Abstract
An important step in translational research is the validation of molecular findings from in vitro experiments using tissue specimens. However, tissue specimens are complex and contain a multitude of diverse cell populations that interfere with the molecular profiling data of a specific cell type. Laser capture microdissection (LCM) alleviates this issue by providing a valuable tool for the enrichment of a specific cell type within complex tissue samples. However, LCM and molecular analysis from tissue specimens can be complex and challenging due to numerous issues related with the tissue processing and its impact on the integrity of biomolecules in the specimen. The intricate nature of this application highlights the essential role a pathologist plays in translational research by contributing an expertise in histopathology, tissue handling, tissue analysis techniques, and clinical correlation of biological findings. The present review examines key practical aspects in tissue handling, specimen selection, quality control, and sample preparation for LCM and downstream molecular analyses that are a primary objective of the investigative pathologist.
Molecular analysis of tissue specimens and its correlation with pathological and clinical data is an important step in assessing the translational value of biological research. Current evidence suggests that the molecular profile of cell lines obtained from in vitro experiments may differ from cells in tissues due in part to the tissue microenvironment. 44 However, working with tissue specimens presents several challenges concerning the proper handling and preservation of the samples aside from the complexity of the tissues. 44,45 Indeed, tissues are an intricate tridimensional meshwork composed of multiple cell populations interacting on each other and the extracellular matrix. Each cell type has its own molecular signature that can be influenced by neighboring cells in the microenvironment and endocrine signals from distant organs. The analysis of biomolecules extracted from the whole tissue will include elements of the signatures from all cells, which can influence the molecular profiling data and contaminate the analysis. 15,28,35,47 There is significant evidence suggesting that for certain molecular studies, particularly quantitative studies such as gene expression analyses, the application of cell enrichment techniques like laser capture microdissection (LCM) can increase the specificity and sensitivity of cell-specific molecular profiling from tissue sections in biological and clinical-focused studies. 6,13,15,28,35,47 LCM requires the knowledge and understanding of pathology for proper tissue handling, selection, preparation, and analysis for molecular analysis, including the correct identification of the target cells under the LCM microscope. The pathologist has a pivotal role in translational research by contributing the knowledge of pathological variables and clinical correlation that increases the translational relevance of biological studies. In this context, LCM represents an important tool for the investigative pathologist working with tissue specimens for molecular profiling.
There are many excellent articles covering the technical aspects of LCM 21,23,24 and also downstream techniques from LCM samples, including RNA, proteomics, and microRNA analysis. 17,18,22,27,33,41,54 In the present article, our focus is to review the selection and use of tissue specimens for LCM and molecular analysis, which are part of the expertise expected from the pathologist in the research team. The consideration of these important factors can decide the feasibility of a project using LCM on tissue specimens. These considerations are presented in a step-by-step approach with the goal to help the pathologists starting a new LCM project (Table 1).
Recommended Quality Controls (QC) Steps for Laser Capture Microdissection (LCM).

Finally, despite that many examples of LCM include references to human tissues, we hope that this article may help investigative veterinary and human pathologists working with both animal and human specimens. For instance, veterinary pathologists can also work with human tissue specimens in research settings; therefore, we believe that the inclusion of references about the procedures and handling of human tissue specimens may be useful to the veterinary pathologist. Pathology in the genomic era requires the integration of morphology, molecular biology, natural history, etiology, and clinical context in both human and animal models of disease, as one pathology, one medicine. 5
Sampling, Preservation, and Storage of Tissue Specimens for Research
Before the selection of the right tissue samples for a particular molecular assay, the pathologist must consider several factors that can affect the success of the entire LCM study. Some of the main factors to consider in the tissue selection for LCM are the sampling, the technique used for tissue preservation, and the storage of the tissue specimens.
Tissue Sampling and Ischemia Time
Pathologists are well trained on gross sampling techniques for histological analysis. For molecular analysis of tissues, the time of ischemia is perhaps one the most important factors that can irreversibly affect the quality of the biomolecules. 2,3,10,11,38,51 The time of ischemia starts as soon as the blood perfusion to the tissue is interrupted due to clamping of the main artery. Warm ischemia is the time of ischemia while the organ or tissue is still in the body, whereas cold ischemia is the time after the extraction or removal of the tissue or organ from the body until its preservation by snap freezing or fixation. In animal specimens, the pathologist can control the ischemia time; however, in human specimens, the time of ischemia depends on the surgical or biopsy procedures. The effects of ischemia times on the molecular profiling of each cell type in tissues are still under research, but some evidence suggests that gene expression data may be affected by prolonged ischemia compromising the integrity of molecules such as messenger RNA (mRNA) in a time-dependent fashion based on the cell and tissue types. 3,19,38,51 From a practical standpoint, the shorter the times of ischemia, the better the preservation of the molecular profile in the tissue specimen. The ischemia times should be recorded for each specimen as part of the tissue tracking records. In the case of autopsy specimens from a human cadaver or necropsy from animal specimens, the time of ischemia should be recorded, including the time of death and the time of extraction of the tissue specimen. This information may be relevant for the interpretation of specific molecular data from the tissue specimens as certain tissues are pointedly affected by prolonged times of ischemia, especially in autopsy specimens. In the gastrointestinal tract, pancreatic and colonic tissues usually show histologic signs of autolysis that may be correlated with irreversible degradation of molecular data integrity depending on the type of downstream analysis. In summary, the effect of the ischemia time on the molecular data from tissue specimens is still under research, and the best recommendations are as follows: (1) the shorter the ischemia and autopsy/necropsy time, the better molecular and histological quality; (2) keep records of the times of ischemia for each specimen; (3) perform a histology quality control of the specimens, including tissue and cell integrity; and (4) perform a molecular quality control of the integrity of the biomolecules extracted from each tissue specimen selected for a particular study.
