Abstract
The accurate distinction of reactive and neoplastic lymphoid proliferations can present challenges. Given the different prognoses and treatment strategies, a correct diagnosis is crucial. Molecular clonality assays assess rearranged lymphocyte antigen receptor gene diversity and can help differentiate reactive from neoplastic lymphoid proliferations. Molecular clonality assays are commonly used to assess atypical, mixed, or mature lymphoid proliferations; small tissue fragments that lack architecture; and fluid samples. In addition, clonality testing can be utilized to track neoplastic clones over time or across anatomic sites. Molecular clonality assays are not stand-alone tests but useful adjuncts that follow clinical, morphologic, and immunophenotypic assessment. Even though clonality testing provides valuable information in a variety of situations, the complexities and pitfalls of this method, as well as its dependency on the experience of the interpreter, are often understated. In addition, a lack of standardized terminology, laboratory practices, and interpretational guidelines hinders the reproducibility of clonality testing across laboratories in veterinary medicine. The objectives of this review are twofold. First, the review is intended to familiarize the diagnostic pathologist or interested clinician with the concepts, potential pitfalls, and limitations of clonality testing. Second, the review strives to provide a basis for future harmonization of clonality testing in veterinary medicine by providing diagnostic guidelines.
Keywords
The Purpose of Clonality Assays
The accurate distinction of reactive and neoplastic lymphoid proliferations can present challenges. Given the different prognoses and treatment strategies, a correct diagnosis is crucial. Molecular clonality assays assess lymphocyte antigen receptor gene rearrangement diversity and can help differentiate reactive from neoplastic lymphoid proliferations if microscopic evaluation is inconclusive. Molecular clonality assays are commonly used to assess atypical, mixed, or mature lymphoid proliferations; small tissue fragments that lack architecture; cell smears; and fluid samples such as cerebrospinal fluid. In addition, clonality testing can be utilized to track neoplastic clones over time or across anatomic sites. Molecular clonality assays are not stand-alone tests but useful adjuncts that follow clinical, morphologic, and immunophenotypic assessment. Clonality testing should not be used as a primary lineage assignment tool, that is, as a surrogate for immunophenotyping. 51 Clonality testing is unnecessary in obvious lymphoid neoplasms. Clonality assays are species specific and in veterinary medicine, currently exist for dogs, cats, and horses.
The Objectives of the Review
Even though clonality testing provides valuable information in a variety of situations, the complexities and pitfalls of this method, as well as its dependency on the experience of the interpreter, are often understated. In addition, a lack of standardized terminology, laboratory practices, and interpretational guidelines hinders the reproducibility of clonality testing across laboratories in veterinary medicine. The objectives of this review are twofold. First, the review is intended to familiarize the diagnostic pathologist or interested clinician with concepts and potential pitfalls and limitations of clonality testing. Second, the review strives to provide a basis for future harmonization of clonality testing in veterinary medicine, influenced by the Biomed-2/EuroClonality (B2/EC) guidelines and adapted for use in animals. 26 Important, frequently used terms are italicized in the text and defined and further elaborated in the Glossary.
The Working Principle of Clonality Assays
Adaptive immunity hinges on specific antigen recognition by lymphocytes. 11 Antigen recognition is facilitated by 2 types of antigen receptors, the B and the T cell receptor (Fig. 1). Antigen receptors are composed of receptor chains, each of which is encoded in a locus, a distinct region of a chromosome. Unlike other genes, lymphocyte antigen receptor genes do not have a constant nucleotide sequence. Instead, a novel composite antigen receptor gene with unique sequence and ligand specificity is created in every lymphocyte through rearrangement of multiple smaller genes (formerly referred to as gene segments). 12,48 Lymphocyte antigen receptor genes are classified into variable (V), diversity (D), joining (J), and constant (C) genes (Fig. 2). During antigen receptor rearrangement, one each of multiple V, D, and J genes are joined by removal of the intervening DNA, and nucleotides are added to or deleted from the ends of the genes that are joined. The resulting junctional region is referred to as complementarity determining region 3 (CDR3). It is unique for every lymphocyte clone and is the major determinant of antigen receptor specificity. Rearrangements can occur on 1 chromosome (mono-allelic rearrangement) or both chromosomes (bi-allelic rearrangement). 26
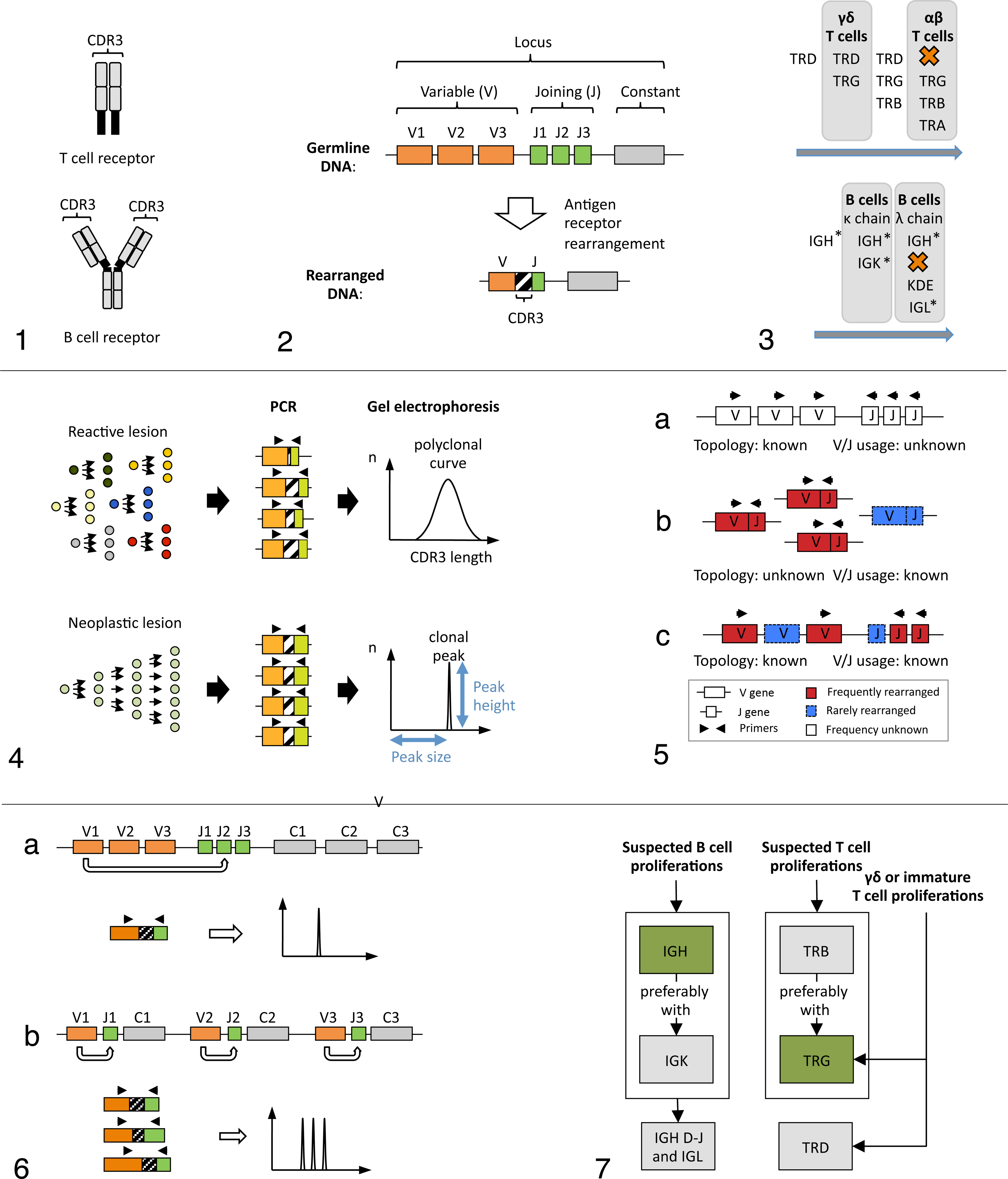
