Abstract
Histopathology remains an important source of descriptive biological data in biomedical research. Recent petitions for enhanced reproducibility in scientific studies have elevated the role of tissue scoring (semiquantitative and quantitative) in research studies. Effective tissue scoring requires appropriate statistical analysis to help validate the group comparisons and give the pathologist confidence in interpreting the data. Each statistical test is typically founded on underlying assumptions regarding the data. If the underlying assumptions of a statistical test do not match the data, then these tests can lead to increased risk of erroneous interpretations of the data. The choice of appropriate statistical test is influenced by the study’s experimental design and resultant data (eg, paired vs unpaired, normality, number of groups, etc). Here, we identify 3 common pitfalls in the analysis of tissue scores: shopping for significance, overuse of paired t-tests, and misguided analysis of multiple groups. Finally, we encourage pathologists to use the full breadth of resources available to them, such as using statistical software, reading key publications about statistical approaches, and identifying a statistician to serve as a collaborator on the multidisciplinary research team. These collective resources can be helpful in choosing the appropriate statistical test for tissue-scoring data to provide the most valid interpretation for the pathologist.
Cells and tissues are commonly studied in biomedical research to offer biological perspective that can clarify and complement clinical and molecular data. At its fundamental level, histopathological evaluation and description of tissues can be summarized by images in a figure to demonstrate group differences. While morphologic descriptions can serve an important function, these have inherent limitations for distinguishing differences between treatment groups. To combat this, as well as to increase the rigor and repeatability of tissue studies, group changes can be enumerated through semiquantitative and/or quantitative scoring. 3,10 Tissue-scoring data can then be analyzed by appropriate statistical tests to provide a more rigorous level of confidence in the interpretations and conclusions. The aim of this article is to identify 3 common pitfalls of tissue-scoring analysis and offer approaches for the pathologist to avoid these issues.
Shopping for Significance
For many people entering into biomedical research, there is an immediate and broad exposure to many different approaches and tools for investigational studies. It does not take long to quickly become aware (from reading journal articles and attending lab meetings) that statistical significance is a vital component of most analyses. While true on many levels, this concept can become dangerous when it mistakenly assumes that any form of statistical significance is a good thing. In this frame of mind, selection of a statistical test could become like window shopping to find one that produces the greatest significance (eg, smallest P-value). This is a flawed approach.
Statistical tests typically have underlying assumptions about the data that should be met to have confidence in the resulting analysis. If the assumptions for a statistical test are not met, the analysis may be prone to incorrect interpretations. Therefore, the best approach is to select a statistical test that fits the experimental design and data. For instance, one common question is whether the data fulfill the assumptions of parametric (eg, continuous data, normal distribution) or nonparametric (eg, discontinuous data or lack of normal distribution) tests, to help guide the selection of a statistical analysis.
