Abstract
Observational studies are a basis for much of our knowledge of veterinary pathology, yet considerations for conducting pathology-based observational studies are not readily available. In part 1 of this series, we offered advice on planning and carrying out an observational study. Part 2 of the series focuses on methodology. Our general recommendations are to consider using already-validated methods, published guidelines, data from primary sources, and quantitative analyses. We discuss 3 common methods in pathology research—histopathologic scoring, immunohistochemistry, and polymerase chain reaction—to illustrate principles of method validation. Some aspects of quality control include use of clear objective grading criteria, validation of key reagents, assessing sample quality, determining specificity and sensitivity, use of technical and biologic negative and positive controls, blinding of investigators, approaches to minimizing operator-dependent variation, measuring technical variation, and consistency in analysis of the different study groups. We close by discussing approaches to increasing the rigor of observational studies by corroborating results with complementary methods, using sufficiently large numbers of study subjects, consideration of the data in light of similar published studies, replicating the results in a second study population, and critical analysis of the study findings.
Keywords
Observational studies are the most frequent study type published in the pages of Veterinary Pathology and fundamentally guide the daily practice of veterinary pathologists. Although advice is available from other disciplines, it may be difficult for veterinary pathologists to apply these guidelines to the types of observational studies they typically conduct. The first article of this series focused on design and development of observational studies. 6 In the current article, we—editors and editorial board members of Veterinary Pathology and their colleagues—focus on principles of method validation and quality control. These are discussed in the context of 3 methods commonly used by veterinary pathologists: histopathologic scoring (grading), immunohistochemistry, and polymerase chain reaction (PCR). We emphasize that this article is neither intended as a ‘cookbook’ for validating these methods nor as a list of requirements for publishing in Veterinary Pathology. Instead, we focus on using these examples to illustrate considerations and principles that are relevant to any investigative method (Table 1).
General Principles for Valid Research Methods.

Many of these principles apply to subjective analyses such as histopathologic grading or immunohistochemistry scoring as well as to objective analyses such as machine-based measurements of analytes. Validation and quality control of the latter are beyond the scope of this article and reviewed elsewhere. 9,14
Innovative methods can provide unexpected answers to longstanding and important questions. However, the methods should not be the basis for the study, but instead are used to address a hypothesis, question or objective backed up by a clear rationale. If the study objective is this journey’s roadmap, then the methodology is the vehicle’s engine. Studies with a clear objective can cruise toward their destination while still having resources to spare for stops at points of interest along the way. Alternatively, investigations built only on the novelty of a method tend to pleasantly ramble through an interesting landscape but never find a destination.
Investigators are also directed to the reporting guidelines available for many analytic methods, which are also relevant to study design and method validation (Minimum Information for Biological and Biomedical Investigations; https://fairsharing.org/collection/MIBBI). Of particular relevance are those for veterinary observational studies (STROBE-Vet 36,44 ), animal studies (ARRIVE: https://www.nc3rs.org.uk/arrive-guidelines), diagnostic test accuracy (STARD 9,15 ), immunohistochemistry and in situ hybridization (http://mged.sourceforge.net/misfishie/), flow cytometry (http://flowcyt.sourceforge.net/miflowcyt/), and reverse transcription-quantitative PCR (RT-qPCR; http://www.rdml.org/miqe.php).
Microscopic Assessment
Diagnosis, classification, and scoring (grading) of microscopic lesions are mainstays of pathology research. Because these methods are subjective and sometimes challenging to quantify, effective study design and validation of methods are particularly critical. Errors in measuring lesions result in imprecision and variability of the data, thereby reducing the likelihood of detecting a difference between study groups. Furthermore, systematic errors that differ between the study groups may cause differential information bias that could lead to a spurious outcome of the study. Guidelines for histologic scoring and grading have been published 16,23,30,32,45 including a series in Veterinary Pathology on scoring histologic lesions in comparative pathology research, 22,24,25,31,52 and guidelines and insights on cancer grading studies are also available. 29,55
Validate Information Obtained From Others
At the beginning of a new study, at least one of the investigators with appropriate training and experience should validate all of the original pathologic diagnoses by reevaluating the histologic sections or descriptions of the gross postmortem findings using a single set of criteria. Do not rely only on the original diagnosis in the pathology report because differences in diagnostic criteria, individual tendencies, and trends over time are likely to introduce variation and perhaps error into the study.
Obtain data from the primary source, if the pathology records are likely to contain errors. We all recognize the frequency of omissions and mistakes in the clinical histories submitted to pathology laboratories. Thus, as an example, the clinical diagnoses and clinical findings should be obtained directly from the clinical case records and not based only on the pathology record. Even then, it should be recognized that variation among clinicians or inconsistency in recording clinical findings may affect the quality of the data.
Definition of the Outcome
Clear, objective and reproducible criteria are essential for defining the outcome. For example, studies of equine sarcoma and canine liposarcoma provide examples of clearly defined characteristics for each neoplasm. 1,12 Another study included a schematic, microscopic images, and detailed description of cytologic grading criteria (Fig. 1). 27 Visual guides such as these are highly useful to clearly define the cut points between grades and thus reduce both intra- and inter-observer variation.