The rearrangement of antigen receptor loci is dependent on the lineage of a lymphocyte and follows a defined sequence (Fig. 3). 42 T cells first rearrange their T cell receptor (TR) delta (TRD) locus followed by their TR gamma (TRG) locus. A T lymphocyte then either assumes a γδ phenotype or proceeds to rearrange its TR beta (TRB) and TR alpha (TRA) loci, resulting in an αβ phenotype. In the latter instance, TRD is deleted on at least 1 chromosome. The rearranged TRG locus is retained in both αβ and γδ T cells, which makes this locus the prime target for T cell clonality assays. B cells first rearrange their immunoglobulin (IG) heavy chain locus (IGH), followed by the IG kappa light chain (IGK) locus. The lymphocyte then either expresses a B cell receptor with kappa light chains or proceeds to rearrange its IG lambda light chain (IGL) locus. In this case, the IGK locus is deleted by rearrangement of the kappa-deleting element (KDE). Since the IGH locus is rearranged in B cells regardless of light chain usage, it is the main target for B cell clonality testing but is subject to somatic hypermutation during immune responses. This leads to reduced sensitivity in clonality assays utilizing IGH. Lambda chain dominant species (eg, canine, feline, and equine) frequently have inactivation of the IGK locus. This occurs via KDE, which can also be targeted for clonality testing. Targeting KDE rearrangements avoids the problem of somatic hypermutation encountered in the IGH locus, since they occur during B cell development, and the locus is silenced prior to antigen encounter.
While lymphocyte antigen receptor gene rearrangements are typically lineage associated, in some instances, B cells rearrange T cell loci or T cells rearrange B cell loci. This phenomenon is referred to as cross-lineage rearrangement. Additionally, myeloid cells may rearrange B or T cell loci. 5,41,45,46 Therefore, lineage assignment based on clonality results is not recommended and is more accurately done by immunophenotyping. 51
Clonality assays visualize the diversity of antigen receptor gene rearrangements across lesional lymphocytes by polymerase chain reaction (PCR) based amplification of the CDR3, followed by gel electrophoretic size separation of amplicons (Fig. 4). CDR3 amplification is done by designing forward primers for V genes and reverse primers for J genes. 51 Amplification is only possible in cells that have rearranged the locus of interest because V and J genes are otherwise too distant. In reactive processes, lymphocytes are derived from multiple cells that differ in CDR3 length, resulting in amplicons of variable size. In neoplastic processes, lymphocytes are derived from a single cell and hence have identical CDR3s resulting in amplicons of identical size. Size separation of amplicons is either done by traditional slab gel electrophoresis or by capillary electrophoresis, a high-resolution variant of traditional slab gels. With capillary electrophoresis, results are output in the form of a histogram with amplicon size on the x-axis and relative fluorescence (corresponding to the abundance of amplicons) on the y-axis. Reactive lesions generate a normally distributed (Gaussian) curve, while neoplastic lesions generate a distinct peak.
Clonality Assays in Veterinary Medicine
When designing clonality assays, identification of all possible rearranged V and J genes is required for optimal sensitivity. PCR-based amplification of a nucleotide sequence relies on the successful annealing of a forward and a reverse primer to a template. If a gene is not covered by any primer of an assay but utilized by a neoplastic clone, the assay will yield a false negative result. There are 2 strategies for identifying all rearranged V and J genes of a locus (Fig. 5). First, genes can be searched for in whole genome sequencing data on a DNA level, preferably in the form of a high-quality genome assembly. This approach additionally identifies the physical localization of genes on a chromosome (topology) but not the frequency at which the genes are rearranged (V/J usage). Second, genes can be identified by high-throughput sequencing of the expressed repertoire at the mRNA level. This approach yields the V/J usage but not the locus topology. Knowledge of the locus topology ensures that all genes are covered by primers, and it is required for the correct interpretation of assay results. Knowledge of V/J usage allows limiting the primer design to genes that are rearranged at an appreciable rate. This reduces the complexity of the primer mix and improves assay performance. For optimal assay design and interpretation of clonality assays, knowledge of both locus topology and V/J usage are therefore crucial.
When PCR-based clonality assays were introduced for the dog and the cat, the respective genome assemblies had not been released and high-throughput sequencing was unavailable. 9,38,50,53,58 To obtain antigen receptor sequence data for primer design, cDNAs were cloned and sequenced. Owing to the laborious nature of this strategy, only a relatively small number of clones could be assessed. As a result, the obtained sequence data were limited, and only the most frequently rearranged genes could be identified.
Improvement of clonality assays subsequently focused on the identification of additional genes to increase primer coverage. Weiss et al 55 –57 and Henrich et al 18 developed clonality assays targeting the feline TRG and IGH locus, respectively. Their assays were based on sequence data obtained from diligent mining of the TRACE archive and sequencing of cDNA clones. Mochizuki and colleagues 36,37 described feline clonality assays that were tailored for use with capillary electrophoresis and were based on additional cDNA sequence data. In dogs, clonality assays have been developed or modified by Yagihara et al, 60 Kisseberth et al, 22 and Gentilini et al. 16 However, all of these assays were based on a limited number of cDNA clones and did not take into account locus topology. As a consequence, false negative results due to inadequate primer coverage were still possible.
