Abstract
The field of anatomic pathology has changed significantly over the last decades and, as a result of the technological developments in molecular pathology and genetics, has had increasing pressures put on it to become quantitative and to provide more information about protein expression on a cellular level in tissue sections. Multispectral imaging (MSI) has a long history as an advanced imaging modality and has been used for over a decade now in pathology to improve quantitative accuracy, enable the analysis of multicolor immunohistochemistry, and drastically reduce the impact of contrast-robbing tissue autofluorescence common in formalin-fixed, paraffin-embedded tissues. When combined with advanced software for the automated segmentation of different tissue morphologies (eg, tumor vs stroma) and cellular and subcellular segmentation, MSI can enable the per-cell quantitation of many markers simultaneously. This article covers the role that MSI has played in anatomic pathology in the analysis of formalin-fixed, paraffin-embedded tissue sections, discusses the technological aspects of why MSI has been adopted, and provides a review of the literature of the application of MSI in anatomic pathology.
Keywords
The field of anatomic pathology has a long history in modern medicine, dating back to Giovanni Morgagni (1682–1771), who is considered its father, and Rudolf Virchow (1821–1902), who is considered the father of microscopic pathology. From those early days, the use of microscopy in anatomic pathology has evolved slowly. The use of formaldehyde and hematoxylin and eosin was in place by the beginning of the 20th century. The first immunohistochemistry (IHC) study was reported in 1942 by Coons et al, 56 who used FITC-labeled antibodies to identify pneumococcal antigens in infected tissue. Since their development in the 1960s, fluorescence-based IHC methods (typically termed immunofluorescence [IF]) have generally been supplanted by chromogenic brightfield methods using hematoxylin counterstain and 3,3'-diaminobenzidine (DAB) as a chromogen in pathology, 283 although not in research where IF is still common. Although improvements were made in protein conjugation, tissue fixation methods, antigen retrieval, detection labels, and microscopy, the basic practice of the analysis of formalin-fixed, paraffin-embedded (FFPE) tissue in anatomic pathology remained relatively static. The advent of digital imaging and computer analysis methods started a new era in pathology, beginning in research and moving slowly but steadily into clinical practice.
The tissue samples studied in anatomic pathology are complex; multiparameter detection methods such as gene/protein expression arrays and flow cytometry make this apparent. It is increasingly important to measure not just the average expression of molecules in homogenized tissue but also their spatial distribution while preserving cellular and tissue architectural features. Such high-resolution molecular imaging is technically challenging, especially when signals of interest are colocalized. In IF, sensitivity and quantitative reliability can be compromised by the presence of autofluorescence, which is commonly present in formalin-fixed tissues. In brightfield microscopy, problems of overlapping chromogenic signals pose similar imaging difficulties. These challenges have been addressed using multispectral imaging* (MSI) technologies, which enable the simultaneous imaging and quantitation of multiple analytes, even in the presence of spatial and spectral overlap.
For samples that do not require 3-dimensional information or submicron-depth resolution (such as thin tissue sections), confocal microscopy is not necessary. In addition, many pathology applications utilize brightfield chromogenic methods, which are not generally handled by laser-based confocal systems. MSI systems that function in wide-field mode on the microscope and acquire sequential images of a sample can work in both brightfield and fluorescence and therefore have more utility in imaging of FFPE tissues.
Since the launch of the first Landsat satellite in 1972, MSI has played a key role in a wide range of satellite and airborne remote sensing systems. 106,232,261 The technology and concepts involved spilled over into a range number of multispectral hardware and analysis software technologies and have been applied in a diverse range of applications, from process control in food 11,111 or pharmaceutical tablet analysis 60,171,263 to the noninvasive imaging of works of art 13,91,109,155 and astrobiology. 207,246
MSI is best suited to those samples that have a high degree of spectral and spatial information, qualities that biological samples have in abundance. For this reason, MSI has proved useful in a range of biological imaging applications. In particular, fluorescence microscopy has a long history of incorporating MSI to be able to resolve multiple overlapping fluorophores. 164 All major microscopy companies (Zeiss, Leica, Olympus, Nikon) manufacture and sell multispectral confocal microscopy systems. In addition, there are a number of other imaging systems that add onto epifluorescence microscopes to create a multispectral platform. These systems and their applications have recently been reviewed. 100,356 Live cell imaging is one of the more common applications because of the need to resolve overlapping fluorophores and the desire to obtain 3- and 4-dimensional data about cellular functions. 368
Quantitation is becoming increasingly important in pathology in terms of area measurements and the measurement of intensity of chromogenic or fluorescence labels, 3,243,244,265,334 both of which are prone to error in a simple visual assessment. Morphologic segmentation software, which divides an image into regions of tissue types through an assessment of its morphology, can greatly assist the estimation of tissue areas. 93 MSI can help address intensity measurement issues, particularly where the separation of multiple colocalized chromogens in brightfield and tissue autofluorescence are key issues. Combining multicolor staining and MSI with software for tissue and cellular segmentation can enable the measurement of the per-cell amount of each of one or many markers from only those cells within a particular tissue compartment or morphology. 166 This kind of per-cell multiplexed measurement is not generally obtainable in any way other than through the use of MSI. 128
MSI is also the foundation of the cytogenetic technique known as spectral karyotyping, which is used to simultaneously visualize all the pairs of chromosomes in an organism in different colors. 114,259,275,276,353 The spectral karyotyping technology, also known as chromosome painting, has been widely applied since its inception, with over 500 publications, primarily in the field of cytogenetics; a full coverage is beyond the scope of this article, and recent reviews of this technology can be found elsewhere. 24,130 In addition, MSI has been used a great deal for decades for the imaging of cultured cells and the imaging of cell lines, live and fixed. While these methods are sometimes (though rarely) utilized in anatomic pathology, a review of them is beyond the scope of this article and can be found elsewhere. 120 There are a number of microscopy applications for MSI in microscopy outside of pathology that will not be covered here, including the ex vivo validation of in vivo imaging. 347,359–361 There has also been a recent review article on MSI in preclinical research and pathology. 163
This article reviews the uses of MSI in the imaging of sections of FFPE tissues in brightfield and fluorescence as broadly applied to the field of human and animal pathology research.
MSI of FFPE Tissues: Technological Aspects
There are a number of technological aspects to MSI as it applied to the analysis of FFPE tissue sections that need to be considered, and these issues differ depending on whether the sample has been chromogenically labeled and is being imaged in brightfield (white light) or it has been labeled with fluorophores and is being imaged in a fluorescence mode.
In brightfield, the most critical aspect is to account for the evenness of illumination of the lamp and the changes in light intensity at various wavelengths in the lamp output, as even the best light-emitting diode lamps require a per-wavelength intensity calibration. This is most easily done by performing a pixel-by-pixel and wavelength-by-wavelength optical density (OD) conversion, which is accomplished by acquiring an image from a blank part of the microscope slide and then doing a Beer-Lambert law transformation for each pixel. 191 This not only provides flat fielding to correct for uneven illumination and adjusts for the spectral variability of the lamp intensity but, probably more important, makes for a better correlation between measured chromogen intensity and antigen concentration because it provides data that are in the correct units for the Beer-Lambert law. 233 In addition, using OD-converted data means that spectral shape for chromogens that follow the Beer-Lambert law is independent of intensity, which is important for correct spectral library generation and good spectral unmixing.
In fluorescence, the most critical aspect in the imaging of FFPE tissues is the correct elimination of tissue autofluorescence. Autofluorescence is an intrinsic fluorescence generated directly from the tissue itself and is far brighter in formalin-fixed tissues than in frozen tissues (this is the major reason that clinical kidney IF is performed on frozen rather than FFPE tissues). One of the properties of tissue autofluorescence is that it is present regardless of the wavelength of excitation, although ultraviolet excitation and blue excitation cause more autofluorescence than red excitation. Because the autofluorescence emission is extremely broad regardless of excitation wavelength, it is virtually impossible to find one emission filter for a given excitation wavelength for which there is no autofluorescence present. There have been attempts to reduce autofluorescence using chemical methods, 227 but even the best attempts only manage a 50% to 75% reduction in autofluorescence. MSI approaches can result in a 99% reduction in autofluorescence and a concomitant reduction in limits of detection and increase in signal to noise 37,169,189,191,192,298
The most common data analysis method applied to microscopy-based MSI data is linear unmixing. 76 Linear unmixing, † also known as multispectral unmixing, is a least squares fit, or linear regression, of a number of given spectral shapes (basis functions; often termed a “spectral library”) into a spectrum acquired from the sample. 191 Obtaining correct and quantitative results from this process relies on having accurate examples of the spectral shapes of the fluorophores that will be found in the sample.
A critical aspect of MSI and an essential component of obtaining quantitative results is the generation of correct spectral signatures (a spectral library) for each chromogen or fluorophore, including any background or intrinsic signal, such as melanin or tissue autofluorescence. There have been a number of publications detailing the use of automated spectral decomposition methods to find the correct signatures, 37,167,189–191,349 and these can be useful, especially for exploring unknown samples. However, since it is easy to generate the correct “control” samples from which accurate spectral signatures can be generated, automated decomposition methods are not the preferred methodologies. What is preferable is to create a set of singly stained spectral control samples from which the signatures can be created. For example, for a dual-color chromogenic IHC staining with a hematoxylin counterstain, one can stain a section with DAB only (and no counterstain), a Fast Red section, and a section stained only with hematoxylin. From these, the correct spectral signatures can be obtained. In fluorescence, one also needs a section that does not contain any added fluorophore: a negative autofluorescence control. However, even a singly stained fluorescence section (eg, FITC only) will likely still contain significant autofluorescence in those areas that contain FITC. Even if that amount of autofluorescence is low (less than 10% of the total signal), using a mixed spectrum of 90% FITC and 10% autofluorescence will still affect the unmixing of unknown samples and make it difficult to detect any FITC signal that is less than 10% of the autofluorescence intensity. In these cases, it is necessary to use computational method to determine what the spectral signature of the fluorophore (eg, FITC) was in this particular sample. 37,167,189–191,193 This “compute pure spectrum” methodology is what enables the accurate quantitation of fluorescence signals in FFPE tissues.