Tissue Preservation Techniques and Molecular Integrity
The preservation technique is one of the irreversible points during tissue processing, so if the tissue is poorly preserved, it can become irrevocably damaged for molecular analysis or even for histological analysis. Basically, there are 2 common ways to preserve a tissue specimen: chemical fixation or snap freezing. Both systems have unique advantages and disadvantages to exploit.
Chemically Fixed Tissues
Chemical fixation is a procedure in which the tissue specimen is treated with a chemical solution for preservation of the tissue morphology by avoiding tissue decay or decomposition by bacteria or fungi. 37 The fixative solution penetrates the tissue and chemically reacts in different ways with the biomolecules. Generally, after chemical fixation, the tissue specimen is embedded in paraffin for storage and cutting on a microtome. Historically, fixatives have been used to preserve morphology regardless of molecular integrity. 37 Recently, more attention has been focused on the molecular integrity of the tissues after fixation, leading to the development of new fixatives with the goal to preserve both morphology and molecular integrity. 1,25,39,42,45 This discussion will begin with the 2 main chemical fixatives for morphology and molecular analysis due to their almost overwhelming presence in pathology: formalin and ethanol.
Formalin (10% solution) is a 4% formaldehyde-buffered solution. Formalin-fixed, paraffin-embedded (FFPE) tissues are prevalent in diagnostic pathology because the method provides excellent morphological quality. 44,45 Unfortunately, formaldehyde induces damage to the integrity of biomolecules via chemical reactions during fixation, including the formation of covalent bonds or cross-linking between proteins. 45 It has been suggested that the chemical alterations induced by formalin are time dependent; therefore, prolonged formalin fixation will accentuate molecular alterations induced by the formalin. 29,45 One important aspect of the process is that formalin penetrates tissues at a rate of 1 mm per hour; therefore, placing large tissue samples or entire organs in formalin will result in poorly fixed tissue at the center of the sample, affecting the molecular and histological integrity of the specimen. For this reason, whole organs should be gross sectioned by the pathologist before placing them in formalin so the formalin can penetrate the tissue. The standard fixation procedure is an overnight incubation at room temperature in a 10% formalin-buffer solution in a flask with a diameter larger than the tissue specimen and ideally containing 10 times the volume of the tissue. 29 After fixation, the tissues are processed and embedded in paraffin blocks.
The advantages of formalin fixation include that it is a low-cost procedure, paraffin blocks can be stored at room temperature for a long time in a dry environment, 53 and it provides excellent quality for histology. 44 Despite the chemical alterations induced by formalin, biomolecules can still be recovered and analyzed using specific molecular extraction protocols and downstream analyses optimized for FFPE tissues. 8,9,41 For example, nucleic acids can still be extracted using proteinase K to digest the cross-linked proteins. A typical LCM sample can provide between 50 and 300 ng total of DNA; however, the DNA will be fragmented due to the formalin fixation. For polymerase chain reaction (PCR) techniques using DNA extracted from LCM samples, our recommendation is to use amplicons <200 bp. Some DNA arrays such as the Illumina Golden methylation array (Illumina, San Diego, CA) for DNA methylation analysis can be used with DNA from microdissected FFPE cells. In total, 500 ng or more of DNA is usually needed for these assays, which requires large LCM samples (or multiple LCM samples combined). 12,13 RNA is difficult to retrieve from formalin-fixed microdissected cells. Recently, technical improvements in mRNA extraction techniques and downstream molecular techniques allow for gene expression analysis from LCM FFPE samples (see below). Finally, it must be noted that formaldehyde is a human carcinogen, so its manipulation must be done with protective procedures and using a fume hood.
A second important chemical fixative is 70% ethanol. Ethanol is a coagulative fixative that does not induce the cross-linking of proteins; therefore, the integrity of biomolecules such as DNA is better than in FFPE tissues. 1,25,42 For instance, PCR products larger than 150 to 200 bp can be efficiently amplified from microdissected ethanol-fixed, paraffin-embedded (EFPE) tissues. 46 Another advantage of ethanol is that immunohistochemical detection (IHC) of certain markers can be easier to perform using EFPE tissues since antigen retrieval may not be needed. For example, IHC on EFPE tissues has been successfully used for immunoguided LCM for DNA methylation analysis. 46 Unfortunately, there is no clear evidence of an appreciable improvement in the recovery and analysis of RNA and proteins from EFPE tissues compared with FFPE tissues. Overall, the recovery of biomolecules from EFPE tissues is not comparable to frozen tissues, and the morphological quality is not as optimal as FFPE tissues. 42
Currently, novel formalin-free fixatives are being developed by combining diverse chemical solutions, including acetone, ethanol, and buffer solutions. 36,39 The goal of these new fixatives is to improve the quality of biomolecules compared with traditional formalin fixation while preserving the histological quality. These novel fixatives could become a better option than traditional FFPE tissue processing. However, snap freezing is still the optimal procedure for biomolecule preservation in tissue specimens. 42,45 To supplant formalin and gain acceptance for use with clinical tissues, these fixatives would have to meet 3 important criteria: (1) strong scientific and clinical evidence demonstrating that the efficiency of the pathology diagnosis is not affected by the new fixative on clinical samples compared with the FFPE standard, (2) the quality of the long-term storage must be comparable or better than FFPE blocks (for human samples, the tissue block storage time is 10 years as defined by the College of American Pathologists), and (3) the cost of the fixative for clinical use must be comparable to the FFPE standard. Nevertheless, these novel techniques should gain in popularity due to the advent of personalized medicine, which involves molecular testing for DNA mutations, gene expression, or posttranscriptional changes in proteins that require better tissue preservation methods. For instance, the employment of phosphorylated protein detection to generate biomedical markers from tissue biopsies has led to the development of a novel fixative for research and clinical medicine that better preserves the proteome and its posttranscriptional changes. 39 In the future, pathologists may need to evaluate and select novel fixatives that will provide good histological, immunohistochemical, and molecular quality for clinical pathology diagnoses and personalized medicine tests.