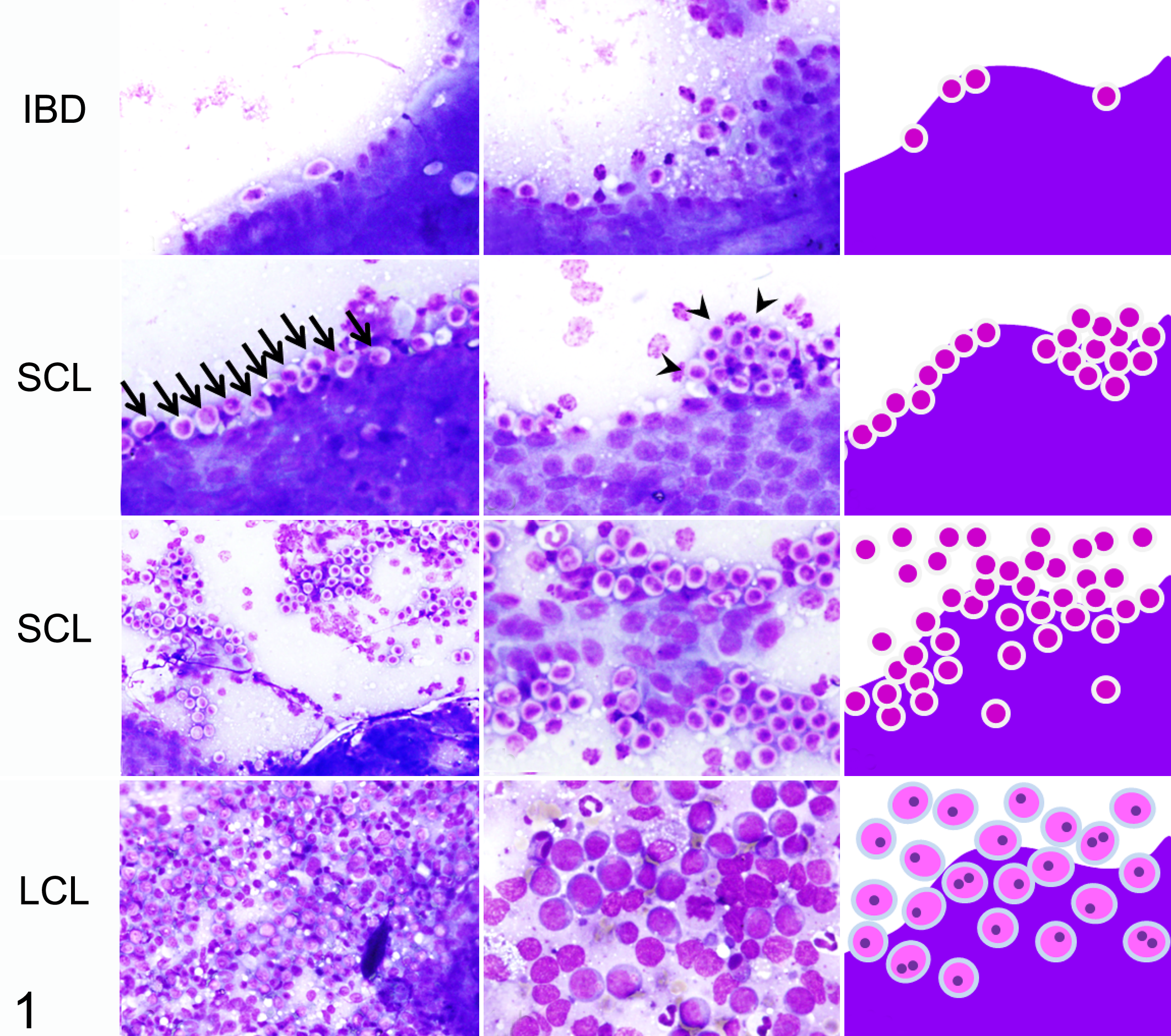
A visual guide is useful to clearly define a microscopic scoring scheme. This example illustrates key criteria for cytologic diagnosis of inflammatory bowel disease (IBD), small cell lymphoma (SCL), and large cell intestinal lymphoma (LCL). The left and center columns show microscopic images and the right column summarizes the key criteria in a schematic. The criteria for small cell lymphoma include a linear alignment of lymphocytes (≥5 cells per 400× field) (arrows), nests of lymphocytes (≥5 clustered cells per 400× field) (arrowheads), and dense infiltration of small lymphocytes. Wright Giemsa. Credit: Shingo Maeda, University of Tokyo; 27 used with permission.
Diagnostic criteria are more clear and reproducible if they are based on objectively observed lesions instead of the inferred morphologic diagnosis. This is particularly important when the diagnostic terms have subtly different meanings among pathologists. For example, “interstitial pneumonia” is used differently among pathologists, whereas the descriptive term “the presence of lymphocytes within pulmonary alveolar septa” should be uniformly understood.
A clear, objective, and precise definition of the outcome is expected to reduce variability of the data, improve understanding of the study methods and results, and make it easier for readers to apply the study findings to their own caseload. 29 Some of our current cancer grading schemes lack clarity or require subjective assessments or interpretations. “Severity of nuclear pleomorphism” and “degree of differentiation” are examples of criteria that cannot be consistently applied to tumor grading by different individuals. Even “percentage of necrosis” is of dubious consistency because the amount of necrosis depends on which parts of the tumor were examined. 29 For studies of new grading schemes, investigators should measure the interobserver variability, using both novice and experienced pathologists. Ideally, each evaluator should be given only the grading criteria used in the study and not provided with additional training, in order to test whether the criteria are sufficiently clear and can be reliably applied to daily practice. 47 If collaborators cannot use the published methods to replicate the results, then it is unlikely that others will be able to validly apply the grading scheme to their clinical case material. 29,47
Defining the Scope of Analysis
The breadth of the histologic analysis should be considered. One approach maximizes the breadth of assessment in order to more comprehensively analyze each study subject. Multiple samples of the lesion are studied, multiple assessments or measurements are made on each sample, and multiple nontarget tissues are examined. This approach is expected to improve the understanding of each individual case and may reduce technical or within-animal variance. The converse approach is to conduct a more limited assessment of each animal, so that more individuals can be included in the study. This approach is expected to more accurately reflect biologic or between-animal variance and may thereby increase statistical power: if systematic biases are avoided, then minor inaccuracies may be compensated for by an increased number of subjects. The balance between these two approaches depends on the number of available cases and the study objectives. A broad-based investigation might be appropriate in an exploratory or descriptive study. However, a focused analysis allows for a more directed interrogation of a specific question. This can allow the investigators to triangulate on testing a specific hypothesis, spend more time and energy on validating the results, and streamline the story that will become the manuscript. Detailed investigation of a single case or very few cases might reveal something new, but a more focused analysis of a large number of cases will increase statistical power and is generally needed to ensure the findings are consistent and applicable to the larger animal population.
