Abstract
To characterize the clinicopathologic features of recently described genotypes of Newcastle disease virus (NDV), 1 representative strain of genotype XIV and 2 of genotype XVII, all isolated from West Africa, were used to infect groups of ten 4-week-old specific pathogen-free chickens. The pathobiology of these 3 strains was compared to a South African NDV strain classified within genotype VII. All chickens infected with the 4 viruses died or were euthanized by day 4 postinfection due to the severity of clinical signs. Gross and histologic lesions in all infected chickens included extensive necrosis of lymphoid tissues (thymus, spleen, bursa of Fabricius, cecal tonsils, gut-associated lymphoid tissue), gastrointestinal necrosis and hemorrhages, and severe hemorrhagic conjunctivitis. Immunohistochemical staining revealed systemic viral distribution, and the most intense staining was in the lymphoid organs. Results demonstrate that the 3 West African strains from the previously uncharacterized genotypes XIV and XVII are typical velogenic viscerotropic NDV strains with lesions similar to the South African strain. Under experimental conditions, QV4 and LaSota NDV vaccine strains successfully protected chickens from morbidity and mortality against the genotype VII and one genotype XVII NDV strain, with no significant differences in the amount of virus shed when 2 vaccine schemes were compared.
Keywords
Newcastle disease (ND) is a highly contagious avian disease that occurs worldwide in poultry and other domestic and wild bird species. 5,6,10 The causative agents of ND are virulent strains of ND virus (NDV), also known as APMV-1 (avian paramyxovirus serotype 1), an enveloped, negative-sense, single-stranded RNA virus of the Avulavirus genus and Paramyxoviridae family. 26 The clinical signs produced upon infection with different strains of NDV are extremely variable and not pathognomonic for ND. ND has to be ruled out from other severe avian diseases, such as highly pathogenic avian influenza, or fowl cholera. 5,6 The wide array of clinical disease depends on both strain virulence and host-related factors, such as the host species infected, immunologic status, and the presence of copathogens in the host. 5,6,10
Historically, based on the clinical disease produced in chickens upon infection, NDV strains have been classified into 4 virulence groups: velogenic (up to 100% mortality with visceral hemorrhages), mesogenic (intermediate virulence, characterized by respiratory signs and lower levels of mortality), lentogenic (very low mortality and clinical signs limited mostly to young birds, usually in association with copathogens), and avirulent strains (asymptomatic). 10 Velogenic NDV strains can be further divided into viscerotropic, which cause severe intestinal and visceral hemorrhages, or neurotropic, which cause severe neurologic clinical signs and encephalitis. 6,10,18,42 One of the main determinants of NDV virulence is the cleavage site of the fusion protein. 6,14,34 Virulent NDV strains have a polybasic amino acid configuration that allows cleavage of the fusion protein by furin-like proteases found ubiquitously in the organism, allowing for systemic viral spread. In comparison, nonvirulent strains have an fusion protein that can be cleaved only in trypsin-rich tissues, confining the infection to the intestine or respiratory tract. 11,34 A widely accepted system used to implement virulence classification is the intracerebral pathogenicity index (ICPI) in day-old chicks, which yields a numeric score on a 0–2 scale, with scores close to 2 being typical of very virulent strains. 5,36 According to the World Organization for Animal Health (former Office International des Epizooties [OIE]), virulent NDV strains (notifiable to the international community) are those viruses that have an ICPI equal to or greater than 0.7, or an amino acid sequence of the fusion protein cleavage site with at least 3 arginine or lysine residues between positions 113 and 116 and a phenylalanine residue at position 117. 36
Although all NDV strains belong to a single serotype (serotype 1), there is large genetic variability among NDV isolates. 1,29,31 A recent implementation of this genetic classification has divided NDV into 2 classes (I and II), with class I composed of only 1 genotype (class I, genotype I) and with class II divided into 18 genotypes (class II, genotypes I–XVIII). 13,15,39 While class I encompasses only avirulent NDV strains (except for 1, APMV-1/chicken/Ireland48/90 4 ) found mainly in waterfowl, class II includes both virulent and avirulent strains. 1,29
In rural Africa, chickens typically scavenge for the majority of their diet, are raised under a range of production conditions, 19,28 and are often referred to as village poultry. Chicken products are an important mean of economic and dietary subsistence for the poor, with an estimated 70% of all poultry products and 20% of human protein intake consumed in Africa derived from poultry. 24 ND is rampant in various parts of Africa and poses a serious threat to poultry production, causing, in rural areas, death of up to 80% of backyard chickens. 41
In West Africa, ND is the main poultry disease in commercial and rural chickens. Recent reports from West and Central Africa described the presence of novel virulent NDV strains belonging to new genetic lineages closely related to genotype VII (mostly circulating in South Africa 22 )—namely, genotypes XIV, XVII, and XVIII. 9,37,39,40,45 Many of these isolates were obtained from live bird markets and village poultry, and in several cases the absence of apparent clinical signs (subclinical infection) was reported. Vaccination against some of these strains with the LaSota vaccine conferred protection from disease and death, but virus shedding was not prevented and proved to be higher when compared with other challenge NDV strains of a genotype evolutionarily closer to LaSota (eg, Texas GB). 37 Furthermore, genome analysis of some of the NDV strains from the Togo region showed a unique 6-nucleotide insert within the intergenic regions between the HN and L genes, which is a novel genomic arrangement of NDV. 23,37
Although characterized genetically, to date there are no studies detailing the clinical and pathologic characteristics of the virulent NDV strains that have been circulating recently in West Africa. The aim of the present study was to characterize the ability of 3 West African NDV strains (1 of genotype XIV and 2 of genotype XVII) to cause disease, induce lesions, and replicate in experimentally infected chickens. These characteristics were compared with a South African strain representative of the previously identified genotype VII. Additionally, the efficacy of commonly deployed vaccine NDV strains (strains QV4 and LaSota) was evaluated for protection against morbidity, mortality, and shedding against genotype VII and XVII NDV strains.
Materials and Methods
Viruses
The NDV isolates Burkina Faso/2415-580/2008 (BF/08) and Niger/1377-7/2006 (Niger/06) were obtained from the Istituto Zooprofilattico Sperimentale delle Venezie (OIE/FAO reference center for NDV, Padua, Italy). These 2 strains have been recently classified as genotype XIV (Niger/06) and genotype XVII (BF/08) NDVs. 39 The NDV strains Nigeria/228-7/2006 (Nigeria/06), Chicken/South Africa/08100426/2008 (SA/08), LaSota/46, and QV4 were obtained from the Southeast Poultry Research Laboratory (SEPRL) repository. Nigeria/06 was isolated and identified as NDV from samples recovered from sick and dead birds in Nigeria during a survey study for H5N1 avian influenza. 2 SA/08 was isolated in October 2008 from an outbreak of ND in chickens from the Paarl area of the Western Cape Providence of South Africa and was obtained by SEPRL from Dr James Kitching of the Western Cape Provincial Veterinary Laboratory. All the viruses were grown in the allantoic cavity of 9- to 10-day-old specific pathogen-free (SPF) embryonating chicken eggs. Egg passage 2 virus stocks were used to perform RNA extraction and all the pathogenesis experiments.