The release of genome assemblies for various species within the past decade allowed detailed descriptions of antigen receptor loci and subsequently the design of clonality assays that cover all genes of a locus. In 2009, a complete locus description by Massari and colleagues 34 showed that the canine TRG locus is composed of 7 complete V-J-C clusters (cassettes) as opposed to 1 in humans. Using a multiplex assay that covered all potentially rearranged genes, Keller and Moore 20,21 showed that canine T cells routinely rearrange multiple cassettes, resulting in more than 1 rearrangement per chromosome or 2 rearrangements per lymphocyte (Fig. 6). This finding was novel because more than 2 clonal rearrangements per sample had previously been attributed to more than a single clonal expansion (oligoclonality). Furthermore, it emphasized the importance of knowing the locus topology when developing and interpreting clonality assays.
While coverage of all rearranged genes significantly decreased false negative results, the multiplex TRG assay remains suboptimal. Since the assay did not take into account large-scale V/J usage, it contains primers for genes that are rearranged infrequently. The primer mix is therefore more complex than necessary, likely resulting in less robust amplification. Furthermore, amplification of all possible rearrangements increases the “noise” effect of non-neoplastic lymphocytes. This makes the assay more susceptible to false negative results in lesions with a significant degree of polyclonal background (Keller, Moore, and Vernau, unpublished data). Further modification of this primer set based on high-throughput sequencing data to establish large-scale V/J usage is therefore desirable.
In summary, existing clonality assays in veterinary medicine are based on limited sequence data and largely a lack of consideration of locus topology and hence prone to false negative results. Genome assemblies to elucidate locus topology are increasingly available for all major companion animal species, and high-throughput sequencing has become sufficiently affordable to be used for assessment of V/J usage. Therefore, the requirements for a new generation of improved clonality assays exist.
The Clonality Testing Workflow
The clonality testing workflow can be divided into a laboratory phase that is composed of sample preparation, target selection, PCR amplification, and gel electrophoresis and a subsequent interpretational phase.
Sample Preparation
Clonality testing is generally done on genomic DNA since DNA is more robust than RNA and can be obtained from lower quality sources such as formalin-fixed and paraffin-embedded (FFPE) tissues. 30 Other sample materials include stained or unstained cytological preparations, frozen specimens, and fluids.
In some instances, it may be indicated to selectively target portions of a sample for DNA extraction. A common scenario is exclusion of tissue components in a multi-tissue block that contain large numbers of normal lymphocytes (eg, lymph node), which may obscure the signal from the neoplastic lymphoid population. Another scenario is exclusion of tissue components that don’t contain lymphocytes (eg, kidney) that are combined on the same paraffin block as the tissue of interest. Large amounts of non-lymphoid DNA will lower the relative abundance of lymphoid DNA and hence decrease assay sensitivity. Selective DNA extraction of a sufficient number of sections to provide adequate DNA quantity increases the likelihood for detection of clonality but also requires microscopic screening prior to DNA extraction. In addition, selective DNA extraction may necessitate re-embedding if unmounted paraffin scrolls are used. Otherwise, scrapings from a sufficient number of mounted slides allow exclusion of unwanted tissue components. DNA should be extracted from the same slide or block that is used for microscopic evaluation to allow correlation of morphological, immunophenotypic, and molecular findings.
Assessment of DNA yield and purity is most commonly done using spectrophotometry. This method is sufficiently precise as clonality PCR is robust with respect to small variations in input DNA. The goal for input DNA is 100 ng per 50 microliter reaction.
Insufficient DNA quantity is primarily seen with low cellularity aspirates, small histopathology samples such as pinch biopsies, or as a consequence of technical errors during DNA extraction.
DNA integrity, in contrast, is not evident from yield, and purity measurements and assessment require amplification of a DNA sequence comparable in size to the clonality amplicon. Since this is cost intensive if done for every sample, it is often omitted in routine diagnostics and only done if electrophoresis results are suspicious for poor DNA integrity. The latter can be suspected if amplification is poor despite large numbers of lesional lymphocytes of the respective immunophenotype. Compromised DNA integrity is primarily seen in FFPE samples after prolonged formalin fixation. 30
Target Selection
The target selection is based primarily on the lineage of the suspected neoplastic population. 26 Theoretically, any of the 7 antigen receptor loci that code for B and T cell receptor chains can be targeted for clonality testing. 51 The investigation of multiple loci across lineages will result in the most comprehensive information. However, in veterinary medicine, choices are limited by assay availability and cost. Immunophenotyping prior to clonality analysis is hence essential not only for appropriate target selection but also for correct interpretation of clonality results. 52 For B cell proliferations, all published assays target the IGH locus. Forward primers anneal within conserved regions of the V gene (framework regions; FRs). Framework region 2 and FR3 assays are generally suited for any sample type. Framework region 1 assays are less broadly applicable, as their larger amplicon size frequently precludes amplification from FFPE tissues that may have compromised DNA integrity. 30 Assays targeting different framework regions are run concurrently to provide redundancy for false negative results due to somatic hypermutation, which only occurs in B cells. For T cell proliferations, all published assays target the TRG locus. This is primarily because the locus is rearranged in both αβ and γδ T cells. 19 The absence of somatic hypermutation in T cells obviates the need for multiple primer sets targeting different FRs.
Multiple assays targeting the IGH and TRG loci are available in veterinary medicine. As previously described, these assays were developed at different institutions, employing different sequence data, PCR strategies, and electrophoresis systems. Given the lack of studies that directly compare assay performance, evidence-based assay selection is currently difficult. This is compounded by the lack of a “gold standard” for lymphoma diagnosis.
To provide redundancy for the shortfalls of IGH and TRG assays, additional loci are targeted in human medicine (Fig. 7). At least 2 loci are assessed simultaneously, namely, IGH and IGK for B cell proliferations and TRB and TRG for T cell proliferations. 26 The combination of 2 independent test results has been shown to significantly improve the accuracy of diagnoses. 52 However, clonality assays targeting alternate loci have not been published in veterinary medicine.
PCR Amplification and Gel Electrophoresis
Using conventional PCR, the CDR3 of rearranged antigen receptor genes is amplified to levels that allow visualization by gel electrophoresis. Reactions are run at least in duplicate to facilitate identification of pseudoclonality. 26 For size separation of PCR amplicons, capillary electrophoresis is increasingly replacing traditional polyacrylamide slab gel electrophoresis due to its superior resolution, higher throughput, and ease of use. 4 High-resolution electrophoresis is especially helpful when using assays that target multi-cassette loci due to the frequent occurrence of multiple, often closely spaced clonal peaks. 21 The endpoint of the laboratory phase is a gel electrophoresis profile that is reflective of the CDR3 lengths and hence the diversity of antigen receptor gene rearrangements of lesional lymphocytes (Fig. 4).
