Abstract
A fatal combined infection with canine distemper virus (CDV) and orthopoxvirus (OPXV) in Asian marmots (Marmota caudata) is reported in this article. A total of 7 Asian marmots from a small zoological garden in Switzerland were found dead in hibernation during a routine check in the winter of 2011. The marmots died in February 2011. No clinical signs of disease were observed at any time. The viruses were detected in all individuals for which the tissues were available (n = 3). Detection of the viruses was performed by reverse transcription polymerase chain reaction. The most consistent gross lesion was a neck and thorax edema. A necrotizing pharyngitis and a multifocal necrotizing pneumonia were observed histologically. Numerous large intracytoplasmic eosinophilic inclusions were seen in the epithelial cells of the pharynx, of the airways, and in the skin keratinocytes. Brain lesions were limited to mild multifocal gliosis. Phylogenetic analysis revealed that the marmot CDV strain was closely related to the clusters of CDVs detected in Switzerland in wild carnivores during a local outbreak in 2002 and the 2009–2010 nationwide epidemic, suggesting a spillover of this virus from wildlife. The OPXV was most closely related to a strain of cowpoxvirus, a poxvirus species considered endemic in Europe. This is the first reported instance of CDV infection in a rodent species and of a combined CDV and OPXV infection.
Canine distemper virus (CDV) is one of the most significant viral pathogens of carnivores. 37 It is an RNA virus characterized by a single stranded, negative-sense, nonsegmented genome. It belongs to the genus Morbillivirus, within the family Paramyxoviridae. The genome encodes 6 structural proteins (matrix [M], fusion [F], hemagglutinin [H], nucleocapsid [N], polymerase [L], and phosphoprotein [P]). Hemagglutinin is a “multitasking” molecule serving both as attachment protein and for cell-to-cell transmission. 36,39 This molecule is also under strong immunological pressure, which translates into a high mutation rate, making it an ideal candidate both for strain comparison and phylogenetic investigation. 20,22 –24,41
Orthopoxviruses (OPXVs) are large double-stranded DNA viruses with a genome ranging from 175 to 225 Kb. 19 The genome encodes for hundreds of genes comprising all the machinery for the de novo synthesis of viral DNA and for the host immune evasion. 1 This genus comprises, among others, several human pathogens such as smallpox, cowpox, vaccinia, variola, and monkeypox. 19 Other OPXVs cause disease in rodents and lagomorphs and include ectromelia virus (ECTV), volepox (VOPX), and rabbitpox (RPXV). 9,14
CDV infection presents as an acute and often fatal disease. Respiratory and neurological clinical signs are the most frequently observed. 4,8,35 Postmortem findings are typified by gross (lung) and histological (lung and brain) tissue changes. An interstitial to a bronchointerstitial pneumonia is the most common finding in the lung, regardless of the infecting strain. Brain lesions can range from a common demyelinating leukoencephalomyelitis to rare instances of a polioencephalitis and appear to be strain dependent. 4,24,32
CDV is a pathogen that is able to infect multiple species of carnivores worldwide. 3,4,15,28,30,38 It has been documented in noncarnivorous hosts such as nonhuman primates 27,33,40 and collared peccaries (Tayassu tajacu). 2 No infection has ever been reported in rodents until now.
OPXV infections can be followed by localized or systemic disease. Intermediate patterns are also known. 7 The disease can range from hyperacute to chronic and is generally associated with respiratory and cutaneous lesions. 7,9 Rodents are considered the natural hosts of cowpoxvirus, although this agent has been shown to be able to infect humans, cats, and zoo animals. 1,9 Cowpox infection in rodent has been shown to be associated with respiratory and cutaneous lesions and can be fatal. 9 No specific marmot OPXV has been detected up to date.
Here we report a fatal combined infection with CDV and OPXV in a group of captive Marmota caudata housed in a small zoological garden in Switzerland. Although dual infection with poxviruses and other agents is known in the literature (Pox and Chlamydia), 17 this is the very first example of combined OPXV and CDV infection in the same host.
Our findings suggest that: (1) CDV may expand to novel species traditionally considered not susceptible to this virus; (2) CDV can occur with other viral agents such as poxviruses; (3) CDV and OPXV could act synergistically, leading to hyperacute death.
Materials and Methods
Animals
Seven marmots (Nos. 1–7) of the species M. caudata were submitted for routine necropsy. They were imported from central Asia 3 years prior to their death and were kept in a small zoological garden at 2000 meters altitude in Switzerland. All individuals were housed in the same enclosure and bred successfully starting 1 year after their arrival. In February 2011, a caretaker found them dead in hibernation during a routine check. None of the marmots exhibited any clinical signs of disease prior to hibernation. The suspicion of a potential predation was mentioned by the zookeeper at the time of submission.