Eggs and Chickens
The source of embryonating chicken eggs and chickens was the SEPRL, SPF White Leghorn flock. Birds (10 for each cage) were housed in negative-pressure isolators under biosafety level 3–enhanced conditions at SEPRL and provided food and water ad libitum. Embryonating SPF chicken eggs were inoculated for viral propagation, isolation, and titration. 5,36
ICPI Test
The ICPI test was performed with the 4 NDV strains of this study, according to standard protocols. 5,36 Briefly, 10 chickens per group were inoculated intracerebrally at 1 day of age with 0.05 ml of a 1:10 dilution in phosphate buffered saline of infective allantoic fluid, or phosphate buffered saline only (control group). Chickens were monitored daily and scored as normal (0), sick or paralyzed (1), and dead (2) to compile an averaged score over an 8-day observation period.
RNA Isolation and Sequencing
Total RNA from the allantoic fluid of Nigeria/06- and SA/08-infected eggs was extracted with TRIzolLS (Invitrogen, Carlsbad, CA) following the manufacturer’s instruction. The fusion gene was amplified by reverse transcription polymerase chain reaction and sequenced with standard Sanger method. Primers for the full fusion gene are available upon request. The sequences of BF/08 and Niger/06 were published elsewhere. 9,21
Phylogenetic Analysis and Alignment
The sequences of the entire open reading frame of the fusion gene (“full fusion”) of BF/08, 9 Niger/06, 21 Nigeria/06, and SA/08 were used for phylogenetic analysis as previously described. 13,16 A phylogenetic tree was inferred to localize the 4 African strains among representative class II reference strains (total n = 56) according to the maximum likelihood method based on the general time-reversible model as implemented in MEGA 5 software, 35,44 including in the analysis the first, second, third, and noncoding positions and the “complete deletion” option for positions with gaps in the final alignment. Initial tree for the heuristic search was obtained by applying the neighbor-joining method to a matrix of pairwise distances estimated via the maximum composite likelihood approach.
Additionally, evolutionary distances were tested between the full fusion of Nigeria/06 and representatives of the 18 NDV genotypes of class II. To accomplish this, strains representing all 18 class II NDV genotypes (I–XVIII) were analyzed (total n = 614; genotype I, n = 71; genotype II, n = 102; genotype III, n = 9; genotype IV, n = 6; genotype V, n = 45; genotype VI, n = 57; genotype VII, n = 233; genotype VIII, n = 4; genotype IX, n = 19; genotype X, n = 18; genotype XI, n = 4; genotype XII, n = 6; genotype XIII, n = 8; genotype XIV, n = 9; genotype XV, n = 5; genotype XVI, n = 4; genotype XVII, n = 7; genotype XVIII, n = 6; and Nigeria/06 strain, n = 1). Distances were shown as the average (per each group pair) number of base substitutions per site and inferred through the maximum likelihood method with a gamma distribution (shape parameter = 1) of rate variation among sites (MEGA 543,44). Criteria for the assignment of genotypes were based on the recently proposed nomenclature by Diel et al, 15 and this system is adopted throughout the present work.
Pathology Assessment in Chickens
Fifty 4-week-old SPF White Leghorn chickens were assigned randomly into 5 experimental groups of 10 birds each, respectively consisting of BF/08-, Niger/06-, Nigeria/06-, and SA/08-infected birds and 1 mock (control) group. Birds were inoculated with 0.1 ml of viral suspension in brain–heart infusion (BHI) broth containing a target dose of 105.0 embryo infectious dose 50% (EID50) of virus. Half the inoculum (0.05 ml) was administered in the left conjunctival sac and half into the choanal slit. BHI media was used for the noninfected (mock) control birds. Back titers of the inocula, determined in embryonating eggs, were as follows: 105.3 EID50 for BF/08, 104.5 EID50 for Niger/06, 105.9 for Nigeria/06, and 105.3 for SA/08. Birds were monitored daily for clinical sings of disease, and 2 birds of each group were scheduled for euthanasia at days 2, 5, 10, and 14 postinfection (pi) to perform necropsy and collect tissues for histopathology (2 birds per group were extra). Birds whose condition became critical were euthanized and tissue samples collected regardless of the sampling schedule.
Twenty-five tissues were collected for each animal. Based on the scheduled and unscheduled sampling activity (birds euthanized in extremis), the following numbers of tissues were examined by HE per each group: eyelids, Harderian gland, spleen, bursa of Fabricius, thymus, proventriculus, small intestine, large intestine, cecal tonsils, air sac, trachea, lung, heart, esophagus, pharynx, larynx, crop, brain, liver, pancreas, kidney, comb, tongue, head of left femur including bone marrow, and turbinates—n = 5 for BF/08, Niger/06, and Nigeria/06 and n = 6 for SA/08. After collection, all tissue samples were fixed by immersion in 10% neutral buffered formalin for 52 hours. The sections of femur and turbinate were decalcified in 5% formic acid for 3 to 4 hours. All sampled tissues were processed into paraffin, and 3-μm sections were cut for hematoxylin and eosin staining. Histologic grading of lesions was carried out as specified in Table 3. When organs had different scoring, the value was averaged and, if needed, rounded to the next integer.
Immunohistochemistry
In each group, all sampled tissues from both birds euthanized on day 2 pi and from 2 birds euthanized or dying on day 4 pi were examined by immunohistochemistry to detect NDV nucleoprotein. Briefly, after deparaffinization, tissue sections were subjected to antigen retrieval by microwaving for 20 minutes at minimum power in Vector antigen unmasking solution (Vector Laboratories, Burlingame, CA), followed by application of a universal blocking reagent (Biogenex, San Ramon, CA) as recommended by the manufacturer. The primary antibody, raised in rabbit, was directed against a NDV nucleoprotein synthetic peptide (TAYETADESETRRIC) and used at 1:8000 dilution. 25,42 The detection system was an avidin–biotin–alkaline phosphatase system (Vector Laboratories). Chromogen was a naphthol-based dye (Fast Red, Dako, Carpinteria, CA). Sections were counterstained lightly with hematoxylin and coverslipped with Permount for a permanent record. Grading was carried out as specified in Table 3. When organs had different scoring, the value was averaged and, if needed, rounded to the next integer.
Vaccination Experiment
The vaccination experiment was carried out to assess if commonly used vaccine strains and vaccination schemes offer a good protection for SPF chickens against a representative of the newly described West African NDV genotype XVII (BF/08). The level of protection and virus shedding was also compared to another challenge virus isolated from Africa (SA/08), classified in genotype VII. Based on how the 2 vaccines, LaSota and QV4, were administered, 3 vaccination groups were used: LaSota/LaSota, QV4/LaSota, and QV4/QV4.
Eighty day-old SPF White Leghorn chicks were placed in groups of 10 birds each into plastic molded isolators. Food and water were provided ad libitum. Serum samples from hatchmates were collected to ensure negative status for NDV antibodies. The primary live vaccine was administered by inoculating 0.05 ml of vaccine into the left eye and 0.05 ml of vaccine into the choanal slit. The second live booster vaccination was administered in the same way 16 days later. Direct vaccine administration was preferred to delivery through drinking water to decrease the possibility of nonhomogeneous vaccine delivery and to avoid the chance of suboptimal vaccination. On day 22 after the first vaccination (5 days after the boost), serum samples were collected to assess hemagglutination inhibition (HI) titers against the first vaccine and the challenge virus, and half the birds (n = 10) per vaccination regimen group were infected with either 1 of the 2 challenge viruses. Nonvaccinated controls received 2 doses of 0.1 ml of uninfected BHI delivered the same route as the vaccines. The challenge viruses BF/08 and SA/08 were administered as the vaccines at a dose of 105.7 EID50 or 106.3 EID50, respectively, as shown by back titration. Birds were monitored clinically and cloacal swabs collected at 2 and 4 days postchallenge. Serum from infected birds at day 4 postchallenge was collected to assess HI titers against the challenge virus.
