Abstract
Bovine papillomaviruses (BPV1/BPV2) have long been associated with equine sarcoids; deciphering their contribution has been difficult due to their ubiquitous presence on skin and in the environment, as well as the lack of decent techniques to interrogate their role in pathogenesis. We have developed and characterized an in situ hybridization (ISH) assay that uses a pool of probes complementary to portions of the E5, E6, and E7 genes. This assay is highly sensitive for direct visualization of viral transcript and nucleic acid in routinely processed histopathologic samples. We demonstrate here the visualization of BPV nucleic acid in 18 of 18 equine sarcoids, whereas no detectable viral DNA was present in 15 of 15 nonsarcoid controls by this technique. In nearly 90% (16/18) of the sarcoids, 50% or more of the fibroblastic cell nuclei distributed throughout the neoplasm had detectable hybridization. In the remaining 2 cases, fewer than half of the fibroblastic cells contained detectable hybridization, but viral nucleic acid was also detected in epithelial cells of the sebaceous glands, hair follicles and epidermis. A sensitive ISH assay is an indispensable addition to the molecular methods used to detect viral nucleic acid in tissue. We have used this technique to determine the specific cellular localization and distribution of BPV in a subset of equine sarcoids.
Equine sarcoids are the most common neoplasm in equids, including horses, donkeys, and mules. 6,17,18,28,30 Clinically, the term refers to a spectrum of locally aggressive nonmetastasizing skin tumors originally described as biphasic tumors with epithelial and mesenchymal components. 18 A viral etiology has been suspected since the original description of sarcoids, 18 and in the subsequent eight decades, considerable evidence has demonstrated that bovine (Bos taurus) papillomavirus (BPV) 1 and BPV2, and rarely BPV13, are associated with the development and maintenance of equine sarcoids. 2,19,22,27
The initial evidence for the role of BPVs in equine sarcoids relied on transmission and hybridization studies. 26,29,34 In subsequent work, Southern blots were used to identify BPV1 or BPV2 genome(s) within tumor tissue, 1,2,19,38 and polymerase chain reaction (PCR)–based surveys routinely demonstrate that up to 100% of sarcoids contain BPV DNA. 12 –14,23,24,27,38,39 A limited number of studies have been designed to investigate the role of BPV in cellular transformation in equine tissue. These studies have identified early and late gene transcripts in at least a subset of naturally occurring sarcoids indicating the initiation of viral transcription and replication. 9,25,36 Further evidence indicates that viral proteins, including E2, 36 E5, 11,36 E7, 11 and L1, 10,11 are also produced in naturally occurring equine sarcoids. While examination of sarcoids for BPV has identified viral DNA, viral transcripts, and even viral proteins, to date there has been no visualization of viral particles in sarcoids, and infection is thus generally considered nonproductive. 1,19 The lack of visualization of virions has perhaps only encouraged scientists to take different approaches to determine if the viral life cycle is occurring. The most recent focus is on studies targeting examination of BPV specifically in the epithelium, where productive infections occur in natural hosts. Laser microdissection and PCR studies have identified BPV DNA in the epidermis in a subset of equine sarcoids, 8,11,37 although at a lower rate than in the dermis 8,37 and at a lower copy number. 8,11 To complement these studies and visualize the location of viral nucleic acid within equine sarcoids, we developed a sensitive colorimetric in situ hybridization (ISH) assay that allows single-molecule visualization by hybridization-based amplification. This allowed for the more precise localization of BPV nucleic acid within equine sarcoids.
Materials and Methods
Cases and Tissues
Retrospective samples were selected from the William R. Pritchard Veterinary Medical Teaching Hospital’s Anatomic Pathology archives at University of California (UC), Davis. Samples were formalin fixed and paraffin embedded for routine histological processing. Table 1 contains a summary of each group of cases and results. Four classic bovine fibropapillomas were selected from the archive as controls for the assay (Suppl. Table S1). From 25 horses, we included 15 nonsarcoid equine tissues and 18 sarcoids (Suppl. Table S2). The sarcoids were classified based on the documented clinical presentation and gross morphology. The nonsarcoid tissues were broadly grouped into 3 categories—hyperplastic and neoplastic lesions (n = 11), inflamed skin (n = 3), and normal skin (n = 1)—and then further classified by lesion (Suppl. Table S2).