To show how selection of a statistical test is important for the final analysis, we will review and then deconstruct an experiment comparing 2 groups of data. In this mock example, we show both a bar graph and dot plot of the same ordinal tissue-scoring data between wild-type (WT) and knockout (KO) groups of mice (Fig. 1a, b). If Fig. 1a were the only figure shown, it would be very difficult for the reader/reviewer to know the number of mice, the study design, and the meaning of the error bar (in this case, it is the margin of error). Transparency is one way to avoid errors, and in this case, one can show the data in a dot plot to be more transparent to reviewers and readers (Fig. 1b). Assuming someone were shopping for a statistical test, they might screen several tests (eg, Mann-Whitney U-test, Kolmogorov-Smirnov, unpaired t-test, paired t-test, etc) to find that the paired t-test was significant and choose it (Fig. 1; Table 1). However, the data from this experiment are ordinal, not continuous. Ordinal data are the most common type of semiquantitative scoring data in pathology research, and in this case, they were assigned from a grading system that included 5 grades: 0, 1, 2, 3, and 4. When using ordinal data, nonparametric tests are often recommended. 4,7,23
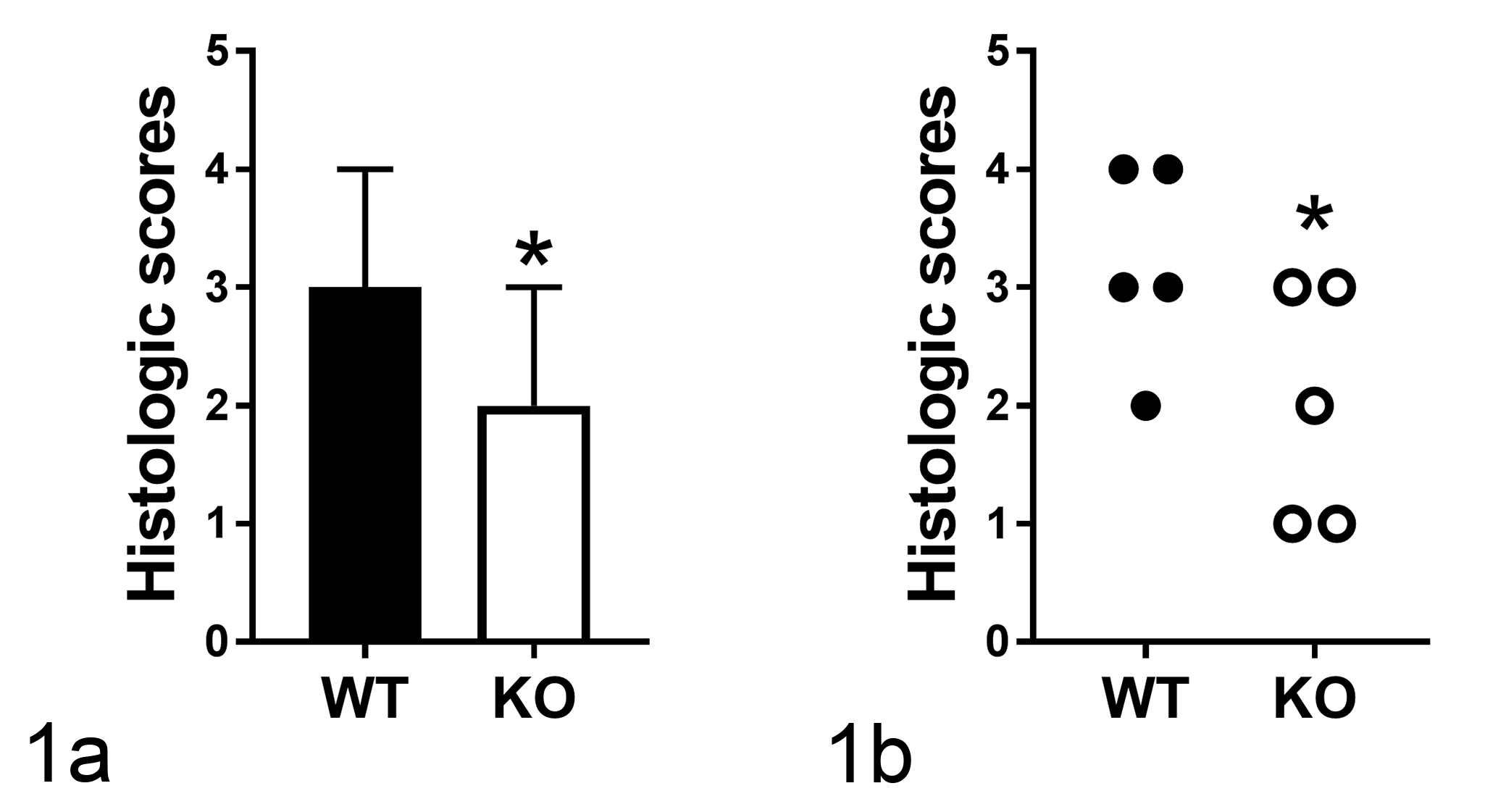
Mock ordinal tissue-scoring data comparing wild-type (WT) and knockout (KO) groups in a bar graph (a; bar = median with 95% confidence interval) or as a dot plot (b). *P = .004, paired t-test. GraphPad Prism Software, v7.03 (GraphPad Software, La Jolla, CA; www.graphpad.com) was used for all statistical analysis in this article.