Virus Isolation and Titration of Swabs
Oral and cloacal swabs were obtained from each bird and placed into separate tubes containing 1.5 ml of BHI containing antibiotics (2000 U/ml penicillin G, 200 μg/ml gentamicin sulfate, 4 μg/ml amphotericin B; Sigma Chemical, St Louis, MO). Swab sample tubes were centrifuged at 1000 × g for 20 minutes and the supernatant removed for virus isolation and titration. Virus infectivity titers were calculated from the result of inoculation (single passage) in the chorio-allantoic cavity of 9- to 10-day-old SPF embryonating chicken eggs with serial 10-fold dilutions in BHI containing antibiotics (100 U/ml penicillin G, 50 μg/ml gentamicin sulfate, 4 μg/ml amphotericin B; Sigma Chemical). 5,36 After 7 days postinoculation, NDV from infected dead or surviving embryos was identified by hemagglutination activity in amnioallantoic fluid harvested from chilled eggs. NDV was confirmed in hemagglutination positive samples by HI test with NDV-specific antiserum.
Statistics
Virus isolation arithmetic means for days 2 and 4 postchallenge were calculated for each vaccine group, including shedding and nonshedding birds, and group means were analyzed by analysis of variance with Tukey post hoc test. Additionally, arithmetic means were calculated for only those birds that shed virus from each group along with the number of birds shedding. Note that the analyses for virus isolation data for day 4 postchallenge compared only the vaccine groups to one another, as there were no survivors for the nonvaccinated control group. Fisher exact tests were performed on data indicating number of birds shedding at day 2 postchallenge. For all tests, significance was reported at P ≤ .05.
Results
ICPI Test and Fusion Sequence
The ICPI scores for the tested NDV strains were as follows: BF/08 = 1.69, Niger/06 = 1.84, Nigeria/06 = 1.90, SA/08 = 1.91. These scores confirm that all the strains are virulent and therefore notifiable. 36 The full fusion sequences for BF/08 (GI: 259879781) and Niger/06 (GI: 374256165) have been already published, and they showed a polybasic configuration between amino acid residues 113–116 and a phenylalanine at position 117, 9,21 which is consistent with virulent NDV strains. 36 Additionally, the configuration of the fusion cleavage site for Nigeria/06 and SA/08 was, respectively, 113RQRR*F117 and 113RQKR*F117 (asterisk represents the cleavage site) consistent with virulent NDV. 36 The full fusion sequences for Nigeria/06 and SA/08 have been submitted to Gene Bank with the following accession numbers: KF442614 and KF442615, respectively.
Phylogenetic Analysis and Alignment
The inferred phylogenetic tree showed that Nigeria/06 and BF/08 clearly clustered with other genotype XVII NDV strains and are clearly distinct from clusters encompassing other NDV genotypes (bootstrap value, 100%; Fig. 1). Niger/06 clustered among isolates of genotype XIV (bootstrap value, 100%). SA/08 clustered with other known genotype VII NDV isolates (bootstrap value, 100%). Based on the maximum likelihood method, the evolutionary distances between the full fusion sequence of Nigeria/06 and representatives of the 18 NDV genotypes of class II were compiled as the number of base substitutions per site when averaged for each genotype. The computed distances showed that Nigeria/06 is significantly closer to NDV genotype XVII (0.041 average base substitution), with distances higher than 10% with all other considered genotypes (Table 1). The most distant genotypes from Nigeria/06 were genotypes II and XI. Together, these data show that Nigeria/06 belongs to NDV genotype XVII. 15
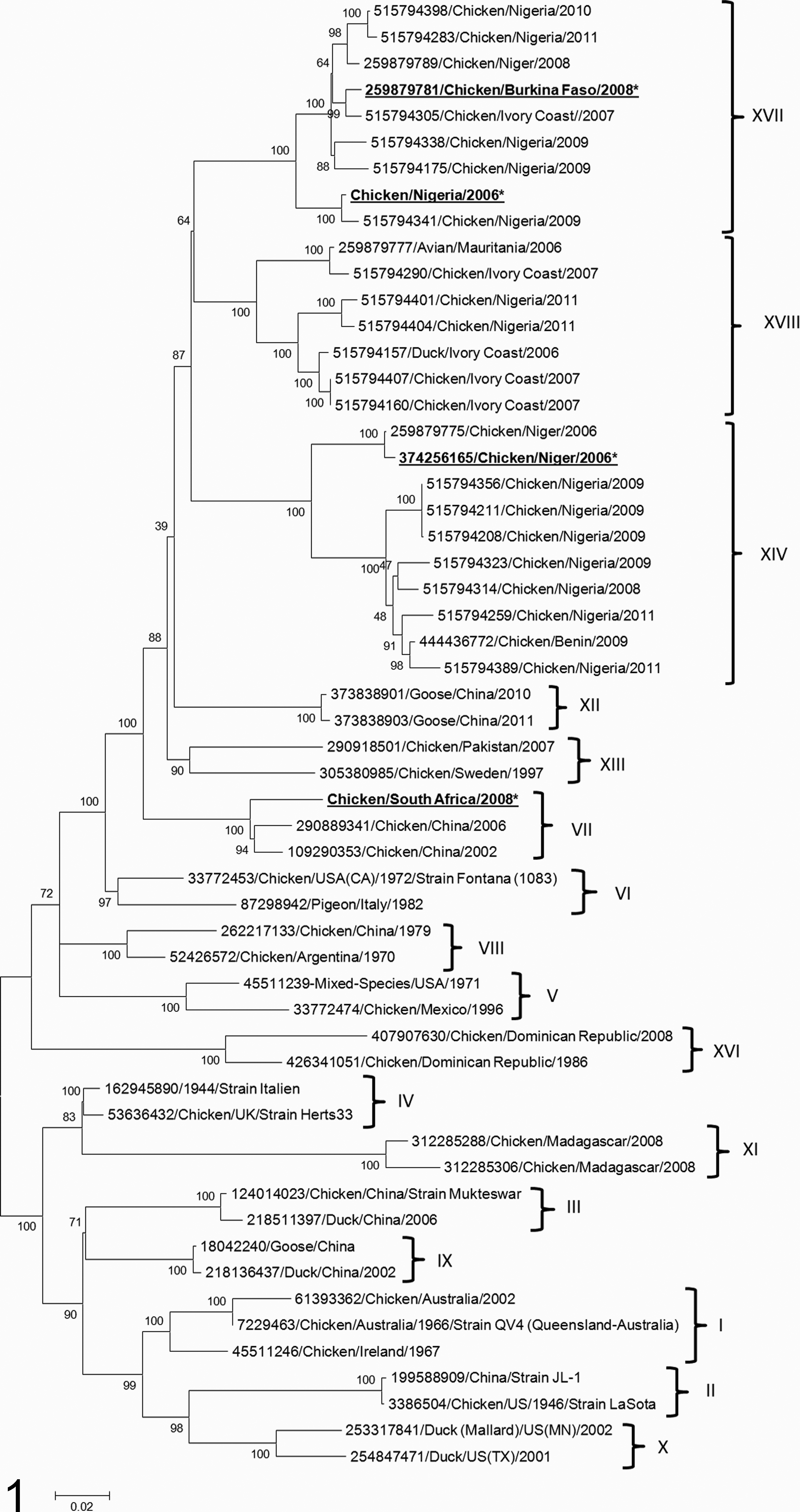
Phylogenetic analysis of the full coding region of the fusion gene for the African isolates in comparison to sequences from viruses representative of different Newcastle disease virus (NDV) genotypes. The evolutionary history was inferred on 56 nucleotide sequences according to the maximum likelihood method based on the general time-reversible model. The percentage of replicate trees in which the associated taxa clustered is shown next to the branches (100 replicates). The tree shows that NDV strains characterized in the article (marked with asterisks) cluster with representatives of genotypes XIV (Niger/06), XVII (Nigeria/06 and BF/08), and VII (SA/08).
