Abstract
Cloning and sequencing of the progesterone receptor gene in dogs have revealed 2 isoforms, A and B, transcribed from a single gene. Distribution of isoforms A and B in canine mammary lesions has hitherto been investigated only by Western blot analysis. This study analyzed progesterone receptor and its isoforms in formalin-fixed, paraffin-embedded tissue samples from canine mammary lesions (4 dysplasias, 10 benign tumors, and 46 carcinomas) using 1-step SYBR Green quantitative real-time polymerase chain reaction (RT-qPCR). Progesterone receptor was expressed in 75% of dysplasias, all benign tumors, and 59% of carcinomas. Carcinomas, and particularly simple epithelial-type carcinomas, displayed the lowest levels of expression. A high rate of agreement was recorded between RT-qPCR and immunohistochemical labeling. Isoforms A and B were successfully amplified, with correlation coefficients of 0.99 and amplification efficiencies close to 2, and were expressed in all lesion types analyzed. Predominance of A over B expression was observed in carcinomas and complex adenomas. Low-grade tumors exhibited higher progesterone receptor messenger RNA (mRNA) levels, but no difference was observed in the expression of isoform A versus B. Analysis of progesterone receptor mRNA isoforms by RT-qPCR was successful in routinely formalin-fixed, paraffin-embedded tissue samples and enabled the distribution of isoforms A and B to be identified for the first time in dysplasias, benign tumors, and malignant tumors of the canine mammary gland. These findings will facilitate future research into the role of progesterone receptor isoforms in the progression of canine mammary tumors.
Keywords
Epidemiologic, clinical, and experimental data indicate that canine mammary tumors (CMTs) are hormone-dependent; that is, they are strongly influenced by ovarian hormones, mainly estrogens and progesterone (P). 35 P acts through binding to its cognate P receptor (PR), a member of the nuclear steroid receptor family. 12 PR expression is currently measured by immunohistochemical (IHC) methods, which have shown that all benign CMTs and two-thirds of malignant CMTs express PR. 7,13,25 IHC expression of PR is a favorable prognostic indicator 25,34 and a predictive marker of positive response to neoadjuvant administration of antiprogestins 14 in canine mammary carcinoma.
The PR exists as 2 isoforms, PRA and PRB, which are expressed from a single gene in both humans and rodents. 19 PRA and PRB have been shown to have different functions as well as different levels of expression in breast cancer. 33 Human PRB is a stronger transcriptional activator than human PRA, due in part to a third activation domain (AF-3) located within the 164-amino acid N terminal. 34 Human PRA acts a repressor capable of inhibiting other receptors, including estrogen receptor and human PRB. 37 PRA and PRB are generally expressed at similar levels in normal mammary tissue, but in breast tumors their ratio is altered, with a predominance of PRA and loss of PRB. 6,19,28
In dogs, cloning and sequencing of the PR gene have confirmed the presence of isoforms PRA and PRB. 23 Western blot analysis has revealed the predominance of PRA over PRB staining in normal, hyperplasic, and neoplastic canine mammary tissue samples. 11 PRA expression is reportedly more marked in carcinomas than in normal and hyperplastic tissue. 11