Case Information and Detection of Bovine and Equine Papillomaviruses in Bovine Fibropapillomas, Equine Sarcoids, and Equine Nonsarcoid Lesions.a
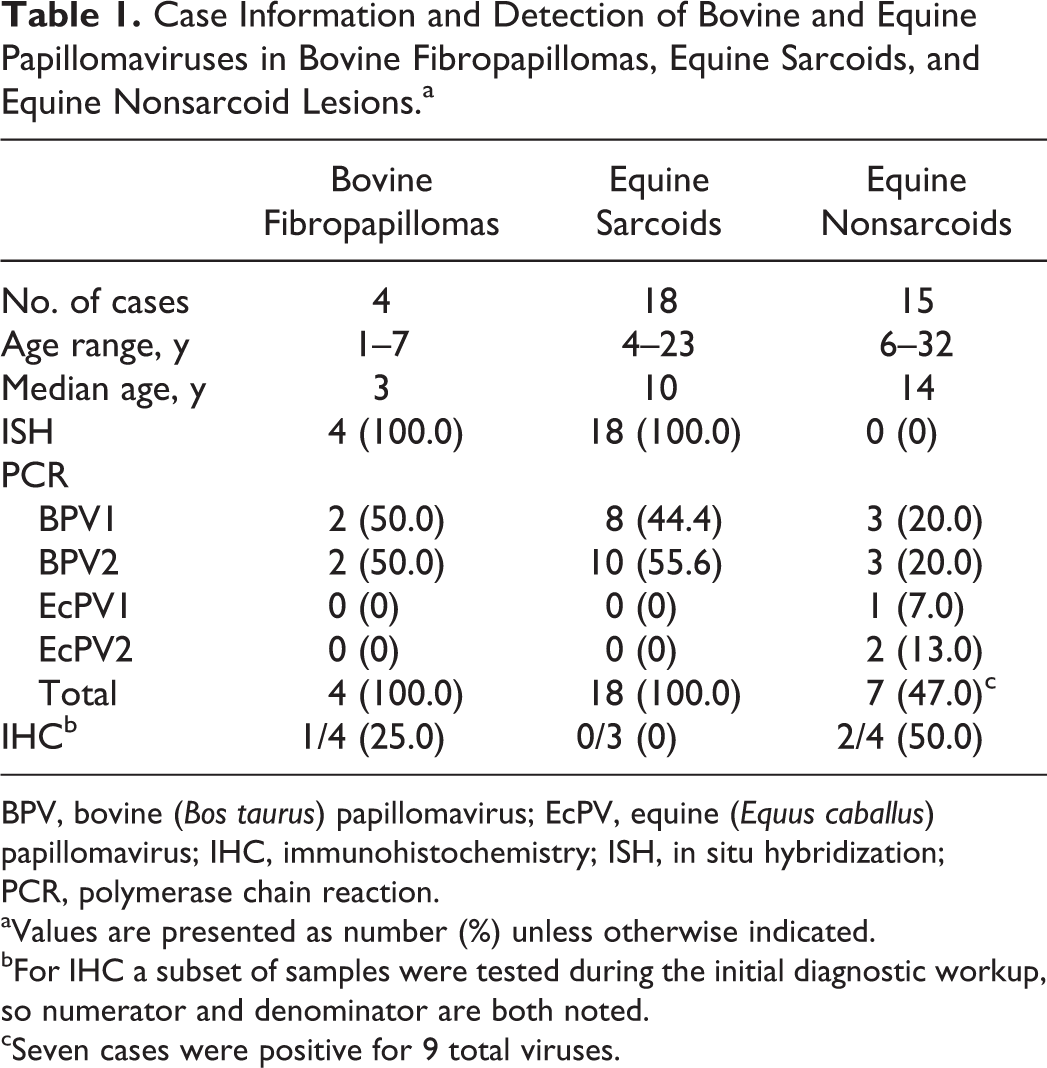
BPV, bovine (Bos taurus) papillomavirus; EcPV, equine (Equus caballus) papillomavirus; IHC, immunohistochemistry; ISH, in situ hybridization; PCR, polymerase chain reaction.
aValues are presented as number (%) unless otherwise indicated.
bFor IHC a subset of samples were tested during the initial diagnostic workup, so numerator and denominator are both noted.
cSeven cases were positive for 9 total viruses.