Estimates of the Net Evolutionary Distances Between the Full Fusion Gene of Nigeria/06 and Other 18 Class II NDV Genotypes.
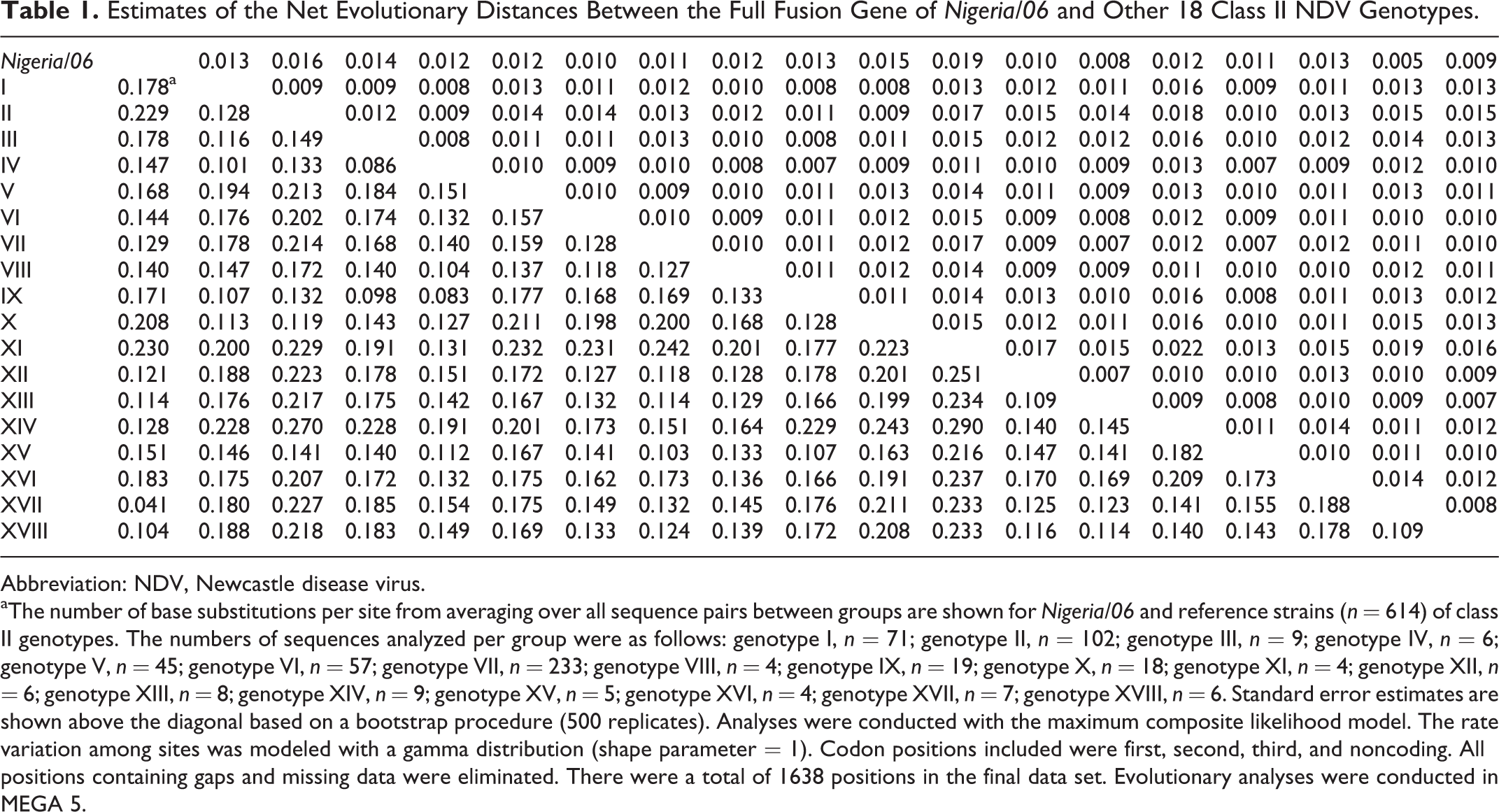
Abbreviation: NDV, Newcastle disease virus.
aThe number of base substitutions per site from averaging over all sequence pairs between groups are shown for Nigeria/06 and reference strains (n = 614) of class II genotypes. The numbers of sequences analyzed per group were as follows: genotype I, n = 71; genotype II, n = 102; genotype III, n = 9; genotype IV, n = 6; genotype V, n = 45; genotype VI, n = 57; genotype VII, n = 233; genotype VIII, n = 4; genotype IX, n = 19; genotype X, n = 18; genotype XI, n = 4; genotype XII, n = 6; genotype XIII, n = 8; genotype XIV, n = 9; genotype XV, n = 5; genotype XVI, n = 4; genotype XVII, n = 7; genotype XVIII, n = 6. Standard error estimates are shown above the diagonal based on a bootstrap procedure (500 replicates). Analyses were conducted with the maximum composite likelihood model. The rate variation among sites was modeled with a gamma distribution (shape parameter = 1). Codon positions included were first, second, third, and noncoding. All positions containing gaps and missing data were eliminated. There were a total of 1638 positions in the final data set. Evolutionary analyses were conducted in MEGA 5.
Clinical and Gross Findings
A summary of the macroscopic lesions, together with the number of birds that died spontaneously or were euthanized for each day, is compiled in Table 2. Overall, the type and severity of lesions, as well the temporal course of the disease, were similar for all the infection groups. At day 2 pi, all birds appeared clinically normal. Shortly after, the clinical picture quickly worsened, and clinical signs consisted of ruffled plumage, conjunctivitis, inability to stand, and prostration. At day 4 pi, all birds infected with BF/08, Niger/06, Nigeria/06, or SA/08 died spontaneously or were euthanized in extremis. Gross findings were first detectable at day 2 pi in all sampled birds and worsened as time progressed. At day 2 pi, lesions consisted mainly of conjunctivitis, splenomegaly, petechial hemorrhages in thymus, and cecal tonsils. At day 4 pi, lesions were severe and consisted of extensive conjunctival edema and hemorrhages (Fig. 2), proventricular and intestinal hemorrhages (duodenum, cecal tonsils, and large intestine), necrosis of the cecal tonsils, splenomegaly and multifocal splenic necrosis (Fig. 3), multifocal hemorrhages and necrosis of the pancreas, atrophy and hemorrhages of the thymus, and comb cyanosis. Birds sacrificed in extremis were markedly dehydrated and showed multifocal accumulation of urates in the kidney. Control animals remained clinically and grossly normal.
