRENI (https://www.goreni.org). Large data repositories such as the Mouse Tumor Biology (MTB) Database (http://tumor.informatics.jax.org/mtbwi/index.do) and the European Mutant Mouse Pathology Database (http://www.pathbase.net) provide standardized tumor classification, information on tumor frequency, latency, genomic data, tumor pathology reports, and images for several mouse models of human cancers. 36,38
Anatomic Considerations
When scoring mouse models, it is critical to understand the normal anatomy of the mouse and how each organ system differs from human. Many models of human cancer have varying patterns of growth and disease and may have differences in the rates at which lesions progress in specific parts of the target tissue. For tissues with nuanced anatomic considerations (eg, prostate), the challenge involves grading each lobe. In the case of prostate, each lobe may illustrate very different grades of precancerous/preneoplastic and malignant lesions, and this may vary according to the specific genetically engineered model or background strain and age. It is, therefore, recommended to score cross-sectional cohorts or assess lesions at specific time points and, in the case of prostate, grade each lobe separately to accurately evaluate and validate the biologic behavior of the model. 23,37
Necropsy, Tissue Trimming, and Artifacts
Standard necropsy techniques should be used in accordance with several published recommendations. The RENI guides (https://reni.item.fraunhofer.de/reni/trimming) provide general tips on proper necropsy and tissue trimming for rodent models. 24,30,35 Refer to the NCI Mouse Models of Human Cancer Consortium consensus papers when devising necropsy and trimming protocols for specific models of neoplasia and to be aware of specific anatomic considerations for modeling of particular organ systems. 7,13,20,23,32,37,45 For example, the mouse prostate has 4 lobes and associated accessory sex glands. GEM models of prostate cancer may develop proliferative lesions in any of the 4 lobes and in the accessory sex glands; thus, it is recommended to prepare the entire prostate and associated accessory sex glands en bloc for histologic examination. 23,37 This allows for accurate evaluation of all prostate lobes and helps differentiate primary prostatic lesions from those extending from the accessory sex glands. The same trimming protocols often work well for standardized sectioning of orthotopic xenografts.
When phenotyping a new model, it is critical to collect and evaluate a complete set of tissues. This allows for examination of all tissues for off-target effects that often can lead to new and unexpected phenotypes. A comprehensive list of organs for morphologic phenotyping may be found in the literature. 9
Artifacts may occur at necropsy and when trimming tissues. Tissue artifacts may impede grading and scoring of models. Uniform, pristine sections are also a necessity for any automated digital imaging analysis of tissue. The pathologist should be aware of common artifacts that may “mimic” invasion or malignancy. The most common artifacts are tangential sections and folds in tissue epithelium. 44 Tangential sections may give the appearance of malignancy or areas of invasion when, in fact, deeper sections may reveal folds in the epithelium, portions of adjacent normal glands, or extension of a lesion from an adjacent gland, rather than true malignancy of the particular gland being evaluated. Deeper sections help differentiate between invasive malignant epithelial cells and herniated epithelial cells, and they help define early lesions. Thus, a specific number of serial sections or step sections will allow one to evaluate if any artifact such as a tangential section occurs, to help aid in definitive diagnosis, and to have serial unstained slides available for immunohistochemical staining for tumor biomarkers.
Histochemical Stains and Immunohistochemistry
Immunohistochemistry (IHC) and histochemical stains are useful for the identification of preneoplastic/precancerous lesions, diagnosis of tumor type, and identification of neoplastic invasion and metastasis in tissue sections. Quantification and scoring of IHC and histochemical stains are necessary for the analysis and comparison of mouse models to the human disease. For example, the use of the Alcian blue histochemical stain and the IHC marker CK19 highlight (PanIN) precursor lesions in GEM models of pancreatic ductal adenocarcinoma. 19 –22 Both markers are specific for PanIN lesions and thus enable identification and quantification within sections. An understanding of the biology and neoplastic spectrum of particular mouse models and an awareness of histochemical stains and specific IHC markers for lesions in each model enable accurate identification and scoring. Careful consideration of the model is necessary to determine whether selection of a human-directed or mouse-directed primary antibody is appropriate for the question at hand, particularly in xenografted and/or humanized mice.