Validation of the Methods
Repeated analysis of all or a subset of the same samples (separated by a “wash-out” period to reduce recall) is an important validation of the histopathology data and measures the technical variability or the consistency of scores assigned by a single individual. 16
If assessment of interobserver variation is an objective, each parameter should be independently evaluated by 3 or more of the investigators. The variability among observers is quantified by kappa statistical analysis for binary data, or by other approaches for continuous data. For example, canine mast cell tumors were independently graded by 3 anatomic pathologists and 3 clinical pathologists, with high (72-77%) agreement for the various grading schemes. 4 In contrast to some diagnostic tests, there may not be another “gold standard” for validating histopathologic data, but measuring the consistency of diagnoses among different pathologists can serve a similar purpose.
Avoiding Technical Error, Bias, and Confounding
Pathology data can have poor intraobserver and interobserver consistency. As discussed above, variability is minimized if grading criteria are precisely defined, based on observed changes instead of inferred diagnoses, and illustrated by examples or a visual grading key. Ensure that the individual who analyzes the samples has sufficient training, experience and knowledge, as the results are expected to be more uniform and more accurate than those assigned by nonexperts. 42 Conversely, inadequate training and experience with the grading method may reduce the ability to detect a difference between groups.
The different study groups must be analyzed in the same way. All samples should ideally be analyzed by the same individual. Alternatively, diagnoses or scores can be assigned by consensus among a team that evaluates each sample in a standardized way. If it is necessary that different cases are evaluated by different individuals, be sure the assignments are randomized, and incorporate a statistical comparison of the different individuals into the study design. Histologic sections from the study groups should be examined concurrently in random order, instead of sequentially analyzing the different study groups. Evaluation of all subjects should be done within a narrow time period to avoid “diagnostic drift”. 45
It is imperative to ensure blinding (masking) of the person making the histologic diagnosis, or assigning a histologic score, or making other subjective assessments of outcomes. These outcomes should be measured without knowledge of the study group or other data; that is, without prior knowledge of either the exposure or the outcome. Most observational studies employ a 2-step process. First, tissues are examined in a nonblinded manner to identify the important criteria to be measured and to establish cut-points between the different histologic grades, and then those few specific lesions that are most relevant to the study objectives are scored in a blinded formal analysis. Finally, recognize that blinding may be impossible in pathology-based studies if the observations defining the exposure and the outcome cannot be made independently. For example, immunohistochemistry findings cannot be assessed independently of tumor subtype if the morphology of the tumor is evident on the immunolabeled glass slide. In such instances, consider how this might cause a differential information bias, and how the impact can be minimized by ensuring the data are collected as objectively as possible.
It is critical to ensure that the study groups are comparable in order to avoid selection bias, information bias or confounding. This issue is particularly relevant for studies of archived samples or data not acquired for the purpose of the study. Analysis of the pathology records may not be sufficient to ensure that the 2 study groups were drawn from similar populations; information from clinical hospital records, owner interviews, or knowledge of laboratory protocols may be required. Further, consider if there might be differences between cases and controls in the details of how samples were obtained, processed or analyzed. Failure to ensure that the study groups are comparable and investigated in a consistent manner may result in spurious findings, as discussed in the first article of this series. 6
These factors argue in favor of systematic methodology for routine diagnostic investigations, especially for those involving case material that is likely to be used for future observational studies. This approach has been particularly useful in investigations of wildlife disease. For example, routine mortalities of harbor seals had been investigated in a standardized manner including recording of life history data, necropsy methods, and sampling of tissues and sera. These provided an effective control group for comparison to harbor seals that died during a phocine distemper epidemic, with respect to analysis of antibody titers, morbillivirus infection, gross and microscopic lesions, and bacterial isolates from lung and other organs. 41 Similarly, consistent collection of demographic data (sex, body weight, geographic location, etc) and standardized necropsy methods including evaluation of the vascular system and collection of muscle samples, were key to analytic studies of muscle pathology and verminous arteritis in stranded cetaceans. 10,49
Immunohistochemistry
Immunohistochemistry is a cardinal method in veterinary pathology, yet it is a technically complex procedure. The many variables in the method allow the possibility of technical error unless assays are optimized and validated for the species, tissue and application of interest. Once validated, quality control procedures are necessary for reliable analysis of samples used in the study. The details of immunohistochemistry validation and quality control are described elsewhere. 17,20,30,38,53,54
Antibody Validation
The sensitivity and specificity of the primary antibody is fundamental to the immunohistochemistry method. Analytic sensitivity reflects the smallest amount of the intended target antigen that can be detected. For example, a low-affinity antibody can cause weak labeling that is not easily differentiated from background staining, or an antibody may fail to react with the expected antigen in a nontarget species. Conversely, analytic specificity reflects whether the antibody reacts only with the intended target antigen. Reactivity with a nontarget antigen can lead to false-positive labeling that may be difficult to recognize (Figs. 2–3). Note that these terms—analytic sensitivity and specificity—are related to but different from diagnostic sensitivity and specificity as used in the context of epidemiology studies (ie, the proportion of truly positive and truly negative animals that are correctly classified by the test, respectively).