Quantitative real-time polymerase chain reaction (RT-qPCR) is currently considered the most sensitive method for PR isoform detection, given that both Western blot and IHC methods provide limited detection sensitivity. 19,27 This has prompted growing interest in using RT-qPCR for the retrospective analysis of the vast archives of formalin-fixed, paraffin-embedded (FFPE) tissue samples available in veterinary pathology laboratories.
This study examined total PR, PRA, and PRB expression in FFPE dysplasias, benign tumors, and malignant tumors of the canine mammary gland using a RT-qPCR method. Total PR expression was also analyzed by IHC for purposes of comparison.
Materials and Methods
Samples
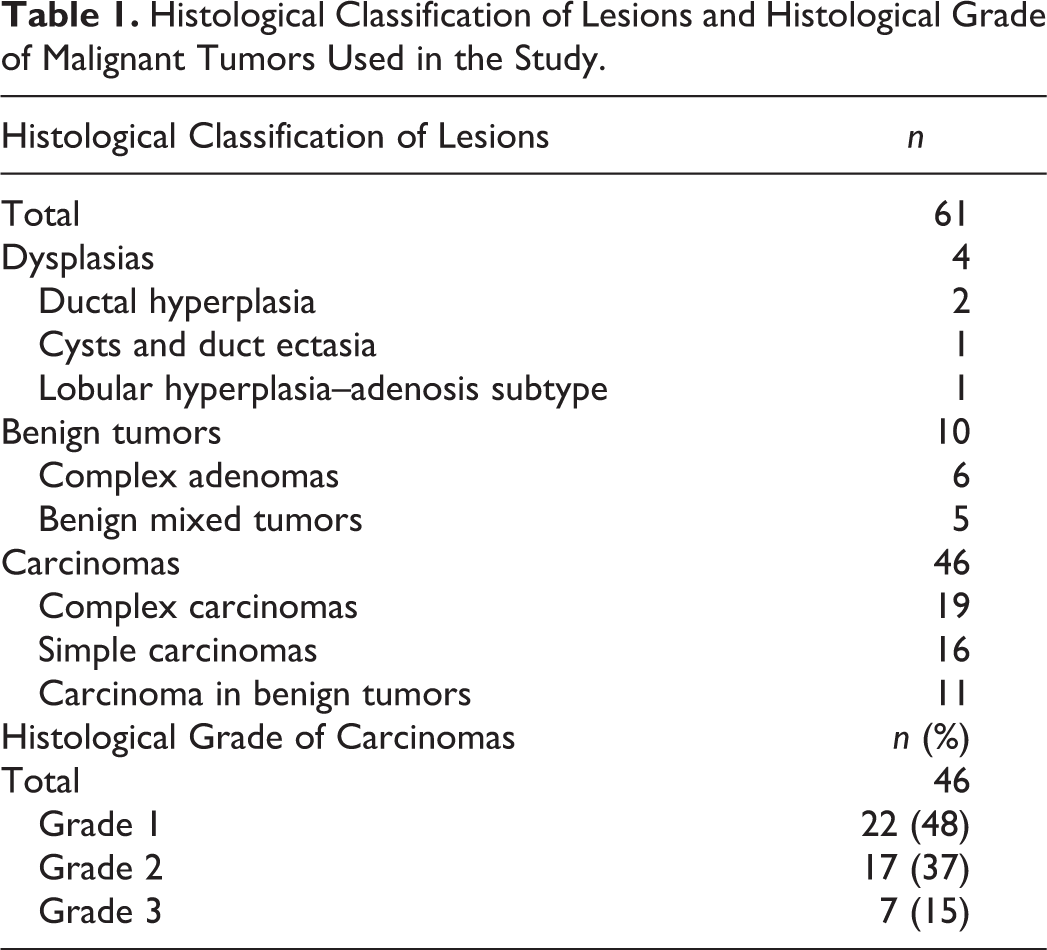
FFPE tissue samples from 61 lesions of the canine mammary gland were retrieved from the archives of the Department of Comparative Pathology at the University of Córdoba (Spain). Samples had been obtained for an earlier prospective study and had been fixed for 24 to 48 hours in 10% buffered formalin and stored as paraffin blocks at 4°C between 0.5 and 2 years prior to this study. The histological classification 26 and grading 21 of lesions are shown in Table 1. In carcinomas, most malignant-appearing areas were selected for both immunohistochemistry and RT-qPCR. Tissue samples from 2 fresh canine mammary tumors were used as controls for validating RT-qPCR and then fixed for 24 hours in 10% buffered formalin and routinely embedded in paraffin wax.
Histological Classification of Lesions and Histological Grade of Malignant Tumors Used in the Study.
Extraction, Quantification, and Quality Assessment of Messenger RNA
RNA was isolated from FFPE and fresh samples using the RNase FFPE kit and the RNeasy Protect Mini kit (Qiagen, Copenhagen, Denmark), respectively, according to manufacturers’ recommendations, and stored at –80°C until use.
RNA yields were determined by measuring spectrophotometric absorbance at 260 nm (A260) using the NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, Delaware). A ratio of absorbance at 260 nm and 280 nm of 1.8–2.0 was accepted as “pure.” Total RNA integrity was checked by denaturing agarose gel electrophoresis and ethidium bromide staining, which shows respective messenger RNAs (mRNAs) as sharply defined bands.
RT-PCR Assay
Primer design
The canine PR genomic sequence was obtained from NCBI GenBank database under the gene ID NM_001003074. Two primer pairs were designed specifically to target the coding regions of isoforms A and B previously reported by Lantinga-van Leeuwen et al 23 using Primer3Plus. Two canine housekeeping genes, hypoxanthine phosphoribosyltransferase 1 (HPRT1: NM_001003357.1) and canine ribosomal protein L32 (RPL32: NM_001252169.1), were selected for the study, and 2 primer pairs were designed using the same tool. These 2 reference genes have proved suitable as internal controls for RT-qPCR in CMTs. 10,18
Sequences of the forward and reverse primers for PR isoforms and housekeeping genes are summarized in Table 2. Primers spanning or flanking 1 intron were chosen wherever possible to minimize inaccuracies due to genomic DNA contamination. The 100% specificity of the different primers was verified by a BLAST search.
Primer Sequences and Product Length for Quantitative Real-Time Polymerase Chain Reaction Amplification.
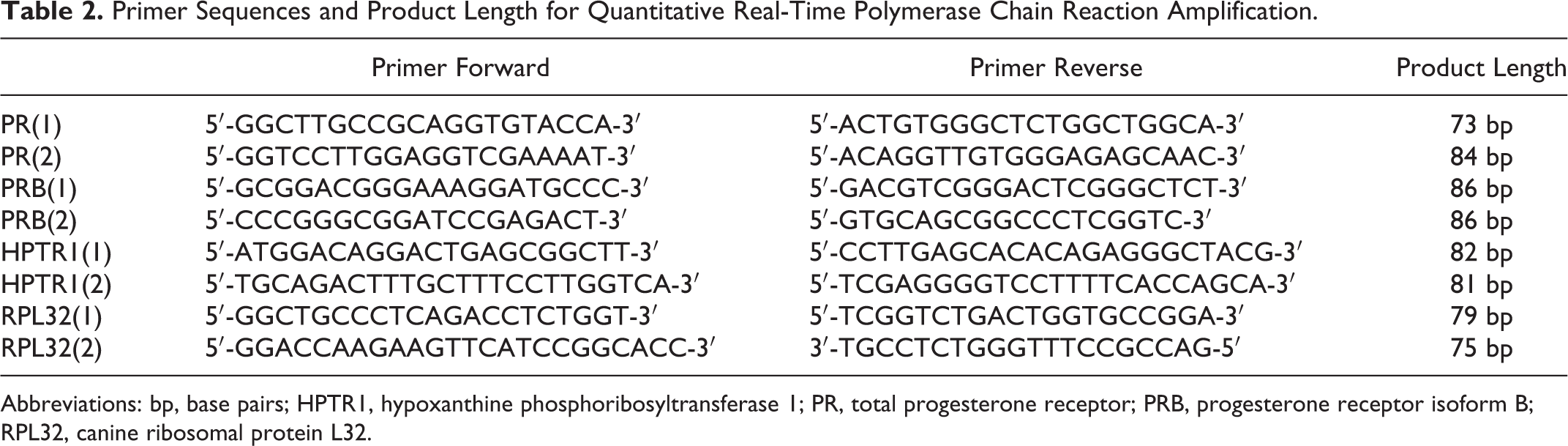

Abbreviations: bp, base pairs; HPTR1, hypoxanthine phosphoribosyltransferase 1; PR, total progesterone receptor; PRB, progesterone receptor isoform B; RPL32, canine ribosomal protein L32.