IHC markers help discern true metastases from primary tumors in organ systems other than that which is being evaluated. For example, primary lung tumors are common and must not be mistaken for metastases. IHC for lung-specific markers such as thyroid transcription factor 1 (TTF-1) and pulmonary-associated surfactant protein C (SPC) can be used to confirm pulmonary origin. 23 IHC markers of transgene expression may be helpful for primary lesions and metastasis identification; however, it is important to note that transgene expression does not necessarily prove a distant lesion is metastatic. Tissue-specific promoters may have transcriptional activity in nontarget cells that is sufficient to transform such cells. Global expression or germline expression may lead to primary tumors at distant sites, which can confound interpretation of lesions. 23 The use of nontissue-specific promoters (eg, keratins) has similar limitations. 23 Thus, the use of an IHC marker for expression of the transgene may not suffice to truly identify metastasis and for characterization and scoring of the model. Based on the biologic activity of specific models and the spectrum of neoplastic lesions, a panel of IHC markers may be necessary to aid in the identification and scoring of mouse models.
Xenograft Studies
Nuances of the Model
Xenografts are typically established in the flank subcutis or an orthotopic site. Sometimes an intravenous, intracardiac (left ventricle), or intraosseous injection may be performed to model metastasis. Cell line–based xenografts, such as those derived from the NCI-60, 1 have very low molecular and phenotypic heterogeneity but are well characterized and reported in the literature. For xenograft studies, online repositories curate biological, clinical, and drug response data pertaining to the tumor donor, such as the NCI Patient-Derived Models Repository (https://pdmr.cancer.gov), the NCI DTP MicroXeno Project (https://dtp.cancer.gov/docs/microxeno/download.htm), the EuroPDX Consortium (http://europdx.eu/pdx-collection.html), the Public Repository of Xenografts (https://www.proxe.org), 40 and many other institution- or company-specific shared resources. Patient-derived xenograft (PDX) models are supplanting this study design, as they more faithfully recapitulate human biological processes. Conventional cell line xenografts could have value in the early stages of evaluating in vivo efficacy of interventions and may in fact perform as well as PDX models for some cancers. 11
In either setting, a panel approach—using multiple tumor lines with a common feature (eg, deficiency in the same tumor suppressor)—is more powerful than testing just one cell line or tumor. In PDX models, there is high fidelity of phenotype and gene expression profile in early passages, although the stroma is typically replaced by murine elements rapidly in the first passage, then completely in passages 3 to 5. 18,31 Growth kinetics and clonal evolution within a PDX are reportedly more akin to a metastasis than a primary tumor. 11,33 Xenografts newly derived from patient specimens (P0) should undergo careful characterization, passaging, cryobanking, and validation before ever being used in the experimental setting. 11,42 To this end, PDX biobanks and commercial services that cover hundreds of tumor subtypes and provide biological, clinical, and drug response data continue to be built. Variations in both the cell line–based and patient-derived xenograft models might include coinjection with basement membrane material, injection of a second cell type such as transformed fibroblasts or stem cells to influence the growth of the tumor cells, or transformation of the tumor cells for longitudinal bioluminescent imaging. True tumoral homogeneity at the microscopic level can facilitate microarray construction, which further eliminates variability in biomarker analysis between slides.
Common Questions: à la Carte Analysis
Generally, xenograft study questions fall into 1 of 2 categories: tumor biology or antitumor efficacy. Commonly queried end points are centered on tumor death (ranging from single-cell death to extensive coagulation necrosis of tumor and stroma), cell cycle/proliferation, identity (eg, markers of metabolic pathways, cell surface markers, differentiation), or stroma (eg, immune infiltrate, vascularization, matrix composition). Scoring schemes for xenograft studies are largely tailored to the question at hand. To a certain extent, extremes of a scoring spectrum are calibrated to the relative magnitude of a change within a given study.