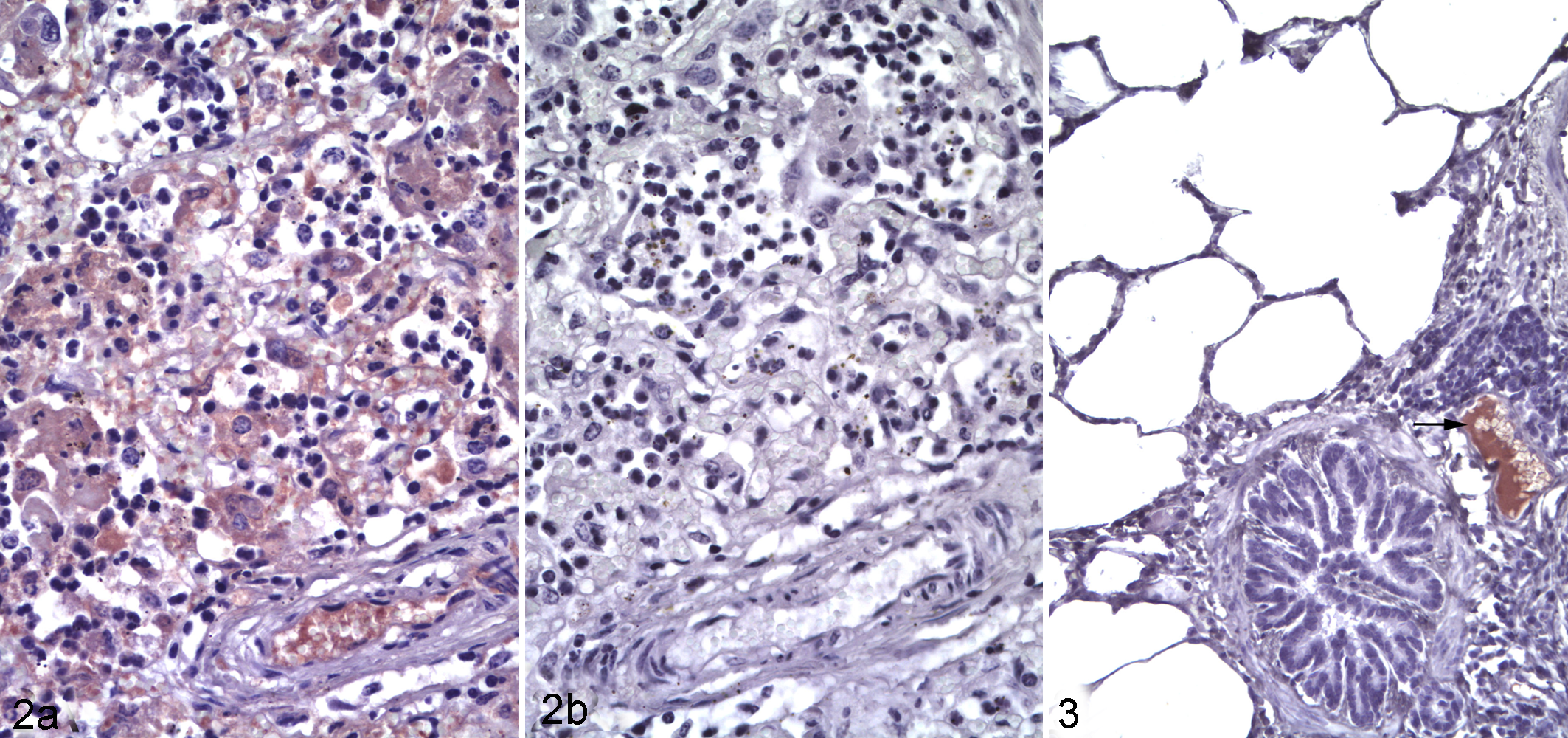
Mannheimia haemolytica pneumonia, lung, steer. Fig 2a. Immunolabeling of leukocytes and intravascular plasma within the lesions. Immunohistochemistry using a rabbit antibody to malondialdehyde (a marker of lipid peroxidation). Fig 2b. Lack of immunolabeling in the same tissue, probed with normal rabbit serum as negative control.
Where possible, veterinary pathologists should use antibodies that have been previously validated for immunohistochemistry in the species of interest. For example, an antibody raised in rabbits against a human protein may not react with the corresponding canine protein, and may react with other unrelated canine antigens. It has been estimated that 75% of commercially available antibodies are nonspecific. 56 Data sheets from the manufacturer may provide validation data. When it is claimed that antibodies have been validated in the dog, consider if the supporting data are available and if they are compelling. If not, investigators must be aware that antibody validation is not trivial, yet it is essential to the credibility and truthfulness of the results.
There are several approaches to validating the analytic sensitivity and specificity of an antibody. 53 One approach is to show immunolabeling in cells or anatomic locations expected to express the antigen (internal positive control), and absence of immunolabeling in cells expected to not express it (internal negative control). For example, immunolabeling is compared in epithelial vs connective tissues of the skin, or in T- vs B-cell areas of lymph node, or in cortex versus medulla of the adrenal gland. However, this is of limited value for antigens that do not have cell- or tissue-specific expression patterns. The subcellular pattern of immunolabeling may also reveal a problem with antibody specificity, if this does not match the expected membranous, cytoplasmic or nuclear location of the protein.
A second approach is western blot analysis of the tissue or tumor being studied (eg, canine lymphoma). An alternative is to use a normal tissue from this species that is known to express the target protein (eg, canine lymph node). In the western blot, the antibody should label a band of the expected molecular weight for the target protein. If there are additional bands, these could be analyzed by mass spectrometry to determine if they are a nontarget protein, or variants or isoforms of the target protein. These additional bands are highly relevant: while target and nontarget antigens can be differentiated on western blots based on molecular weight of the protein bands, these signals are indistinguishable by immunohistochemistry. An important caveat that limits the value of this approach is that the labeling pattern in western blot analysis of fresh tissue may differ from that in formalin-fixed tissues. The proteins in a western blot are denatured, and the process of formalin fixation and paraffin embedding may alter the protein conformation and thus affect the epitope recognized by the antibody. Consequently, immunolabeling of the target antigen or of a cross-reacting antigen in the western blot does not necessarily mean the same reaction would occur in fixed tissues, and vice versa. 38
An example of validation by western blot is provided by a study of canine liposarcoma that used an antibody validated for detection of mouse and human MDM2. A western blot of normal canine testicular tissue probed with this antibody showed a single band of 90 kDa molecular weight as expected for canine MDM2. 1 Conversely, when normal and neoplastic canine mammary tissues were probed with an antibody approved for HER2 assessment in human breast cancer, the western blot did not label any protein at the expected molecular weight of canine HER2 (138 kDa). In this analysis, the antibody labeled 2 other bands (50-60 kDa), and mass spectrometry showed that these bands did not contain HER2. This cross-reactivity may have been responsible for the unexpected cytoplasmic immunolabeling of neoplastic cells and labeling of non-neoplastic cells, which were observed when canine tissues were probed with this antibody. 3,40 These are disruptive findings: they challenge the validity of previously published studies, and question the use of canine mammary tumors as a model of HER2 overexpressing breast cancer. 3
A third approach to validating antibody specificity is to compare cells that are engineered to express or not express the target protein. One method is to transfect nonexpressing cells with the gene of interest (Fig. 4). A second method is knockout or knockdown of the gene of interest in cells that naturally express it. Expressing and nonexpressing cells are pelleted, fixed in formalin and embedded in paraffin, then immunoreactivity is compared in histologic sections of the different cell pellets. Note that this approach may not detect immunoreactivity against cross-reacting antigens, because complex tissues include a greater diversity of proteins than are expressed by the negative control cell line.