Once correct spectral signatures are present, the individual images corresponding to the intensity contributions of each chromogen/fluorophore are extracted from the multispectral data set using linear unmixing. 54,76,165,191,301
Until the advent of MSI methods and their use in pathology, researchers were limited in what chromogen combinations were useful in multiplex IHC. There are a number of useful dual-color IHC staining combinations (red and blue, brown and blue) that show a generally easy-to-visualize colocalization color; that is, one can see the red stain and the blue stain, and there is a recognizable purple color present when the 2 chromogens are colocalized. However, these dual-color IHC combinations preclude the use of a hematoxylin counterstain, as that would interfere with the visualization of the blue chromogen. With the implementation of MSI methods, a range of colors and color combinations becomes possible, limited only by the ability to perform the multicolor IHC. 316–318,321 Most companies that provide chromogens for IHC have kits for dual and even triple IHC, and although a review of the literature for multicolor IHC is beyond the scope of this article, there are a number of good general and review articles available. 318,319,321
An example of multicolor IHC can be seen in Figure 1. This is a section of gut which has been stained for beta-catenin in red (Fast Red) and Ki-67 in brown (DAB) and then counterstained with hematoxylin (blue). The overexpression of beta-catenin in the nucleus in colorectal cancer has been associated with an APC gene mutation, 270 and Ki-67 is an important proliferation marker. It can be extremely difficult using visual analysis to determine which nuclei are expressing both beta-catenin and Ki-67, as double-positive nuclei will contain red, brown and blue together (see Figure 1a). Using a MSI and unmixing approach, it is simple to obtain individual unmixed images of each of the markers separately (see Figures 1b, 1c and 1d). In order to visualize the co-localization better, these unmixed images can be recolored and layered together as composite images, rendering either a simulated brightfield view (see Figure 2b) or a simulated fluorescence view (Figure 2c). Additionally, these unmixed images can be used as input data for more complicated analyses using software designed for the analysis of multispectral data in pathology. 162,166,193
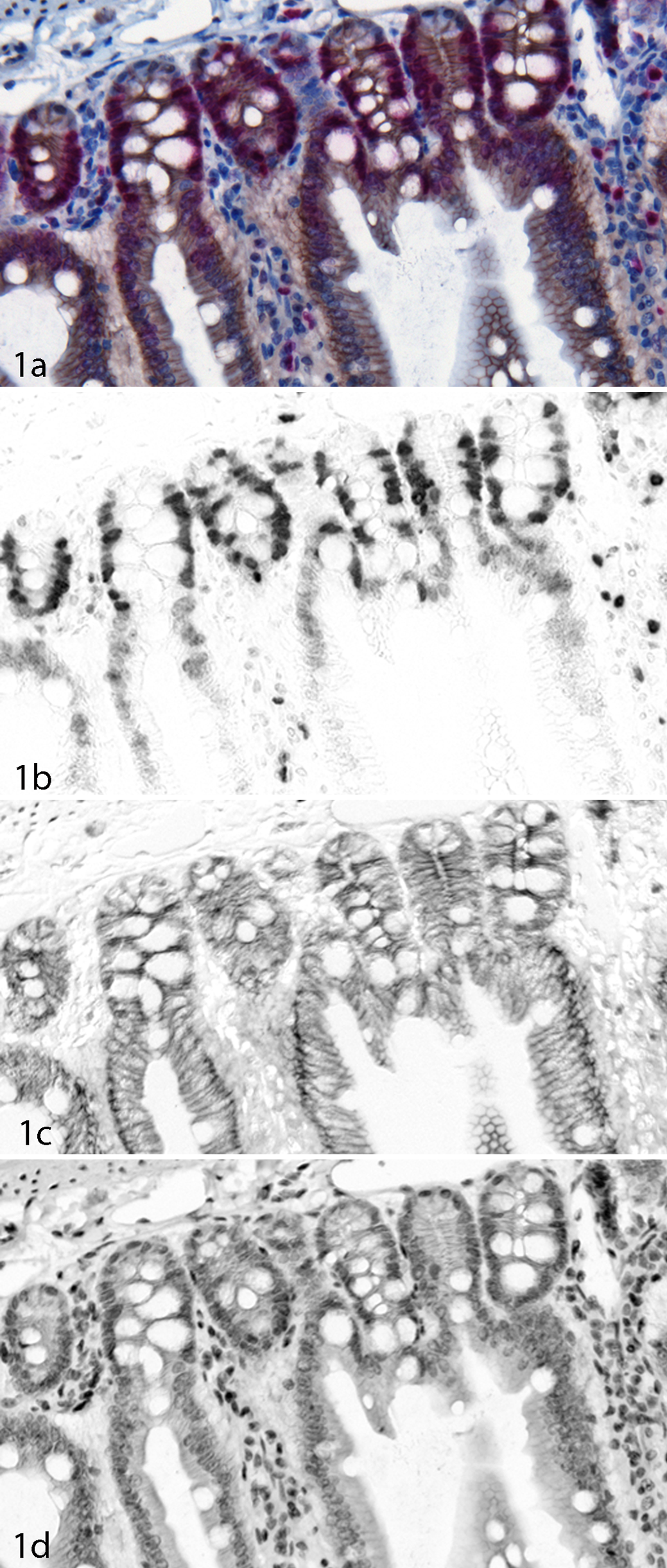
Beta-catenin (DAB) and Ki-67 (Fast Red) in a gut section. (a) Color (RGB) image of the sample. (b) Unmixed Fast Red image showing Ki-67 expression. (c) Unmixed DAB image showing beta-catenin expression. (d) Unmixed hematoxylin image.
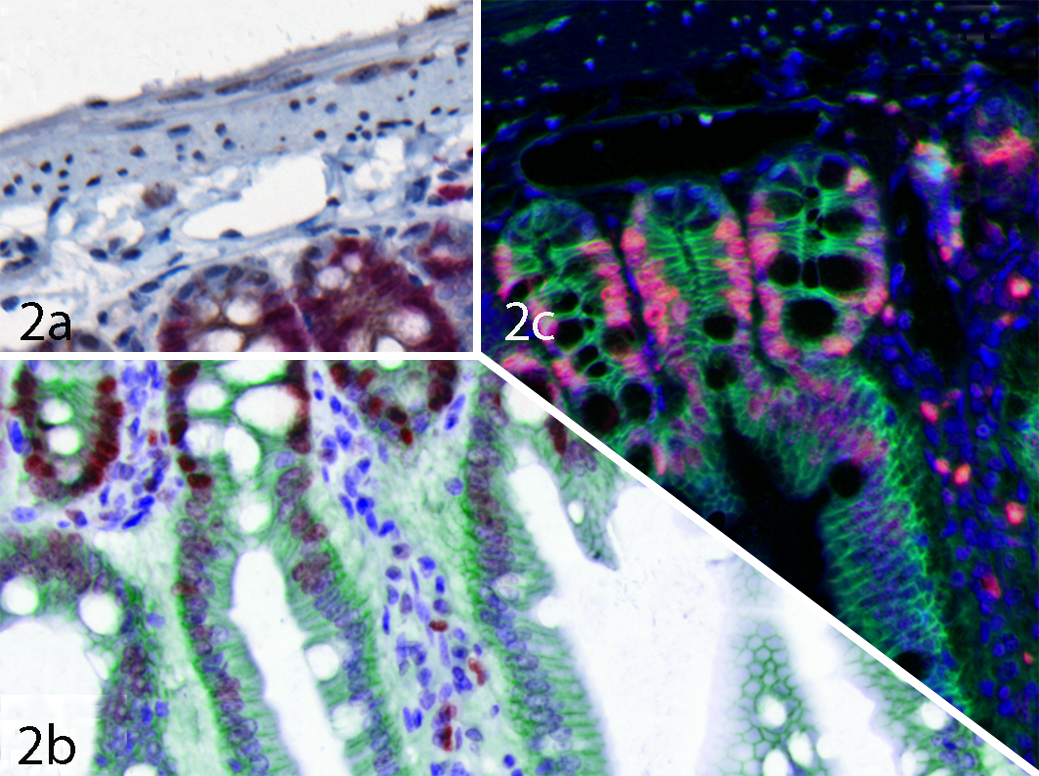
Composite images of beta-catenin/Ki-67 sample from Figure 1. (a) Color (RGB) image. (b) Simulated brightfield composite image with beta-catenin in green, Ki-67 in red, and hematoxylin in blue. (c) Simulated fluorescence composite image with beta-catenin in green, Ki-67 in red, and hematoxylin in blue.
There is considerable debate in the pathology community about how well the amount of color measured (the OD of the chromogen) or viewed in an image relates back to the amount of antigen found in that region on the microscope slide, with both sides having good points. One of the problematic points is that the majority of IHC that was performed for visual assessment is overstained for use in quantitative image analysis. Once a chromogen becomes too dark, it is unusable for the quantitative measurement of intensity. However, if the staining level is kept low (with a total OD of less than 0.6 OD units), it is quite possible to make quantitative intensity measurements. Comparing the intensity of the stain to the concentration of the antigen in the tissue is another matter, one often confounded with preanalytic variables, such as fixation time, warm time before fixation, the age of the sample, and so on. In general, however, there is a good correlation between antigen concentration and the measured intensity of a chromogen in a microscope slide.