Mortality and Macroscopic Lesions in 4-Week-Old Chickens Infected With BF/08, Niger/06, Nigeria/06, SA/08, and Controla.

Abbreviation: pi, postinfection.
aNewcastle disease virus strains: BF/08, Burkina Faso/2415-580/2008; Niger/06, Niger/1377-7/2006; Nigeria/06, Nigeria/228-7/2006; SA/08, Chicken/South Africa/08100426/2008. Numbers in parentheses indicate birds showing the lesion/total number of birds necropsied that day.
Histopathologic Findings
The severity of the microscopic changes seen in selected collected organs is presented in Table 3. The character and distribution of the lesions were similar with all 4 strains and progressed to reach a peak of severity by day 4 pi. At day 2 pi, lesions were minimal in most tissues and consisted of conjunctival edema and scattered lymphocytic apoptosis in the lymphoid organs. By day 4 pi, lesions became severe and bilaterally targeted the eyelids, the lymphoid organs (eg, thymus, bursa, and spleen; Figs. 4, 5), and the mucosa-associated lymphoid tissue (MALT), such as the lymphoid patches in the ileum (Fig. 6) and the cecal tonsils. In the eyelid, lesions consisted of diffuse subcutaneous edema, multifocal necrosis, hemorrhages, and pleomorphic infiltrate of heterophils and macrophages. The affected eyelids consistently demonstrated marked lymphangiectasia. In the lymphoid organs, lesions consisted of severe lymphoid depletion, accumulation of necrotic debris occasionally associated with mats of fibrin, and infiltration with prominent macrophages and fewer heterophils. In the intestines, destruction of the MALT resulted in necrosis and ulceration of the overlying epithelium (Fig. 6). With all 4 strains, the respiratory tract had minimal lesions overall: in the lungs, there was generalized mild to moderate depletion of bronchus-associated lymphoid tissues with concomitant infiltrate of histiocytes and fewer heterophils. No lesions in the trachea or the respiratory parenchyma were observed. Lesions in the brain consisted of randomly scattered necrotic foci, minimal to mild acute hemorrhage, focal gliosis, and/or mild perivascular histiocytic infiltrate and vascular reactivity.
Immunohistochemistry
The distribution, timing, and intensity of immunostaining for nucleoprotein were similar for all 4 NDV strains; immunohistochemical findings are summarized in Table 3. In the infected tissues, the intracellular immunolabeling for NDV was intracytoplasmic and finely to coarsely granular, with occasionally extracellular signal (fine granular staining) observed in necrotic areas. For all viruses, by day 4 pi, antigen was widely distributed in the tissues, with total of 23, 22, 22, and 22 positive tissues out of 25 for BF/08, Niger/06, Nigeria/06, and SA/08, respectively. Intensity and distribution of positive immunohistochemical signal were highest at day 4 pi for all 4 strains. Strongest signal was observed in the eyelids, the lymphoid organs, the mucosa-associated lymphoid aggregates (MALT) in multiple organs, and the bone marrow. In the lymphoid organs, immunolabeled cells consisted of lymphocytes and macrophages. In the spleen, positive signal was observed in the fixed macrophages surrounding the penicilliary arteries (Fig. 7). In the cecal tonsils, immunolabeling was prominent within the lamina propria and submucosa and scattered within few cells in the epithelium (Fig. 8). In the bursa of Fabricius, immunolabeling was prominent within the medulla and cortex (Fig. 9). In the respiratory system, positive cells were mainly confined to the lymphoid aggregates present in the larynx (laryngeal tonsils), trachea, and the secondary and tertiary bronchi (bronchus-associated lymphoid tissues). In the intestine, scattered myenteric plexuses showed positive staining in cells consistent with neurons and glia. In the brain, staining was observed in rare neurons. Scattered Kupffer cells were also positive in the liver.
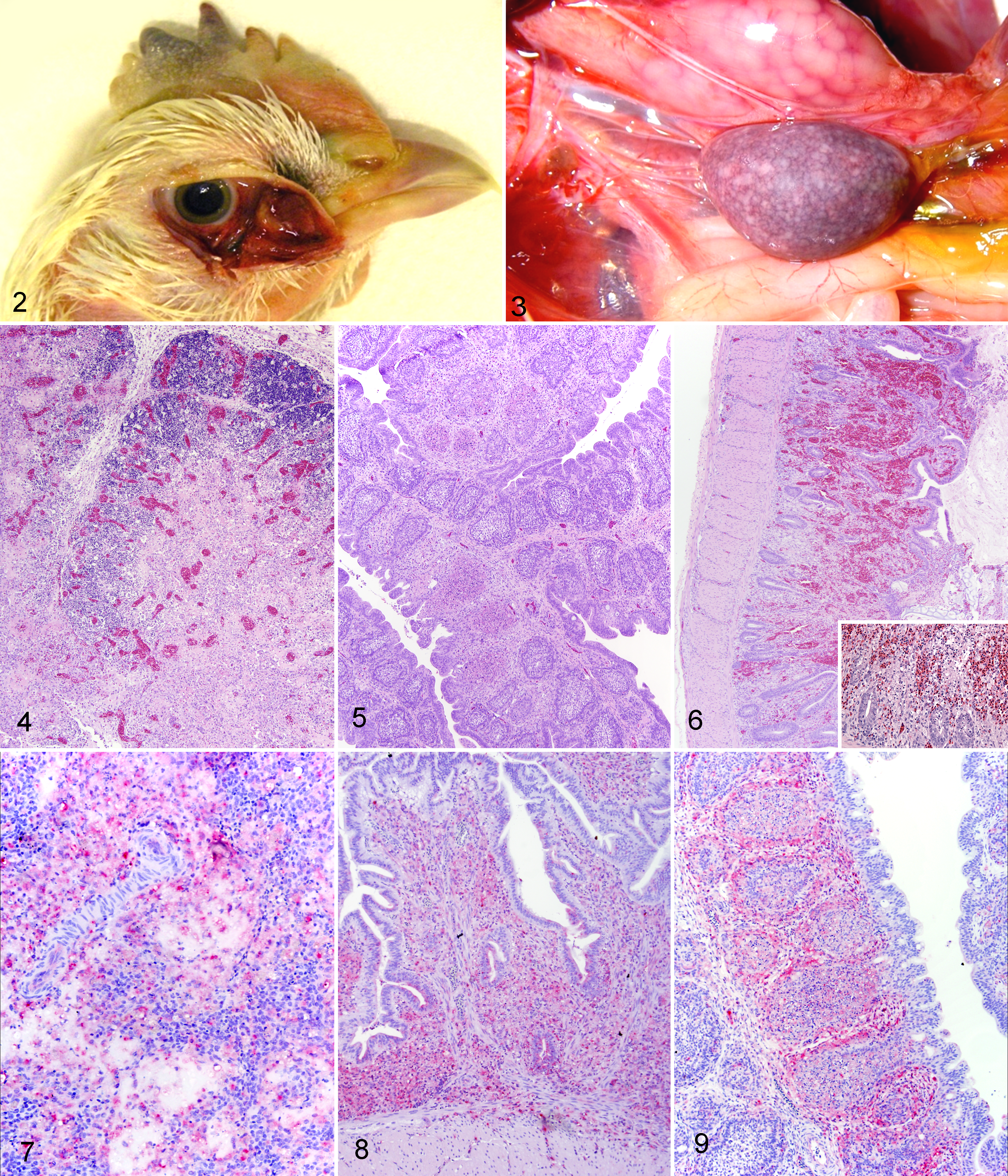
Eyelid; 4-week-old White Leghorn chicken infected with Niger/06 strain, 4 days postinfection (dpi). Hemorrhages in the eyelid and periocular soft tissue, associated with moderate cyanosis of the comb.