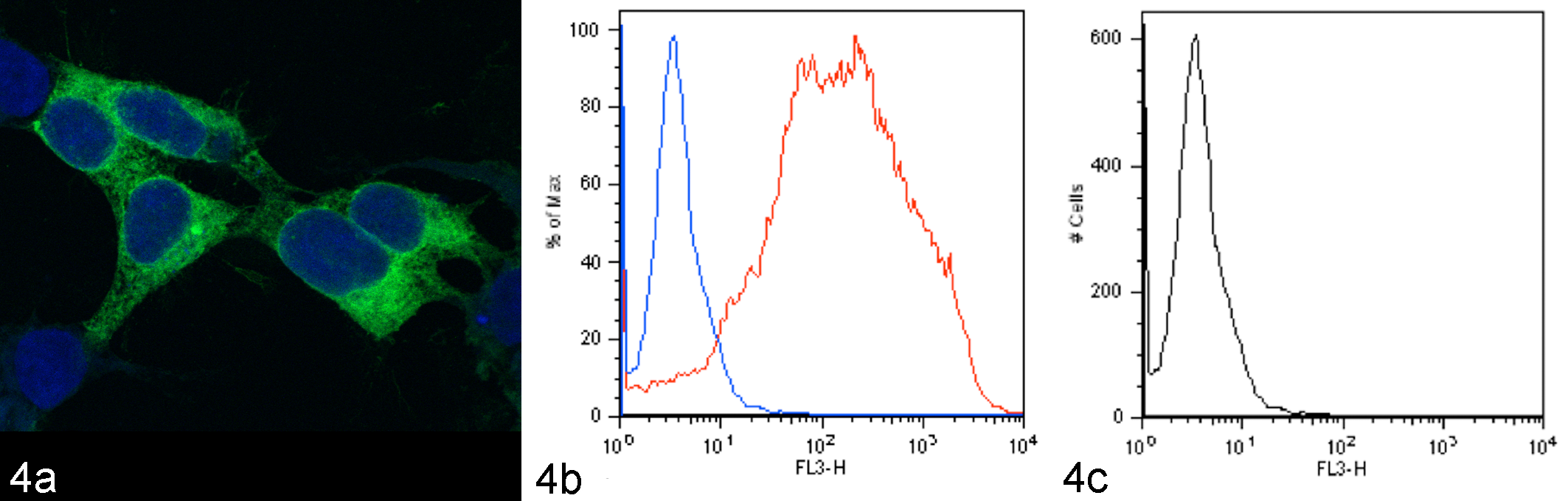
Validation of antibody reactivity against canine CD1a. The canine CD1a gene was cloned into an expression vector expressing green fluorescent protein (GFP), and transfected into human 293T cells. Green fluorescence of transfected cells confirms expression of the recombinant protein (Fig 4a). With flow cytometry, transfected cells labeled with the antibody to canine CD1a have positive fluorescence (Fig. 4b, red line). In contrast, low (negative) fluorescence is present in transfected cells labeled with the negative contol antibody (Fig. 4b, blue line) and in nontransfected cells labeled with the antibody to canine CD1a (Fig. 4c, black line). Credit: Mette Schjaerff, University of Copenhagen; 46 used with permission.
A fourth approach is competitive inhibition or adsorption. In this procedure, antibody is preincubated with an excess of the purified antigen (or adsorbed using antigen-bound beads) before applying it to the tissue section, and the resulting neutralization of the antibody’s binding site is expected to abrogate immunolabeling. 2,48 However, this method may fail to detect cross-reactivity if the non-target protein contains an epitope similar to that of the target protein, or if the antibody has greater affinity for the target antigen compared to the nontarget antigen. 17,53
A fifth approach is in silico comparison of the amino acid sequence of the target protein among species. This can be used to infer cross-reactivity, particularly when the epitope targeted by the antibody is known. However, it does not directly confirm antibody specificity.
Additional approaches include (a) a comparison of immunolabeling patterns using 2 or more antibodies that bind different epitopes of the target protein, (b) a correlation of immunohistochemistry findings with those of another analytical method such as in situ hybridization, ELISA or RT-qPCR, and (c) immunocapture followed by mass spectrometry. 17,53
Assay Validation
In addition to validating the primary antibody, it is necessary to validate the assay as a whole: the combined effectiveness of sample preparation, reagents, buffers, antigen retrieval, blocking, antibody avidity and specificity, batch-to-batch variability of antibodies, incubation conditions, detection platforms, and the ability of individual observers to interpret the findings. This is done by demonstrating positive reactivity in tissue known to express the protein of interest, and negative reactivity in tissue known to not express the protein of interest. Here we take a qualitative approach to assay validation; quantitative measurement of test accuracy, sensitivity and specificity is described elsewhere. 9,13,15
The positive control is used to detect a false negative outcome. For an infectious agent, tissues could be obtained from samples known to be infected or not infected, such as from an experimental challenge study or based on another method such as RT-qPCR. For tumor markers, positive controls are normal tissues known to express the antigen and negative controls are those known to not express the antigen. Cell lines that naturally express the protein or are engineered to do so are alternatives. 19 Ensure that immunolabeling is not only present in the positive control, but that it has the expected extent and distribution of the antigen within the tissue, considering the cell types that are labeled and whether the labeling is nuclear, cytoplasmic, or membranous.