IF has long been used in microscopy as a means of multiplexing markers, as the combination of red, green, and blue fluorophores provides for easy-to-visualize colocalization colors. However, for FFPE tissues, which are nearly ubiquitous in pathology, there is considerable tissue autofluorescence that interferes with the ability to see the fluorescence of specific markers. Figure 3 shows an example of C4d (a transplant damage marker) labeled with FITC in an FFPE kidney section. From the morphology and pattern of the green color in Figure 3a, one could assume that the brighter areas that appear to be vessels are specific C4d label. However, it is not possible from this image to determine whether there is a strong nonspecific FITC signal or whether the nonvessel signals are autofluorescence or FITC. Even using a narrow-band monochrome emission filter (which is generally what is recommended by microscope companies for autofluorescence problems) does not solve the problem. In the monochrome image in Figure 3b, the contrast is possibly improved slightly, but there is still no way to tell what is autofluorescence and what is FITC. MSI combined with proper spectral library generation can unmix or separate the FITC from the autofluorescence, 168,169,190 giving the FITC-only image shown in Figure 3c. This drastic improvement in contrast clearly shows which structures contained FITC and which did not. Often, the contrast improvement is so drastic that the autofluorescence “landmarks” are no longer present in the image, making it difficult to orient oneself. One solution to this is to present the unmixed images as a simulated composite image with the FITC shown in green and the autofluorescence shown in a highly contrasting gray color (see Figure 3d). The contrast improvement given by unmixing can give upward of a 100-fold increase in signal to noise and a concomitant 100-fold decrease in limits of detection. 298 The improved image quality and contrast is important, but overall it is the drastic improvement in the quantitation of the intensity of fluorescent markers in FFPE tissue that is important: in Figure 3b, over 90% of the measured signal is simply autofluorescence with only a small amount of FITC, while in Figure 3c, the signal is solely that of FITC. Not only does this improve quantitative results, but in cases where autofluorescence appears brighter than the real signal, MSI and the unmixing of autofluorescence can prevent obtaining erroneous or misleading results.
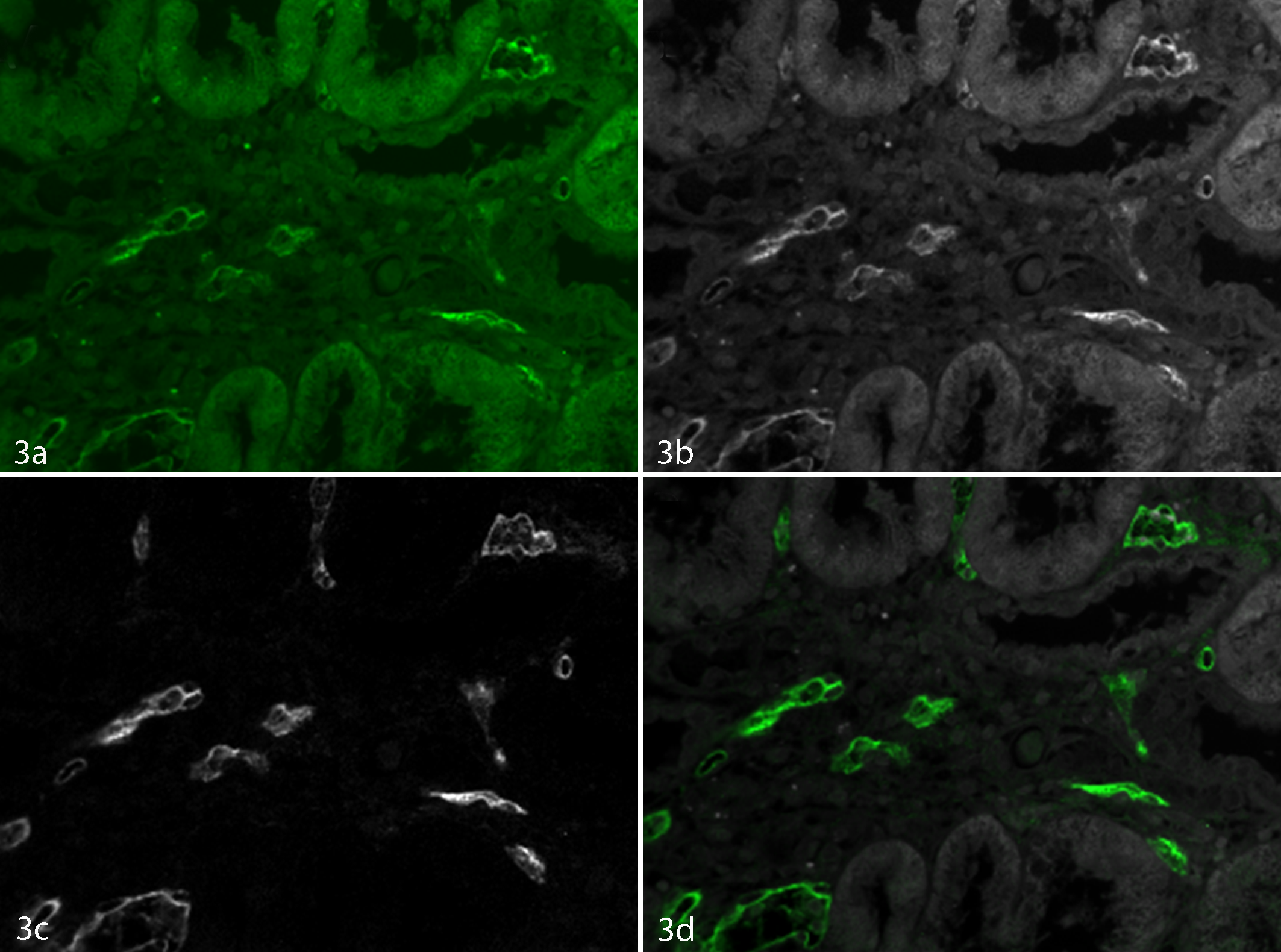
FFPE kidney section stained with FITC-labeled C4d. (a) Color (RGB) image of the sample. (b) Monochrome narrow-bandpass emission filter image at 520 nm. (c) Unmixed FITC image. (d) Composite image with C4d shown in green and autofluorescence shown in gray.
The improvement in signal to noise using MSI and unmixing can be easily extended from 1 marker to many. Figure 4 shows images from a breast cancer section that has been labeled with DAPI, Alexa Fluor ‡ 488 (estrogen receptor [ER]), Cy3 (Ki-67), Alexa Fluor 594 (progesterone receptor, PR), and Alexa Fluor 647 (HER2), combining in 1 section the ASCO/CAP breast panel. Figure 4a shows the color (RGB) representation. Figure 4c–4h shows the individual unmixed images for these markers, with each one separated from autofluorescence and one another. Figure 4b shows a simulated fluorescence composite image. In addition to the problems with autofluorescence, putting this many fluorophores in a sample typically results in a significant level of cross talk, or bleed-through, of one fluorophore into another’s channel. MSI, through its unmixing methodology, accounts for fluorophore cross talk, providing cleanly separated and quantitative channels/images for each marker/fluorophore.
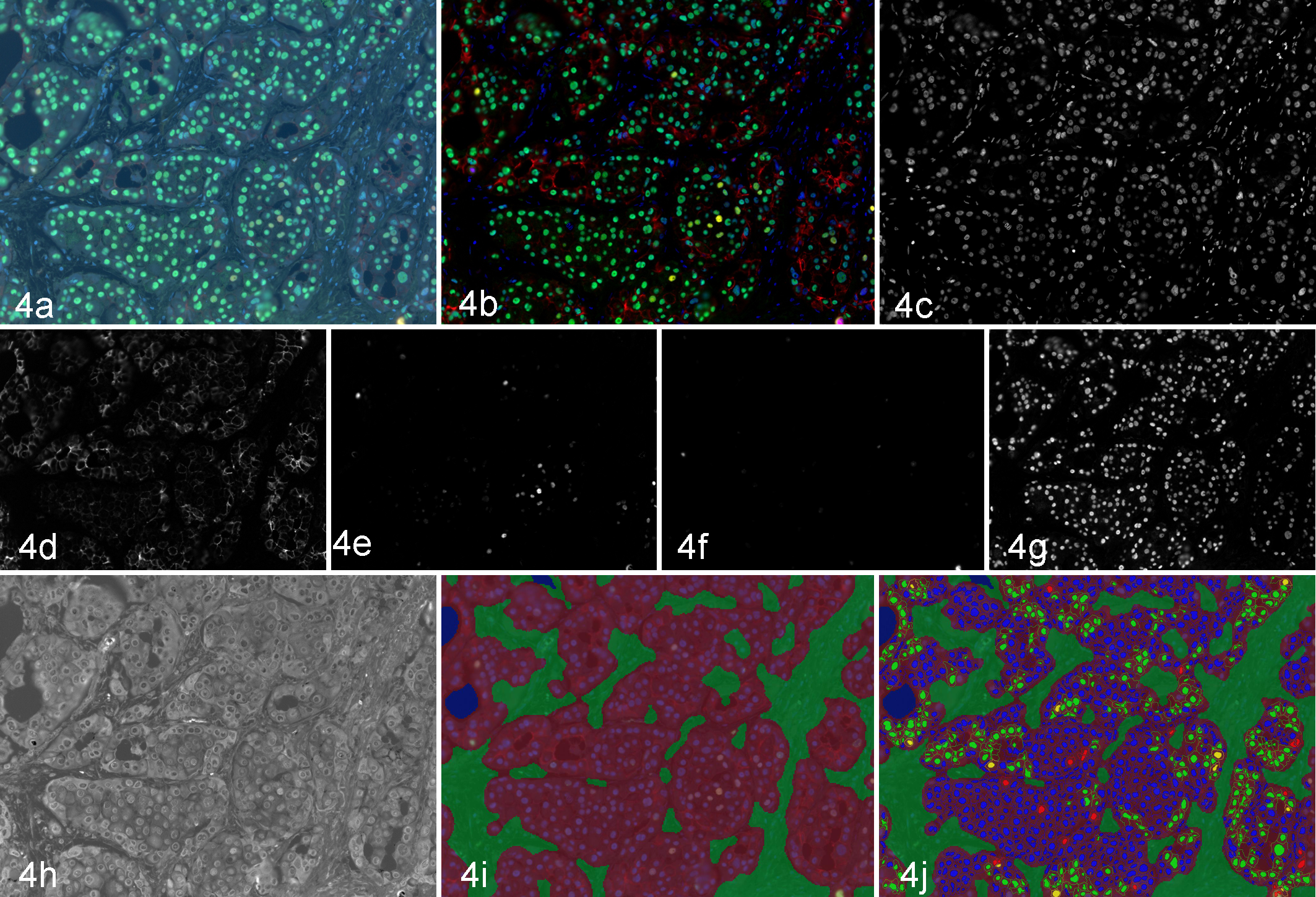
Breast cancer section labeled with ER (AF488), Ki-67 (Cy3), PR (AF594), HER2 (AF647), and DAPI. (a) Color (RGB) image. (b) Composite image. (c–g) Unmixed DAPI, HER2, PR, Ki-67, and ER images, respectively. (h) Unmixed autofluorescence image. (i) Tissue segmentation (red = tumor; green = stroma). (j) Double positivity scoring (yellow = HER2+/Ki-67+; blue = HER2–/Ki-67–; red = HER2–/Ki-67+; green = HER2+/Ki-67–).