For example, an investigator may ask the pathologist to score tumor stroma or more specific stromal elements of interest (eg, collagen isoform, immune cell marker, adipokine), in which case the pathologist guides the investigator toward the appropriate histochemical/immunohistochemical stains to use. But often, it requires sitting down with the slide set and assessing what biological processes are principally at play, to make the determination which stains may discriminate among individuals or groups. For example, if the fibromyxomatous stroma appears more or less dense in intratumoral vs junctional/invasive foci, the pathologist may proceed by separately scoring the stromal abundance in each location using the tools of his or her choosing, such as hematoxylin and eosin (HE) stain or markers of collagen and mucin. Furthermore, the pathologist may apply his or her understanding of a biological process to suggest other in situ biomarkers; in this example of stromal remodeling, matrix metalloproteases might be of interest.
Consider the simple example of tumor necrosis. A semiquantitative approach would exist on a 5-point scale of none (0), minimal (1), mild (2), moderate (3), or marked (4) for HE-stained slides. The deliverable is a set of exemplary photomicrographs accompanied by descriptions and tabular data showing which animals fall into each category. The quantitative approach involves calculating the total 2-dimensional necrotic area as a proportion of total tumor area. The numbers and sizes of necrotic foci within a tumor may be of interest to the investigator, so the deliverable in this case is a spreadsheet containing the area of each necrotic focus within a tumor and the summary data for a whole tumor. Algorithms exist for automated calculation of total stained area and detection of necrotic foci, but some tumors may require manual delineation of regions of interest (ROIs) to segment types of cell death (Fig. 1). Building upon this, employing histomorphometry, repeated analyses across serial sections could enable 3-dimensional reconstruction and volumetric analysis of tumor necrosis.
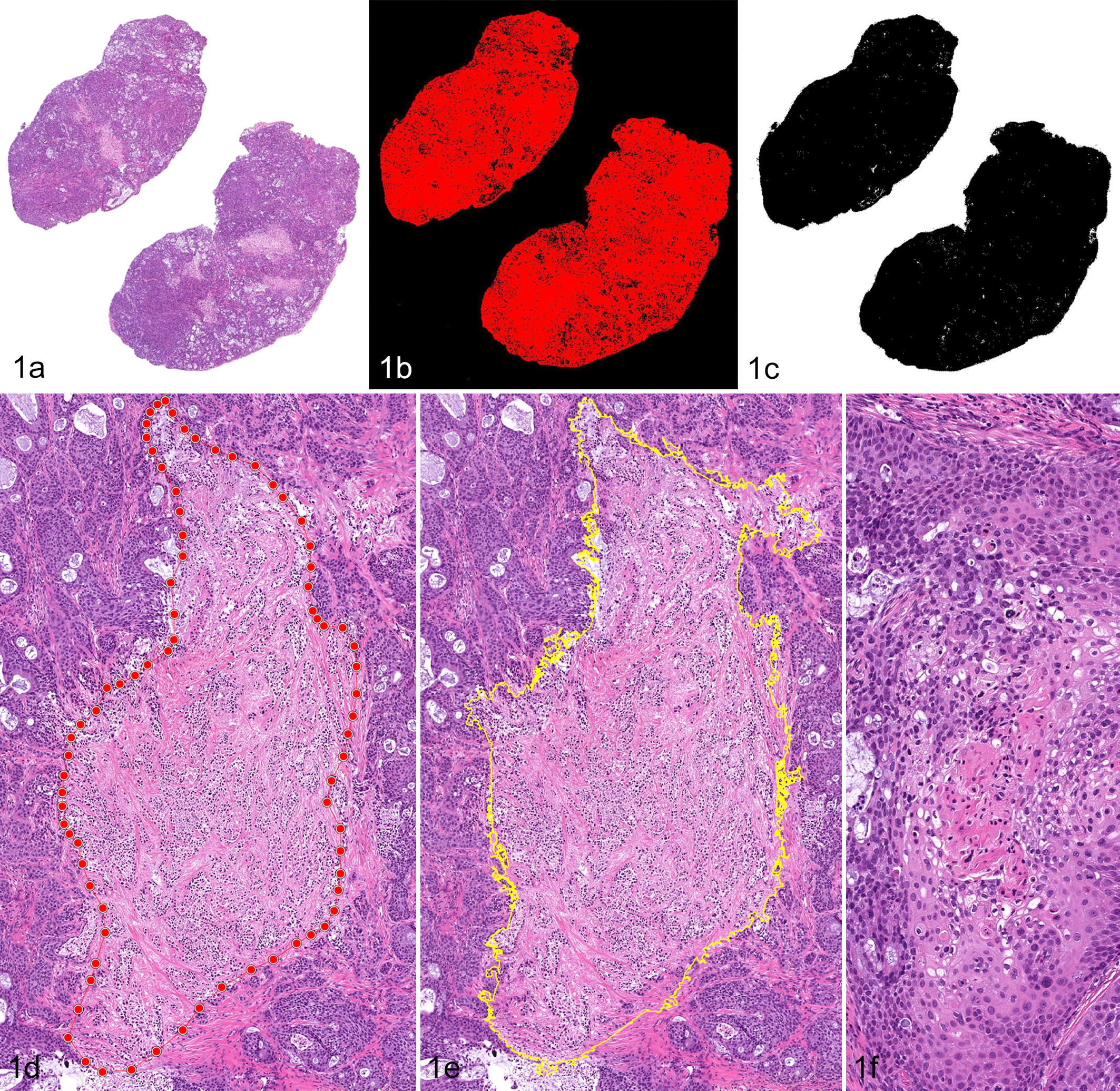