Negative controls identify nonspecific labeling and thereby identify false positives. The first negative control—the technical negative control—uses the positive control tissue but replaces the primary antibody with a non-antigen-specific substitute. If the primary antibody is a monoclonal antibody, the negative control should be an irrelevant monoclonal antibody of the same concentration and isotype. In the same way, purified immunoglobulin is a technical negative control for purified polyclonal antibodies, and preimmune serum or normal serum would be appropriate if immune serum were used as the primary antibody. Omission of the primary antibody is often used as a negative control for diagnostic case material once the method has been well-validated and is in routine use. However, in our opinion, this is insufficient for assay validation or in assays that have some nonspecific background staining. It is important to note that the technical negative control does not assess nonspecific labeling with the primary antibody; if the antibody cross-reacted with an unrelated antigen, the staining would be abrogated by replacement or omission of the primary antibody. Additional technical negative controls are useful for troubleshooting problems when validating a new assay.
The second negative control—the biologic negative control—should normally include both lesional tissue samples as well as normal tissues. Sometimes, a single organ may contain different tissues that can serve as positive and negative control (such as epidermis adjacent to a tumor). Tissues from 10 positive and 10 negative individuals have been suggested as a minimum for analytic validation and can be inexpensively analyzed in tissue microarrays or multitissue blocks. 13 Tissues used for validation must be processed in the same way as would be done for clinical specimens, including fixation and decalcification.
These guidelines apply to qualitative analysis of the presence or absence of immunolabeling, and do not measure test sensitivity and specificity in an epidemiologic context. Furthermore, additional validation is needed for quantitative immunohistochemical analysis, if the goal is to compare the immunolabeling scores with pathogen load, or to compare the findings of two different antibodies.
Quality Control
When using a validated assay to analyze samples from the study, one aspect of quality control is to ensure the immunohistochemistry procedures remain effective. These use the same approaches as described above and can be considered as external positive and negative controls.
The second aspect of quality control is to provide sample-specific validation of every specimen, and can be considered as internal positive and negative controls. False negative findings might result from a problem with an individual sample such as prolonged fixation or inadequate fixation, excessive decalcification, prolonged exposure to air or light, absence of the lesion within the sample, necrosis of the tissue, or errors in sample processing. Such problems are ideally identified by use of an internal positive control, such as a tumor sample with the expected positive labeling of adjacent normal tissue. Alternatively, positive immunolabeling of a different antigen provides some reassurance that the sample is suitable for analysis. However, antigens differ in their susceptibility to the above problems, so positive labeling of one antigen does not rule out a false negative test for the antigen of interest. Knowledge of how the samples were processed can also be helpful in this regard.
Conversely, false positive findings represent nonspecific staining and are detected by an internal negative control (labeling of an area of tissue not expected to contain the antigen; Fig. 5), and second by applying an isotype-matched irrelevant antibody to a different section of the same sample.
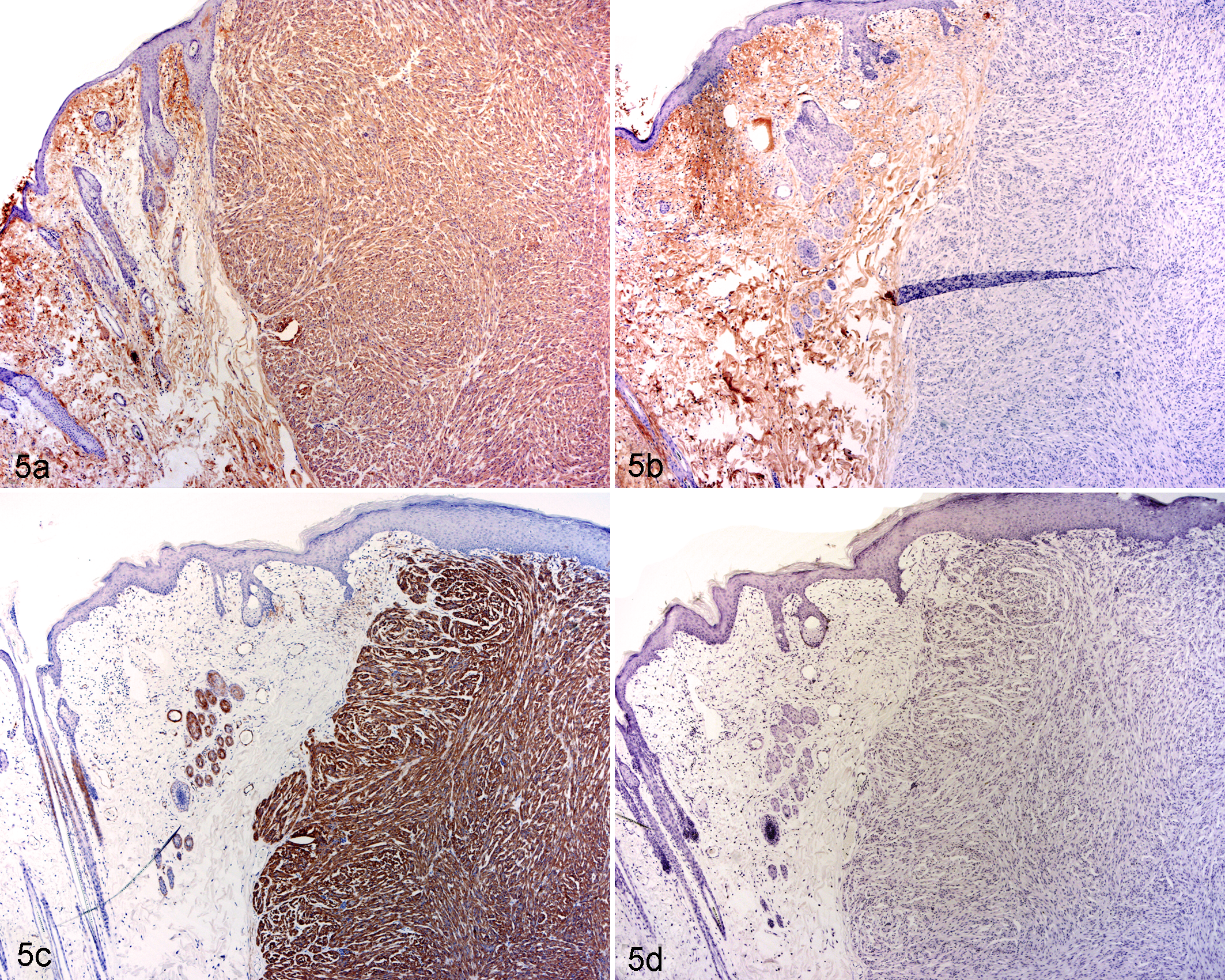
Leiomyoma, skin, rabbit. Fig. 5a. Labeling of both the tumor and the adjacent dermis, using an immunohistochemistry assay developed to detect a-smooth muscle actin. Fig. 5b. With omission of the primary antibody (negative control), labeling of the dermis remains strong while labeling of the tumor is abrogated. Review of the procedure identified that the detection system reacted against both anti-rabbit and anti-mouse immunoglobulins (rather than only against mouse immunoglobulin), thus immunolabeling of the dermis presumably represents rabbit immunoglobulin in the dermis. Fig. 5c. Use of an anti-mouse detection system resulted in specific labeling of the tumor as well as smooth muscle in the normal skin. Fig. 5d. With this detection system, labeling was not present in the negative control. Credit: Josepha Delay, University of Guelph; used with permission.