Higher levels of multiplexing have been a goal of a number of development projects in pathology, and one means of trying to increase the level of multiplexing is to use quantum dots. Since their first development in 1998, 42 semiconductor nanocrystals, often known as quantum dots, have been used in a range of bioimaging applications. 98 Quantum dots offer a unique combination of brightness and high levels of multiplexing using direct conjugation when used with antibodies or a complementary DNA or RNA strand. Some of their first uses in pathology were for the imaging of the in situ hybridization of mRNA, 45,304 although this was quickly followed by numerous general IF applications. 47,97,99,296,306,348 The majority of the utility of quantum dots in pathology has been for their quantitative and multiplexing imaging aspects, and they have been used in a range of quantitative IF applications, § even down to the single molecule level. 370 Most recently, quantum dots combined with MSI have been used as a part of prognostic “signatures” of multiple proteins or genes in the tumor and tumor microenvironment. 52,237,238,305 Quantum dots can also be combined with organic fluorophores, as there are trade-offs in labeling methods. In the end, testing and validation are required for any multiplex staining methodology.
Putting the “More” Back in Morphology: Image Analysis Software and Multispectral Data
Although image analysis in pathology tends to be primarily that of a visual assessment of a sample or image by a pathologist or other domain expert, the visual assessment of multimarker IHC or IF samples, even when aided by MSI, is difficult, especially as the number of markers being used simultaneously in one sample grows. In addition, advances in molecular pathology and multicolor flow cytometry have increased the need for image analysis methods that provide quantitative, per-cell measurements from these multilabeled IHC and IF samples. To this end, there have been a number of image analysis programs developed—open source and commercial—the references for which are included in the literature review below.
There are 3 key aspects to performing quantitative, per-cell analysis of multimarker samples: first, the image data must be sufficiently separable to support independent quantitation of each marker; second, there should be an automated means of selecting which morphologic region (or regions) in the tissue section contains the cells of interest; third, there needs to be a means of performing cellular segmentation to be able to quantitate the intensity of one or more markers in a cell or subcellular compartment (eg, nucleus, membrane, cytoplasm). The sample shown in Figure 4 is an illustrative example of what can be achieved by combining MSI methods with advanced image analysis.
Figures 4i and 4j shows the sample as it goes through the stages of image analysis. Figure 4i shows the result of an automated tissue segmentation, achieved by having the user train the software to recognize tumor (shown in red) versus stroma (shown in green) versus blank slide (in blue; not shown in this image) using a method that has been previously described. 78,128,162,166,193 One critical aspect of this user training for use with clinical samples is that a sufficient number of training images be used (anywhere from 5 to 30) to ensure that the training set spans the range of morphologic variability that will be encountered in subsequent samples. Following tissue segmentation, a cellular segmentation is performed on those cells in the morphologic region of interest (in this case, the tumor region), often using DAPI, hematoxylin, or other counterstain as a means of determining where each cell is. Once each cell’s nuclear region has been determined, the cytoplasmic and membrane subcellular compartments can then be determined and the amount of each chromogen/fluorophore from each cell’s subcellular compartment extracted. After the per-cell (and per–subcellular compartment) amounts for each marker have been determined, the cells can be scored using a range of standard pathology scoring systems (percentage positivity, H-score, 0/1+/2+/3+, etc) and/or phenotyped (marker A positive, “A+”; marker B negative, “B–”; A+/B+, A–B+, etc) in a manner analogous to flow cytometry. Figure 4j shows double positivity scoring, or phenotyping, for the HER2 and Ki-67 markers with cells labeled with yellow indicating double-positive HER2+/Ki-67+ cells, green indicating HER2+/Ki-67– cells, red indicating HER2–/Ki-67+ cells, and blue indicating double-negative HER2–/Ki-67– cells. As can be seen in Figure 4j, only those cells in the tumor region (shaded red) are scored, which is analogous to the manner in which visual pathology scoring is performed. There are a total of 4 IF markers in this sample, so similar quantitation and scoring methods can be applied to any combination of the 4 markers, up to and including all the combinations of the 4-marker ER/PR/HER2/Ki-67 phenotypes.
To help show how MSI can be combined with microscopy automation, morphologic tissue segmentation, and cellular segmentation, 3 articles are reviewed in detail. These were chosen to be representative of the majority of the publications on MSI in pathology; 2 are cancer applications, one in brightfield and the other in fluorescence, and the third is in immunology.
The first article is entitled “Sex Steroid Receptor Expression and Localization in Benign Prostatic Hyperplasia (BPH) Varies With Tissue Compartment” and is a tissue microarray (TMA) study of the differential expression of androgen and estrogen receptor (AR and ER, respectively) status in stromal and epithelial cells.
213
TMAs are an important tool in clinical cancer research and are a convenient means of having tissue from potentially hundreds of subjects in a single tissue block.
14,49
In this study, a section of a prostate TMA of 104 duplicate cores from 52 patients was stained with multiplexed chromogenic stains for AR, ER, and smooth muscle alpha-actin and counterstained with hematoxylin. Multispectral brightfield images were acquired from each TMA core using an automated MSI system (Vectra, PerkinElmer, Waltham, MA), and then each was unmixed into their 4 individual component images. To automatically divide each image into epithelial and stromal regions, a pathologist trained morphologic software (inForm, PerkinElmer) to segment each image into epithelial, stromal, and blank regions. This training was done by drawing representative training regions on 18% of the images in the cohort and allowing the software to “learn” to recognize the morphologies of interest. This is an iterative process (draw regions, train, view resulting segmentation, redraw regions, etc) that is done until the pathologist is satisfied with the results on a blind test set (viz, images that were not included in the training set). The resulting image segmentation algorithm is then applied to each image. Once the images were segmented, a nuclear segmentation was performed using the unmixed hematoxylin image to find each nucleus. Each image’s nuclei were divided into those within the epithelial region and those within the stromal region. Within each nuclear region, the per-cell total and average amount of AR and ER was calculated by extracting the total OD of each chromogen from its unmixed image within that nucleus’s area. Following that, statistical analyses were performed on all the samples, comparing the stromal and epithelial expression of AR and ER between subjects with normal prostate and BPH. Among several other conclusions, this study found that
double positive (AR and ERα) epithelial cells were more prevalent in BPH, and fewer double negative (AR and ERα) stromal and epithelial negative cells were observed in BPH. These data underscore the importance of tissue layer localization and expression of steroid hormone receptors in the prostate. Understanding the tissue-specific hormone action of androgens and estrogens will lead to a better understanding of mechanisms of pathogenesis in the prostate and may lead to better treatment for BPH.
This conclusion can easily be generalized to a range of other disease states where it is important to understand the tissue-specific expression of one or more protein or genetic markers.
The second article is entitled “Multiplexed Quantum Dot Labeling of Activated c-Met Signaling in Castration-Resistant Human Prostate Cancer” and is a whole-section fluorescence study of signaling proteins that shows the utility of using automated morphologically based tissue sampling combined with highly multiplexed IF quantitation. 125 The use of multiplexed panels of markers for the diagnosis, prognosis, and therapeutic monitoring of cancer and other diseases is a rapidly growing field of research and one that likely will lead to specific multiplexed clinical tests in the future. This article shows how, through a carefully designed study with controls and single-stain validation, one can monitor the per-cell expression of several proteins involved in a signaling chain simultaneously and investigate—in this case, the expression level of 1 protein (c-Met) specifically in cells that are undergoing epithelial-to-mesenchymal transition (EMT). The samples in this study were stained for neuropilin-1, p-c-Met, VEGF, p-p65-NFkB, and RANKL and then counterstained with DAPI. An automated MSI system (Vectra, PerkinElmer) was used to acquire images from the samples, and it addresses one of the issues in fluorescence imaging—namely, that acquiring fluorescence images, particularly multispectral images, can be time-consuming, taking many minutes or hours for even the fastest slide scanners. In this study, the imaging system was “trained” by the users using morphologic analysis software (inForm, PerkinElmer) to recognize the tissue of interest at low magnification (4×)—in this case, prostate epithelial cells; then, a specified number of high-resolution (20×) multispectral images were automatically acquired only from regions of the sample that contained the tissue of interest. This automated “survey and drill” methodology can greatly speed up the acquisition of high-resolution images. The resulting MSI data were unmixed into their individual component images, including an autofluorescence channel, thereby preventing the autofluorescence from interfering in intensity quantitation. The researchers then trained the morphologic software to find epithelial cells, stroma and blank, in each 20× image; then, the epithelial cells were segmented using the DAPI unmixed image to find the cells. Each epithelial cell was further segmented into subcellular regions (nucleus, cytoplasm, and membrane), and the intensities of each fluorescent marker in each subcellular region for each cell were calculated. This per-cell protein expression data from multiple markers were used to determine that the “activation of c-Met signaling and subsequent induction of EMT is probably a common feature of prostate cancer cells, xenograft models and clinical prostate cancer specimens exhibiting an increased propensity for lethal progression.” This kind of multimarker study can be applied to nearly any combination of 2 to 6 markers in just about any kind of FFPE tissue and greatly expands the information that can be obtained from tissue sections.