(a) Scanned whole-slide image of a bisected BxPC-3 pancreatic xenograft in a high-intensity focused ultrasound study in which the proportion of total necrotic area per tumor was measured. Hematoxylin and eosin (HE). (b) Mockup of the same image using a software-based total stained area image analysis algorithm to determine tumor area. (c) Mockup of the same image using an open-source platform for thresholding and hole-filling of a grayscale image to determine tumor area. (d) Manual annotation of a focus of intratumoral coagulation necrosis to calculate area. HE. (e) Automation-assisted annotation of the same focus of intratumoral coagulation necrosis. HE. (f) Comedo pattern of necrosis that was excluded from calculation of total necrotic area in this study. HE.
Complementary Data
Histopathologic scoring is reinforced by interpretation alongside other study data. Imaging modalities such as ultrasound, magnetic resonance imaging, bioluminescence, micro–computed tomography, or positron emission tomography may provide a noninvasive way to monitor tumor development before tissue collection. Genetically based imaging methods such as bioluminescence and fluorescence use reporters such as green fluorescent protein (GFP) or luciferase for tracking and imaging tumors in GEM and xenograft models. 16 Imaging modalities may assist in tumor identification, responses to therapy, and tracking early disease. Comparing in vivo data with histopathologic scoring is becoming increasingly important in therapeutic studies. Similarly, ancillary data from the same tumor specimen, such as RNA sequencing or proteomics, could provide valuable guidance to the pathologist when drawing mechanistic links between tumor attributes.
Histopathologic Evaluation
The histopathologic evaluation of tissues is a crucial part of the evaluation and validation of mouse models of cancer. Standard histopathologic evaluation in GEM is often a descriptive evaluation of morphology; however, as clinical trials are increasing and as the effectiveness of new chemotherapeutic drugs and other treatment modalities are studied, it has become necessary to devise quantitative and semiquantitative scoring systems that measure treatment efficacy and outcomes for specific types of cancer. On the other hand, features of malignancy that pathologists are trained to recognize and describe are of little or no interest in studies where the tumor is already well characterized and its behavior is expected to be identical in all animals on a study. Thus, for xenogeneic cancer models, histologic interpretation is largely centered on measurable features of an intervention.