Snap-Frozen Tissues
Snap freezing is a procedure in which the tissue specimen is fast frozen and immediately stored at –80°C. The best way to snap freeze a tissue is by using isopentane and liquid nitrogen to freeze the tissue quickly and homogeneously to prevent ice crystals forming from the water molecules naturally present in tissue. 45 The tissue section should be smaller than the size of the cryomold with a thickness that does not exceed 5 mm. The specimen should be frozen as soon as it is retrieved from the necropsy or biopsy. Also, it is recommended to freeze the tissue in Tissue-Tek Optimal Cutting Temperature (OCT) embedding medium (Sakura Finetek, Torrance, CA), or a similar compound from other sources, to avoid tissue desiccation. 17,45 However, OCT must be removed from the tissue section before LCM and molecular extraction. 17,45 The main advantage for snap freezing tissues is the superior quality and yield of the biomolecules that can be extracted and employed in a variety of molecular assays, including high-throughput platforms for large-scale molecular profiling. 45 In general, the morphological quality is lower than FFPE tissues; however, when the freezing procedure is performed under optimal conditions, the histology quality can be very good, allowing for the identification of general histologic features, including the cells of interest. 45 The main disadvantages of snap freezing tissues include that freezing and storing tissues at –80°C is more expensive than FFPE tissue banking; access to isopentane and dry ice or liquid nitrogen is required, which are dangerous to manipulate; and an emergency electrical supply for storage freezers is necessary to protect frozen specimens in case of power outage. Occasionally, RNAlater (Life Technologies, Grand Island, NY) has been used for RNA preservation in frozen tissue samples, but there are several reports that the morphological quality required for LCM in tissues treated with RNAlater is compromised, and therefore it is not routinely used for tissue specimens employed for this purpose. The overall effect of RNAlater on histological quality still needs to be fully clarified and tested by the pathologist. 17,30
In summary, the 2 main procedures for tissue preservation are chemical fixation and snap freezing. The choice of the procedure depends on the goals of the study and the resources available. However, for molecular analysis, frozen tissues present a clear advantage because the integrity of all types of biomolecules is better preserved, particularly RNA and proteins. If large samples are available, it is possible to create mirror samples (ie, cut 2 sections from the specimen, formalin fix one for histopathology and conventional immunohistochemical analysis, and snap freeze the other for molecular analysis). When a choice must be made between tissue fixation and snap freezing, and the main goal for the specimens is molecular analysis, snap freezing is the method of choice. If the snap-freezing procedure is performed in an optimal way, 45 the biomolecules have a much better integrity than formalin, ethanol, or any current fixative, and the histopathology quality is sufficient to identify the cell types when performing LCM. 45
Tissue Storage
FFPE tissue blocks should be stored at room temperature, protected from direct sunlight, and placed ideally in a dry environment. 53 If these conditions are met, the blocks can be stored for decades since the proteins are locked due to the cross-linking induced by the formalin. While the tissue is embedded in paraffin, there is not much decay in the integrity of the molecules. A scrape test (see below) should be conducted to assess the quality of the sample before inclusion in an LCM study. Precut tissue sections already mounted on slides seem to be more affected due to exposure to the environment, particularly humidity and oxygen, which induce oxidation reactions within the molecules, leading to decay, a phenomenon that may affect certain epitopes for IHC and DNA extraction more so than others. 53 Therefore, the recommendation is to keep the tissue protected in the paraffin block and avoid cutting the sections until they are ready to be used in a study.
Snap-frozen tissues should be immediately stored at –80°C and transported temporarily on dry ice to avoid thawing the specimen. Freeze/thaw cycles are detrimental to the quality of the biomolecules, especially RNA, and cannot be reversed. 4 For long-term storage, the tissues should be protected from desiccation by embedding them in OCT or at least by wrapping the tissue in aluminum foil and storing it in a closed cryovial. The potential effect of time of storage on the integrity of biomolecules, especially RNA, is not yet well characterized, so it is highly recommended to perform a quality control analysis on all tissue samples to test the integrity of the biomolecule type of interest. 17
Sample Selection for LCM
Selecting the tissue specimens for use in a study involving LCM is an important step. This process will depend largely on a histological quality control that includes evaluation of the presence and abundance of the target cells and a molecular quality control test to assess the integrity and yield of the biomolecules present for compatibility in downstream analyses (Table 1).
Histological Quality Control of the Tissue Specimens for LCM
A pathologist should perform a histopathological evaluation and quality control of the tissues to be selected for LCM and molecular profiling. In studies involving animal tissues, the pathologist should ensure the tissue has been obtained following the established procedures and ethical regulations for animal work. 34 For human tissue specimens, the pathologist must check the status of the approval by an institutional review board (IRB) for each sample before it can be used in research. A basic pathological evaluation of the tissue block begins with evaluating a hematoxylin and eosin (HE)–stained section. If the identification of the target cell requires special techniques such as IHC, then an additional slide prepared with the specific technique should also be evaluated. The determination of inclusion in the study will be dependent on the presence of target cells, the amount of target cells, and any other conditions that may interfere with molecular profiling analysis such as necrosis, autolysis, inflammation, and so on. Making a table with an estimation of the relative percentages of target cells, inflammatory cells, fibrosis, necrosis, and so forth can be very helpful during the selection process and interpretation of the molecular data. During the histological evaluation, the pathologist may also pay special attention to certain conditions required for a particular molecular assay such as the minimum amount of target cells necessary or the purity of the enriched target cell population. In this regard, it is useful to include an evaluation of the distribution pattern of the target cells, noting if they are present in solid clusters that can be easily microdissected, or are isolated and sparse, making the LCM procedure more challenging and time-consuming. Finally, the standard HE coverslipped slide is useful for guidance during the microdissection session as the identification of cells may be more difficult in the sections prepared for LCM due to the lack of a coverslip and variations in the staining protocol for LCM (Figs. 1, 2).