The third article is entitled “Quantitative, Architectural Analysis of Immune Cell Subsets in Tumor-Draining Lymph Nodes from Breast Cancer Patients and Healthy Lymph Nodes,”
279
which highlights the interest in the imaging of immune cells and in the counting and determination of the distribution and density of specific phenotypes in lymphoid tissue, a methodology that is easily extended to tumors and other tissues. The analysis of the tumor microenvironment is important in a range of cancers and, when combined with an analysis of immune cell subsets, in the burgeoning new field of cancer immunotherapy. This article combines a brightfield MSI approach with automated image acquisition, morphologic tissue segmentation, and cellular segmentation leading to per-cell quantitative expression data from a number of immune system biomarkers and a study of the distribution, or density, of the immune cells across the tissue section. In this study, tumor-draining lymph node (TDLN) sections from 25 breast cancer patients and healthy lymph nodes from 7 healthy subjects were stained for CD20 (B cells), CD3 (T cells), and cytokeratin (epithelial/tumor cells) and counterstained with hematoxylin. As with the other in-depth reviews, multispectral images were acquired using an automated imaging system (Vectra, PerkinElmer), images unmixed into their individual component images; a morphologic tissue segmentation was performed; and then a cellular and subcellular segmentation was performed to obtain per-cell quantitative values for each marker in each cell in each tissue region (inForm, PerkinElmer). With these data, this study found that the spatial grouping patterns of T and B cells differed between TDLNs and healthy lymph nodes and that TDLNs and healthy lymph nodes have different B-cell localization patterns. While some assessment of these distributions could have been done the traditional way (with singly stained samples and a visual assessment), multiplexing these 3 markers together in the same tissue section allows the automated assessment of tumor invasion (via the cytokeratin) and T- and B-cell distributions simultaneously. In addition, adding further markers for colocalization (eg, FOXP3) or markers for proliferation or cellular signaling (eg, Ki-67 or HER2) would enable the distribution of cells with multimarker phenotypes to be investigated, and with the rapid growth of cancer immunotherapy, this is an important methodology. As the authors state,
this integrated image analysis approach can also be applied to study interactions between cancer and immune cells within tumors.…In addition to immunohistochemistry, this approach can also be adapted for immunofluorescence and in-situ hybridization techniques. More complex staining combinations using various immune cell markers, as well as their functional markers, are being developed.…Preliminary results showed that spatial relationships between immune cells in TDLNs may have an intriguing potential to be developed into novel prognostic tools beyond numerical changes and tumor invasion status.
Literature Review of MSI in Anatomic Pathology
Some of the earliest applications of MSI in pathology were in using the spectral signatures present in each pixel of hematoxylin-and-eosin images to classify or categorize that pixel as being of a particular tissue type—for instance, in the assessment of human melanomas and precursors, 92 the classification of urothelial cells in urine cytology, 135 or the detection of hyperchromicity in nuclei. 200 It has also been used to investigate metamerism 61 and eosinophilic esophagitis. 20 There are a number of early general articles describing the imaging and unmixing of multiple chromogens in multicolor IHC, 183,317,318 which were among the first to utilize MSI to separate colors difficult to differentiate visually. Because of its ability to remove autofluorescence and eliminate cross talk between fluorophores, MSI has been used for the multiplexed analysis of proteins 17 and the automated localization and quantification of proteins 303 in tissue sections.
There has been a comparative performance study characterizing breast TMAs using standard RGB and MSI, 247 a description of the use of MSI in pathology, 62 and that of the use of multispectral texture analysis for colon tissue biopsy classification. 197,198 There has been a review of imaging solutions for integrated quantitative IHC in routine pathology 265 and a description of the role and impact of quantitative discovery pathology. 244 MSI can also be used as an enhancement of regular staining, in a move toward “digital staining.” 21 Work has been done to combine MSI with advanced software as a part of learning histologic patterns 140 and for cell-based quantification of molecular biomarkers in histopathology specimens. 3,4 There are some general articles on combining the spectral–spatial analysis and MSI for the analysis of multiple biomarkers 9,162,166 and some discussions of how to move beyond morphology in whole slide imaging, computer-aided detection, and other techniques. 3,93
The primary use of MSI in pathology has been in cancer and immunology; however, MSI has been used as a technique in pathology topics, from arthritis and atherosclerosis to liver disease and biomaterials applications. This section attempts to reference a majority of the publications that have utilized MSI in some way in a particular application.
Joints, liver, and respiratory studies
In arthritis, MSI has been used for the measurement of the degeneration of articular cartilage, 147,148 vascular engineering, 63 quantitative imaging of Herovici’s polychrome for the assessment of collagen content and tissue remodeling, 314 and scaffolding of skeletal muscle extracellular matrix. 343 In liver disease, MSI has been used to investigate the effects of ximelagatran in human hepatocyte cultures, 2 as a means of evaluating multiplexed quantum dot imaging in liver and allograft biopsies, 131 to investigate the effects of estrogen on female biliary epithelial cell IL-6 expression, 133 to investigate liver ischemia/reperfusion-induced coagulation in rats, 151 and to investigate the cross-beta-sheet conformation of keratin 8 in Mallory-Denk bodies compared with other hepatocyte inclusions. 186 Combined with automated tissue and cell segmentation software, MSI has been applied to accurately quantitate Ki-67+ proliferating hepatocytes in rabbit liver. 320 In pulmonary research, MSI has been used to determine that acute respiratory distress syndrome leads to reduced ratio of ACE/ACE2 activities and is prevented by angiotensin-(1-7) or an angiotensin II receptor antagonist 345 and in the monitoring and assessment of ventilator-induced injury in lung. 344,346
Neuroscience studies
Fluorescence imaging in brain tissue in particular is often done using confocal microscopy to produce 3-dimensional images of complex neural structures and to aid in determining 3-dimensional colocalization. 369 However, for highly autofluorescent samples (typically, FFPE tissue), it can be advantageous to use MSI on thin sections that do not require 3-dimensional analysis. In addition, there are many multicolor IHC problems best addressed by MSI. It has been used in a number of Alzheimer disease applications, including in the design, synthesis, and evaluation of new fluorescent probes for the detection of tau fibrils in Alzheimer brain, 112 to investigate cell injury and premature neurodegeneration in cortical development malformations, 134 to show that reactive glia show increased proteasome activity, 230 that Rab6 is increased in Alzheimer brain, 273 to investigate how CCL affects neurocognitive dysfunction, 149 and how concerted changes in transcripts in the prefrontal cortex precede neuropathology in Alzheimer disease. 36
MSI has also been used in neuroscience to investigate the differential expression of histocompatibility complex class I in developmental glioneuronal lesions, 245 to test glycogen synthase kinase-3 activation through lysosomal degradation, 215 to determine the melanocortin 4 receptor distribution in the hypothalamus, 285 to determine that hyperforin attenuates brain damage in a transient middle cerebral artery occlusion, 176 and to investigate the critical role of the MyD88-dependent pathway in non-CNS MPTP-mediated toxicity. 59 In Parkinson disease research, MSI has been used to show ultrasensitive in situ visualization of active glucocerebrosidase molecules 342 and to determine the activation of unfolded protein response in the endoplasmic reticulum 122 and changes in selenoprotein P and glutathione peroxidase 422,23 in disease progression.