Histology and Immunohistochemistry
Formalin-fixed tissues were routinely processed and embedded in paraffin within 1 day of collection, sectioned at 5 μm, and routinely stained with hematoxylin-eosin (HE) for histologic evaluation. Immunohistochemistry was performed on formalin-fixed, paraffin-embedded tissue sectioned at 4 μm and mounted on charged slides. Slides were routinely deparaffinized and blocked with Dako Serum Free Protein Block (X0909; Dako, Carpinteria, CA). Anti-BPV1 antibody (rb-antiBPV1; Dako B0580) reactive to the L1 capsid protein was diluted 1:800 in Dako Antibody Diluent (S0809) with no antigen retrieval. Antibody was visualized with Rabbit-on-Canine HRP-Polymer (RC542; Biocare Medical, Concord, CA) per the manufacturer’s instructions. Sections were counterstained with Mayer’s hematoxylin. Normal serum or isotype-matched nonrelevant primary antibodies were used as negative controls and to evaluate nonspecific background staining. The positive control for the antibody was a section of a bovine esophageal viral papilloma validated by the histology laboratory at the William R. Pritchard Veterinary Medical Teaching Hospital’s Anatomic Pathology Service at UC Davis.
ISH
ISH probes were designed to detect both BPV1 and BPV2. BPV1 (X02346.1) and BPV2 (M20219.1) share high sequence identity across the genome (87%). To detect both viruses equally, the 2 genomes were aligned and 15 probe pairs were designed complementary to regions of the E5 (n = 2), E6 (n = 6), and E7 (n = 7) coding regions, covering approximately 1 kb of total sequence. The probes are designed to hybridize to both messenger RNA (mRNA) transcripts and viral genomic DNA, and no discrimination between these was made in this study.
Colorimetric ISH was performed manually on Superfrost Plus slides (Fisher Scientific, Pittsburgh, PA) using the RNAscope kit (Advanced Cell Diagnostics, Hayward, CA) according to the manufacturer’s instructions. 35 The sequences of the preamplifier, amplifier, and label probes are proprietary (Advanced Cell Diagnostics). Each 5-μm section of formalin-fixed, paraffin-embedded tissue was pretreated with heat and protease prior to hybridization with probe. A set of pooled probes designed to detect equine (Equus caballus) papillomavirus 2 (EcPV2) (E6 and E7 coding sequence) was used as a control. 40 BPV1 and BPV2 each have 41% pairwise identity with EcPV2, and therefore probes to EcPV2 should not detect these bovine papillomaviruses (Suppl. Table S3). A horseradish peroxidase–based signal amplification system was then hybridized to the target probes followed by color development with 3,3′-diaminobenzidine (DAB; Advanced Cell Diagnostics). Slides were counterstained with hematoxylin and mounted with xylene-based SHUR/Mount (Triangle Biomedical Sciences, Durham, NC). Positive staining was identified as brown, punctate dots in the nucleus. ISH-processed sections of lesions were scored on a scale of 0 to 3 for number of positive fibroblastic cells (nuclear localization) in 10 high-power 400× fields by a board-certified pathologist (P.A.P.) blinded to all case-related information and testing (Suppl. Table S2). The cells were scored in a binary fashion (positive or negative) without consideration for the total amount of deposit. A score of 0 indicated no positive cells, a score of 1 indicated low numbers (20%–50%) of positive cells, a score of 2 identified an intermediate number (50%–80%), and a score of 3 indicated that 80% to 100% of cells were positive in 10 high-power 400× fields.