Immunohistochemistry is a wonderful tool for colocalization of lesions and antigens, but careful attention to method validation is essential to avoid misleading results. As discussed below, confirmation of immunohistochemistry findings by a second analytical method can provide additional rigor.
Polymerase Chain Reaction
Assay Validation
While it is true that PCR assays provide more objective data than the subjective interpretations sometimes required for histopathology or immunohistochemistry, effort is nonetheless required to ensure PCR data are valid. A first step is to ensure that the assay is measuring the intended target. For either conventional or quantitative PCR (qPCR), the primer and probe specificity should be confirmed by BLAST search against the target and nontarget species, and by sequencing the amplified products. For quantitative PCR, additional confirmation is provided by analysis of the amplification curve, product melting curve and melting peaks for each reaction (Fig. 6). Use of labeled probes can be valuable to provide additional specificity.
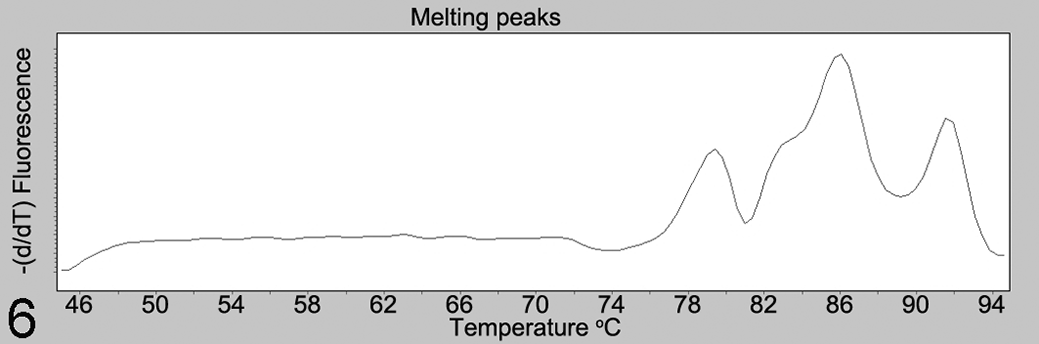
Melting peaks analysis, for quantitative PCR analysis of a fungal pathogen. The melting curve contains 3 peaks at approximately 78.5, 86, and 92°C, instead of the expected single peak. Sequence analysis (data not shown) of amplicons corresponding to the 3 peaks revealed a lack of primer specificity, with amplification of nontarget fungal species.
False-positive outcomes are further avoided by use of negative controls, and these may include 3 forms that address different potential problems. First, using water in place of DNA template from the sample distinguishes positive from negative results. Second, for reverse transcription PCR (RT-PCR), a positive sample processed without reverse transcriptase would detect false positive results caused by sample contamination. Third, cross-reactions should be further ruled out with samples representing alternative disease conditions and using the same sample matrix as clinical samples. 50 Examples include testing an assay for Mannheimia haemolytica against a panel of other bacteria, 18 testing an assay for detection of ovine herpesvirus-2 infection in cattle and sheep against other species of animals infected with closely related herpesviruses, 37 and measuring the sensitivity and specificity of RT-qPCR for bovine respiratory syncytial virus compared to an established assay. 51
Quality Control
Following validation of the method, quality control is required in each run of the analysis to ensure valid results. Negative controls may include water in place of DNA template, as well as a positive sample processed without reverse transcriptase as described above. False-positive results due to cross-contamination were prevented in 1 investigation by using a new microtome blade for each sample, and forceps cleaned with detergent and alcohol. 12
The positive control—analysis of a known positive sample or a synthesized nucleic acid target—would detect a general failure of the assay. However, this would not detect sample-specific false negative results such as those caused by sample degradation, poor yield or poor quality of nucleic acid, inadequate deparaffinization, the presence of PCR inhibitors, or polymorphisms or splice variants at primer binding sites. Thus, nucleic acid integrity and purity should be measured at least for a subset of samples. Additional sample-specific quality control measures were previously illustrated for an assay detecting fungal nucleic acid: a conserved mammalian gene was also amplified as a sample-specific positive control to ensure sample quality, and all test-negative samples were spiked with positive control DNA to investigate the possibility of PCR inhibitors. 28
For quantitative PCR, additional standard quality control procedures include a dilution series (standard curve) of the positive control, establishing the crossing point at which samples are considered reliably positive, documentation that values for samples fall within the dynamic range of the assay, analysis of amplification efficiency for each gene targeted, consideration of the method used to normalize data obtained from different runs, establishing the repeatability and reproducibility of the assay, and validation of reference (“housekeeping”) genes including changes induced by the stimulus being studied. Readers are further directed to technical guidelines for analysis of gene expression (miqe.gene-quantification.info).