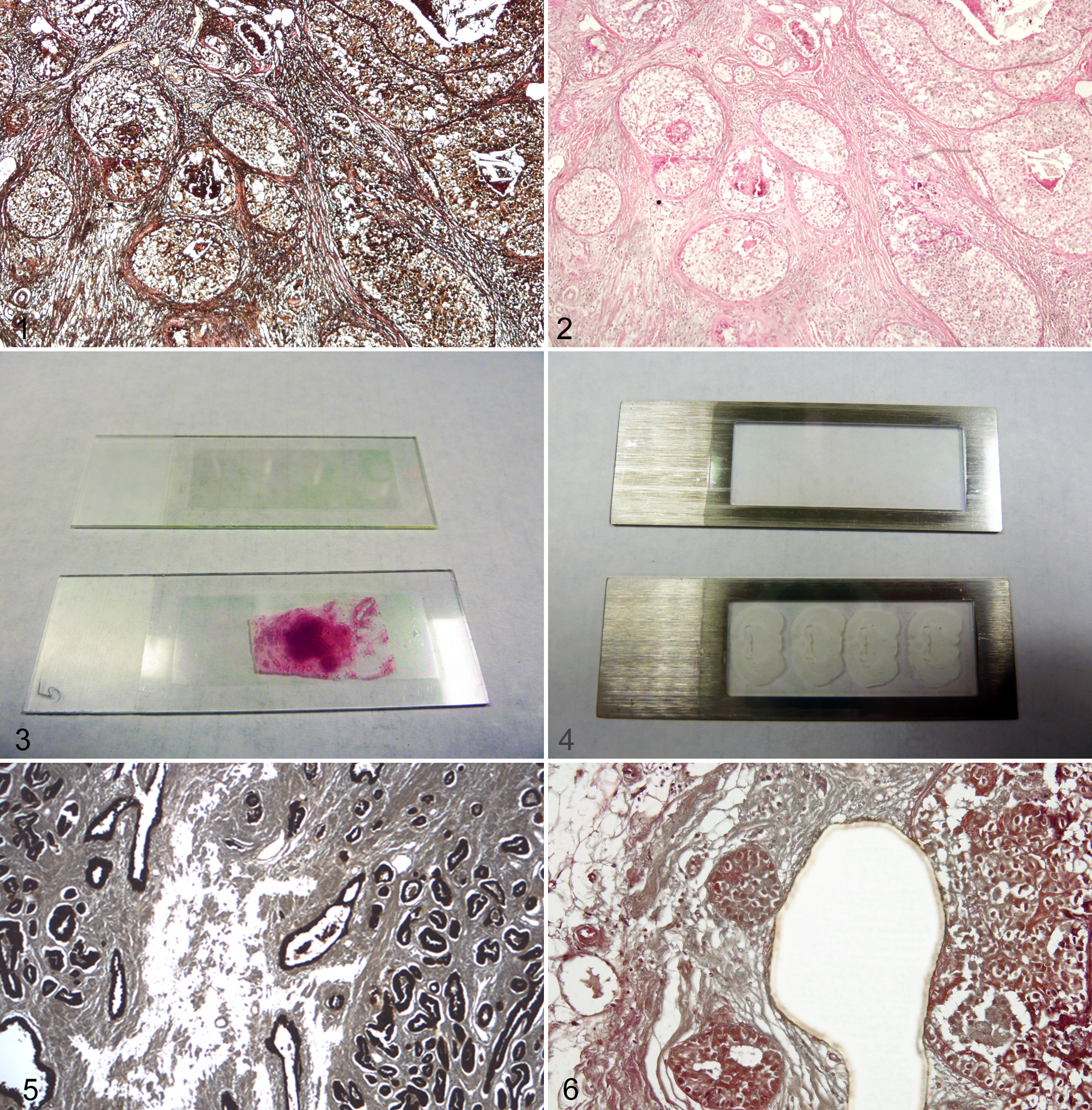
Human breast cancer prepared for laser capture microdissection (LCM). The image was taken from a membrane slide prepared for LCM without a coverslip, as it is actually visualized in the LCM microscope. Fine histological features are lost due to the lack of coverslip, making it difficult to recognize the cell types. Hematoxylin and eosin (HE).
Molecular Quality Control of the Tissue Specimens for LCM: Scrape Test
After the histological quality control and selection of the specimens containing the target cells for LCM, a second quality control should be performed to evaluate the yield and integrity of the target biomolecules from each selected block. For instance, there can be variations in the DNA integrity between different paraffin blocks due to variations in fixation or storage conditions. In frozen tissues, there can also be variations in the RNA integrity depending on the original freezing conditions as well as the storing and thawing events, which can affect the integrity of the RNA. If the biomolecule of interest is partially degraded in a particular block, then the results from LCM will be poor using that block. The lack of molecular quality control of the tissue blocks before LCM will create difficulties in the identification of the cause of technical issues from LCM samples. For this reason, investigators need to perform a molecular quality control on each block before LCM using the scrape test. The scrape test consists of the extraction of the molecules from a whole tissue section (Table 2). Importantly, it is recommended to use the same protocol or kit for molecular extraction that will be used on the LCM samples. The extracted molecules should be tested for the amount and integrity of molecules. For example, with RNA extracted from frozen sections, the amount can be tested using a spectrophotometer assay such as NanoDrop (Thermo Scientific, Wilmington, DE), and the integrity can be evaluated in a Bioanalyzer (Agilent Technologies, Santa Clara, CA) to assess the RNA integrity number (RIN). Several RNA assays such as microarrays will require a minimum quantity of RNA and a minimum RNA quality as assessed by the RIN number. The evaluation of the total amount of biomolecules recovered from the whole tissue section correlated with the relative percentage of target cells identified during the histological evaluation on the same tissue may also help to estimate the size of the LCM sample needed to obtain a certain amount of biomolecules from that particular tissue section. As a general rule, a typical LCM sample from one slide will provide 50 to 300 ng of total RNA or DNA depending on the number of microdissected cells (usually around 500 to 10,000 cells per sample). Also, if the RIN number is below the minimum required for a particular assay, then the LCM from that sample will not provide a better RIN, and the sample or the assay may need to be reevaluated. As a general idea, optimal RIN numbers for LCM samples from frozen tissues are between 7 and 8 (with a maximum optimal RIN of 10). 17,18 However, many LCM samples can show RIN numbers between 5 and 7, especially in tissues stored for a long time and used many times for previous projects. The optimal RIN number needed for a particular platform can vary, but in general, it is accepted that RIN >6–7 are acceptable for high-throughput platforms while RIN <5 can still be used for Taqman quantitative PCR (qPCR) assays (Life Technologies). There are several ways to test the amount and concentration of DNA. Purified DNA after extraction can be also tested in the NanoDrop and integrity can be tested using sizing PCR. 12 Finally, the molecules extracted from the scrape test can be also used to run a test on the actual assay platform to evaluate the feasibility of the assay using molecules extracted from tissues. Importantly, it must be considered that both the molecular extraction protocol or kit and the downstream analysis should be compatible with microdissected samples from frozen or FFPE tissues. For example, if the assay platform requires optimal biomolecular integrity and amounts in the order of micrograms, then the LCM project may not be feasible with that particular platform. Testing the molecules extracted from the scrape test on the assay platform can give an idea of the feasibility of the project using that particular assay.