RT-qPCR amplification
RNA was amplified using the LightCycler 480 Real-Time PCR System. One-step RT-qPCR was performed using the QuantiFast SYBR Green RT-PCR (Qiagen, Denmark) for a total volume of 10 μl and a template concentration of 5 ng/μl according to the manufacturer’s recommendations. Thermal cycling conditions were 50°C for 10 minutes (RT step) and 95°C for 5 minutes, followed by 40 cycles of 95°C of 10 seconds and 60°C for 30 seconds. A melting curve analysis was performed following every run to ensure a single amplified product for every reaction (1 cycle of 95°C, with continuous acquisition mode and ramp rate of 0.1°C per second). RT-qPCR products were analyzed by agarose gel electrophoresis. Reverse transcription negative controls and nontemplate controls were included. All reactions were performed in triplicate in 96-well reaction plates (Applied Biosystems, Foster City, California).
Standard curves generated from series of dilutions (1–20 ng/μl) of a known sample to encode the canine PR gene were used to determine qPCR efficiency (E = 10(-1/slope)) and to establish the linear range of the assay.
The reliability of the RT-qPCR was defined by calculating the coefficient of variation (CV) of replicates for each analyzed sample (intra-assay variability) and for each plate (interassay variability). CV was calculated as follows: (SD/Ct average) × 100, where SD is standard deviation and Ct is threshold cycle. Relative mRNA expression was defined as 2-▵Ct, where ▵Ct = CtTARGET – CtRPL32/HPTR1, and CtRPL32/HPTR1 is the average of the Ct values of the 2 housekeeping genes for each sample. The amount of PRA was calculated by subtracting the relative amount of PRB from that of total PR. 15,36 The PR positive-status cutoff was set at 0.04, since this value agreed best with IHC results (see below).
IHC Assay
To detect total PR by IHC, commercial mouse monoclonal anti-human PR antibody (clone 10A9, Immunotech, Marseille, France) diluted 1:500 and the avidin-biotin-peroxidase complex technique (Vector, Burlingame, California) were used as previously reported. 7 This antibody is raised against the recombinant hormone-binding domain of human PR located in the C-terminal region common to PRA and PRB. 20 For quantitative analysis of total PR expression, digital images pictures were captured at 40× magnification from 15 randomly selected, neighboring, nonoverlapping fields of each sample labeled with anti-PR antibody. The number of positive and negative cells was counted with ImageJ software (ImageJ 1.43, National Institutes of Health, Bethesda, Maryland). A minimum of 1000 tumor cells were counted per case. PR expression was expressed as the percentage of positive cells with respect to the total number of cells. The cutoff for the determination of PR positive-status was set at 10%. 14
Statistical Analysis
Statistical analysis was performed using the GraphPad Prism 5 software package version 5.01 (GraphPad Software Inc, San Diego, California). The D’Agostino-Pearson test was used to assess normality of data and the Mann-Whitney U-test to analyze RT-qPCR (total PR, PRA, and PRB) and IHC (total PR) results as a function of histological types and subtypes and histological grade. The agreement between RT-qPCR and IHC findings was estimated by Cohen’s κ coefficient. A P value <.05 was regarded as statistically significant.
Results
RT-qPCR Validation
A total of 61 FFPE CMTs were processed. Mean RNA content was 46 ng/μl (range 9.7–189.3 ng/μl) and mean purity obtained at 260/280 nm was 1.85 (1.6–2.1). Only 1 sample (<2%) histologically classified as a benign mixed tumor was excluded due to low quantity and poor quality. For canine fresh tissues and their equivalent FFPE tissues, mean values for RNA content and purity were 242 ng/μl and 1.8 and 180.7 ng/μl and 1.7, respectively.
All RT-qPCR amplification plots displayed adequate amplification curves with an exponential phase followed by a nonexponential phase, ending in a plateau. Melting analysis showed curves with a single peak and adequate melting temperatures (Tm) for housekeeping genes and PR. For PRB, however, the melting curve was characterized by a sharp peak at 91°C, although multiple extra peaks were observable at lower Tm, this being consistent with agarose gel data. Nontemplate controls and reverse transcription negative controls showed no amplification.
RNA isolated from the 2 fresh CMT samples yielded Ct values 4 to 6 cycles lower than those obtained with the same RNA isolated from equivalent FFPE tissues (Table 3). In both cases, however, a clear peak in the melting curve at the same temperature confirmed the purity and specificity of amplified PCR fragments (Table 3).