Histologic and Immunohistologic Findings in Chickens Infected With 4 African Strains of Newcastle Disease Virus.
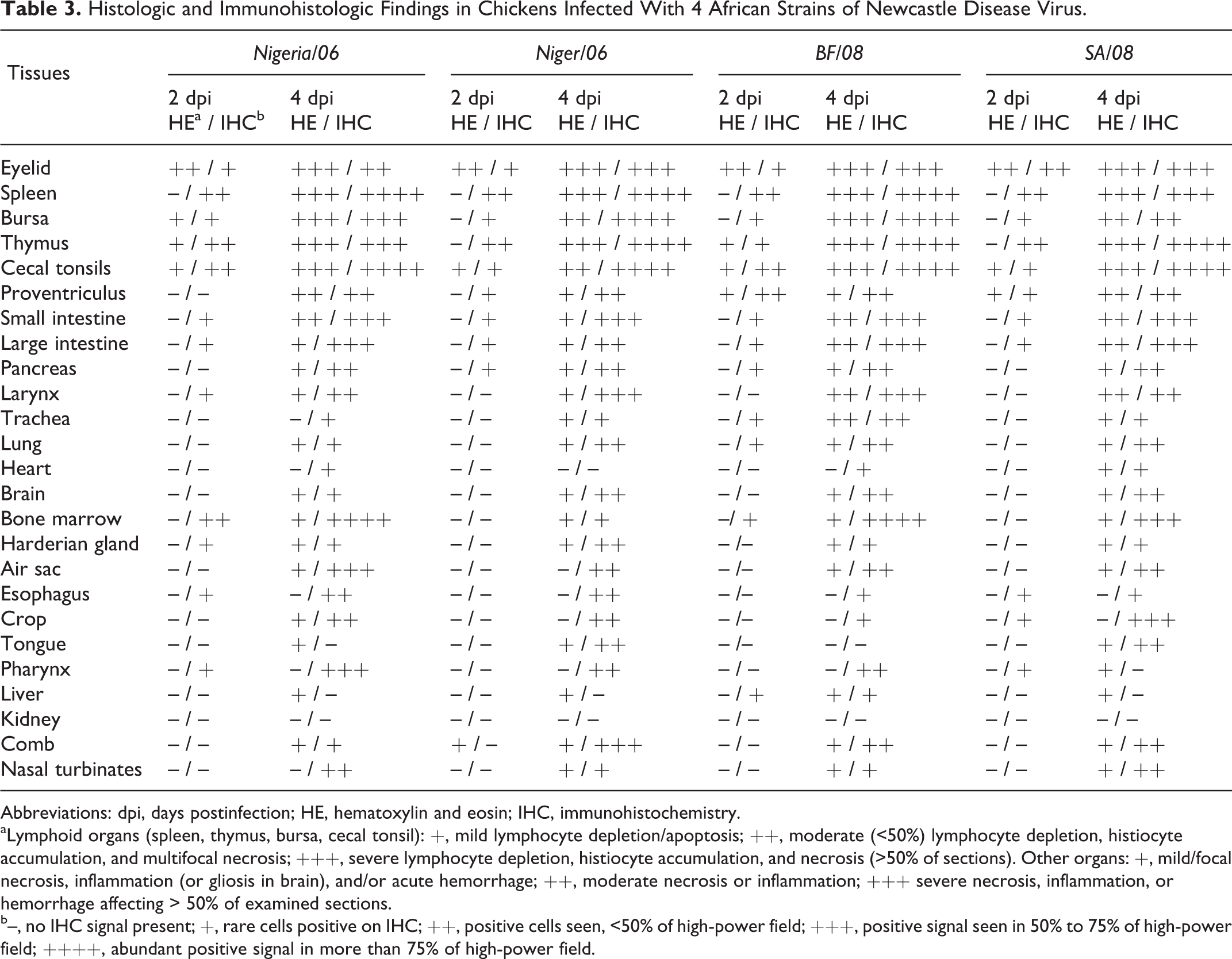
Abbreviations: dpi, days postinfection; HE, hematoxylin and eosin; IHC, immunohistochemistry.
aLymphoid organs (spleen, thymus, bursa, cecal tonsil): +, mild lymphocyte depletion/apoptosis; ++, moderate (<50%) lymphocyte depletion, histiocyte accumulation, and multifocal necrosis; +++, severe lymphocyte depletion, histiocyte accumulation, and necrosis (>50% of sections). Other organs: +, mild/focal necrosis, inflammation (or gliosis in brain), and/or acute hemorrhage; ++, moderate necrosis or inflammation; +++ severe necrosis, inflammation, or hemorrhage affecting > 50% of examined sections.
b–, no IHC signal present; +, rare cells positive on IHC; ++, positive cells seen, <50% of high-power field; +++, positive signal seen in 50% to 75% of high-power field; ++++, abundant positive signal in more than 75% of high-power field.
Vaccination Experiment
Results of vaccination are summarized in Tables 4 and 5. All vaccinated birds survived the challenge without any noticeable clinical illness. Nonvaccinated animals (mock) died, on average, at 3.5 days postchallenge for BF/08 and 3.8 days for SA/08. Average arithmetic mean of HI antibody response after each vaccination for each group was similar among the groups, except for the prechallenge antibody titers for the QV4/QV4 group after the first vaccination, which were twofold higher. However, when the geometric mean was calculated removing the high and low outliers for these 2 groups (in parentheses, Table 4), the values are consistent with the other prechallenge HI values to homologous antigen. The absent or low increase in HI titers between the first and second vaccine doses is consistent with the short time passed between the first vaccination and boost, only 5 days (Table 4). Prechallenge HI antibody titers to the group challenge showed that all 3 vaccine schemes gave positive values (>16) for SA/08, and there was higher response to the SA/08 antigen than to the BF/08 antigen. The prechallenge HI titers to the BF/08 antigen were negative (<16) for all 3 vaccine schemes. The postchallenge HI titers to the homologous challenge antigens were elevated after challenge, suggesting viral replication with both BF/08 and SA/08.
Mortality and Hemagglutination Inhibition Test Titers During the Vaccination Experiment.