Molecular Characterization
Nucleic acids were extracted from 2 × 25–μm paraffin scrolls following deparaffinization with the DNeasy Blood and Tissue Kit (QIAGEN, Valencia, CA). To detect BPV1 and BPV2, a single primer pair was used that amplifies a 244-bp (BPV1) and 248-bp (BPV2) product spanning a portion of the E2 gene, an intergenic region, and the E5 gene. 37 The full protocol is provided in the Supplemental Materials. To detect equine papillomavirus, EcPV1 (AF498323.1) and EcPV2 (EU503122.1) specific primers were designed to amplify the E1 region of the genome. Controls were performed to detect contamination and included no template, as well as forward and reverse primer alone. Normal skin and inflamed skin were also used as internal negative controls. PCR amplicons were purified and Sanger sequenced with a BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Carlsbad, CA). Sequenced fragments were analyzed for high-quality sequence and then analyzed using BLASTn against the Nationa Center for Biotechnology Information nucleotide collection (nr/nt). All samples that were negative for viral nucleic acid contained amplifiable DNA as tested by a universal mitochondrial DNA primer set 33 (data not shown).
Results
Validation of the ISH Assay
We selected bovine fibropapillomas from 4 distinct anatomic locations, and in all cases, the masses were comprised of a prominent mesenchymal component with an overlying, variably hyperplastic epithelium (Suppl. Table S1; Figs. 1, 3). Two cases were BPV1 positive (cases Bov1, Bov3) and 2 were BPV2 positive (cases Bov2, Bov4) based on 100% identity of the PCR amplicons to published sequences. In all cases, ISH and immunohistochemistry (IHC) for BPV were performed on serial sections. IHC for BPV L1 protein identified viral antigen in only 1 case (Bov1) with the antibody binding restricted to mature epithelial cells of the cornified layer (data not shown). By ISH, BPV nucleic acid was present within nuclei of the fibroblastic population in all cases (Figs. 1–3). Detectable BPV nucleic acid within the epidermal layer was quite variable, from extensive and strong hybridization (Figs. 1, 2) to only scattered cells (Fig. 3), and 1 case had no detectable viral nucleic acid within the overlying epithelium (data not shown). As a negative control for these tissues, a nucleotide-matched, scrambled probe was used in parallel, which was negative in all cases (Fig. 3c).