Threshold Cycle Values and Melting Temperatures of Fresh and FFPE Samples of the 2 Housekeeping Genes (RPL32 and HPTR1), PR, and PRB.a
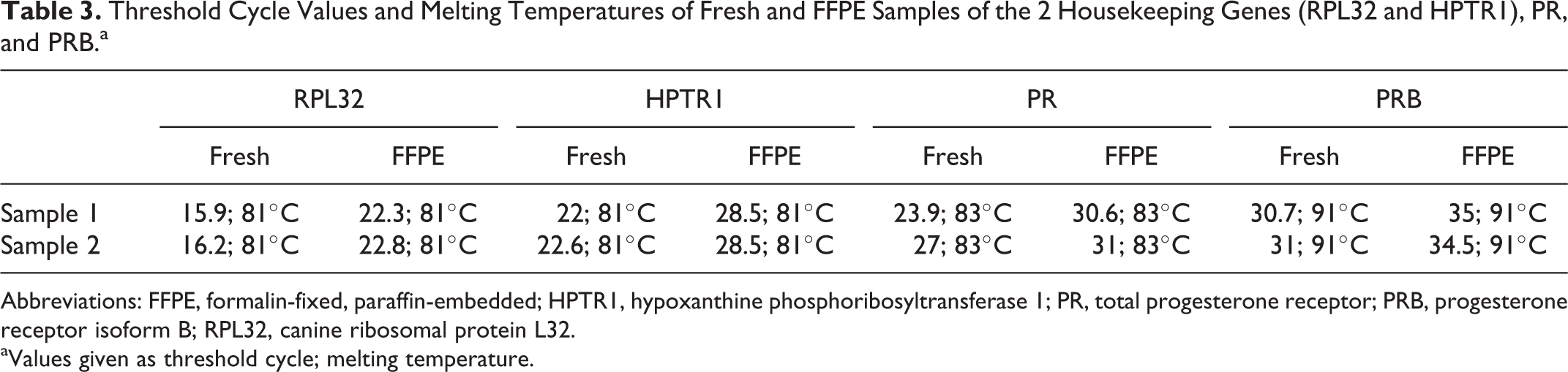
Abbreviations: FFPE, formalin-fixed, paraffin-embedded; HPTR1, hypoxanthine phosphoribosyltransferase 1; PR, total progesterone receptor; PRB, progesterone receptor isoform B; RPL32, canine ribosomal protein L32.
aValues given as threshold cycle; melting temperature.
The molecular weight of the PCR products arising from primer pairs was verified on agarose gels; findings (Table 2) confirmed their specificity. Correlation coefficients (R 2 ) between 0.98 and 0.99 were recorded for all qPCR assays, while E values lay between 2.19 and 2.89. Primer-pair total PR (1), PRB (2), HPTR1 (2), and RPL32 (1) (Table 2) were selected on the basis of agarose gel analysis and better R 2 and E values.
Intra-assay variability was below 5% in all cases, and CV values for most samples ranged between 0.03 and 2.64. CV values for interassay variability ranged from 0.78 to 1.2%.
Total PR, PRA, and PRB Analysis by RT-qPCR
Seventy-five percent of dysplasias, 100% of benign tumors, and 59% of carcinomas were positive for total PR mRNA expression by RT-qPCR (Table 4). Benign tumors and dysplasias presented very similar RT-qPCR values (0.07 ± 0.04 and 0.07 ± 0.06, respectively) (Fig. 1). PR expression was weakest in carcinomas (0.05 ± 0.008) (P = .02), and especially in simple epithelial-type carcinomas (0.03 ± 0.02) (Fig. 1). Grade 1 and 2 carcinomas displayed significantly stronger expression of total PR than did grade 3 carcinomas (P = .02) (Fig. 2).
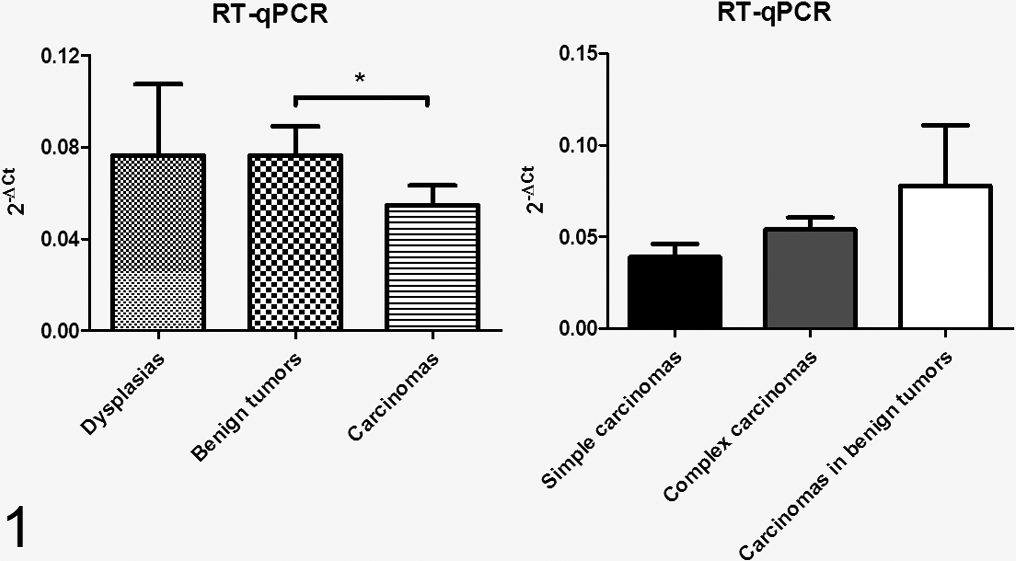
Expression of total progesterone receptor (PR) in histological tumor types and histological carcinoma subtypes by RT-qPCR. Bars with asterisk differ significantly (P < .05).