Scoring for tumor biomarkers labeled via IHC or immunofluorescence is important in translational research. In human medicine, identification of the expression of tumor biomarkers is often key for diagnosis and essential for prognosis and prediction of therapeutic response. The study pathologist should identify and understand human tumor scoring systems that are commonly used for the specific type of cancer being studied. Thus, expression and quantification of biomarkers is an essential piece when using mouse models of human cancers. Markers of interest include tumor suppressor proteins (eg, PTEN, p53), markers specific to activation of distinct signaling pathways (eg, Akt), proliferation markers (eg, Ki-67, PCNA, BrdU), and endothelial cell markers for determination of angiogenesis or microvessel density (eg, CD31, CD34). Other markers important to particular models include p63, specific cytokeratins, smooth muscle actin, and basement membrane constituents such as type IV collagen and laminin. 13
In the setting of immunohistochemistry, the semiquantitative approach for a novel marker could also exist on a 5-point scale of none (0), minimal (1), mild (2), moderate (3), and strong (4) staining intensity. The deliverable includes a set of exemplary photomicrographs accompanied by descriptions—which for IHC should include descriptors not only of intensity but also localization (nuclear, cytoplasmic, membranous) and possibly texture—and the desired quantitative measurements (eg, neovessel density, nuclear size, proportion of positive cells) at the individual and group levels. There are few examples of widely accepted scoring systems for IHC, perhaps the most notable of which is the HercepTest for HER2, which is a vertically integrated immunohistochemical approach resulting in scores of 0, 1, 2, or 3 in human breast cancers. 4,43 Building upon this, the H-score and Allred score are composite approaches that take into account not only IHC staining intensity but also percentage or proportion of cells at a given intensity. These methodologies are now combined with in situ hybridization (ISH) for comprehensive evaluation of patients with breast cancer. 14 For quantitative outputs, perhaps the most basic techniques are cell counts or pixel counts based on single-chromogen (traditionally DAB) staining. The most complex of cases could involve automated analysis using custom-designed algorithms for multiplexed staining protocols that are difficult for even the most experienced human eye to parse.
Quantitative and Semiquantitative Scoring
Semiquantitative and quantitative scoring schemes enable lesions to be numerically scored. Quantitative scoring is based on continuous, measured values and yields ratio data, whereas semiquantitative approaches assign numeric scores and yield ordinal data. 17 When scoring mouse models of neoplasia, a combination of approaches may be selected for a single animal.
The gold standard for quantitative scoring is image analysis (IA). Whole-slide scanned images are obtained and analyzed by IA software systems. Digital pathology applications are extremely powerful in eliminating interobserver variability when a trained pathologist is involved. IA software is useful for identification or segmentation of an image into normal, neoplastic, and even preneoplastic lesions, which can then be counted or measured for area. The software separates the images based on set parameters such as shape and color. Tumor area can be measured via pattern recognition software, and other parameters can be measured such as the area labeled by a specific marker for the tumor, total mitotic cells, and total number of cells. Once the morphology is defined and any regions of interest measured, IA algorithms may be devised to score tumor biomarkers labeled via IHC or immunofluorescence. Manual and digital methods can be used in parallel to reinforce one another. IA is beneficial in models with coalescing lesions or where scoring via semiquantitative methods is not achievable. It is also robust in simultaneous analysis of multiple parameters, for example, microvessel diameter and density, nuclear size and staining intensity, and type and location of immune cell infiltrates. For IA applications, the pathologist is involved in biomarker validation, ROI selection, algorithm tuning, data analysis, and interpretation. IA methods have been previously described and reviewed in detail elsewhere. 2,15,27,41
Incidence scoring reports the number affected by certain types of neoplasia or preneoplastic/precancerous lesions within a specific group or cohort. 17 For carcinogenesis studies, groups may be represented as treated or nontreated, or in the case of GEM, they may be organized by genotype. Data may be reported as total number of tumors within a category or the percent affected by a type of neoplasia. The results are typically reported in a graph or contingency table. Examples are available in the literature. 17
To account for differences in grade and distribution, grading schemes have been devised and termed adjusted lesion grades. Many mouse models of human cancer have varying patterns of growth, and may have differences in the rates at which lesions progress in specific parts of the target tissue. A good example is mouse models of prostatic neoplasia where, depending on the model, there may be differences in the grade of lesion and progression of lesions in each prostatic lobe. Suttie et al 39 devised a grading scheme whereby tissues were graded as hyperplasia (grades 1 to 3), adenoma (grades 4 to 5), or adenocarcinoma (grade 6). A distribution was estimated and described as focal (2 or fewer lesions), multifocal (less than 30% of the lobe involved with 3 or more lesions), or diffuse (greater than 30% of the lobe affected). 6,39 An adjusted lesion grade was calculated by combining the lesion and distribution grades. This grading scheme is referred to as a “distribution-adjusted lesion grade.” The adjusted grades can be collated for statistical analysis and comparison among genotypes and groups.