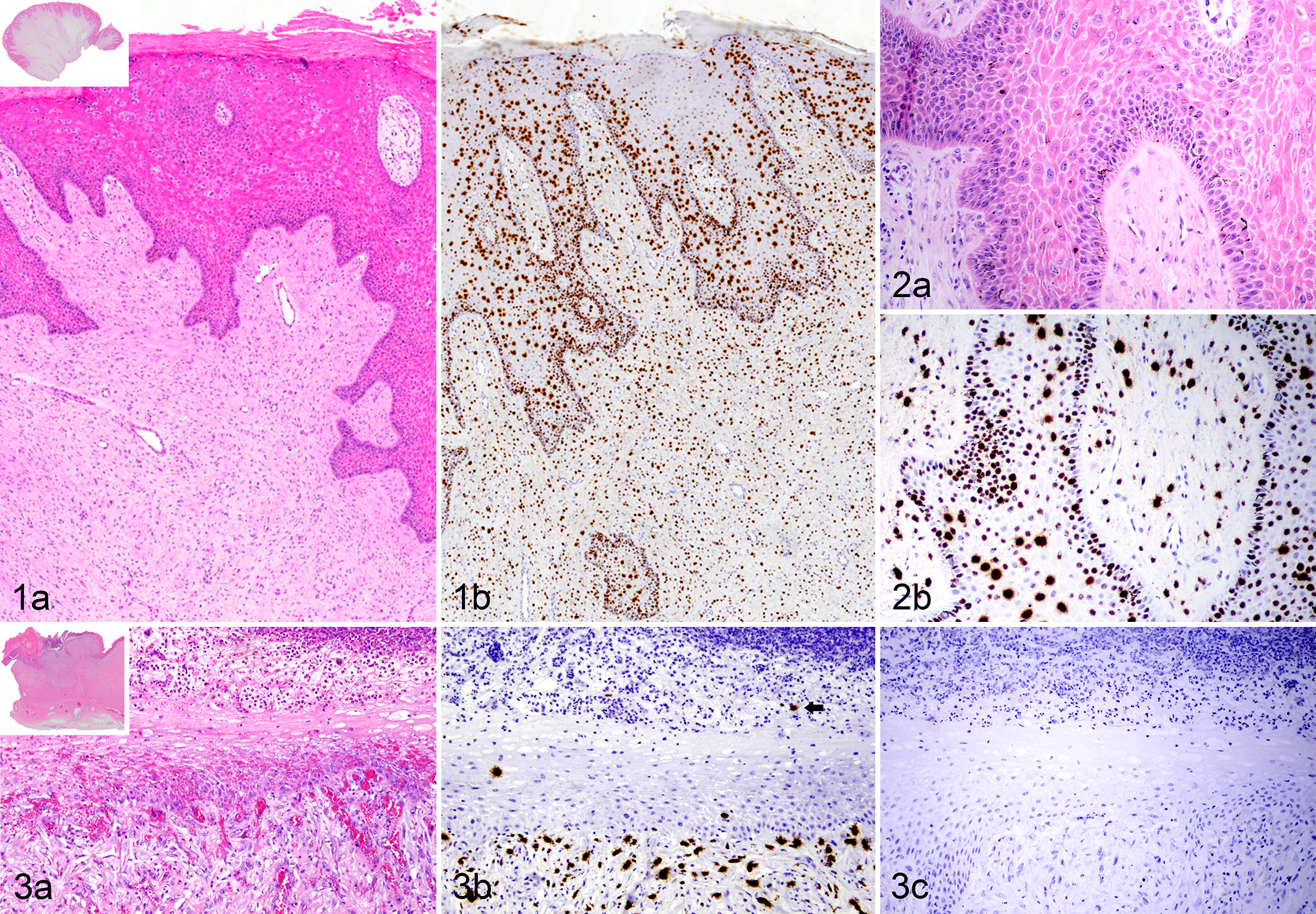
Fibropapilloma, bovine.
Examination of Equine Sarcoids
To visualize the presence and distribution of BPV in equine sarcoids and to evaluate whether virus was present within epithelial cells, we examined 18 cases, of which several tumors had broad contact with an intact overlying epithelium (Figs. 4–6). All sarcoids (18/18) had visible hybridization with BPV probes using the RNAScope ISH technique (Table 1; Figs. 5, 6, 7b, 9). In 12 of 18 cases, normal skin was present within the section, adjacent to the sarcoids; in normal regions, we were unable to visualize any hybridization of the BPV probes (Fig. 5). To categorize the spectrum of hybridization in the sarcoids and to determine whether clinical types had different patterns of hybridization, a simple scoring scheme was used (Suppl. Table S2). Sixteen of 18 cases (89%) were scored as a 2 or 3 (≥50% positive fibroblastic nuclei), and in half (9/18) of the cases, >80% of the nuclei were positive, scored as a 3 (Figs. 5, 6). There were no significant associations with clinical type. Most sarcoids were composed of the canonical spindle cells, but in 1 case (Eq11), there was high anisocytosis and anisokaryosis with a prominent population of multinucleate cells (Fig. 10a). The nuclei of both the fibroblastic cells and the multinucleate cells hybridized robustly with the BPV probe (Fig. 10b). We were not able to visualize any hybridization to control probes (EcPV2) in serial sections run in parallel (Fig. 10c). The remaining 2 cases, Eq4 and Eq7 (Figs. 9, 10), had fewer than 50% of the fibroblastic nuclei positive. They were also more difficult than the solid sarcoids to assess since the tumor population dissected along the epithelial border and among the dermal fibroblasts beneath fronds of hyperplastic epithelium (Fig. 8).