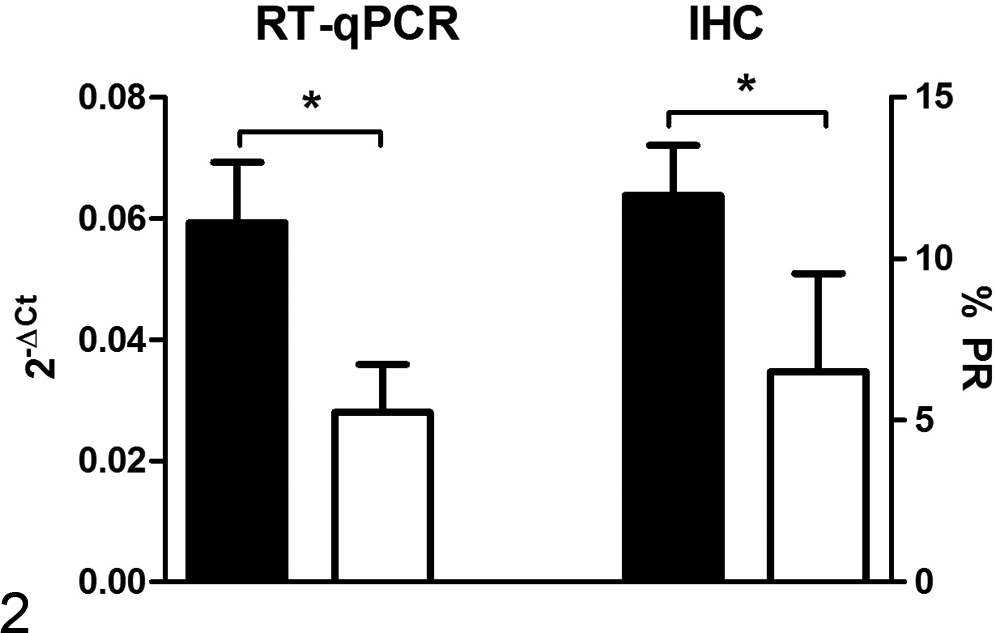
Expression of total PR by RT-qPCR (left Y axis) and by immunohistochemistry (IHC) (right Y axis) as a function of histological tumor grade. Black columns represent low-grade (grade 1 and 2) tumors and white columns high-grade (grade 3) tumors. Bars with asterisk differ significantly (P < .05).
PR-Positive Dysplasias, Benign Tumors, and Malignant Tumors by RT-qPCR and IHC.a
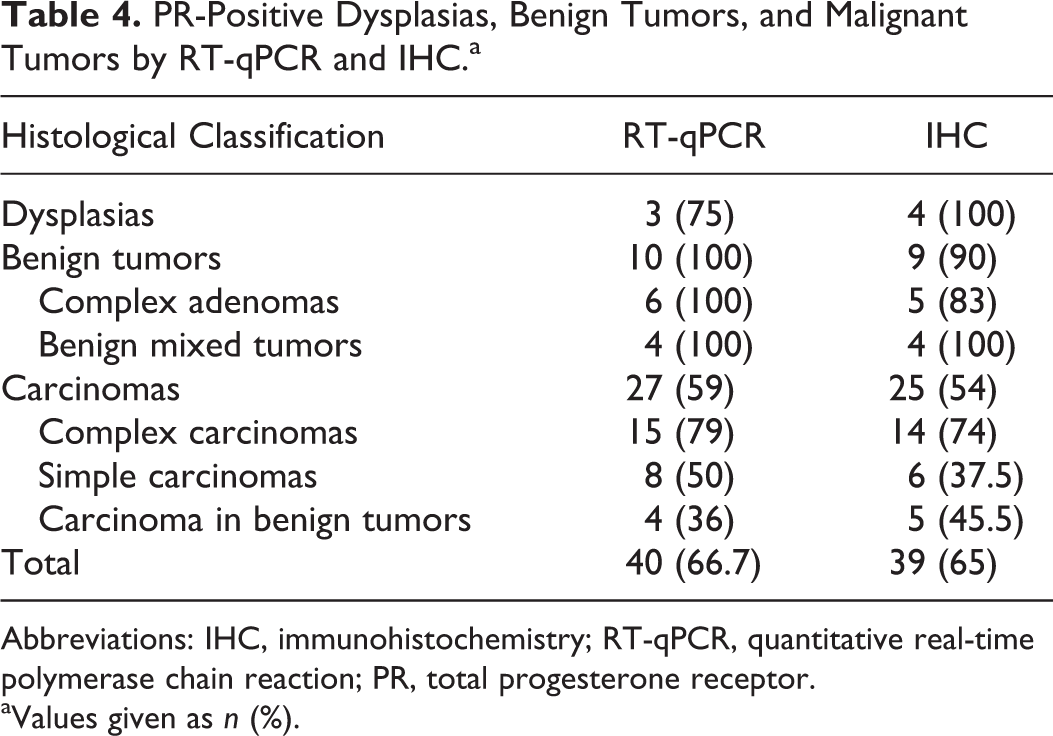
Abbreviations: IHC, immunohistochemistry; RT-qPCR, quantitative real-time polymerase chain reaction; PR, total progesterone receptor.
aValues given as n (%).
While PRA and PRB expressions were similar in dysplasias and benign tumors, PRA expression in carcinomas was greater than PRB expression (0.03 and 0.01, respectively; P = .0006) (Fig. 3). Differences between PRA and PRB were more marked in complex carcinomas (P < .0001) and simple carcinomas (P = .07) than in carcinomas in benign tumors (Fig. 3). Separate analysis of benign-tumor histological subtypes showed that PRA expression was stronger than PRB expression in complex adenomas but not in benign mixed tumors (P = .01) (Fig. 3). No differences in PRA and PRB expression were found as a function of histological carcinoma grade.