Interpretation of Electrophoresis Results
Electrophoresis profiles alone are not pathognomonic of a reactive or a neoplastic process and have to be interpreted in the context of clinical, morphologic, and immunophenotypic data. 26,51 Hence, clonality testing alone is not as definitive as often portrayed and is reliant on the interpreter’s training, experience, and integrative efforts. 27
Electrophoresis profiles are classified into several categories represented by archetypal patterns (Fig. 8). 26 These patterns are also referred to as technical descriptions. In general, a clonal electrophoresis profile is considered indicative of a neoplastic process, and a polyclonal electrophoresis profile is indicative of a reactive process. However, polyclonal results can occur in the presence of neoplasia, and clonal results can occur in the absence of neoplasia. 51 In addition, clones can occur within a polyclonal background resulting in peaks of uncertain significance. The technical descriptions pseudoclonal (PSC) or no specific product (NSP) may be indicative of either a reactive or a neoplastic process among other things. Regardless, both profiles can be very informative if interpreted in the appropriate biological context.
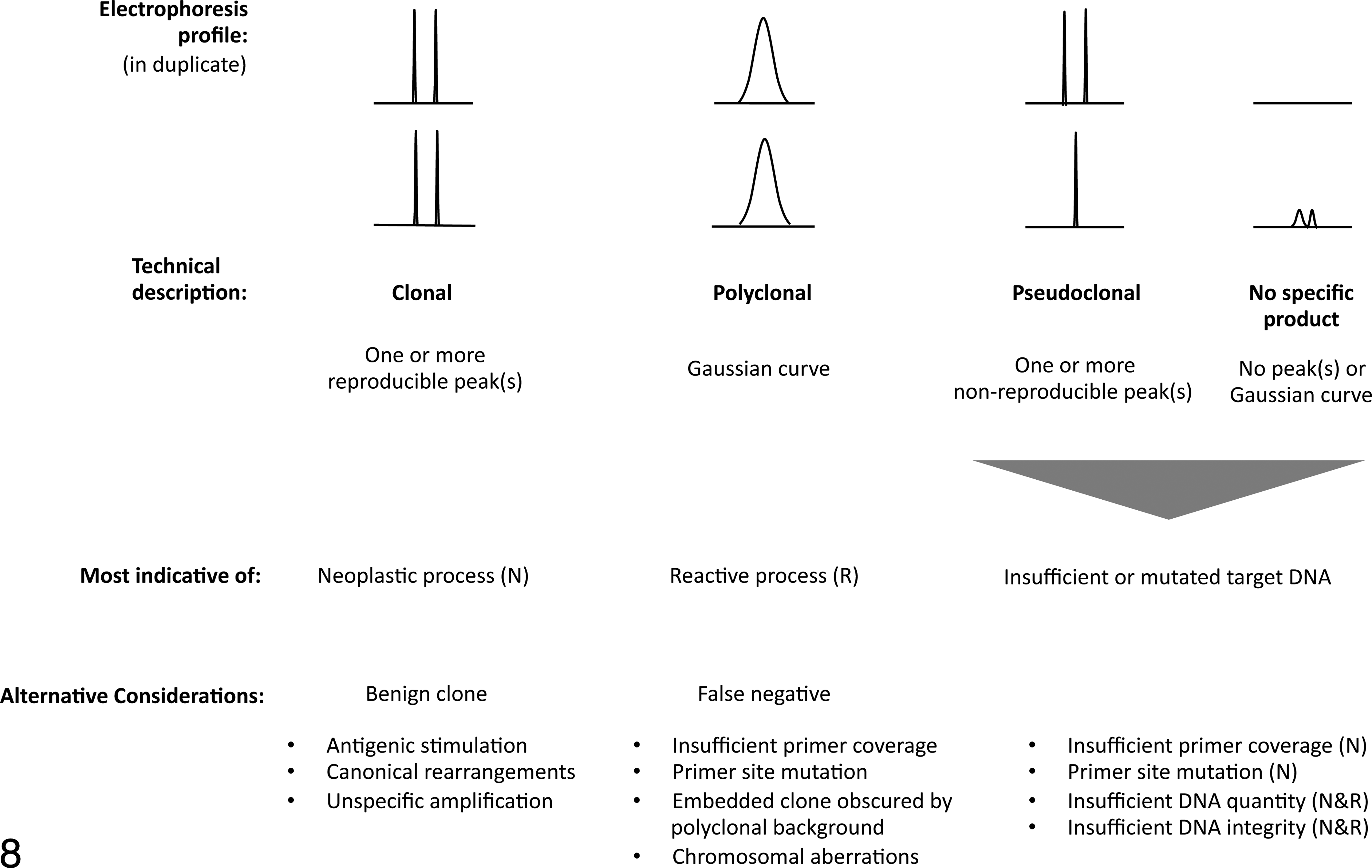
Archetypal electrophoresis patterns, their technical description, interpretation, and interpretational pitfalls.
The correct interpretation of electrophoresis patterns requires the consideration of the following pitfalls: Failure to detect clonality in the presence of neoplasia.
Inadequate primer coverage. Successful amplification of a clonal rearrangement is dependent on the recognition of both the V and the J genes by primers of the assay. If a neoplastic clone utilizes a gene that is not detected by any of the primers in the mix, the assay will yield a false negative result (Fig. 9a). A false negative result due to inadequate primer coverage can be suspected if negative clonality results conflict with clinical, morphologic, and/or immunophenotypic data and the assay is known to have incomplete primer coverage. The prevalence of false negative results due to inadequate primer coverage varies between assays and will decrease over time as existing primer sets are replaced by more comprehensive multiplex primer sets.
Primer site mutation. As part of a normal immune response, B cells increase antibody affinity by mutating nucleotides of antigen receptor genes of B cell loci, a process referred to as somatic hypermutation. Mutations of primer binding sites can impair or preclude priming (Fig. 9b). A false negative result due to primer site mutation can be suspected if negative clonality results are in conflict with clinical, morphologic, and/or immunophenotypic data and the potential neoplastic population is of germinal center or post-germinal center B cell origin. A false negative result due to primer site mutation cannot be completely resolved by improvement of existing primer sets and requires targeting of additional B cell loci to increase sensitivity.