Scrape Test General Protocol.

Sample Preparation for LCM
Once the tissue blocks have been selected after histological and molecular quality control (QC), the next step is to prepare the selected samples for microdissection. The main factors to be considered are the type of molecule to be extracted, the microdissection platform available and type of slide, and the visualization of the target cells in the sample.
Type of Molecule and Sample Type
It is possible that this is the main factor to be considered for the sample preparation for LCM.
RNA From Frozen Sections
If the molecule to be extracted is RNA, then the bench work needs to be done under RNA precaution conditions, in an RNAse-free environment. The article by Erickson et al provides the best protocol for the LCM of snap-frozen tissues for RNA extraction. 17 In summary, snap-frozen tissues must be kept at –80°C until sectioning in the cryostat and transported on dry ice without thawing the tissues. The slides used for LCM must be brand-new uncharged glass slides or membrane slides prepared for LCM (see below). After cryosectioning, the tissue section must be quickly mounted on the slide and immediately placed in a slide box on dry ice. The slides should then be transported on dry ice stored at –80°C and stored until being stained for LCM. Ideally, the sections should be microdissected within 2 weeks after cutting with the cryostat. 17 After staining, it is suggested that LCM and RNA extraction be performed in no more than 30 minutes (longer times can be used in certain tissues such as brain, while shorter times may be needed for gastrointestinal tissues). The optimal procedure is to stain the slides one by one for each LCM sample, while keeping the rest of the slides on dry ice until microdissection. 17
RNA From FFPE Tissues
As with frozen sections, it is recommended that RNA precautions are employed for RNA extracted from FFPE tissues even during the sectioning of the paraffin blocks with a microtome. These blocks can be stored at room temperature. The molecules are better protected while the tissue is in the paraffin block, so it is advisable to cut the slides a short time before LCM. Tissue sections mounted on LCM slides should be stored away from light in a dry environment without dewaxing the section. 29 As with frozen sections, FFPE slides should be stained just before LCM. We have observed differences in the integrity of nucleic acids from slides that were dewaxed and stained several days or weeks before LCM. It is important that the slides are kept dry after staining because humidity and water can further damage the already partially degraded macromolecules. 29,53 Recently, commercial kits have been introduced to the market for RNA extraction from FFPE microdissected tissues that include preamplification steps; however, working with RNA from FFPE microdissected samples is still a technically challenging procedure, and its feasibility for a particular project should be carefully evaluated. Overall, it is always better to use snap-frozen tissues for RNA extraction. 17
DNA
The sectioning and staining procedures for DNA extraction are similar to the ones described for RNA, but the main difference is that the slides from snap-frozen or FFPE tissues can be stored for several hours or even a few days before LCM due to the innate stability of DNA. The slides should be temporarily stored in a dry environment, protected from direct light. 53 It is also highly recommended to work with nuclease-free reagents and molecular-grade water. As mentioned before, frozen sections will provide better quality DNA than FFPE sections. For example, using frozen microdissected samples, it is possible to amplify amplicons >300 bp, while assays for amplicons obtained from FFPE should be designed for segments <150 to 200 bp.
Other Biomolecules
MicroRNAs can be extracted from both frozen microdissected tissues as described by Zhu et al 54 as well as from FFPE microdissected tissues as described by Nonn et al. 40 Proteins can be extracted from frozen microdissected tissues for proteomics analysis as described by Johann et al. 33 Silvestri et al 47 have also performed analysis of phosphoproteins extracted from frozen clinical tissues using LCM. Despite the effects of formalin fixation on proteins, attempts to extract and analyze the proteomic profile from FFPE microdissected samples have been made, particularly using mass spectrometry, as described by Patel et al. 41
Type of Slide and Microdissection System Available
There are 3 main types of LCM slides. LCM with a near-infrared (IR) laser, such as the Arcturus systems (Life Technologies), can microdissect samples mounted on regular glass slides, similar to those used for routine histology. In general, it is better to use uncharged glass slides for LCM. However, certain tissue types such as lung and brain may require the use of charged or “plus-coated” slides, which are positively charged to facilitate tissue adhesion to the slide. Charged slides are advantageous in histology and immunohistochemistry since the tissue will remain on the glass after staining or antigen retrieval, but it may represent an obstacle for LCM. Tissues mounted on glass slides are also the best for scrape tests and manual dissection using a needle. Use brand-new glass slides for each study. The slides can be precleaned with RNaseZap (Life Technologies) if the microdissection is for RNA extraction. The cleaned slide must be dried at room temperature before mounting the tissue section.