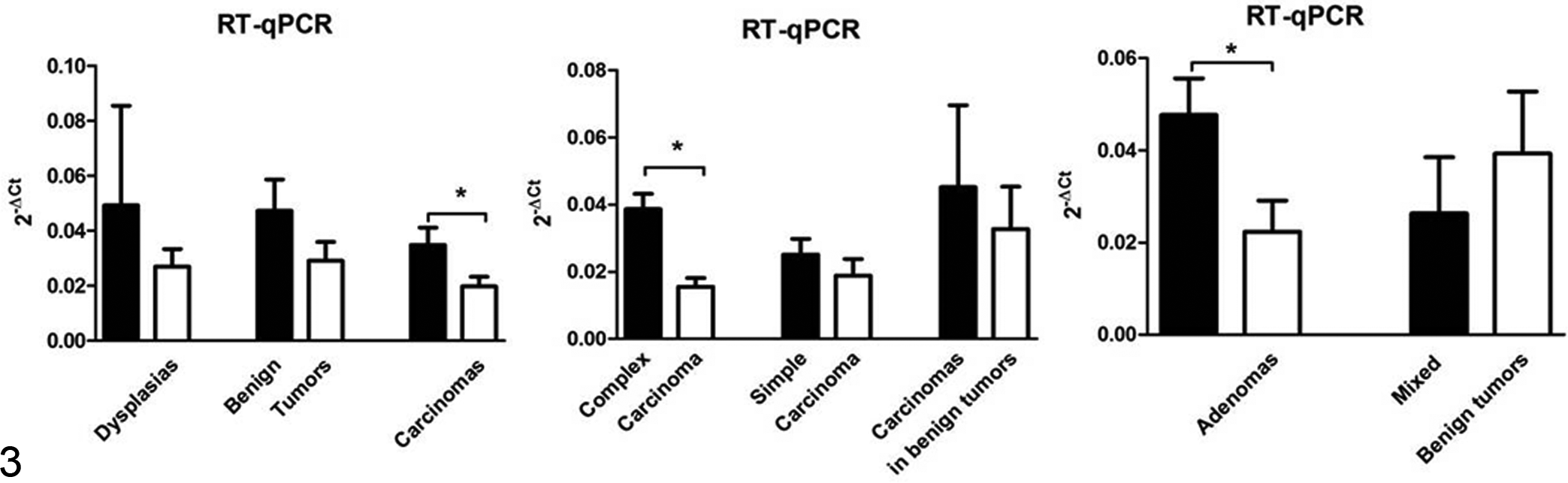
Expression of progesterone receptor isoform A (PRA) (black column) and progesterone receptor isoform B (PRB) (white column) in dysplasias, benign tumors, and carcinomas and in histological subtypes of carcinomas and benign tumors, by RT-qPCR. Bars with asterisk differ significantly (P < .05).
PR Expression by IHC
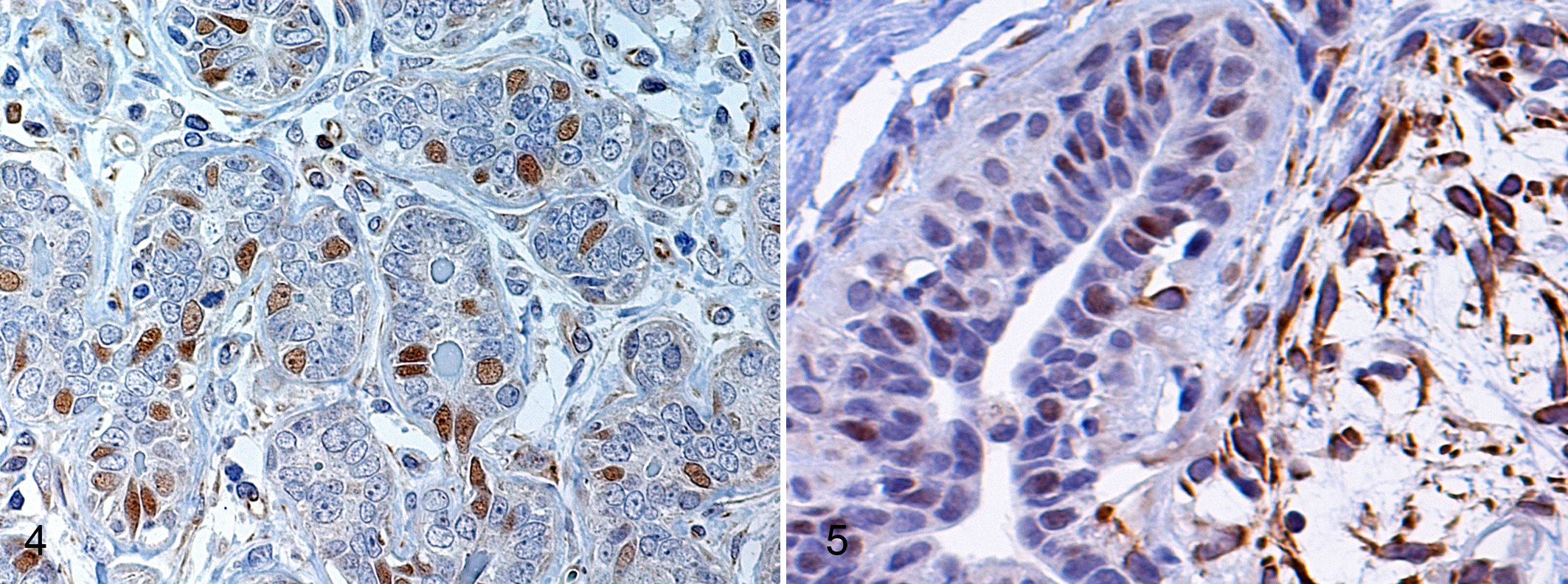
All dysplasias, 90% of benign tumors, and 54% of malignant tumors exhibited PR labeling in luminal-type epithelial cell nuclei (Fig. 4). Myoepithelial cell cytoplasm was also labeled in complex and mixed tumors (Fig. 5). PR expression was weakest in simple carcinomas. Grade 1 and 2 carcinomas displayed stronger PR labeling than did grade 3 carcinomas (P = .045) (Fig. 2).