2,15,23,39
Polyclonal background. Most lymphomas contain some proportion of non-neoplastic lymphocytes, either because the tumor arose in a lymphoid organ or occurs in an inflammatory background. If the proportion of non-neoplastic lymphocytes is significant, a clonal signal can be obscured by the noise provided by the polyclonal population (Fig. 9c). A false negative result due to polyclonal background can be suspected if negative clonality results conflict with clinical, morphologic, and/or immunophenotypic data and significant numbers of non-neoplastic lymphocytes are present.
Chromosomal aberrations. Chromosomal aberrations such as translocations can eliminate or displace antigen receptor genes precluding their amplification. Well-known examples in humans are t(11;14) and t(14;18) translocations in mature B cell proliferations. Although chromosomal aberrations have been documented in lymphoid neoplasms in animals, their effect on clonality testing has not been investigated.
7,59
Additionally, the possibility of an amplifiable rearrangement in a second allele may reduce the incidence of this cause of false negative results. Detection of clonality in the absence of neoplasia (benign clonal expansion). Identification of benign clones cannot be done based on the electrophoresis profile but requires integration of clinical, morphologic, and immunophenotypic data. Benign clones can be seen in the following circumstances:
Clonal expansions in response to antigenic stimulation. Following antigenic stimulation, lymphocytes undergo clonal expansion as part of a normal immune response. In most cases, this will affect numerous lymphocyte clones with variable CDR3 lengths resulting in a polyclonal electrophoresis profile. However, in some instances, few clones can proliferate disproportionally and appear as distinct peaks over a variably distinct polyclonal background. In humans, this phenomenon has been documented in response to infectious agents, drug responses, neoplasia, or autoimmune diseases.
1,6,33
In veterinary medicine, benign clonal expansions are suspected but rarely documented. Conditions that have been associated with benign clones are regressing canine cutaneous histiocytoma (Gericota, Affolter, Kass and Moore, manuscript in preparation), hepatitis associated with drug hypersensitivity reaction (Moore, personal observation, single case), Ehrlichiosis, and Leishmaniasis.
9,16
Canonical rearrangements. In humans, certain γδ T cell clones utilize V/J combinations without significant CDR3 nucleotide additions, resulting in a minimally diversified CDR3.
8,13
These rearrangements are referred to as canonical rearrangements and can mimic a clonal population in a polyclonal background. Canonical rearrangements have not been investigated systematically in domestic animals. However, V/J usage assessment of canine T cells by high-throughput sequencing showed a minimally diversified CDR3 for Vγ5-2/Jγ6-1 rearrangements (Keller and Moore, unpublished data) (Fig. 10). The significance of this finding for clonality testing in dogs remains to be determined.
Unspecific amplification. Amplification of DNA sequences other than rearranged antigen receptor genes is referred to as unspecific amplification. The peak size can be within or outside the expected size range.
25
Unspecific amplification is more likely to occur in samples with low target DNA because unspecific targets are usually outcompeted by specific targets if present.
26
Unspecific amplification can be suspected if clones of identical size recur in multiple samples over time, especially if they are outside the expected target size range. In human medicine, a list of unspecific targets has been compiled by the B2/EC initiative.
26
Given the lack of standardized primer sets in veterinary medicine, the identification of nonspecific amplifications is reliant on the interpreter’s experience with a specific assay. Finally, it should be noted that a clonal peak outside the expected size range can represent specific amplification (containing insertions or deletions) but should be sequence-confirmed to avoid false positive diagnoses.
43
Peak(s) of uncertain significance. Electrophoresis profiles may be mixtures of archetypal patterns or patterns that do not fit any established category. An interpretational challenge is an electrophoresis profile with 1 or more peaks in a Gaussian curve (polyclonal with minor clones). This can be indicative of a neoplastic process with a concomitant reactive component, or it could represent a dominant immune response following antigenic stimulation. Differentiation of neoplastic from reactive clones has been attempted based on the relative height of a peak (absolute peak height divided by the height of the polyclonal curve).
24,28,32,44
However, the relative peak height of a neoplastic clone is primarily determined by the abundance of lesional reactive lymphocytes and the location of the peak respective to the polyclonal population (Fig. 11), both of which are unique for every lesion. In addition, the relative height of a peak is influenced by assay properties such as differences in amplification efficiencies between V/J primer combinations and cycling conditions. Simply put, a neoplastic clone might have a small relative peak height, and a reactive clone might have a large relative peak height. In our opinion, it is therefore pointless to identify neoplastic clones solely based on relative peak height criteria. The rejection of cutoff values is in line with the B2/EC initiative’s perception that “cutoff values … create a false sense of accuracy and might even lead to false- positive or false-negative interpretation.”
26
While it is desirable to minimize the subjectivity of clonality testing, we believe that interpretation of clonality results cannot be accomplished by mathematical algorithms but requires consideration of clinical, morphologic, and immunophenotypic data. Pseudoclonal profiles/no specific product. These electrophoresis profiles may be indicative of a neoplastic or a reactive process and result from either a lack of primers covering the rearranged genes, mutation of primer binding sites, or a paucity of target DNA (the proportion of input DNA available for amplification). Antigen receptor genes are unique in that PCR amplifiable target DNA sequences do not exist unless a cell has rearranged the locus of interest. Even though the total amount of input DNA per PCR reaction is standardized, the amount of target DNA is dependent on the cellular composition of a sample (Fig. 12). The 2 main reasons for insufficient target DNA are primer site mutation during somatic hypermutation of B cell loci and low numbers of lesional B/T cells. In both cases, total DNA concentration and purity measurements may appear adequate. To interpret PSC/NSP profiles, it is essential to assess the abundance of lesional lymphocytes of the respective immunophenotype, namely, the abundance of B cells when interpreting IGH profiles and the abundance of T cells when interpreting TRG profiles. For example, a lesion with a PSC/NSP IGH electrophoresis profile containing low numbers of B cells is likely not a B cell lymphoma. However, there are exceptions such as T cell–rich B cell lymphoma, in which reactive T cells outnumber B cells, or in situations of B cell antigen loss and further V gene mutation with lymphoma progression. If a lesion contains abundant B or T cells, a PSC/NSP result can be attributed to multiple reasons with different implications for result interpretation. First, poor amplification can be due to insufficient total DNA quantity or integrity, which can be identified by DNA quantification and amplification of a control gene, respectively. In this case, the clonality results are not interpretable. If total DNA quantity and integrity are sufficient, a PSC/NSP profile may actually be suggestive of a neoplastic process. This is most commonly seen in B cell neoplasms due to mutation of primer sites during somatic hypermutation or in B or T cell neoplasms due to inadequate primer coverage.