In other neuroscience applications, MSI has been used to investigate the role of an intrathecal cannabinoid agonist in spinal anti-inflammatory cytokine responses in neuropathic rats exhibiting relief from allodynia 339 and the relationship between mitochondria and α-synuclein in substantia nigra neurons. 250 It has also been used to investigate central nervous system dysfunction in a murine model of FA2H deficiency 242 and to show the reversal of markers of unhealthy brain aging by exercise in mice 159 and the role of chronic oxidative stress as a mechanism for radiation nephropathy. 160 Proteomics and MSI have been combined to implicate extracellular matrix proteins and protease systems in neuronal development induced by astrocyte cholinergic stimulation 203 and to show that acupuncture at ST36 prevents chronic stress-induced increases in neuropeptide Y in the rat. 86 MSI has been used to show that there are morphologic and functional abnormalities in mitochondria associated with synaptic degeneration in prion disease, 286 and it has been used to study neuronal CXCR4 expression and function and its disruption by opioids. 241,284
Transplantation studies
MSI has been applied to the prevention of diabetic microangiopathy by prophylactic transplant of mobilized peripheral blood mononuclear cells 366 and for determining that enhanced mobilization of bone marrow cells does not ameliorate renal fibrosis. 292 It has been used in a study of the molecular regulation of hepatic dendritic cell function and its relation to liver transplant outcome 295 and the accumulation of intrahepatic islet amyloid in a nonhuman primate transplant model. 179 MSI was helpful in an histologic graft assessment after clinical islet transplantation 310 and in a study of persistent hyperinsulinemic hypoglycemia of infancy to determine that constitutive activation of the mTOR pathway is associated with exocrine-islet transdifferentiation and their therapeutic implications. 5 MSI was used to learn that hematopoietic stem cell migration to the ischemic damaged kidney is not altered by manipulating the SDF-1/CXCR4-axis 294 and that intragraft tubular vimentin and CD44 expression correlate with long-term renal allograft function and interstitial fibrosis and tubular atrophy. 142 Finally, MSI has been compared to other whole slide imaging methods in transplantation biology in looking at combining whole slide imaging, multiplex staining, and automated image analysis. 132
Cardiovascular disease
MSI has been used to monitor the dynamics of human myocardial progenitor cell populations in the neonatal period, 8 to monitor the arterial nanoparticle delivery of siRNA for NOX2 knockdown for the prevention of restenosis in a rat model of atherosclerosis, 173 to monitor smooth muscle homeostasis in human atherosclerotic plaques through IL-15 signaling, 322 in the investigation of the role of prolactin in macrophages in human carotid atherosclerotic plaques, 256 to measure the accumulation of myocardial lipids and accompanying left ventricular function in an acyl-CoA-dehydrogenase knockout mouse model, 15 and to investigate the expression patterns of angiogenic and cell cycle–dependent factors in the proliferation and maturation of microvessels in arteriovenous malformations. 201 MSI has been used to monitor the number of Foxp3-positive regulatory T cells in human atherosclerotic lesions. 69
In situ hybridization and MSI
Fluorescence in situ hybridization (FISH) and its brightfield counterpart, chromogenic in situ hybridization, are cytogenic techniques developed to detect and localize the presence of specific DNA or RNA sequences. These in situ hybridization (ISH) methods are often used for finding specific sequences in DNA or for measuring specific RNA targets. In FFPE tissues and in some cellular applications, autofluorescence can be a limiting factor in visualizing FISH signals, and in looking at multiple FISH signals, fluorophore cross talk and bleed-through can be an issue. MSI can help in separating fluorophores from one another and from autofluorescence 136 and has been used in leukemia 28,66,139 and lymphoma 358 research, as a part of HER2 scoring in a clinical study, 307 in squamous cell carcinoma, 182 for measuring HER2 and CCND1 in ER-positive breast cancer, 267 and in HER2 amplification in urothelial bladder cancer in a clinical study of 1005 patients. 156 Multispectral FISH has also been used in pancreatic tissue, 102,331 to analyze a break-apart ALK probe for various phases of cell division in myofibroblastic tumors, 235 and to assess MED12 mutations in smooth muscle tumors and uterine fibroid and their relationship to cytogenetic subgroups. 194,195 MSI has been used to image chromosome 21 aneuploidies in breast fibroadenomas; 289 in NBS1 mutations, immunodeficiency, lymphoid malignancy, and early T-cell development; 269 in investigations of fusion genes (EWSR1-NFATC1) in hemangioma of the bone; 10 and in PPAR gamma-specific probes for the detection of PAX8/PPAR gamma translocation in follicular thyroid neoplasms 282 or thyroid carcinoma samples. 150
FISH and chromogenic in situ hybridization can also be used in combination with IHC or IF to simultaneously measure DNA or RNA and protein expression, and MSI is very helpful in the multiplexed imaging. It has been used to look at interrelations between gene aberrations and protein expression in gastric adenocarcinoma; for MYC; 58 for TP53 gene deletion, protein expression, and chromosome 17 aneusomy; 144 and for chromosome aneuploidy, protein expression, and CDKN2A gene. 113 It has been used to look at RAB23 gene amplification and protein expression, 124 whether there is a correlation between the loss of MSRT/NCoR2 corepressor and JAG2 overexpression in multiple myeloma 104 and that there is a role of ROC1 in the control of cyclin D1 expression in skin melanomas. 208
One particular subset of ISH analysis for which MSI is important is the imaging and quantitation of messenger RNA (mRNA) and microRNA (miRNA, or, for a single target, miR). miRNA are small noncoding RNA molecules (circa 22 nucleotides long) that function in the transcriptional and posttranscriptional regulation of gene expression. They are thought to be involved in most biological functions, and the aberrant expression of them has been implicated in a number of disease states. An important aspect of the imaging of miRNA expression is that it generally needs to be imaged simultaneously along with IHC or IF for the putative target protein that the miR is thought to encode. 217,218,221 In this manner, one can determine the relationship between the miR and target protein expressions on a cell-by-cell basis. 188 MSI has been used to measure miRNA expression in COPD, 88 inflammation in temporal lobe epilepsy, 12 LKB1/AMPK signaling in metabolic stress in glioma cells, 108 and human papillomavirus DNA in productive infection 222 and in investigating the binding of miRNAs to Toll-like receptors in the induction of a prometastatic inflammatory response. 89 Given its role in controlling gene expression, it is not surprising that a large amount of miRNA imaging and quantitation has been in cancer. MSI of miRNA expression has been used in monitoring that hepatitis C virus proteins modulate miRNA expression in malignant hepatocytes 38 and in an investigation of the mismatch repair and genomic stability of miR-155 in Lynch syndrome and colorectal cancers. 315 It was also used to determine that miR-221 silencing blocks hepatocellular carcinoma and promotes survival 234 and to investigate the role of stromal PTEN-regulated miR-320 in the reprogramming of the tumor microenvironment 41 and the oncosuppressive role of P53-induced miR-205 in triple-negative breast cancer. 240 It has also been used in determining that the reovirus-associated reduction of microRNA-let-7d is related to the increased apoptotic death of cancer cells in clinical samples. 219 MSI has been combined with a particular ISH product (RNAScope || ) for multiplexed imaging of mRNA in brightfield and fluorescence 329 and used to investigate multiple markers in circulating tumor cells. 236 MSI has been used in studies of mRNA in a semiautomated multiplexed quantum dot–based in situ hybridization and spectral deconvolution, 45 in the imaging of multiple mRNA targets using quantum dots in clinical biopsies, 304 and as a part of a quantitative multiplex quantum dot in situ hybridization–based gene expression profiling in TMAs identifying prognostic genes in acute myeloid leukemia. 305
Small interfering RNA (siRNA) is a class of double-stranded RNA molecules, 20 to 25 base pairs in length. siRNA plays many roles, most notably in the RNA interference pathway, where it can interfere with the expression of specific genes with complementary nucleotide sequence, the investigation of which is a relatively new field. Nonetheless MSI has been used to monitor an HIV-1 poly A and transactivator of the HIV-1 LTR-(TAR-)-specific siRNA 184 for therapy and with local arterial nanoparticle delivery of siRNA for NOX2 knockdown to prevent restenosis in an atherosclerotic rat model. 173
Oncology and cancer research
Oncology and cancer research is the largest application category in which MSI has been found useful, because of its usefulness in imaging multiple markers in brightfield and fluorescence and because of its ability to remove autofluorescence and improve quantitation. In general studies, it has been used to investigate a novel multipurpose monoclonal antibody for evaluating human c-Met expression in preclinical and clinical settings, 152 to determine that there is no haploinsufficiency but loss of heterozygosity for EXT in multiple osteochondromas, 252 and that there is enhanced tumor growth after portal vein embolization in a rabbit tumor model. 121
MSI has been used in studies of a range of carcinomas. It has been used to determine that Sp1 and c-Myc regulate transcription of BMI1 in nasopharyngeal carcinoma, 330 to analyze circulating and intertumoral macrophages in hepatocellular carcinoma, 212 and in an analysis of HSPB1 expression in head and neck squamous cell carcinoma. 216 It has been used to look at new variants of epithelial–myoepithelial carcinoma (oncocytic-sebaceous and apocrine), 277 for quantitation to show that reduction in Pten tumor suppressor activity promotes ErbB-2-induced mouse prostate adenocarcinoma formation through the activation of signaling cascades downstream of PDK1, 262 and to show that the regulation and function of the forkhead transcription factor forkhead box O1 is dependent on the progesterone receptor in endometrial carcinoma. 332 A MSI immunohistochemical method identifies lymphovascular invasion in a majority of oral squamous cell carcinomas and discriminates between blood and lymphatic vessel invasion. 223 MSI has been used to show that the inhibition of MAPK kinase signaling pathways suppressed renal cell carcinoma growth and angiogenesis in vivo 126 and that regional differences in intratumoral lymphangiogenesis of oral squamous cell carcinomas can be evaluated by IHC and MSI using D2-40 and podoplanin antibody in comparison with angiogenesis. 225 MSI has been used to show that the distribution of dendritic cell subtypes in primary oral squamous cell carcinoma is inconsistent with a functional response. 224 It has also been used to investigate whether recombinant insulin compounds could contribute to adenocarcinoma progression by stimulating local angiogenesis. 254 Mammalian target of rapamycin-dependent acinar cell neoplasia after inactivation of Apc and Pten in the mouse salivary gland was studied using MSI, with implications for human acinic cell carcinoma. 77 MSI was used for an analysis of HSPB1 expression in head and neck squamous cell carcinoma 216 and to determine that pericyte coverage of differentiated vessels inside tumor vasculature is an independent unfavorable prognostic factor for patients with clear cell renal cell carcinoma. 50
Prostatic cancer
MSI has been used to study seasonal variation in expression of markers in the vitamin D pathway in prostate tissue, 85 what the role of the vitamin D receptor protein expression in tumor tissue and prostate cancer progression is, 119 as a part of gamma tubulin IHC scoring in isolated prostate cells, 309 and to show that a prostatic intraepithelial neoplasia-dependent p27 Kip1 checkpoint induces senescence and inhibits cell proliferation and cancer progression. 187 MSI has also been used to show that interleukin-1-beta promotes progression of metastatic prostate cancer cells 1 and Trop-2 promotes prostate metastasis by modulating beta-1 integrin function. 311 It has been used to investigate whether the androgen receptor is a biologically relevant vaccine target for the treatment of prostate cancer 229 and whether androgen deprivation induces senescence characteristics in prostate cancer cells in vivo. 87 MSI has been used for the identification of the JNK signaling pathway as a functional target of the tumor suppressor PTEN, 326 to show that a dual blockade of PKA and NF-κB inhibits H2 relaxin-mediated castrate-resistant growth of prostate cancer sublines and induces apoptosis, 324 and to show that HP1gamma expression is elevated in prostate cancer and superior to Gleason score as a predictor of biochemical recurrence after radical prostatectomy. 287 MSI has been useful when combined with systems pathology for predictions of recurrence in prostate cancer 57,83 and in a systems-based modeling approach using transurethral resection of the prostate specimens, which yielded incremental prognostic significance to Gleason when predicting long-term outcome in men with localized prostate cancer. 83 MSI was also used to show that overexpression of fatty acid synthase is associated with palmitoylation of Wnt1 and cytoplasmic stabilization of beta-catenin in prostate cancer. 95 In a significant study, MSI was used to show that SMAD4-dependent barrier constrains prostate cancer growth and metastatic progression. 78