Mock Ordinal Scoring Data From Wild-Type and Knockout Animals (n = 5/Group) From Figure 1 With Parametric (Paired t-Test, Unpaired t-Test) and Nonparametric (Mann Whitney U-test, Kolmogorov-Smirnov test) Statistical Analysesa
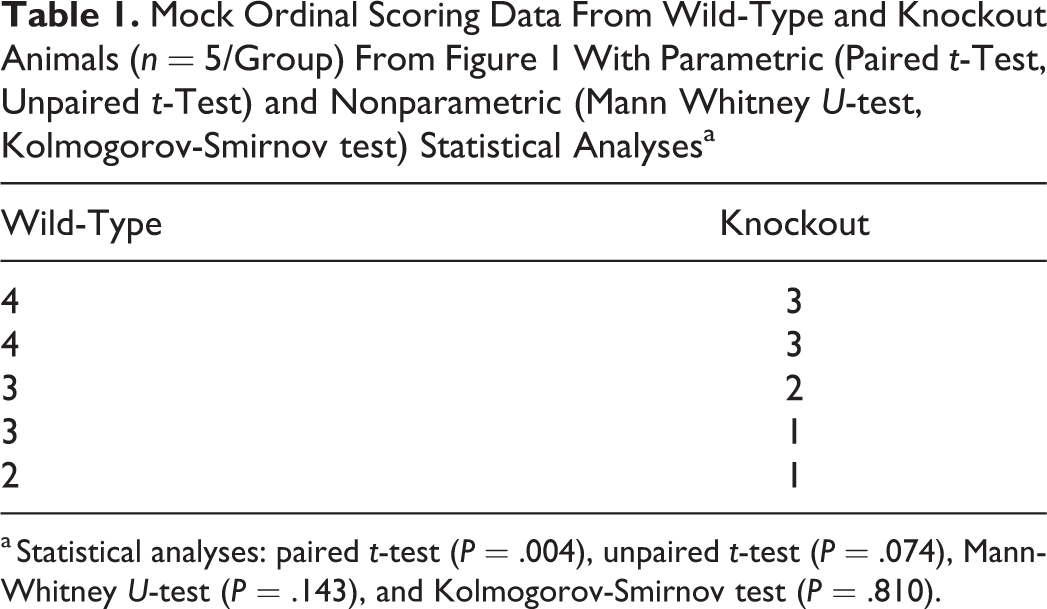
a Statistical analyses: paired t-test (P = .004), unpaired t-test (P = .074), Mann-Whitney U-test (P = .143), and Kolmogorov-Smirnov test (P = .810).
Table 1 illustrates the results of nonparametric tests on the mock data to demonstrate how easy it is to make errors. Notice that the 4 different tests give 4 very different P-values! Had we shopped around and chose the paired t-test because it had a significant difference, we would be ignoring the fact that the different tests are making very different assumptions about how the data were generated and, as noted above, the ordinal nature of the data. In reality, the large P-values for the Mann-Whitney U-test and Kolmogorov-Smirnov test (which are appropriate for these data, whereas the t-tests are not) suggest that these data could have easily been generated randomly. For example, assume that the Table 1 data were instead generated through rolling 2 sets of dice (n = 5 samples/group), using the right hand for one group (see “WT” data) and using the left hand for the other group (see “KO” data). The values in this mock data set are in line with the kind of results that would happen by rolling dice (chance) versus an actual effect. If we used appropriate nonparametric tests, these mock data would have shown no significant differences in rolling dice between the right and left hand, or rather, that the outcome is in line with being a random event.