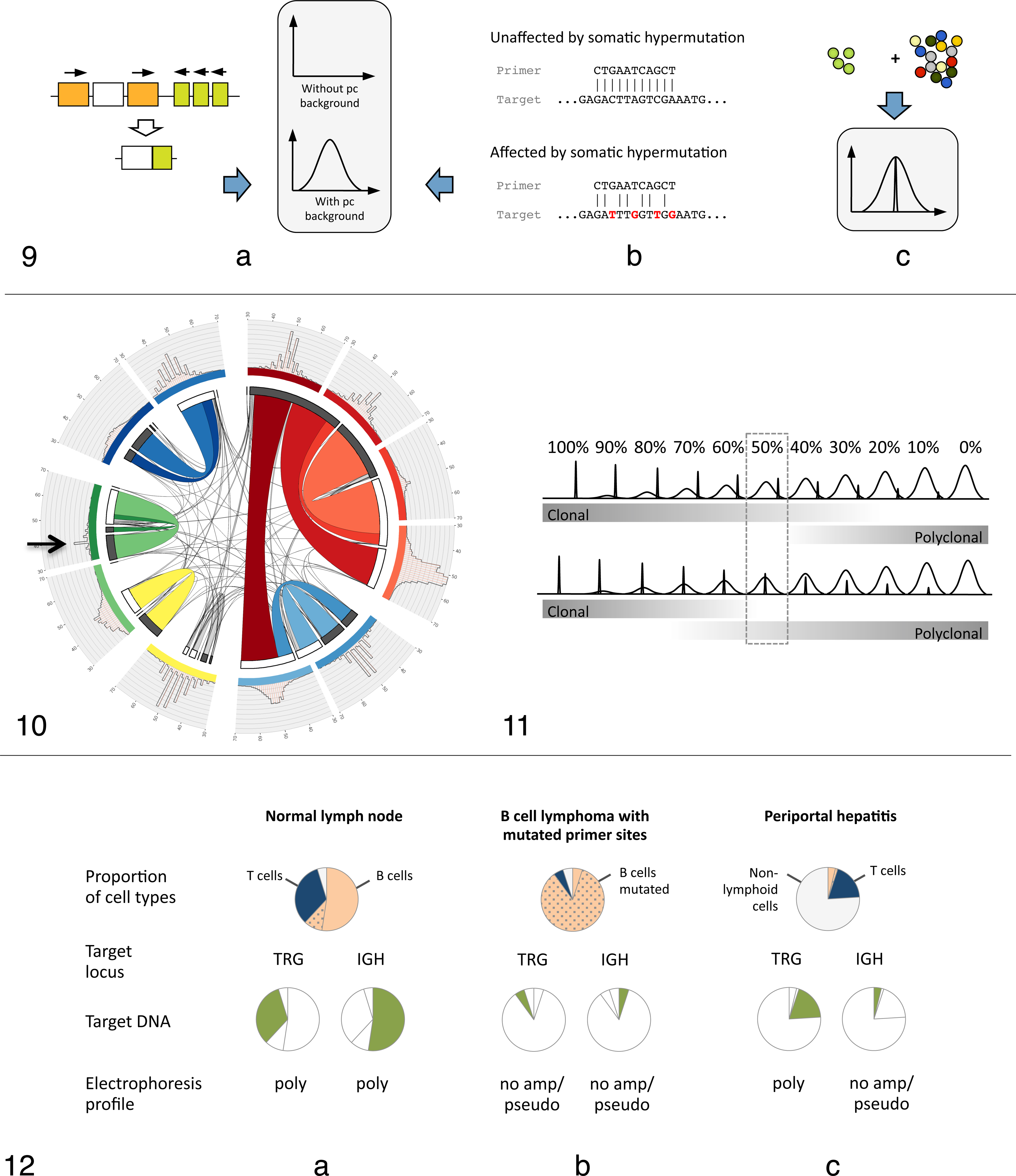
The Importance of Integrating Clinical, Morphologic, and Immunophenotypic Data
The integration of clinical, morphologic, and immunophenotypic data is required during multiple steps of the clonality workflow and is essential to minimize misdiagnoses. Specifically, it is needed for assessment of sample suitability, target selection, adequate interpretation of electrophoresis profiles, and lineage determination. 17 These principles are illustrated in the following with 6 case examples.
Microscopic lesion assessment and immunophenotyping prior to clonality testing is required for appropriate target selection and may facilitate identification of the tumor type. 49,52 Samples requiring selective DNA extraction will also be identified. For example, in Fig. 13, a section of feline intestine intended for clonality assessment has been embedded with a section of mesenteric lymph node. DNA extracted from the entire section was polyclonal for TRG, but selective DNA extraction of intestinal tissue yielded a clonal TRG result indicative of T cell lymphoma, which was supported by histopathology. Clearly, the presence of lymph node on the slide overwhelmed the system with polyclonal DNA, thereby obscuring the presence of a clonal T cell expansion.