Ovarian cancer
MSI was used to study a novel infectivity-enhanced oncolytic adenovirus with a capsid-incorporated dual-imaging moiety for monitoring virotherapy in ovarian cancer 146 and to show that there is an elevation of seprase expression and promotion of an invasive phenotype by collagenous matrices in ovarian tumor cells. 141
Breast cancer
MSI has been used to quantitate progesterone and estrogen receptors in pregnant and premenopausal nonpregnant normal human breast, 302 to show that the detection of truncated HER2 forms in FFPE breast cancer captures heterogeneity and is not affected by HER2 therapies, 153 to show that AMD3465 regulates oncogenic signaling and prevents breast cancer growth and metastasis in vivo, 177 and to investigate the role of the nuclear receptor coactivator AIB1-delta4 splice variant in the control of gene transcription. 53 MSI has been used to show that the PPARgamma antagonist T0070907 suppresses breast cancer cell proliferation and motility via both PPARgamma-dependent and PPARgamma-independent mechanisms, 357 that high IGF-IR activity in triple-negative breast cancer cell lines and tumor grafts correlates with sensitivity to anti-IGF-IR therapy, 178 and to study the effect of mTOR inhibition alone or combined with MEK inhibitors on brain metastasis in vivo in triple-negative breast cancer models. 362 MSI has been used to show that integrin-associated CD151 drives ErbB2-evoked mammary tumor onset and metastasis, 71 that nuclear PARP-1 protein overexpression is associated with poor overall survival in early breast cancer, 264 that there is an oncosuppressive role of p53-induced miR-205 in triple-negative breast cancer, 240 and that the cocaine- and amphetamine-regulated transcript mediates ligand-independent activation of ERα and that ERα is an independent prognostic factor in node-negative breast cancer. 40 MSI has been used as a part of the evaluation of the heterogeneous inflammatory breast cancer microenvironment 154 and in a genetic strategy for combined screening and localized imaging of breast cancer. 333 MSI was used as a part of a quantitative architectural analysis of immune cell subsets in TDLNs from breast cancer patients and healthy lymph nodes 279 and in a randomized phase II trial of soy isoflavone supplementation for breast cancer risk reduction. 143
Melanoma
MSI was used to show that lymphatic invasion is independently prognostic of metastasis in primary cutaneous melanoma, 352 as a part of a phase II trial of Reolysin (reovirus serotype-3-Dearing strain) in patients with metastatic melanoma, 96 to determine GPNMB expression in uveal melanoma and to show its potential for targeted therapy, 341 and to show that lymphatic invasion predicts aggressive behavior in melanocytic tumors of uncertain malignant potential. 1
Sarcomas
MSI has been applied to investigate whether human sarcomas are telomerase dependent and have telomerase-independent telomere maintenance mechanisms, 107 that mitogen-activated kinase signaling promotes the growth and vascularization of fibrosarcoma, 79 and to quantitate COX-2 expression in a study of whether there is a role for celecoxib treatment in chondrosarcoma. 274
MSI has been used in several colorectal cancer studies. MSI played a role in performing a co-occurrence and morphological analysis for colon tissue biopsy classification; 197 in showing that the TNF family member APRIL promotes colorectal tumorigenesis; 158 in showing that ulcerative colitis-associated colorectal cancer arises in a field of short telomeres, senescence, and inflammation; 260 that VEGF receptor signaling links inflammation and tumorigenesis in colitis-associated cancer; 328 and in showing that Toll-like receptor 5 engagement modulates tumor development and growth in a mouse xenograft model of human colon cancer. 257 MSI has also been used to investigate the chemoprevention of BBN-induced bladder carcinogenesis by tamoxifen. 103
Lung cancer samples are prone to extensive autofluorescence, for which MSI is useful. MSI has been used to show the predictive value of intratumoral microvascular density in patients with advanced non–small cell lung cancer (NSCLC) receiving chemotherapy plus bevacizumab; 364 in showing that RRM2 regulates Bcl-2 in head, neck, and lung cancers and is a potential target for therapy; 248 to demonstrate that the EGFR and MET receptor tyrosine kinase-altered miRNA expression induces tumorigenesis and gefitinib resistance in lung cancers; 101 in monitoring the targeting of polymeric nanoparticles to lung metastases by surface attachment of YIGSR peptide from laminin; 272 that Met activation in NSCLC is associated with de novo resistance to EGFR inhibitors and the development of brain metastasis; 25 in a systems pathology model for predicting overall survival in patients with refractory, advanced NSCLC treated with gefitinib; 84 and to show that LKB1 inactivation dictates therapeutic response of NSCLC to the metabolism drug phenformin. 280
MSI has been used to study gliomas and other brain tumors, as an ex vivo validation of quantitative preclinical imaging of TSPO expression in glioma, 300 to show that therapy-resistant tumor microvascular endothelial cells contribute to treatment failure in glioblastoma multiforme, 35 and that the RNA-binding protein HuR promotes glioma growth and treatment resistance. 94
MSI has been used in studies of a range of blood-borne cancers, such as hemangioma, leukemia, multiple myeloma, and lymphoma. It has been used to study the expression of HLA-G in hemangioma and its clinical significance; 281 to show that bortezomib-induced “BRCAness” sensitizes multiple myeloma cells to PARP inhibitors; 211 to investigate the integrin β7-mediated regulation of multiple myeloma cell adhesion, migration, and invasion; 210 that YY1 expression predicts favorable outcome in follicular lymphoma; 209 and that clinical quantitation of immune signature in follicular lymphoma by RT-PCR-based gene expression profiling can be performed. 46 MSI has been used to show that HDAC inhibitors potentiate the apoptotic effect of enzastaurin in lymphoma cells, 32 that for pronounced hypoxia in models of murine and human leukemia there is a high efficacy of hypoxia-activated prodrug PR-104, 26 and that proapoptotic signaling activity of the anti-CD40 monoclonal antibody dacetuzumab circumvents multiple oncogenic transformation events and chemosensitizes non-Hodgkin lymphoma cells. 172
Pancreatic and liver cancer
MSI has been used as a part of the development of an orthotopic model of invasive pancreatic cancer in an immunocompetent murine host 313 and in conjunction with functional magnetic resonance imaging in an animal model of pancreatic cancer. 170 MSI was used to show the effect of transcatheter arterial embolization on levels of hypoxia-inducible factor-1alpha in rabbit VX2 liver tumors, 258 as a part of a study to distinguish reactive urothelium from neoplastic urothelium, 105 and in a comparison of hypoxia-inducible factor-1alpha expression before and after transcatheter arterial embolization in rabbit VX2 liver tumors. 325
In addition to the phase II clinical trials previously mentioned, MSI has been used as a part of clinical trials to determine the synergistic cytotoxicity of oncolytic reovirus in combination with cisplatin-paclitaxel doublet chemotherapy; 266 to show that the death receptor 5 agonist TRA8, in combination with the bisphosphonate zoledronic acid, attenuated the growth of breast cancer metastasis; 297 in a phase II study of MLN8237 (alisertib), an investigational Aurora A kinase inhibitor, in patients with platinum-resistant or -refractory epithelial carcinomas; 199 as a part of a study investigating dual combination therapy targeting DR5 and EMMPRIN in pancreatic adenocarcinoma; 145 as a part of a phase I study of intravenous reovirus and docetaxel in patients with advanced cancer; 55 to show that the antitumor effect of the angiogenesis inhibitor bevacizumab is dependent on susceptibility of tumors to hypoxia-induced apoptosis; 278 and in a randomized phase II study of soy isoflavone supplementation for breast cancer risk reduction. 83
The quantitative analysis of protein expression in tissues through visual assessment, while common for an expert to perform on hematoxylin and DAB samples, is still subjective and has many difficulties, including a lack of reproducibility and potentially drastic differences even among experts. In addition, the traditional percentage positivity or 0/1+/2+/3+ scoring systems do not give enough bins, or levels, to provide good quantitative data for many research applications. To address these deficiencies, a methodology called advanced quantitative analysis, or AQUA, ¶ was developed. 27,48,268 This method utilizes 2 fluorophores, one as a tissue compartment marker (typically cytokeratin, but others are used) and another as an analytic marker (eg, HER2, pAKT) along with a DAPI counterstain. A full description of the methodology can be found elsewhere. 80 Recently, MSI, automated image acquisition and analysis. and AQUA have been combined, enabling more accurate measurements, particularly of weakly expressing markers because of the removal of any residual autofluorescence and expanding the potential for the simultaneous analysis of multiple analytes using the AQUA technology. 82,265,128,265,81
Cancer stem cells
Stem cells and circulating tumor cells are another application that requires the use of multiple markers to phenotype cells, and MSI has played an important role in studying them. The existence of cancer stem cells is still a subject of debate. However, whether putative cancer stem cells actually are stem cells or not, determining which cells in tissue have one of the many different phenotypes put forth as being that of a cancer stem cell is an inherently multimarker problem and one in which MSI has aided. 117 There has been a recent review of technologies and methods used for the detection, enrichment, and characterization of cancer stem cells, including MSI. 340 With MSI, it has been determined that most early disseminated cancer cells detected in bone marrow have cancer stem cell phenotypes, 16 that oncostatin-M promotes phenotypic changes associated with mesenchymal and stem cell–like differentiation in breast cancer, 338 and that methylation of cancer stem cell–associated Wnt target genes predicts poor prognosis in colorectal cancer patients. 70 In addition, MSI has been used for the identification of putative cancer stem cells (CD44+/CD24–) in breast carcinoma 288 and for determining the amounts of putative breast cancer stem cells in lymph node occult metastases of patients with node-negative breast cancer. 118 MSI has been used to look at other kinds of stem cells as well, including stem cell factor expression and hematopoietic stem cell migration after renal ischemia 293,294 and quantum dot imaging for embryonic stem cells. 175 MSI has been combined with a micromagnetic–microfluidic device for rapid capture and culture of rare circulating tumor cells 137 and for viable circulating tumor cell detection using multiplex RNA in situ hybridization predicting progression-free survival in metastatic breast cancer patients. 236
As a result of the importance of flow cytometry and its multimarker phenotyping methodology in immunology, hematology, infectious disease, and virology, there has been considerable effort put into replicating the same kinds of multimarker labeling that is routinely done in flow cytometry in situ in a tissue section. One would then be able to image a sample and look at the distribution and locations of the same kinds of multimarker phenotyped cells that are important in flow cytometry. Because of this inherent multimarker, multicolor aspect of imaging in immunology, it is not surprising that MSI has played an important role in the imaging and analysis of immune cell applications, in immunology and infectious disease, and, increasingly, in cancer and tumor immunogenicity applications.