Glass membrane slides (Fig. 3) are required for UV laser systems such as PALM/Zeiss (Zeiss, Thornwood, NY), Leica LMD (Leica Microsystems, Buffalo Grove, IL), MMI (Molecular Machines & Industries, Haslett, MI), and the Arcturus XT (Life Technologies) equipped with a UV laser. These slides consist of a membrane polymer, such as polyethylene naphthalate (PEN), that is glued onto a regular glass slide. The tissue is mounted on the center of the PEN membrane slide. Then, during microdissection, the UV laser cuts the tissue and the PEN membrane around the cells of interest. It is important to pretreat membrane slides before mounting the tissue sections by irradiating the slides (without tissue) with a UV light (254 nm for 60 min) to make the PEN membrane of the slide hydrophilic so the tissue can better adhere to the membrane. 12 The UV pretreatment will also help to clean the slide and deter potential contamination from nucleic acids.
Frame metal slides can be used with any UV-LCM system but are required by the MMI system (Fig. 4). The membrane polymer is attached on its border to a metal frame, without any glass support. This type of slide is more expensive than regular glass slides or glass membrane slides. Also, the lack of a glass support makes it more difficult to mount tissue sections on the membrane. However, MMI offers a plastic device that can be used to create a support for the membrane.
Selecting the type of slide will largely depend on the type of LCM systems available to the investigator. For example, glass slides can be used for the scrape test, manual microdissection, and LCM only on instruments using an IR laser such as the Arcturus LCM microscopes (PixCell II, Autopix, and the IR-only version of the current Arcturus XT). LCM systems using a UV laser such as the Leica LMD, PALM/Zeiss, and the Arcturus XT with UV laser should use glass membrane slides. Frame metal slides are required by the MMI system and can also be used for other UV-LCM systems. Overall, microdissection using membrane slides with any of the LCM instruments is faster and easier and can be applied to both clusters of cells and isolated cells. In general, glass membrane slides will be a good first option for general LCM applications from tissues using any of the LCM systems. The exceptions are the MMI system, which requires frame metal slides, and the Arcturus systems that only have an IR laser, which only use plain glass slides.
Visualization of the Target Cells
Visualization of histological or cytological features in an LCM system can be challenging due to several factors, including the tissue preparation (frozen specimens usually have a less optimal morphological quality than fixed tissues), the complexity of the histopathology, issues arising from the staining, and so on. However, the main challenge in the visualization of the specimen is the lack of a coverslip. Routine histology slides have a glass coverslip, which improves the microscopic visualization, but LCM slides cannot be covered. In these cases, as mentioned before, a coverslipped, serial section stained with routine HE can be used as a “guide slide” to locate the region of interest (ROI) on the LCM slide. Another technique used to enhance the visualization of an LCM slide is to pipette a small drop (10 μL) of ethanol or xylene onto the slide and observe it through the LCM microscope until the drop evaporates (Figs. 1, 2); however, xylene gases are toxic and should be handled with extreme precautions in a fume hood and well-ventilated room. Another difficulty with the visualization of the target cells under an LCM microscope is the staining for LCM. The staining protocols for LCM are similar to the standard histochemical staining for pathology except that the duration of the staining steps is normally reduced to avoid biomolecule degradation; therefore, the staining is typically paler compared with the standard pathological stain. Importantly, molecular-grade, nuclease-free water and fresh reagents should be used for LCM staining. The most common staining protocols include HE, toluidine blue, Nissl staining, and commercial staining kits.
Hematoxylin and eosin is the most popular staining method for LCM of tissue specimens. Table 3 shows a typical protocol used for LCM. Compared with the traditional HE staining for pathology, the incubation times in this protocol are reduced to avoid degradation of macromolecules, particularly RNA, in frozen tissues. Mayer’s hematoxylin is an advantageous selection because for LCM, it can be used without the bluing step, which reduces the exposure to water and the risk of molecular degradation by endogenous RNases. 45 Toluidine blue is a 1-step staining procedure using a metachromatic reagent that can provide a similar staining quality to a fast HE for LCM. Because it uses only 1 staining reagent, it has a rapid staining time. In some hospitals, toluidine blue is still used for fast staining during intraoperative biopsies. Nissl staining is also a 1-step staining method based on cresyl violet. It is generally used for LCM on brain tissue, but it can be also used as a general stain.
General Hematoxylin and Eosin Protocol for Laser Capture Microdissection (LCM).

Some companies offer ready-to-use staining kits for LCM such as the LCM Staining Kit (Ambion; Life Technologies), which includes cresyl violet, the Histogene Staining Solution (Life Technologies), a blue stain with metachromatic features like toluidine blue, and the Paradise staining kit (Life Technologies) for FFPE tissues, which also contains a blue staining solution similar to toluidine blue. The main advantage of the commercial kits is that they are ready to use and include optimized protocols and molecular-grade reagents, which makes the staining procedure easier and safer for molecular integrity. However, the cost can be more expensive than preparing the stain in the laboratory. For general LCM, the modified HE staining as described by Erickson et al 17 and the protocol presented in Table 3 work well for most applications.
Other visualization systems include detection with IHC, immunofluorescence (IF), or endogenous proteins, such as green fluorescent protein (GFP). IHC and IF have been used for LCM on tissue specimens, mainly for DNA extraction and methylation analysis. 12,13,46,49,50 However, there is evidence that the amount and integrity of biomolecules can be affected by the IHC procedure, particularly RNA. 50 GFP has been successfully used for LCM on animal models with cells that have been labeled with the protein. 52
Molecular Quality Control of the Staining for LCM
Once the stain for visualization of the target cells is selected, a molecular quality control test should be performed to evaluate the effect of the staining on the yield and integrity of biomolecules. To compare how the staining procedure affects the recovery of the biomolecule to be analyzed, a scrape test (as described before) is recommended, comparing an unstained slide as a control with a slide stained using the method to be applied during LCM. After extracting the molecules from the unstained control and the stained test slide, the amount and integrity can be analyzed and compared. If the stained slide provides a poor yield or integrity, then the staining method should be reviewed or a commercially available staining kit should be considered.