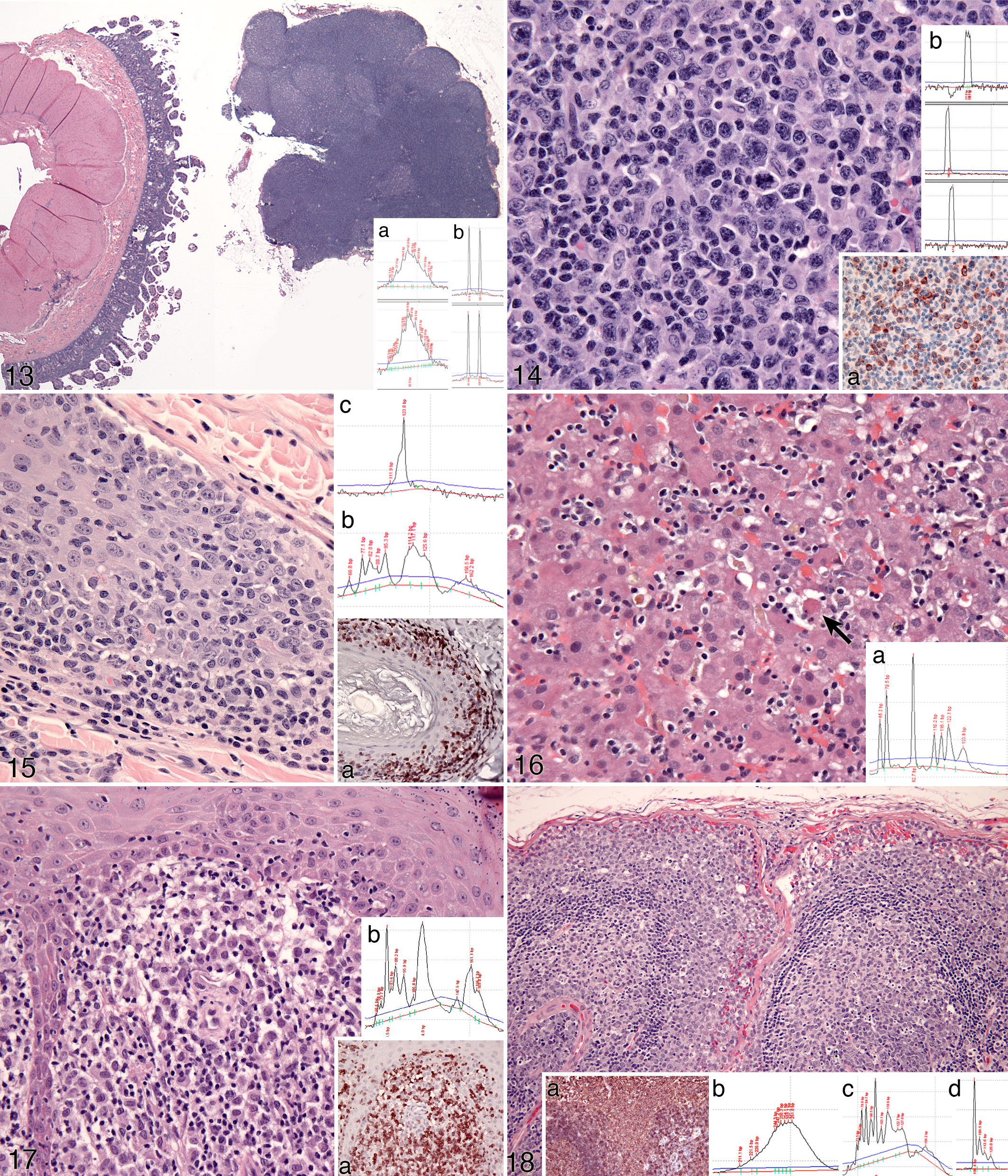
Immunophenotyping is also potentially useful for explaining a pseudoclonal PCR result. Somatic hypermutation of IGH V genes is common in B cell neoplasia. This may lead to failure of primer annealing to target tumor DNA, resulting in either no amplification or pseudoclonality (Fig. 14). There were adequate numbers of B cells in the lesion, which, in conjunction with their morphology, lead to an interpretation of large B cell lymphoma with somatic hypermutation, or use of a V gene not recognized by the primer, as a cause of the pseudoclonal PCR result.
The integration of adjunctive data is also required to identify false negative or false positive clonality results. A common cause of false negative clonality is marked polyclonal background. Polyclonal background can be recognized by a mixed cellular infiltrate, architectural cues such as follicle formation, or the topological distribution of B and T cells as determined by immunophenotyping. In Fig. 15, a skin biopsy from a dog is suspicious for epitheliotropic T cell lymphoma based on histopathology and immunohistochemistry. TRG clonality was polyclonal, but assessment of a complementary T cell locus (TRD) produced a clonal result. TRD is less susceptible to polyclonal background in this instance. The final diagnosis was inflamed epitheliotropic T cell lymphoma.
False positive clonality results are potentially more catastrophic. Benign clonal expansions have been encountered in a limited set of circumstances (already mentioned), which are expected to expand. “Pseudolymphoma” associated with drug hypersensitivity was observed in a dog with Rimadyl administration. This dog developed a lymphocytic hepatic infiltrate and elevated liver enzymes (Fig. 16); molecular clonality assessment revealed clonal TRG. Following drug withdrawal, clinical parameters normalized. Another example is regressing histiocytoma, which is associated with heavy lymphocytic infiltrates dominated by T cells (Fig. 17). In advanced regression, very few neoplastic histiocytes remain, and these lesions have been confused with lymphoma. Molecular clonality assessment of TRG is often clonal, further confounding the diagnosis (Fig. 17). This is believed to be a focused immune response to the neoplastic histiocytes (Gericota, Affolter, Kass and Moore, manuscript in preparation). Consideration of clinical data and follow-up are important in resolving these cases.
Finally, immunophenotyping is essential for lineage determination of a neoplastic population. The most frequently encountered situation in which clonality results are not reflective of a cell’s true immunophenotype is cross-lineage rearrangements of TRG in diffuse large B cell lymphomas (Fig. 18) or plasma cell neoplasms. IGH clonality may not be detected due to inadequate primer coverage, mutation of primer binding sites, excessive polyclonal background, or a combination thereof. Assessment of a complementary B cell locus unaffected by somatic hypermutation (eg, KDE) may provide valuable insight (Fig. 18). Conversely, immunophenotyping may fail due to pan-B or pan-T cell antigen loss. In these instances, antigen receptor gene rearrangement may be useful in detecting these events.
Future Perspectives
Clonality testing in veterinary medicine today is in many ways comparable to the situation in human medicine in the early 1990s. Back then, clonality assays were routinely available for IGH and TRG loci, and false negative results due to inadequate primer coverage were common. 26 In addition, harmonization of primer sets, laboratory standards, and result interpretation did not exist. Since then, clonality testing in human medicine has evolved into an effective, standardized, and broadly accepted tool for the assessment of lymphoid proliferations. 27 The progression of clonality testing in human medicine, especially the B2/EC initiative, can serve as a model for the improvement of clonality testing in veterinary medicine.
The basis for improvement of clonality testing in human medicine was to increase assay sensitivity by development of sophisticated multiplex primer sets that covered all possible rearranged genes. 51 In veterinary medicine, inadequate primer coverage is prevalent because whole genome assemblies were frequently unavailable when assays were developed and sequencing of individual clones only allowed limited assessment of V/J usage. 21 Today, genome assemblies are available for various species of interest, and high-throughput sequencing facilitates large-scale assessment of V/J usage. 14,31,40,54 In addition, genome assemblies allow identification of genes in-silico, which provides the sequence data required for primer design, and the locus structure required for adequate interpretation. Several immunoglobulin and T cell receptor loci of veterinary interest have been fully described, and sequence data for some are available through a public database (IMGT). 3,29,34,35 However, most loci comprise a large number of genes requiring more primers than is feasible for multiplexing in a single tube. Knowledge of the V/J usage informs primer design because non-rearranged genes don’t have to be considered (Fig. 10). This should result in a new generation of primer sets with increased sensitivity.