What are other ways can we increase our confidence in the data besides using valid scoring systems and appropriate statistical analyses? We could repeat the experiment to show that multiple replicates have similar tendencies and thereby also increase the sample size to strengthen our statistical analysis. Another way we can increase our confidence in the data is to corroborate these data to other biologically relevant data collected from the same animals. For instance, the severity of acute inflammatory lesions in lung tissue between 2 groups of animals might be corroborated by changes in a complete blood count (eg, neutrophilia). 8
Overuse of Paired t-Tests
The example above highlights a frequent mistake seen in the comparison of 2 groups: that of using paired t-tests to compare unpaired treatment groups. 22 If treatment groups are independent of each other, meaning that the samples in one group are not linked in any way to samples in the other group, the data are unpaired (Fig. 2a). Conversely, if the treatment groups are dependent, meaning the samples in one group are linked to the samples in the other group, the data are paired. In pathology studies, common examples of paired data include (1) repeated measures taken on the same animal at different times (Fig. 2b) or (2) two different treatments performed on the same animal (or tissue) with similar endpoints (Fig. 2c). Importantly, data for paired samples are linked and must be analyzed in a related fashion. If an accidental switch in data entry order occurs, it can radically influence the analysis of paired t-tests, whereas entry order for unpaired t-tests does not matter because the samples are not linked (Table 2).
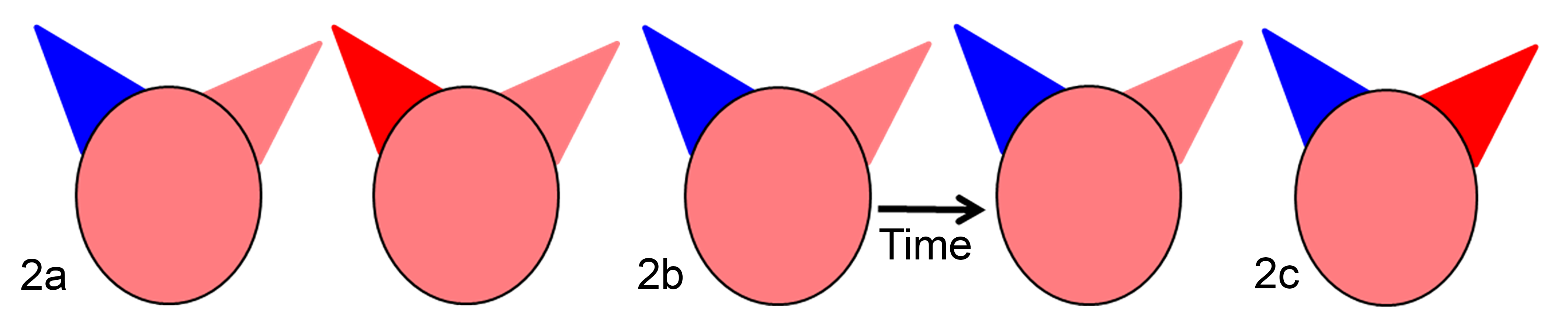
Three examples of study designs to evaluate topical compounds (identified as red or blue) applied on the ears of pigs to assess epidermal injury from biopsied skin. (a) In an unpaired design, 2 groups of pigs were compared. Each group of pigs had a unique treatment (red or blue). (b) In a paired design using repeated measures, each animal had the same treatment, but tissue biopsy data were collected at 2 different times. Each paired comparison (ie, early and late time points) was from the same pig. (c) In a paired design not using repeated measures, 2 distinct treatments were applied to the ears of each pig. Each paired comparison (red and blue ears) was from the same pig.