Agreement Between IHC and RT-PCR
A similar percentage of lesions were classed as PR-positive using RT-qPCR (66.7%) and IHC (65%). Overall agreement between the 2 methods was 75% (κ index 0.4). Seven cases were PR-negative by RT-qPCR but PR-positive by IHC, while 8 cases were PR-positive by RT-qPCR and PR-negative by IHC (Fig. 6). The strongest agreement between methods was found for benign tumors (90%).
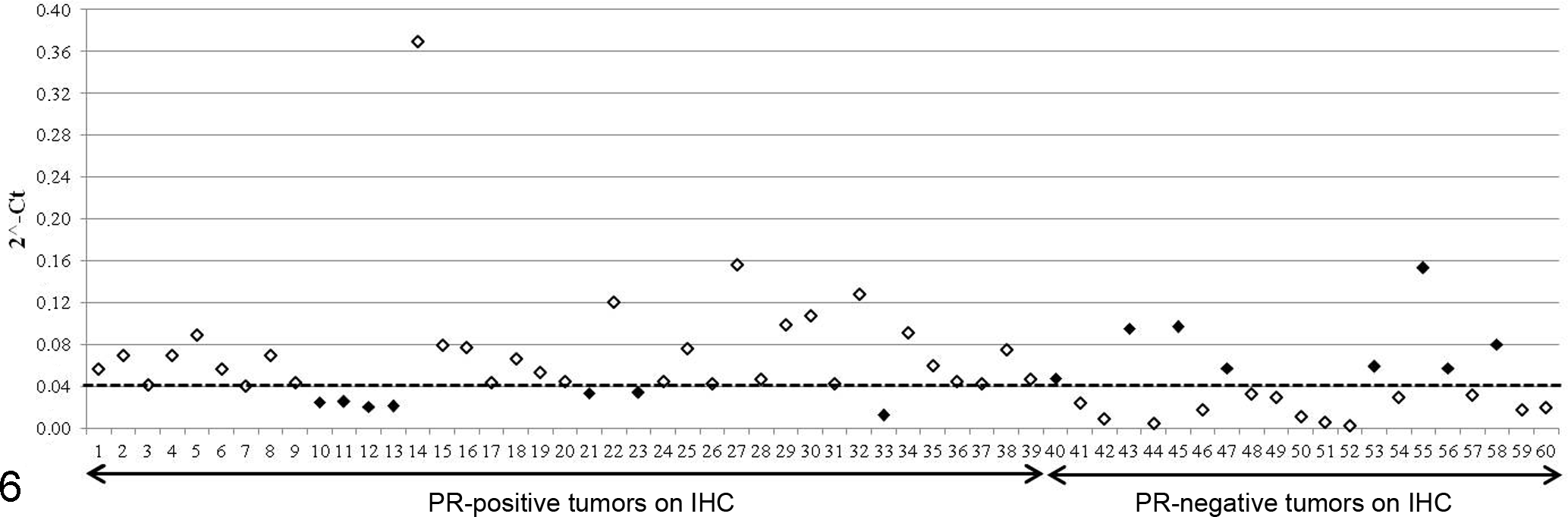
RT-qPCR versus IHC findings. Y axis shows PR expression values by RT-qPCR; X axis shows PR-positive and PR-negative samples by IHC. The cutoff for PR-positive status by RT-qPCR is indicated by the horizontal broken line. Black points indicate discrepancy between the 2 methods.
Discussion
This appears to be the first published report on the analysis of total PR, PRA, and PRB mRNA expression in FFPE tissue samples from canine mammary lesions using an RT-qPCR method. Seventy-five percent of dysplasias, 100% of benign tumors, and 59% of carcinomas were classed as positive for total PR mRNA expression. Findings were similar to those obtained with the gold-standard IHC method, which served as control. PRA and PRB mRNA expression was found in all lesion types, and a predominance of PRA over PRB expression was observed in carcinomas and complex adenomas.