Quantitative Assessment and Statistical Analysis
In general, for analytic studies, quantitative or semiquantitative assessment of pathologic findings is preferred over a descriptive narrative, because it allows statistical analysis of the data and formal comparison of study groups. As an example, cluster analysis of multiple quantified parameters was used for unbiased analysis of glomerular diseases in dogs, which defined and validated their morphologic categorization as membranoproliferative and membranous patterns of immune complex–mediated glomerulonephritis, glomerular amyloidosis, glomerulosclerosis, and normal. 8 When a difference between two groups is identified, the level of certainty beyond chance is estimated by the P value. However, the statistical analysis should not end with the P value. As important, the magnitude of the difference between study groups is summarized by the risk ratio or odds ratio for categorical data or the mean difference between groups for continuous data. Finally, the study data are only an estimate of the truth: the 95% confidence interval considers the variability of the data and the sample size to estimate the range in which the actual (true) parameter likely lies, and is therefore essential for valid interpretation of the results.
Given the effort put into designing and carrying out an observational study, why risk a deceiving outcome caused by inappropriate statistical analysis? Seek professional help in the planning stage to ensure an effective study design, and after the study is completed to have confidence in the analysis. Common pitfalls in statistical analysis are discussed in a commentary in Veterinary Pathology. 31 Readers are also directed to the “Statistics Simplified” series published in 2011 in the Journal of the American Veterinary Medical Association.
Approaches to Increased Rigor
Rigor involves an approach to conducting research that ensures the findings reflect the truth. Casadevall and Fang described 5 pillars of scientific rigor: redundancy in experimental design, sound statistical analysis, recognition of error, avoidance of logical traps, and intellectual honesty. 5 How can we achieve rigor? First, examine your motivations for conducting the study. Surely, a passion for finding the truth motivates us in these endeavors. How far are you willing to go to bring rigor to your investigations and ensure the study outcomes are valid?
One way is to replicate the technical analyses on different occasions. Especially for unexpected or pivotal findings, key analyses should be repeated on a different day, ideally using different samples from the same study subjects. This is done routinely for in vitro experimental studies, but may be impossible for observational case-based studies.
Another way is to use different methods to show the same finding, an approach known as triangulation. 34 Particularly for unexpected outcomes, choose complementary methods that address a different aspect of biology and therefore greatly increase the confidence in the findings. Probe the research question from different angles for “validation of tissue pathobiology” 16 by using combinations of morphologic and molecular pathology, clinical biochemistry, clinical observations, diagnostic imaging, epidemiologic analyses, and microbiology assays, as examples. For example, a study used immunohistochemistry, polymerase chain reaction and in situ hybridization as redundant methods to demonstrate canine papillomavirus within pigmented plaques that progressed to squamous cell carcinoma. 26 Imaging can be used to corroborate the histopathologic findings. For example, X-ray fluorescence microscopy and Raman spectroscopy were used to characterize metal alloy and oxalate crystals within tissues, respectively. 11,33 Bone metastases were verified by both histopathology and computed tomography. 7 Just as you wouldn’t invest retirement savings in a single stock, we shouldn’t base our understanding of a disease on a single methodology.
Critically consider how the study findings concur and differ from those of other investigators. Often this is done superficially with the simple goal of justifying the current results. A more enlightened approach, in those situations where the prior study design is indeed comparable to the present one, is a detailed analysis filled with sparkling insights on what the discrepancies suggest about the true biology of the disease, with additional analyses to investigate these novel possibilities.
The same investigators or ideally an independent group should confirm the results on a second population of study subjects. This is essential for major findings that change routine practice. Cancer grading schemes are an example: a new grading scheme is not rock solid until it has been validated in a second population of study subjects (see Hill’s criterion of consistency 6 ). We have illustrative examples of this in veterinary pathology. 4,21,35,39,43,47 Generally, we as journal editors expect that such confirmatory studies not only corroborate or refute the outcome of the original study, but also include additional results that add novelty to the new investigation.
Finally, develop a mindset of critical analysis and skeptical interpretation of your own findings. Carefully probe a variety of alternative explanations. Consider the possibility of bias and how it might impact the findings. Don’t ignore troublesome details of the data that seem not to make sense. Niggling problems are often a window that can be opened to reveal a new landscape of truth.
Conclusions
The methods that veterinary pathologists use in observational studies are key to advancing knowledge of animal disease. Yet, the most-used methods in pathology are somewhat subjective, inherently variable, not easily quantitated, and susceptible to technical error. Careful attention to validation, quality control, methodologic rigor, and critical analysis of the results are essential for rigorous advancement of knowledge in veterinary pathology.
Footnotes
Acknowledgements
We thank Josepha Delay, Stefan Keller, Doran Kirkbright, Celina Osakowicz, Siobhan O’Sullivan, Lauren Sergejewich, and Geoffrey Wood for their contributions.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Some authors of this commentary are editors of Veterinary Pathology. This editorial commentary was not peer reviewed.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Some aspects of this work were supported by a grant from the Natural Sciences and Engineering Research Council of Canada (RGPIN-2017-03872, J. Caswell).