Similarly, Berman-Booty et al 6 proposed a modified grading scheme for prostatic lesions that provides an adjusted lesion score. This modified grading scheme included an adjusted score for the most severe lesion present, an adjusted score for the most frequent lesion present, and the distribution for each. The distributions of these lesions were determined and described as focal if fewer than 3 foci contained the lesion, multifocal if there were 3 or more foci or less than 50% of the section containing the lesion, or diffuse if greater than 50% of the section contained the lesion. 6,39 Similar to the “distribution-adjusted lesion” grading scheme, an adjusted grade was assigned based on distribution. This scheme differs as it incorporates the most severe and most common lesions, both of which contribute to the pathology present within a section. Each of the 4 lobes of the prostate (dorsal, ventral, lateral, anterior) was assessed individually and assigned 2 grades each. The adjusted scores were added to obtain a sum (total pathology score) that reflected the most severe lesion and its distribution and the most common lesion and its distribution (sum of the adjusted lesion scores). 6
Both “distribution-adjusted lesion” grading schemes are helpful with mouse models of neoplasia in which anatomically specific sites have variation in the severity of lesions, and there is variation among anatomic sites (such as specific glands or lobes) in the rate at which lesions progress. A distribution-adjusted lesion grading scheme may be applied and useful for any model that includes a multistage progression or continuum through preneoplastic and precancerous lesions to malignancy, such as for prostate, mammary, and pancreatic mouse models of neoplasia. Examples are available in the literature. 6
Reporting
The goal of reporting histopathologic scores is to clearly communicate results and foster reproducibility within and between studies. Details about the scoring system adopted for the specific model or study should be included in the Materials and Methods sections of manuscripts for publication. Results should be organized in a table or spreadsheet and include a definition of the scoring scheme used for the study. Include any references from which the scoring scheme was adapted and any references used to define and classify lesions, such as the NCI MMHCC consensus reports or INHAND publications. Tabular data are nicely complemented by representative photomicrographs and a prose summary, including limitations of the study where appropriate. Statistical analysis is typically done by the investigator; however, statistical approaches will vary depending on data type (ie, ratio vs ordinal); therefore, consultation with a statistician is recommended.
Conclusions
Histologic scoring of mouse GEM and xenograft models of cancer may be challenging. It is critical to keep model-specific considerations in mind, and this may guide the choice of the most appropriate scoring system for the particular model. Considerations include a full understanding of the model, from its development and expected phenotype, to the similarities and differences between the model and the human disease and anatomic considerations of mice and human when collecting tissues and grading lesions. The proper accepted nomenclature should be used when evaluating and reporting mouse models of human cancer. Quantification via digital IA should be considered for measuring or quantitating the extent of neoplasia and expression of tumor biomarkers. Semiquantitative scoring schemes have been devised that include adjusted lesion grades to account for severity, frequency, and distribution of lesions.
Pathologist-generated tumor scoring generates meaningful, powerful data, but the process is susceptible to the fallibility of the human mind. Intraobserver variability is perhaps one of the more insidious threats to validity of pathology data, and we must be mindful in this regard. Besides managing our own cognitive traps, 3 we must also be perceptive educators and facilitators, translating molecular biological processes into measurable histopathologic end points and connecting investigators with cutting-edge quantitative pathology applications that will be the bellwether of 21st-century anatomic pathology. Pathologists: 1, sloppy science: 0.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