Although both IHC and Western blot methods have been used to analyze PRA and PRB isoforms in human tissue samples, a number of authors report that some commercially available antibodies lack the specificity required to distinguish between the 2 isoforms. 19,28,32 In the dog, gene expression of PR has previously been reported using fresh or frozen tissue samples, 5,11,23 but no data are available on gene expression of PR isoforms in FFPE canine tissues by RT-qPCR, widely considered the most sensitive method for detecting PRA and PRB. 19 The nucleotide sequence of the canine PR gene and the specific sequences for PRA and PRB were identified from fresh mammary tissues. 23 Both isoforms contain the hormone binding domain, a highly conserved region, and a DNA binding domain; but the less conserved amino-terminal sequence is unique to the longer PRB isoform. 23 Here, total PR, PRA, and PRB mRNA was successfully amplified using canine-specific primers from archival FFPE samples. Fresh and FFPE samples exhibited identical Tm values and agarose gel bands, thus confirming the reliability of the FFPE data. The use of RT-qPCR for gene amplification from FFPE tissue samples may be affected by the process of fixation and embedding, which could exert a negative impact on the quality and usefulness of extracted RNA. 1,32 The samples used here had been fixed for less than 48 hours and stored no longer than 2 years at 4°C. In these conditions, all samples examined were deemed suitable for analysis, with acceptable results for RNA purity, yield, and integrity. A further critical factor for RT-qPCR is primer design. 19 Here, canine specific primer pairs were designed to produce an amplicon smaller than 100 base pairs, in order to ensure that sequences were template-unique, thus improving RT-qPCR efficiency. 29 Investigated transcripts showed acceptable E rates, ranging from 1 to 20 ng RNA input with high linearity. To offset interassay variation, each plate was run with its own calibrators for the standard curve under identical experimental conditions. Inter- and intra-assay variability was very low, and CV values mostly lay below 3% for both isoforms. A CV of up to 5% is considered acceptable and has no negative impact on the interpretation of results. 9 The RT-qPCR assay standardized here was therefore highly reliable and robust in terms of sensitivity, repeatability, and reproducibility for the detection of total PR, PRA, and PRB.
Seventy-five percent of dysplasias, 100% of benign tumors, and 59% of carcinomas were positive for total PR mRNA expression using RT-qPCR. These figures were similar to those obtained with the gold standard IHC method, which served as control: All dysplasias, 90% of benign tumors, and 54% of malignant tumors presented PR labeling in luminal-type epithelial cell nuclei. 25 Nuclear labeling is considered specific for PR in FFPE canine mammary tissue samples, although cytoplasmic staining of myoepithelial cells has been reported. 23,25 PR expression was weaker in carcinomas than in benign tumors and dysplasias, both by RT-qPCR and IHC, a finding also reported by other authors for IHC. 25,35 The present results confirm earlier reports regarding lower levels of PR expression in simple epithelial-type carcinomas than in complex or mixed subtypes and the direct correlation between PR expression and lower grade malignancy. 25 However, results for dysplasias should be viewed with caution, and they require confirmation with a larger number of samples. Further, given the frequency of intratumoral heterogeneity, it would clearly be useful to combine RT-qPCR with other techniques such as laser capture microdissection, which enables detection of the different cell subpopulations comprising a tumor, thus providing fuller information. As in human studies, strong agreement was found between RT-qPCR and IHC for PR expression. 3,17,30 Eight cases that were PR-positive by RT-qPCR were classified as PR-negative by IHC. The IHC expression of PR in myoepithelial cell cytoplasm may have contributed to false-positive results in RT-qPCR, since the presence of cytoplasmic PR staining in myoepithelial cells is suggestive of PRB expression. 8,11 However, loss of tissue in the paraffin block during the procedure may equally be the cause of discrepancies in those cases where IHC detected PR antigen but RT-qPCR did not amplify the mRNA.
Both PR isoforms were found in all types of lesions analyzed. PRA and PRB were expressed at similar levels in canine dysplasias and benign mixed tumors, whereas PRA was more strongly expressed than PRB in carcinomas and complex adenomas. Similar findings have been reported for canine mammary tissue using Western blotting. 11 In normal human breast tissue, levels of PRA and PRB expression are generally similar; in some breast cancers, however, their ratio is dysregulated, with a predominance of PRA over PRB. 4,16,22 It is thought that the coordinated expression of both isoforms is required for the normal P response of the mammary gland and that dysregulation of this ratio appears early in tumorigenesis. 28 The dissimilar expression of PR isoforms may be useful indicator of tumor response to endocrine treatment, a high PRA/PRB ratio being associated with poorer outcomes in patients undergoing hormonal therapy. 16 However, a predominance of isoform A has also been reported in some benign breast lesions, including atypical ductal hyperplasias. 28 In humans and rodents, both PRA and PRB are expressed in the luminal epithelium. 2,28 In the rat, however, PRA is expressed only in that location, whereas PRB is detected in both luminal and myoepithelial cells. 19 Moreover, PRA has been reported in nuclei whereas PRB is found in the cytoplasm. 8,24 The cytoplasmic staining of myoepithelial cells observed here may be linked to PRB expression, while the stronger expression of PRA mRNA in complex tumors may be attributable to the incomplete or aberrant immunophenotype reported in neoplastic myoepithelial cells. 31 Differences may also exist in the distribution of PRA and PRB in human versus canine mammary lesions.
Finally, while expression of PR was significantly stronger in well-differentiated carcinomas (grade 1 and 2) with both RT-qPCR and IHC methods, 25,35 no differences in PR isoform distribution were found as a function of carcinoma grade. In human breast cancer, a predominance of isoform A has been associated with higher histological grades 4 in studies based on protein (IHC and Western blot) rather than mRNA expression of PR isoforms.
Analysis of progesterone receptors by the highly sensitive RT-qPCR method was successful in routinely formalin-fixed, paraffin-embedded tissues. This appears to be the first published report on the distribution of PRA and PRB mRNA in dysplasias, benign tumors, and malignant tumors of the canine mammary gland. The apparent predominance of PRA over PRB in CMTs may be critical for prognosis and therapeutic handling. These present findings may serve as the basis for future research into the role of PR isoforms in the progression of CMTs.
Footnotes
Acknowledgements
This was project no. AGL2011-25553; the research group is PAIDI Group BIO287.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