Cancer immunology
MSI has played an important role in the understanding of immune cell subsets in tumors and lymphoid tissues. A general dual-labeling technique to investigate T-lymphocyte subsets in FFPE rat lymphoid tissue has been developed. 249 MSI has been used to show that tumor hypoxia promotes tolerance and angiogenesis via CCL28 and T-regulatory cells (Tregs); 90 that CD20+ tumor-infiltrating lymphocytes have an atypical CD27– phenotype and that, when combined with CD8+ T cells, promote good prognosis in ovarian cancer; 214 that activated CD69+ T cells foster immune privilege by regulating IDO expression in tumor-associated macrophages; 363 that the expression of macrophage migration inhibitory factor in neuroblastoma leads to the inhibition of antitumor T-cell reactivity; 367 that DEK is a neuroblastoma tumor antigen in the context of Treg inhibition; 365 that the infiltration of T cells into neuroblastoma is interferon-gamma dependent; 251 that tumor-infiltrating macrophages are associated with metastasis suppression in high-grade osteosarcoma, providing a rationale for treatment with macrophage activating agents; 43 and to explore FOXP3+ Treg and epithelial FOXP3 expression in cancers of the breast. 157 In lymphoid tissue, MSI has been used to show that interleukin-1-beta selectively expands and sustains IL-22+ immature natural killer cells in secondary lymphoid tissue 129 and how adoptive transfer of allogeneic tumor-specific T cells mediates effective regression of large tumors across major histocompatibility barriers. 33 It has also been used to investigate the immunoprofile of endocervical and endometrial stromal cells and its potential application in localization of tumor involvement 18 and that T(H)1, T(H)2, and T(H)17 cells instruct monocytes to differentiate into specialized dendritic cell subsets. 6 When combined with automated acquisition and advanced image analysis, MSI has been used to evaluate the heterogeneity of the inflammatory breast microenvironment 154 and in a quantitative architectural analysis of immune cell subsets in entire sections of TDLNs in breast cancer patients. 279
EMT
One of the “hallmarks of cancer,” 115,116 thought to be an important step toward metastasis, and in the investigation of the tumor microenvironment, MSI has been used to prove EMT in biliary atresia fibrosis 75 and portal tract fibrogenesis in biliary atresia, 72 to investigate markers of fibrosis and EMT in field cancerization in normal tissue adjacent to breast tumors, 312 and to investigate TRAIL-transduced multipotent mesenchymal stromal cells and TRAIL resistance in CRC cell lines in vitro and in vivo. 205 It has also been used to investigate cancer stem cells, EMT, and the phenotypic changes associated with oncostatin-M promotion in breast cancer 338 and to measure EGFR and MET receptor tyrosine kinase–altered miRNA expression in lung cancer and the association with gefitinib resistance. MSI has also been used to investigate the effects of cold ischemia and inflammatory tumor microenvironment on the detection of cell signaling markers in IHC, 34 how stromal PTEN-regulated miR-320 affects the tumor microenvironment, 41 and that miRNA binds to Toll-like receptors to induce prometastatic inflammatory response. 89 MSI has been used with quantum dots to map out tumor heterogeneity, 180,237,238 to track the inflammation-induced mobilization of mesenchymal stem cells, 291 and in the evaluation of the heterogeneous inflammatory breast cancer microenvironment. 154 MSI has also been used in quantum dot–based quantification that revealed differences in subcellular localization of EGFR and E-cadherin between EGFR-TKI sensitive and insensitive cancer cells.
MSI and infectious diseases
MSI has been used on the in situ detection of the colocalization of cytokine responses to viral pathogenesis, 7 in HIV to look at human papillomavirus pseudovirion vaccines delivering simian immunodeficiency virus DNA, 110 at the loss of neuronal integrity during HIV infection in humanized mice, 65 at the transmission of the HIV virus across cervical mucosa to tonsil tissue, 290 to investigate the antiretroviral and neuroprotective responses to NanoART in a murine model, 64 to examine the distribution of CD4+/CCR5+ T cells in the gut during mother-to-child transmission of HIV, 44 to investigate the delivery of antiretroviral drugs using nanoparticles, 185 in investigating the loss of a tyrosine-dependent trafficking in SIV and its effect on disease progression, 39 and to look at respiratory insufficiency in rodents infected with West Nile virus. 204 In tuberculosis research, it has been used to investigate the influence of oral lactoferrin on Mycobacterium tuberculosis–induced immunopathology, 335 the role of complement factor C7 in lung immunopathology in tuberculosis, 336 and the increase of Tregs and DEC-205-positive macrophages in cavitary lesions in postprimary tuberculosis. 337 Hepatitis C and occult hepatitis C viral infection is a newly reported pathologic entity associated with increased risk of developing hepatocellular carcinoma and lymphoproliferative disorders, and MSI has been used to investigate whether the hepatitis C virus is potentially lymphotropic, invading and propagating in cells of the immune system. 31 MSI has also been used to investigate how occult hepatitis B viral infection induces DNA damage that can be seen in FISH in peripheral blood lymphocytes 30 and to determine the oncolytic reovirus distribution in human tumors. 138 MSI has been used to investigate novel targets for malarial chemotherapy 206 and the susceptibility of erythroblasts to Plasmodium infection, 299 to investigate the role of SUR7 in Candida albicans infections, 29 to investigate the responses of endothelial cells to pathogenic and nonpathogenic leptospiral infections, 196 and to show the transient detection of chlamydial-specific Th1 memory cells in the peripheral circulation of women with genital tract infections. 323 The role of S100A8/A9 in the host defense during murine urinary tract infections was also investigated using MSI, 74 and MSI was used to determine that hemolysin and the multifunctional autoprocessing RTX toxin are virulence factors during intestinal infection of mice with Vibrio cholerae El Tor O1 strains, 228 that anthrax lethal toxin inhibits growth of and vascular endothelial growth factor release from endothelial cells expressing the human herpes virus 8 viral G protein coupled receptor, 73 for the in situ detection of aspergillus 18 s ribosomal RNA sequences using a terminally biotinylated locked nucleic acid probe, 202 and that APOBEC3 proteins expressed in mammary epithelial cells are packaged into retroviruses and can restrict transmission of milk-borne virions. 226
Inflammation is an important disease response in which immune cells play an important role. MSI has been used in inflammation research to monitor the effects of Dragon’s Blood on the inhibition of chronic inflammatory and neuropathic pain response through blocking the release of substance P in rats, 174 and to look at the apoptosis resistance of mucosal lymphocytes and IL-10 deficiency in Crohn disease, 271 and in inflammatory dermatoses and in normal skin, MSI has been used to phenotype and then enumerate FOXP3/GITR/CD4/CD25 in lymphocytes 67,68 and to look at a range of inflammatory responses in the rat lung. 344–346
As in flow cytometry, immune cell subsets—or specific populations of immune cells gated by a multiple marker phenotype—are central to immunology, and MSI has aided developments in a range of tissues and applications of the imaging and counting of specific immune cell subsets in situ, including investigating altered B-cell development and anergy in the absence of FOXP3, 161 CD8+ effector-memory T cells in decidual tissues, 308 the modulation of relaxin on the proinflammatory secretion of macrophages, 123 the overrepresentation of IL-17A and IL-22 CD8 T cells in psoriatic skin lesions, 255 the distribution of immunomodulatory cells in the lungs of patients with pulmonary fibrosis, 220 and an intraepithelial phenotype of CD8+ T-cell involvement during influenza infection in human lung. 239 It has also been important in bone regarding the effect of UHMWPE particles on macrophage levels and osteolysis 253 and at IRF7-dependent IFN-β production in medullary thymic epithelial cell development. 231
Conclusions
Although the field of anatomic pathology changed only slowly in its first centuries, the past few decades have seen an increasing rate of change as the questions being asked in a biopsy specimen become increasingly complex. IHC is becoming ubiquitous. The need for quantitative accuracy in the assessment of IHC and IF samples and the use of multiplexing to address complicated cell phenotype and expression questions have led to the adoption of MSI methods in a range of research and clinical trial applications. MSI has aided the development of these imaging methods by addressing 2 major issues: in brightfield, MSI allows the separation and independent quantitation of multiple chromogens in the same cellular compartments; in fluorescence, MSI separates interfering autofluorescence from fluorophores of interest, enabling accurate fluorescence intensity quantitation in FFPE samples. In addition, the combination of MSI with software for morphologic tissue area segmentation and cellular and subcellular segmentation to perform quantitative per-cell (and per–cell compartment) intensity quantitation provides researchers with per-cell data about the cells within particular morphologic subregions of an image, which is analogous to the multimarker information found in flow cytometry. As the demands on anatomic pathology grow in response to the advances in anatomic and molecular pathology and genetic testing, MSI, especially when combined with advanced software to obtain multimarker, per-cell quantitative data, will continue to be on the forefront of the development of new methods and new tests.
Footnotes
Acknowledgements
Declaration of Conflicting Interests
The author is an employee of PerkinElmer, Inc, and a former employee of both Caliper Life Sciences and Cambridge Research & Instrumentation, Inc, now wholly owned subsidiaries of PerkinElmer, Inc.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