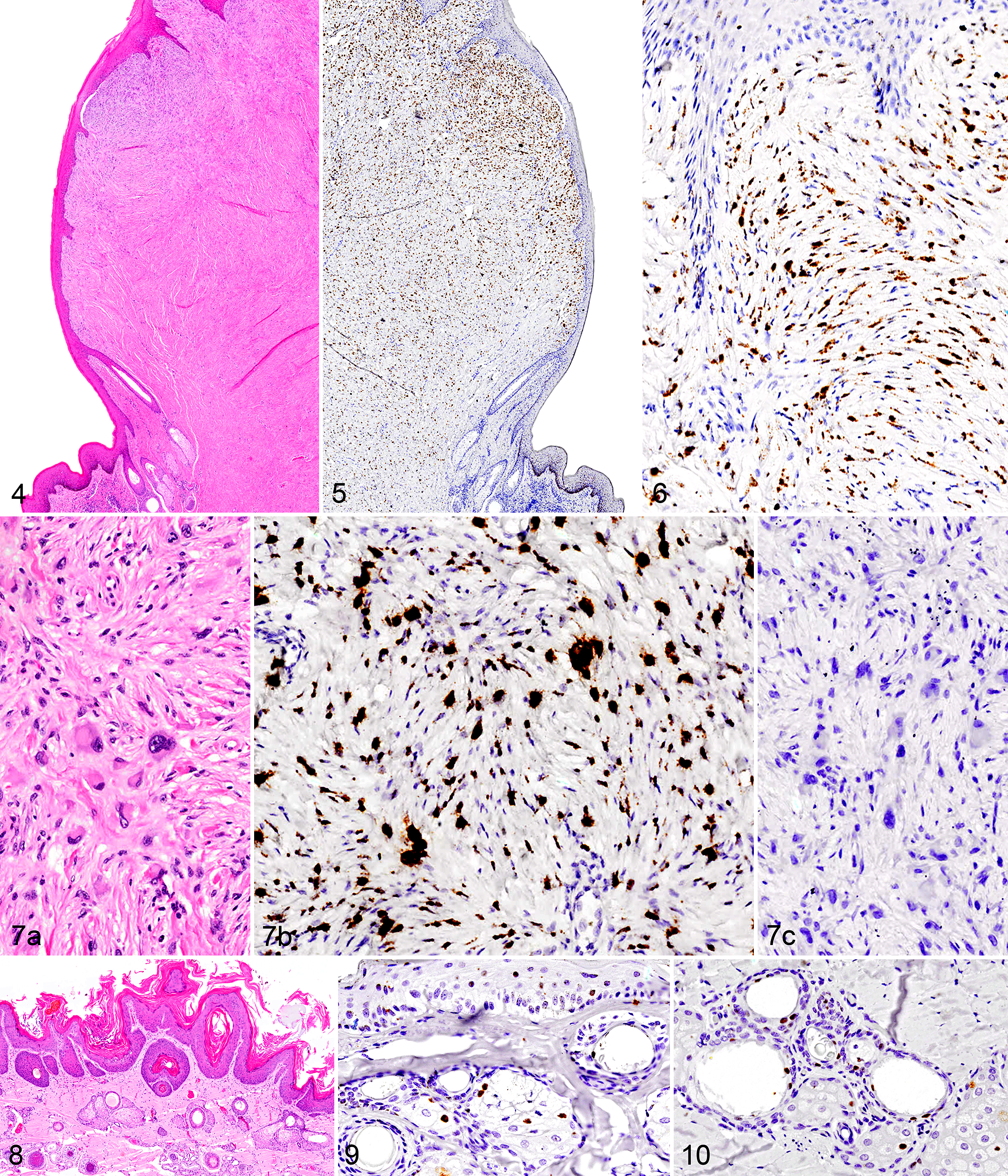
The overlying epithelium and adnexa (including epidermis, exocrine glands, sebaceous glands, hair, ± serous glands) were examined in all cases available (17/18). Hybridization to BPV nucleic acid within epithelial cells, while quite rare, was detected in 2 cases, Eq4 and Eq7 (Figs. 9, 10). Hybridization was detected in the nuclei of sebocytes, hair follicular epithelia, and rarely in the epidermis (Figs. 9, 10). ISH-positive sebaceous glandular and follicular epithelial cells were embedded within the sarcoid, but hybridization also occurred in nuclei of these glands in adjacent histologically normal skin (Fig. 9). No detectable inflammation or alteration of the glands was histologically apparent (Fig. 8). These 2 cases were subjectively considered early lesions based on clinical history, and both were classified clinically as verrucous.