Mock Quantitative Scoring Data for Cohorts 1 and 2a
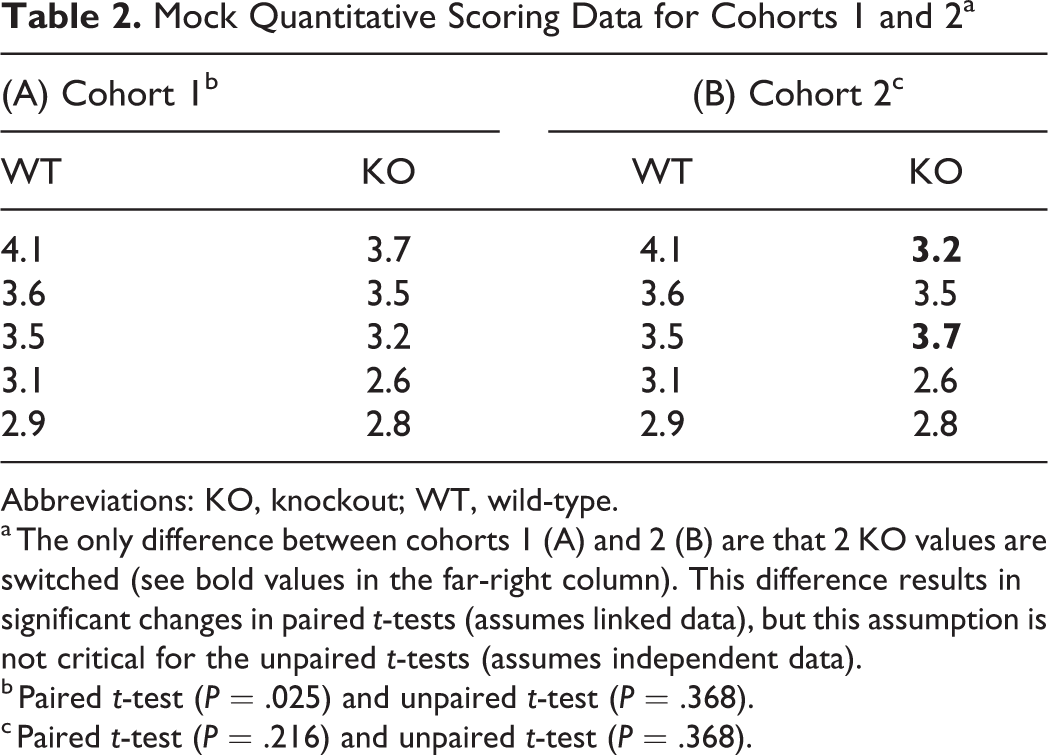
Abbreviations: KO, knockout; WT, wild-type.
a The only difference between cohorts 1 (A) and 2 (B) are that 2 KO values are switched (see bold values in the far-right column). This difference results in significant changes in paired t-tests (assumes linked data), but this assumption is not critical for the unpaired t-tests (assumes independent data).
b Paired t-test (P = .025) and unpaired t-test (P = .368).
c Paired t-test (P = .216) and unpaired t-test (P = .368).
Misguided Analysis of Multiple Groups
While many pathology studies are composed of 2 basic test groups (eg, a control and treated group), as in the above examples, other studies are more complex, such as those involving multiple (3 or more) treatment groups. In these cases, investigators may try to apply a series of t-tests to make group-to-group comparisons. This is not valid because t-tests are designed for studies that compare 2 groups. The added complexity of having multiple groups requires different types of statistical analysis.
For example, if the data support parametric tests, one approach is to evaluate the groups using analysis of variance (ANOVA); this test initially evaluates whether the mean of each group is the same (Fig. 3). If the ANOVA results in significance, the interpretation from the evidence is that the group means are not all equal. A nonparametric correlate to the one-way ANOVA is the Kruskal-Wallis test. After performing an ANOVA or Kruskal-Wallis test and finding evidence that at least one group mean is different from the rest, post hoc evaluations comparing pairs of groups can be conducted (Table 3).