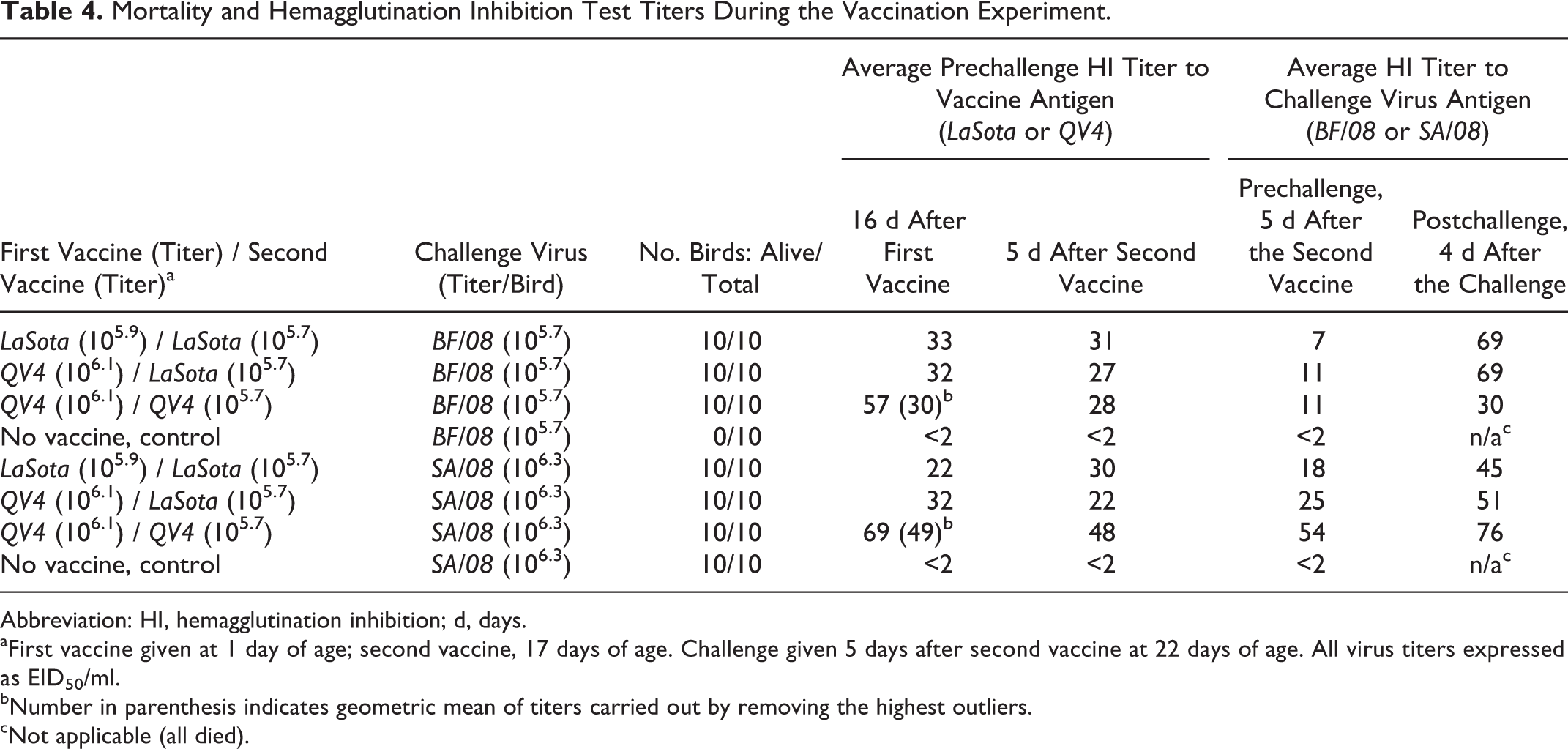
Abbreviation: HI, hemagglutination inhibition; d, days.
aFirst vaccine given at 1 day of age; second vaccine, 17 days of age. Challenge given 5 days after second vaccine at 22 days of age. All virus titers expressed as EID50/ml.
bNumber in parenthesis indicates geometric mean of titers carried out by removing the highest outliers.
cNot applicable (all died).
Nonvaccinated chickens (control) shed significantly larger amounts of either challenge viruses compared to the vaccinated birds in both oral and cloacal swabs at day 2 postchallenge (P ≤ .05, Table 5). Regarding only the vaccinated groups, no significant difference in the amount of shed challenge virus was observed between vaccination schemes or between challenge viruses, either when birds not shedding virus were included in the calculation (data not shown) or when the amounts from only the birds shedding virus were averaged (Table 5). There were significantly fewer birds shedding virus compared to the nonvaccinated groups on day 2 for oral and cloacal swabs, except for the number of birds shedding from oropharyngeal swabs for the LaSota/LaSota vaccine group and challenged with SA/08 (Table 5). No control birds survived for statistical analysiss of number of birds shedding on day 4 postchallenge.
Discussion
New genotypes of NDV are continuously described worldwide. To date, 18 class II genotypes of NDV have been described. 13,15,39 The increasing genomic diversity of NDV poses several problems for the control and surveillance of ND. In fact, genetically distant NDV strains might have the potential for increased virulence, increased host range, or vaccine escape. 1 It has been observed, for example, that genome-heterologous vaccines protect against morbidity and mortality but allow more shedding of challenge virus compared to genotype-homologous vaccination. 30,32 Additionally, reports describing possible outbreaks in vaccinated flocks raise concerns regarding the true efficacy of commonly used vaccines against genetically distant NDV strains. 1,7,20,29,46 The genotypes involved in the most recent outbreaks worldwide belong to genotypes V (Central and South America), VI (Europe), and VII (China, South Africa). 1,29
Shedding of Challenge Virus in Vaccinated and Control Birds.
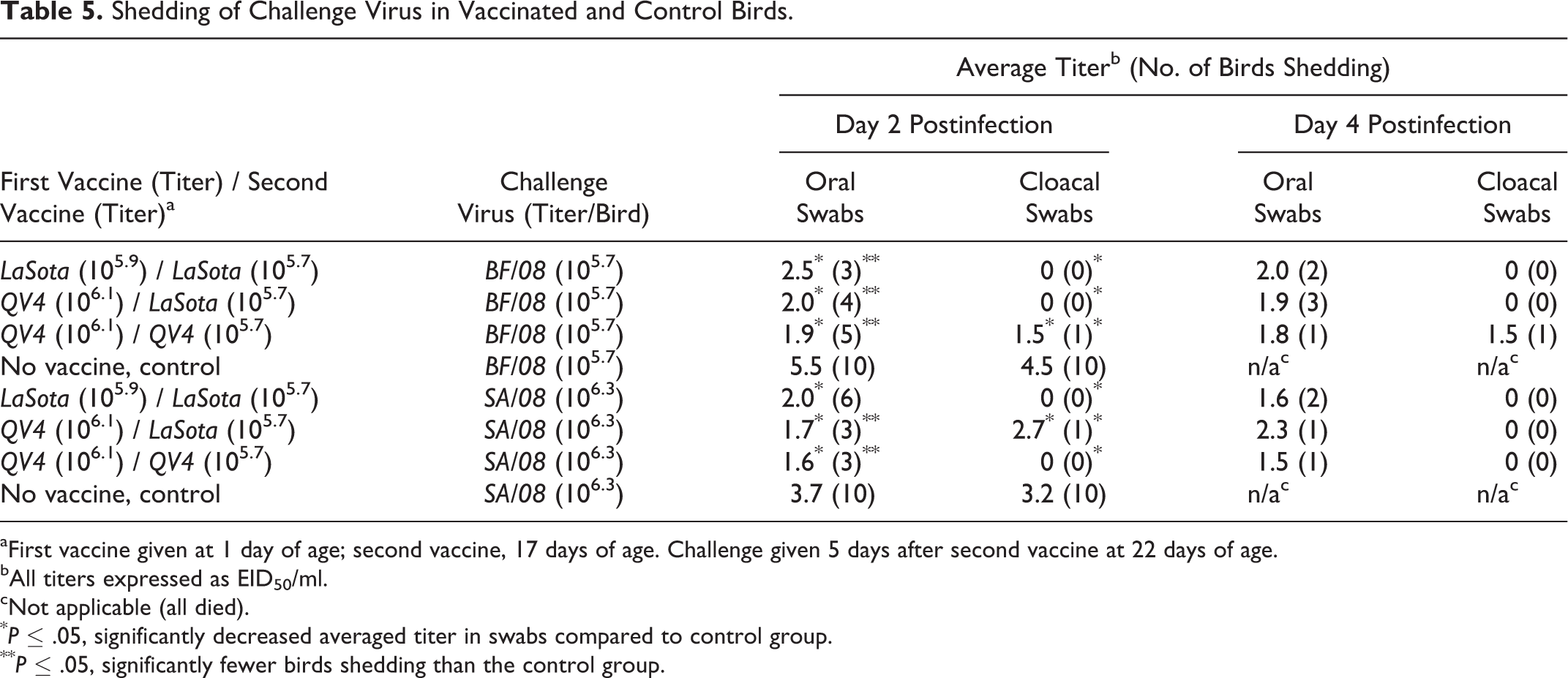
aFirst vaccine given at 1 day of age; second vaccine, 17 days of age. Challenge given 5 days after second vaccine at 22 days of age.