All 18 cases of equine sarcoids also contained BPV DNA detectable by PCR. Sequencing of PCR amplicons established that BPV1 was present in 8 cases and BPV2 in 10 cases, which was based on 98% to 100% identity to previously published sequences (Table 1; Suppl. Tables S2–S3). IHC for BPV performed as part of the diagnostic workup in 3 cases was negative (Table 1; Suppl. Table S2).
Testing of Nonsarcoid Equine Skin
Following several reports that BPV genome and transcript can be amplified by PCR from equine skin (normal or with various dermatologic conditions) and as a control for the sarcoid testing, we examined 15 nonsarcoid skin samples by ISH and PCR (Table 1). BPV nucleic acid was not detected by ISH in any of the 15 nonsarcoid equine skin samples examined in these cases. When examined by PCR, 7 of 15 tissues contained papillomaviral DNA (Table 1; Suppl. Table S2). Three cases (3/15, 20%), previously classified as papillomas by histopathology, contained equine papillomavirus DNA (Eq22, Eq24, and Eq27), and of those, 2 cases (Eq22, Eq24) were also IHC positive with the cross-reactive anti-BPV L1 antibody. The remaining 4 cases (4/15, 27%) were PCR positive for BPV1, BPV2, or both (Table 1; Suppl. Table S2).
Horses With Multiple Tissues
Six of the horses in this study had multiple anatomically separable lesions that were evaluated (Suppl. Table S2). Two horses (H9 [Figs. 4–6] and H11) had 2 sarcoids each; for each horse, the sarcoids were of the same type, had similar scoring on ISH (H9: 3; H11: 2), and contained the same type of viral DNA (H9: BPV1; H11: BPV2). In the other 4 horses (H2, H4, H8, and H13), each contained at least 1 sarcoid and 1 nonsarcoid tissue. In the ISH technique, the sarcoids (Eq2, Eq4, Eq8, Eq15) had visible hybridization to BPV DNA, while the nonsarcoid tissues had no detectable hybridization (Eq20, Eq25, Eq26 Eq29, Eq30, Eq32); however, 3 of 6 did have detectable papillomaviral DNA by PCR (Suppl. Table S2).
Discussion
In this report, we used a sensitive ISH method with signal amplification to determine the localization of bovine papillomaviral (BPV1 and BPV2) nucleic acid within equine sarcoids and nonsarcoid tissue from horses. Verification of the ISH method was based on viral nucleic acid localization within bovine fibropapillomas that were confirmed to contain BPV1 or BPV2. All 18 equine sarcoids, independent of clinical typing, were positive by ISH, and none of the 15 nonsarcoid samples contained detectable viral nucleic acid by this technique. We did not identify any overt difference in hybridization pattern that correlated with clinical type. PCR testing of these samples revealed that 18 of 18 equine sarcoids contained amplifiable BPV DNA, and 7 of 15 nonsarcoid samples contained amplifiable papillomaviral DNA, with 3 of 15 positive for equine papillomavirus DNA and 4 of 15 positive for bovine papillomavirus DNA. These results contribute to the evidence that BPV1 and BPV2 may have a productive life cycle in equine tissues.
ISH has been used to examine equine sarcoids in 2 previous studies, and in those studies, a small subset of neoplastic fibroblasts near the dermal-epidermal junction contained BPV nucleic acid in 42% (10/24) 21 and 100% (20/20) 32 of tested cases. Our results corroborate these results, with 18 of 18 equine sarcoids having hybridization primarily in the nuclei of the fibroblastic cells. However, we detected BPV DNA distributed throughout the entire mass, including all deep borders, and not just at the dermal-epidermal junction as previously reported. The detection of an expanded distribution could be due to differences in the sarcoids that were examined or an increased sensitivity of this assay compared with other ISH techniques. A direct comparison and other controlled studies would need to be performed to demonstrate an increased sensitivity of the RNAscope ISH compared with other nonradioactive ISH methods.