Pathology
A complete necropsy was performed on all the submitted animals. Different subsets of tissue samples, including those with gross lesions, were collected from the marmots for histological investigation. More specifically, the liver, kidneys, urinary bladder, skin, and brain were collected from marmot No. 1, the brain and pharynx from marmot No. 4, and the lung from marmot No. 5. No tissues were collected from marmot Nos. 2, 3, 6, and 7 given the absence of significant gross lesions other than a subcutaneous edema in marmot No. 3. The tissues were fixed in 10% buffered formalin, processed, and embedded in paraffin. Formalin fixation for the tissues described above was prolonged for up to 10 days, in contrast with the normal 24- to 48-hour routine fixation time, for logistical reasons. Following processing, 5-micron-thick sections were obtained from each tissue and stained with hematoxylin and eosin (HE) prior to examination under light microscopy.
CDV-Specific Immunohistochemistry
Immunohistochemistry (IHC) was performed on the lung from marmot No. 5, the pharynx from marmot No. 4, and the brain from marmot No. 1. IHC was performed according to an established protocol using the monoclonal antibody D110 directed against the N protein of the virus. 5
Bacteriology
General bacteriologic examination was performed on the liver, kidney, and spleen from marmot Nos. 1, 2, and 4 and on the lung from marmot No. 5.
Molecular Diagnostic
RT-PCR of CDV N, H and of OPXV DNA Polymerase (poxDNApol) and Hemagglutinin (poxH) Genes and Gene Sequencing
Total RNA extraction was performed on the tissues obtained from the paraffin blocks from marmot Nos. 1 (brain), 4 (pharynx), and 5 (lung). RNA extraction was performed with the Qiagen paraffin tissue kit (Hombrechtikon, Switzerland) following the manufacturer’s instructions. cDNA synthesis was performed using M-MLV reverse transcriptase (Promega, Madison, WI) following the manufacturer’s instructions. An established polymerase chain reaction (PCR) protocol was used to obtain the partial sequence of the N gene from the samples tested (from marmot Nos. 1, 4, and 5). 13
Full amplification of the H gene was performed using the PCR protocol published by Sekulin et al. 31 The cDNA that was used as a template was that obtained from the lung of marmot No. 5.
The cDNA from all the tissues tested for CDV was also tested by PCR for the presence of orthopoxviruses using 2 pairs of “panorthopoxvirus” primers directed to amplify the partial sequence of the poxDNApol and poxH genes, respectively, according to established protocols. 19 Control DNA was obtained from a recombinant vaccinia virus (MVA-T7), which was a kind gift from Dr Philippe Plattet.
The reverse transcription PCR (RT-PCR) products from the CDV N and H genes and from the OPXV poxDNApol and poxH genes were submitted for automated sequencing (ABI Prism 3100 genetic analyzer; Applied Biosystems, Foster City, CA) that was carried out using the BigDye Terminator cycle sequencing kit (Applied Biosystems) following the manufacturer’s instructions. Sequences were obtained using the same primers used for the amplifications of the different amplicons.
Phylogenetic Study
The full amino acid sequence of the CDV hemagglutinin protein obtained from marmot No. 5 (lung), along with the homologous sequences from 56 CDV strains comprising those of 6 novel Swiss strains 24 (accession numbers are listed in the supplemental material) and from a strain of measles virus (used as the outgroup), was used for the phylogenetic analysis (Fig. 1). Multiple alignments were performed with the CLUSTAL W program, 34 available through the Biology workbench (3.2) of the San Diego Supercomputer Center website (http://workbench.sdsc.edu/). Phylogenetic trees were resolved using software applications included in the PHYLIP (Phylogeny Inference Package, version 3.69) program package. 11 For distance matrix analysis, the aligned amino acid sequences were first processed with PROTDIST (Jones-Taylor-Thornton model) and then with the FITCH program (Fitch-Margoliash with global rearrangements, 10 times randomized input of the sequences and 100 data set replicates). For bootstrap analysis, the SEQBOOT program was run before PROTDIST and FITCH. The most probable tree was calculated with the CONSENSE program. The trees were visualized with the TREEVIEW program. 26 For maximum likelihood analysis, the aligned sequences, following SEQBOOT, were analyzed first with the program PROTML and then with CONSENSE. The final tree was visualized with TREEVIEW. For parsimony analysis, the program PROTPARS was run after SEQBOOT. The most probable tree was then calculated with the program CONSENSE and visualized with TREEVIEW.