Laser Capture Microdissection
Once the sample preparation, including the sample preservation medium, LCM slide type, and staining method, is determined, it is important to consider the type of molecule extracted before proceeding with the LCM:
For mRNA and other molecules that degrade quickly after extraction from frozen tissues, the optimal procedure is to thaw and stain the slide immediately before LCM and perform the procedure in about 30 to 45 minutes to obtain the highest possible RNA integrity number. 17
For a stable molecule such as DNA extracted from frozen or FFPE tissues, the slide can be stained a few days prior and then stored in a closed box in a dry, dark environment. These conditions prevent contact with water molecules, which can deteriorate the molecular integrity and affect the efficiency of the LCM. 23,53 A desiccant can also be stored with the slides.
For other molecules such as proteins or microRNAs, advancements in the extraction and analysis of LCM samples are constantly occurring, so the most practical recommendation is to perform a literature search for the most recent techniques. 41,54
LCM Systems Available
The precise steps for LCM will depend on the type of instrument available. Essentially, there are 2 main types of laser systems: near-IR capture laser and UV-cutting laser.
The near-IR capture laser is the original LCM system invented at the National Institutes of Health (NIH) in 1996 by Emmert-Buck et al. 16 During this technique, an IR laser (810 nm) activates a thermo-sensitive polymer (ethinyl-vinyl acetate, EVA), which has a dye that absorbs the IR laser energy, producing a change of the EVA conformation. 26 The activated EVA then attaches to the cells on the slide underneath, embedding and capturing them in the polymer—hence the name laser capture microdissection. Although the laser beam is directed at the target cells, the energy of the laser is not absorbed. The heat produced by the laser in the polymer is transient in the order of milliseconds, and hence the laser does not inflict any damage in the cells or biomolecules. 16,26 Either glass slides or membrane slides can be used with this IR laser (Fig. 5). Currently, the IR capture laser is offered only as a feature of the Arcturus XT LCM system from Life Technologies, which also includes a UV-cutting laser. Life Technologies offers the LCM caps, including the EVA polymer attached to an optically clear plastic cap, which fits in a 0.5-mL microcentrifuge tube.
The UV-cutting laser system uses a high-energy laser in the UV spectrum (355 nm) that is capable of cutting the tissues. Special membrane slides must be used during this application. The UV laser cuts around the target cells (Fig. 6), leaving them intact, compared with an IR capture laser that focuses directly on the cells. After cutting, the cells are retrieved into collection caps through a variety of methods. In the PALM/Zeiss system (Zeiss), the photonic pressure from a second laser shot catapults the cells into the collection cap. Gravity can also be exploited for depositing the cells into a collection cap as occurs in the Leica LMD system (Leica Microsystems). An additional collection technique was developed by MMI Instruments (MMI), in which the cells are glued onto a sticky cap after LCM. Detailed microdissection protocols for each of the commercially available LCM instruments are located in the instruction manuals provided by the manufacturer. Also, the Journal of Visualized Experiments provides videos with examples of various LCM protocols. 14,31,43,48,52
During the LCM process, it is important to document and keep records of the sample size provided by the LCM instrument and photographic documentation of the microdissected cells. All LCM systems incorporate photographic imaging and either a counter of IR laser shots or microdissected area to document the microdissected samples. These tools provide invaluable records that a pathologist reviewer may require to evaluate the microdissected cells and confirm that the molecular profile data actually originated from the target cells.
LCM Pilot Tests: Microdissection Test and Estimation of the Number of Cells Needed for a Particular Assay
Before initiating an LCM project for the first time, it is important to perform 2 pilot tests that can be done in the same session. During the first session using the LCM microscope, it is highly recommended to practice with the instrument using test slides. The goal is to focus on the use of the LCM instrument, visualizing the target cells and microdissecting them without concern for the molecular extraction. As soon as the operator learns to recognize and microdissect the target cells on a particular LCM instrument, then a new pilot test should be performed by microdissecting a test specimen for molecular extraction and quality control of the amount and integrity of the biomolecules. It is important that tissue sections used in this test have been evaluated and passed the histological and molecular quality controls previously described. A relatively large sample (10 000 cells, if possible) should be used initially and scaled down on successive samples (eg, 5000, 3000, 1000, and 500 cells) to determine the quantity of biomolecules that can be retrieved from the tissues and to ascertain the minimum number of cells or the LCM sample size required for a particular assay. During the pilot tests, duplicate samples should be assayed to ensure consistency. Once these tests are completed, the investigator will know how much area or how many cells will need to be microdissected from the actual project specimens. Typically, a subset of samples will need repeated attempts to acquire the amount of macromolecules necessary for downstream analysis.
General Considerations About Molecular Extraction and Downstream Analysis for LCM Samples
The procedures for the extraction of molecules from LCM specimens and the downstream assays should be carefully selected and tested prior to starting the project with study specimens. Currently, many commercially available extraction kits and molecular assay platforms are specially adapted to work with small amounts of molecules from frozen and FFPE tissues. As previously mentioned, depending on the specimen, a typical LCM sample may contain between 50 and 300 ng of total nucleic acids per sample. However, 300 ng of nucleic acids from an optimal snap-frozen tissue will not be the same as 300 ng of the same nucleic acid from FFPE tissues. This is very important to consider before choosing the downstream analysis method.
DNA Extraction and Quality Control
Most extraction methods for DNA from LCM samples are based on proteinase K digestion, but they differ in the purification step. For example, the Picopure DNA Kit (Life Technologies) does not purify DNA after proteinase K digestion to avoid any loss of DNA. This can present a problem when quantifying the DNA using a spectrophotometer-based assay such as the NanoDrop (Thermo-Scientific, Waltham, MA). Nevertheless, the Picopure and other similar proteinase K buffers have been successfully used in research projects. 46 The QIAMP DNA-micro kit (Qiagen, Gaithersburg, MD) includes a purification step to clean the DNA after proteinase K digestion. QIAMP spin columns are optimized for very small amounts of DNA, so the kit is compatible with LCM. The main advantage of purified DNA is the accuracy of the concentration obtained from a NanoDrop or other spectrophotometric or fluorescence-based methods.