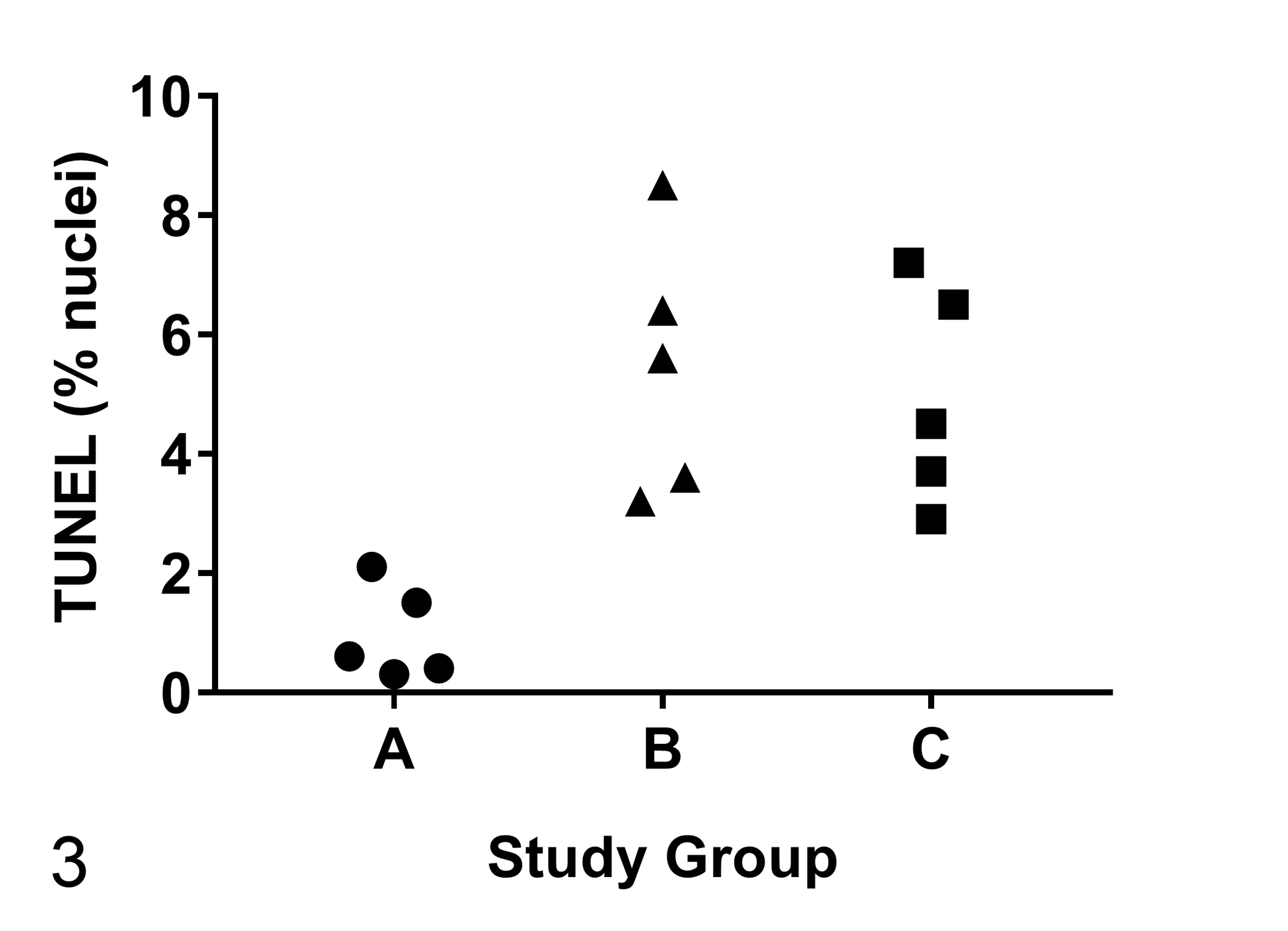
Mock example analyzing a treatment applied to 3 independent groups (A–C). The one-way analysis of variance was significant (P = .0024), suggesting that the means for the groups were not all equal. Further evaluation of post hoc tests (eg, Tukey’s tests) for specific group comparisons showed the following results: A versus B (P = .0034), A versus C (P = .0078), and B versus C (P = .8885).