Improved clonality testing in human medicine resulted from expanding the range of targeted loci to include TRB, TRD, and IGL among others, to compensate for the diminished sensitivity of individual assays. 26,52 In veterinary medicine, assays for additional targets have not been published, and routine assessment of multiple loci is hampered by financial considerations. However, additional assays would be desirable to follow up inconclusive clonality results. Clonality testing beyond the standard IGH and TRG loci could be determined on a case-by-case basis, weighing the need and benefit for additional information against its cost. High-throughput sequencing analysis of V/J usage of all canine T cell receptor loci has facilitated the development of a TRD multiplex assay (Keller and Moore, manuscript in preparation), and an assay targeting the kappa-deleting element (KDE) for dogs, cats, and horses is undergoing testing (Moore and Keller, unpublished data). This assay is not affected by V gene mutation and can yield valuable information if IGH clonality testing fails.
Lastly, a major achievement in human medicine was the harmonization of clonality testing across institutions. A major impetus for this development was the B2/EC initiative, which brought together 47 institutions from 7 countries to establish standardized primer sets and testing guidelines. 51 A similar initiative in veterinary medicine would be desirable but would face multiple challenges. First, antigen receptor gene sequences differ between species and hence require specific primer sets for every species of interest. The effort associated with harmonizing primer sets would therefore be considerably greater than in human medicine. Second, there is a wide range of laboratory practices, primer sets, and technical equipment used across various institutions. 9,10,16,18,21,37,47,53,60 Third, a comprehensive, multi-institutional initiative analogous to the B2/EC initiative would require significant funding. Given these considerations, any multi-institutional study in veterinary medicine will realistically be of smaller scale, require careful planning, and take years to complete.
While the harmonization of primer sets and laboratory practices in veterinary medicine faces considerable challenges, general principles of clonality testing, as discussed in this review, could be harmonized immediately and without the need for multi-institutional studies. The body of literature generated over 2 decades of clonality testing in human and veterinary medicine have highlighted general principles of clonality testing that apply to all species. These general principles include recognition of archetypal electrophoresis patterns and their interpretation, basic laboratory quality control measures, and interpretational pitfalls (Fig. 19). Universal adoption of these principles of clonality testing would allow focusing on unification of those aspects of clonality testing that are species and/or laboratory specific. This would accelerate the improvement of clonality testing in veterinary medicine.
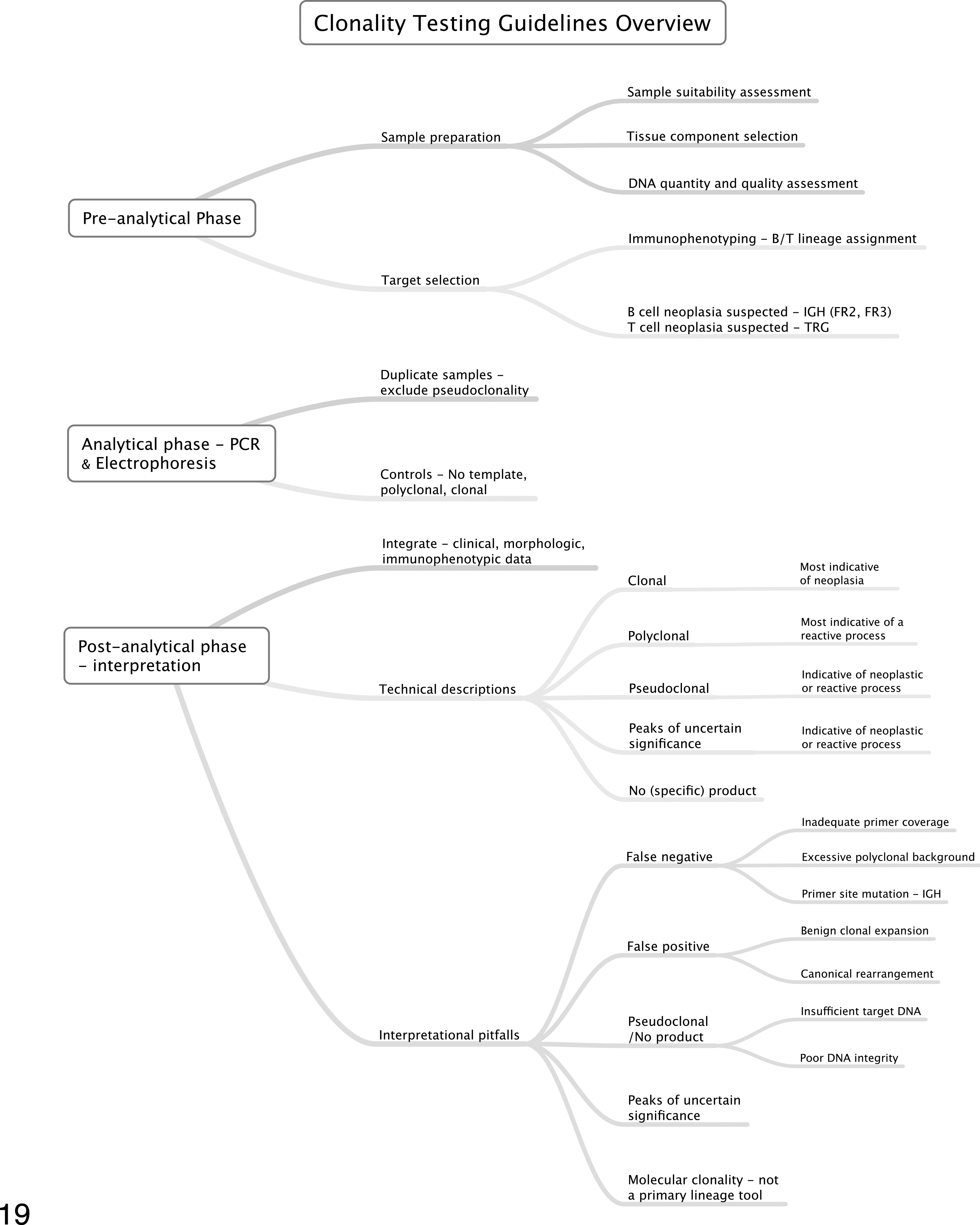
Proposed guidelines for clonality testing in veterinary medicine. Importantly, molecular clonality testing is an adjunctive procedure that requires integration of clinical, morphologic, and immunophenotypic data to produce biologically meaningful results. The flow diagram illustrates the major phases of clonality testing in a skeletonized format.
Footnotes
Glossary
Acknowledgements
We would like to thank the following people for their input on the proposed guidelines: Makoto Bonkobara, University of Tokyo, Japan; Manfred Henrich, Justus-Liebig-Universität Giessen, Germany; Barbara Rütgen and Sabine Hammer, University of Vienna, Austria; Masamine Takanosu, Nasunogahara Animal Clinic, Japan.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