A practical approach to assessing the DNA integrity is either to amplify it using a PCR-based test to evaluate differentially sized amplicons, such as the one employed by Eberle et al, 12 or to perform a technical test on the same assay platform that will be used on the real samples.
RNA Extraction and Quality Control
Many commercially available kits for RNA extraction are also optimized for LCM, among them the Picopure RNA kit (Life Technologies) and the RecoverAll kit (Ambion). As with DNA, a NanoDrop can provide a good estimation of the concentration (and thus amount) of the sample, while the RNA integrity number can be evaluated with a Bioanalyzer 2100 (Agilent Technologies, Inc, Santa Clara, CA). A detailed protocol for RNA extraction, evaluation, and reverse transcription (RT)–qPCR from microdissected samples is provided by Erickson et al. 17
RNA recovery from FFPE tissues is a technical challenge due to the effects of the FFPE on the integrity of biomolecules, particularly mRNA. Many of the kits are based on proteinase K digestion of the FFPE tissue and subsequent purification of the total RNA. There are several commercially available kits specifically designed to recover small amounts of RNA from FFPE tissues that are compatible with LCM, including those offered by Qiagen (RNeasy FFPE Kit,), Ambion (RecoverAll Kit for FFPE), and Life Technologies (Arcturus Paradise Plus FFPE RNA Isolation Kit). The operator should select a protocol or kit by performing technical tests using control FFPE cell pellets or tissue samples prior to starting the study on actual specimens to compare the different methods.
Downstream analysis techniques from mRNA extracted from LCM FFPE specimens should be limited mainly to RT-qPCR due to the loss of RNA integrity suffered as a consequence of the formalin fixation. 32 However, novel technologies for the extraction and analysis of a small number of partially fragmented molecules may lead to the introduction of new kits and assay platforms in the near future. Therefore, the investigator should always review the recent scientific literature to look for novel products that can be applied to challenging LCM projects.
MicroRNAs and Protein Extraction From LCM Samples
MicroRNAs may still be preserved after formalin fixation, and some of the RNA extraction kits for FFPE microdissected cells can be employed to purify microRNAs. For example, Nonn et al 40 have successfully extracted RNA from LCM samples of FFPE biopsies using the RecoverAll Kit (Ambion) and then used TaqMan RT-qPCR (Life Technologies) for the analysis of both mRNA and microRNA. There are also kits optimized for microRNA extraction from FFPE that can be used with LCM samples, like the miRNeasy FFPE Kit (Qiagen). Proteomic analysis from LCM FFPE samples is very limited due to the cross-linking of the proteins induced by formalin fixation; however, mass spectrometry has been successfully used for proteomic profiling. 41
Downstream Analysis From LCM Samples
As mentioned before, many variables will affect the success or failure of an LCM project using tissue specimens. The choice of the downstream analysis technique is one of these factors. LCM must be compatible with both the goals of the study and the type of tissue available. If the samples to be analyzed are FFPE tissues, the downstream analysis should be able to detect a small amount of partially degraded biomolecules. Even if snap-frozen tissues are available, the molecular technique needs to be compatible with biomolecules extracted from tissue specimens, which may not be in the same optimal conditions as those from cell lines.
The second consideration is the minimum amount of biomolecules needed for the analysis technique. The downstream molecular assay should have a minimum input within the ranges of an LCM sample. For example, a microarray platform that needs micrograms of high-quality RNA may not be the proper choice for LCM samples from FFPE tissues. However, these samples could be analyzed with a microarray platform that requires a few hundred nanograms and works with partially fragmented nucleic acids. Also, the small amount of molecules from an LCM sample can be amplified for use in certain downstream platforms. For example, the TaqMan PreAmp Master Mix Kit (Life Technologies) can be used for primer-specific amplification of RNA from microdissected FFPE tissues prior to qPCR for gene expression. Investigators must be careful to avoid introducing artifacts generated from very small and suboptimal molecules. Additional experiments to validate the LCM findings may be necessary, as described by Chuaqui et al. 7
As mentioned before, pilot tests are an essential tool to evaluate the compatibility of a desired molecular analysis platform with the molecules extracted from actual tissue specimens. The pilot tests should include serially diluted molecules extracted from control cell lines to test the lower detection limit of the assay. Also, molecules extracted from actual tissues, including whole sections and LCM samples, should be assayed to determine if the desired platform is compatible with the tissue specimens (frozen or FFPE) and the minimum amount of microdissected cells required for the assay to get reliable and reproducible results (Table 1).
Conclusions
Pathology is a scientific discipline that explores the alterations related with diseases in tissues and cells. Historically, the focus of pathology was first placed on gross alterations, which shifted to microscopic alterations of tissues and cells that later focused on phenotypical expression using special stains and immunohistochemistry. Currently, the study of pathology is in the middle of a revolution that involves the development and application of molecular biology to the study of the mechanisms of disease. In this context, the pathologist is placed in a privileged position to contribute to the multidisciplinary scientific team by applying the knowledge of tissue analysis techniques and clinical correlation to molecular biology studies, thus increasing the translational value of biological findings. LCM is one of the essential tools in the pathologist’s arsenal as a powerful tissue analysis technique that can correlate histopathology with fine molecular findings. However, LCM can represent technical challenges that the investigative pathologist must know and understand. In this regard, the use of pilot tests and a step-by-step approach can make the difference, increasing the probability of success in a challenging project. The use of molecular profiling techniques in personalized medicine will require further advancements in cell enrichment techniques to increase the specificity of the molecular profiling data, which will improve the results of personalized medicine. 20,47 Therefore, we foresee LCM techniques occupying an increasingly strategic role in translational research as a pathologist’s tool.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