Examples of Common Statistical Tests for Various Group Comparisonsa
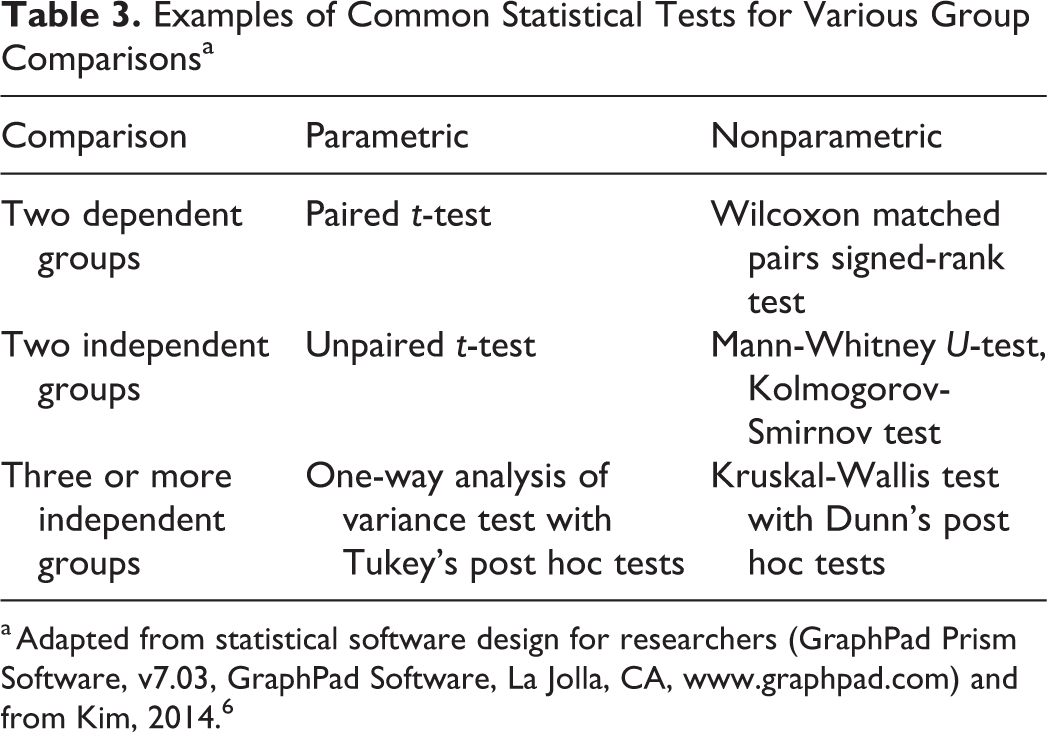
a Adapted from statistical software design for researchers (GraphPad Prism Software, v7.03, GraphPad Software, La Jolla, CA, www.graphpad.com) and from Kim, 2014. 6
While we have briefly discussed the most common approaches to analyze tissue scores, these are not to be viewed as dogmatic recommendations, as it is important to recognize that there are several ways to analyze tissue-scoring data. The approaches we have described are commonly used and published in the literature. Sometimes, the data, experiment designs, or questions being asked can increase the permutations and complexity of the study, so other statistical approaches might be better used and advised by the statistician.
Summary
We have highlighted several approaches to avoid slipping into common pitfalls when analyzing tissue scores. Pathologists who perform tissue scoring should have access to fundamental statistical resources, and several are available. In recent years, statistical software platforms have become increasingly more user-friendly and as such are common tools used in biomedical publications. 4,23 Several published resources (eg, books or articles) are also available to learn more about tissue scoring, experimental design, and statistical analysis. 2,6,7,9,11 –21,23 Last, but not least, pathologists should secure the professional expertise and collaboration of a statistician as a part of the multidisciplinary team. 1,5,24
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