bAll titers expressed as EID50/ml.
cNot applicable (all died).
* P ≤ .05, significantly decreased averaged titer in swabs compared to control group.
** P ≤ .05, significantly fewer birds shedding than the control group.
On the continent of Africa, reports describing NDV circulation are limited. Virulent NDV strains of genotypes VII and VIII have been described in South Africa 22,29 ; genotype XI 27 has been described in Madagascar; and genotype II has been reported to circulate in Egypt. 33 Few data, however, have been reported for West and Central Africa. Only recently, an increasing number of reports have shown that a new lineage of NDV (referred to as lineage 7 according to Aldous classification 3,9 ) is endemic in West and Central African countries (Nigeria, Burkina Faso, Cameroon, Mauritania, Ivory Coast). 9,40,45 A more recent implementation of the classification of these strains according to the criteria proposed by Diel 15 showed that lineage 7 encompasses NDV genotypes XIV, XVII, and XVIII, the last 2 never previously described. 39 To increase the genetic diversity of these strains, some viruses isolated from Togo and Benin showed a particular genomic arrangement (addition of 6 nucleotides between HN and L), which was not described previously. 37
In the present study, 3 representatives of the newly described West African NDV genotype XIV and XVII were characterized by detailed clinicopathologic assessment and compared to a virulent NDV strain of genotype VII (SA/08), isolated from South Africa. Phylogenetic analysis of the full fusion sequence confirmed that Niger/06 and BF/08 belong to genotypes XIV and XVII, respectively, in agreement with the recent work by Snoeck and colleagues. 39 Phylogenetic analysis and distance matrix showed that Nigeria/06 belongs to genotype XVII, whereas SA/08 was included within genotype VII, as expected. 22
All 4 NDV strains tested in this study have a virulent configuration of the cleavage site, and all had an ICPI score greater than 0.7, which is typical of virulent NDV strains. 5,10,36 Based on these characteristics, the 3 West African strains and the South African NDV strain are considered virulent by the OIE standards. 36
Birds inoculated with BF/08, Niger/06, Nigeria/06, and SA/08 displayed severe clinicopathologic disease, with death or euthanasia occurring for all birds (100% mortality) by day 4 pi. Based on numerous pathogenesis studies done in our laboratory in a consistent manner with multiple strains, death of birds by day 4 pi indicates a strain of considerable virulence and is comparable to many other highly velogenic viscerotropic NDV strains. 8,10,13,16,42 Principal gross lesions consisted of severe conjunctivitis, visceral hemorrhages, splenic necrosis, and thymic hemorrhages and atrophy, all of which are consistent with other velogenic viscerotropic NDV strains. 10 Histologically, all strains behaved as a typical highly virulent strain, with extensive necrosis of lymphoid tissues, especially those of the spleen and intestine. 8,10,13,16,42 By immunohistochemistry, animals infected with BF/08, Niger/06, Nigeria/06, and SA/08 strains had 23, 22, 22, and 22 positive tissues out of 25, respectively, indicating a marked systemic pantropism, which is also typical of highly virulent NDV strains. 10 In light of the results presented herein—that is, highly virulent phenotype and clinical protection provided by vaccination—it is hard to reconcile the numerous reports describing isolation of genotype XIV, XVII, and XVIII NDV strains from apparently clinically healthy chickens, 37,38 unless the birds were already immunized against NDV (previous exposure to lentogenic or vaccine strains circulating in the environment) or the samples were collected during the incubation period.
Due to the raising concerns that phylogenetically distant genotypes could escape the protection offered by traditional vaccination programs, we evaluated also the efficacy of traditional vaccines (LaSota and QV4) against a representative of genotype XVII NDV strain (BF/08). SA/08, a genotype VII NDV strain circulating in South Africa, was used as a control for the challenge virus. Results showed that both LaSota and QV4, regardless of the vaccination scheme, proved to protect animals against morbidity and mortality. Furthermore, no significant differences in shedding were observed between the genotype XVII (BF/08) and genotype VII (SA/08) NDV challenge virus. These data show that under experimental conditions, vaccination with LaSota and QV4 NDV vaccines are very efficient in protecting animals against strains of the new genotype XVII and that genomic variation did not affect virus shedding in comparison to other genotypes already circulating in Africa. These results confirm what has been shown by other authors: provided a sufficient immune response (ie, adequate HI titers before challenge), even significant genetic distances between vaccine and challenge viruses do not cause development of clinical signs. 12,17 Even though the data reported here are from highly controlled experimental conditions with SPF chickens lacking maternal antibodies, caution is warranted in considering genetic distance between NDV vaccines and the challenge virus as the sole cause of the reported cases of vaccine escape in the field. The high variability in the field, coupled with inadequate implementation of biosecurity protocols, can cause poor or suboptimal vaccination, ultimately leading to poor flock immunity and increased susceptibility to infections, including virulent NDV.
In conclusion, all 3 West African NDV strains (BF/08, Niger/06, and Nigeria/06) were highly virulent and exhibited a broad tissue tropism. Lesions and antigen distribution in the newly classified viruses of genotype XIV and XVII were equally severe to those observed with the South African NDV strain representing genotypes known to circulate in South Africa (genotype VII). 22 The high virulence of these new genotype underscores the importance of ongoing surveillance and monitoring for newly emerging NDV strains in Africa and the need for experimental infection studies to fully assess potential pathogenicity in poultry and other species.
Footnotes
Acknowledgements
We would like to acknowledge Dr I. Bangana and Dr H. Thiam for submitting samples to IZSVe, Padua, Italy, and the RESOLAB network for its technical support in facilitating samples submission and exchange of information. We also would like to acknowledge Marcelo Pedrotti de Cesaro for assistance in the pathology portion of this work, Tim Olivier, and Dawn Williams-Coplin for technical assistance with the animal experiment and sequencing.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: This research was supported by USDA-ARS CRIS 6612-32000-064.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