We report for the first time the visual localization of BPV DNA in equine epithelium. This finding corroborates 3 previous PCR-based studies that used laser microdissection to selectively isolate cellular compartments such as the fibroblastic population and the overlying epithelium. In these studies, BPV DNA was detected not only at high frequencies in the dermal tissue (fibroblasts), as we saw with ISH, but also, less prominently, in the epidermis. 8,11,37 Similarly, we specifically visualized BPV DNA in the sebocytes, hair follicle epithelium, and rarely epidermal cells. Detection in sebocytes has also been reported in related viruses, including Bandicoot papillomatosis carcinomatosis virus type 1, 7 human papillomaviruses, 16 and Micromys minutus papillomavirus. 31 As terminally differentiated cells, sebocytes accumulate lipids and undergo cell membrane disruption and cell destruction (holocrine secretion), releasing sebum. This natural process would support the dispersal of viral particles in the sebum that coats the skin and hair. If they are infected, sebocytes could serve as a potential reservoir of virus and mechanism of transmission. It is also of note that the concentration of sebaceous glands is highest in the regions where sarcoids are most common. The 2011 study by Brandt et al 11 also detected early (E5 and E7) and late (L1) proteins in the epidermis. The collective data from our study and previous work of the detection and localization of viral nucleic acid (genomic or transcript), as well as viral protein in the epithelium, are indicative of at least a portion of the life cycle successfully executing despite the lack of detection of virions. 1,19 Further work is necessary to understand how and when these events of the viral life cycle occur and their relationship to sarcoid development and/or maintenance. We propose, as have others, 8,37 that early in sarcoid development, there could be an early viral infection in the adnexa or surface epithelium that is undetectable later in the development of the tumor.
Despite overwhelming evidence that BPV is associated with equine sarcoids, the causal role of BPV has been questioned due to the confounding information that bovine papillomaviruses are ubiquitous and, hence, are also present in nonsarcoid lesions and even on normal skin of equids. Bovine papillomaviruses are not unique in this regard; in fact, papillomaviruses are commonly found on healthy skin of humans and animals. 3 –5,15,20 Therefore, prudence is required when interpreting the presence of amplified papillomavirus DNA. Among PCR-based equine studies, reported prevalence of BPV amplified from nonsarcoid (normal or inflamed) tissues ranges from 0% to 80%. 8,37 Efforts to decontaminate the skin have demonstrated the same rate of PCR positivity before and after “decontamination” (clipped hair, scrubbing, disinfecting with ethanol, or tape stripping). 8 Embedded negative controls were in place in our and other PCR-based studies for scrolling, extraction, and processing steps. Despite these efforts, BPV DNA is detectable in fresh, frozen, and formalin-fixed, paraffin-embedded biopsy specimens, as well as swabs, and it is present in several dermatologic conditions and from multiple anatomic locations. We recognize the longstanding controversy that detection of BPV DNA by PCR does not necessarily indicate that the virus has initiated or is involved in a given lesion. However, using a sensitive ISH, we have localized BPV DNA to a subset of neoplastic cells within every equine sarcoid but not in nonsarcoid lesions. The limited studies that have examined nonsarcoid samples by non-PCR methods (Western blot) have not detected BPV. 14 While one could make the case that ISH is simply less sensitive than PCR, we suggest that the ubiquity of BPV on skin surfaces confounds many studies directed at estimating the contribution of the virus to equine sarcoids.
This study demonstrates that BPV nucleic acid is present within bovine fibropapillomas and equine sarcoids but undetectable by ISH in nonsarcoid equine tissues. We further identified BPV DNA in the epithelium of a subset of bovine fibropapillomas and equine sarcoid cases. In 89% (16/18) of the sarcoid cases, there was no detectable BPV DNA in the epithelium, despite the apposition of the sarcoid with the overlying epithelium. However, the identification of adnexal and surface epithelial cells positive for BPV DNA in 2 equine cases represents a potential site of viral production and a mechanism for viral dispersal through the equine sebum and hair.
Footnotes
Acknowledgements
We thank the UC Davis VMTH Histopathology Laboratories for their technical expertise. We also thank Drs Jennifer Luff and John Munday for their helpful discussions and input, as well as Kevin Keel for his assistance with obtaining the images.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is partially funded by grants received for pathogen discovery efforts, including The Bernice Barbour Foundation and Boerhinger-Ingleheim Vetmedica.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