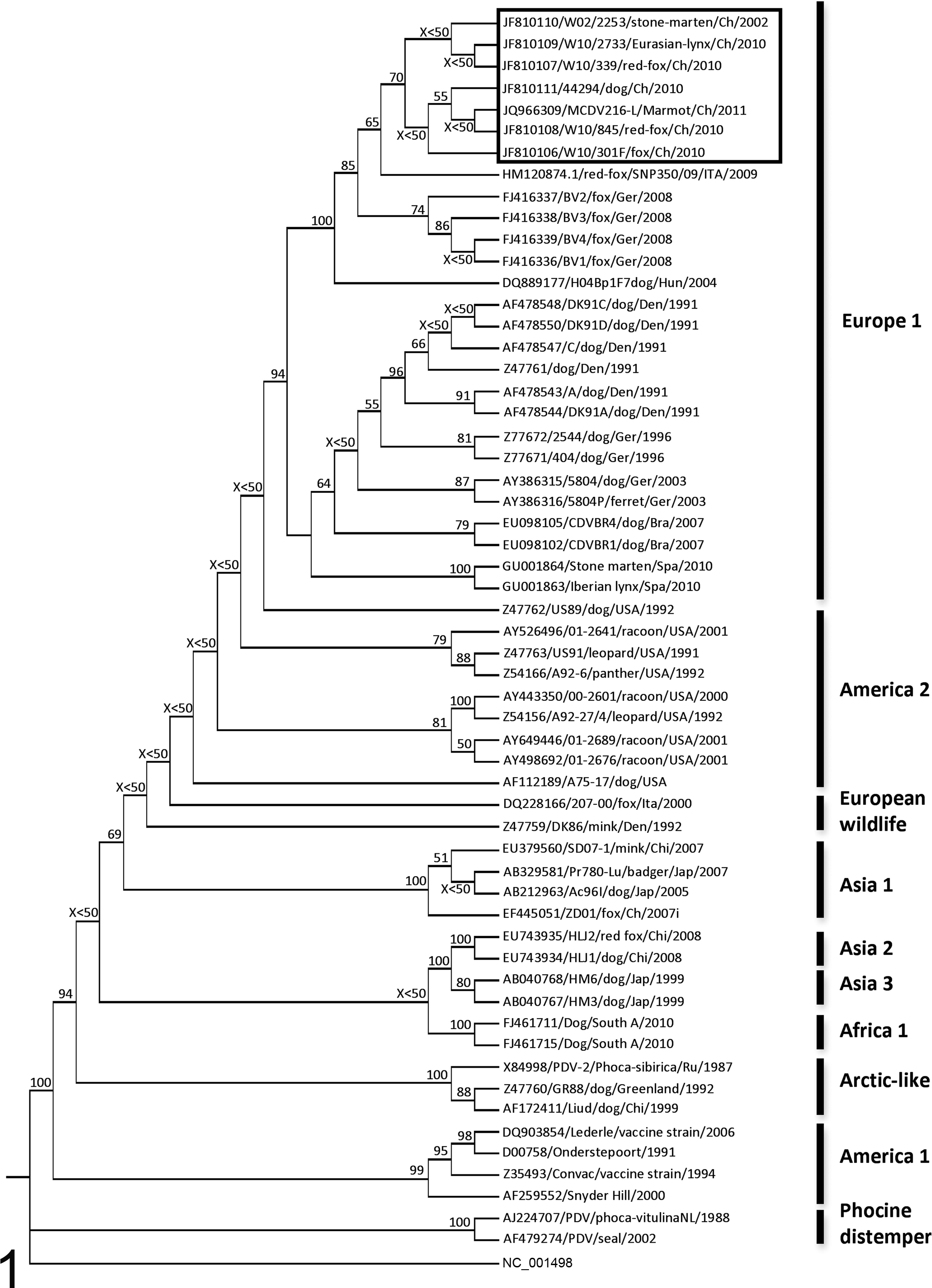
Distance-matrix phylogenetic analysis based on the full amino acid sequence of the H protein. The marmot strain (JQ966309/MCDV216-L/Marmot/Ch/2011) clusters with the other Swiss isolates detected during the 2002 and 2009–2010 canine distemper virus epidemics in wildlife (black frame). Geographical groups are reported on the right side of the tree, whereas the bootstrap values are reported at the respective nodes (bootstrap values lower than 50 are reported as X<50) [Rooted tree; Measles strain (NC_001498) = outgroup] (the amino acid sequences have been derived from the nucleotide sequence accession numbers reported in the tree).
Glycosylation Analysis
Analysis of potential glycosylation sites was performed on the amino acid sequence of the marmot CDV hemagglutinin protein using the online software ``NetNGlyc 1.0 Server'' (http://www.cbs.dtu.dk/services/NetNGlyc).
Results
Pathology
Gross Findings
Marmot Nos. 1, 2, 3, 4, and 7 were well preserved, whereas Nos. 5 and 6 showed advanced autolytic changes and rigor mortis could not be detected. The body condition of all the animals was good, and there was no sign of significant fat depletion. Partial skin and muscle loss over the ribcage of marmot No. 1 and from the abdominal area of marmot No. 7 was also observed, with no associated hemorrhages. The most consistent gross lesion was a marked subcutaneous edema that was observed in 3 animals (marmot Nos. 1, 3, and 4). The edema was present in the region of the head and neck (marmots Nos. 1 and 3) or the thorax (marmot No. 4). Furthermore, there was extensive pulmonary atelectasis in 1 marmot (No. 5) and a pharyngeal hemorrhage with coagulated blood in another animal (No. 4). Mild multifocal hyperkeratosis was observed in marmot No. 1.
Histological Findings
Histological examination of the hemorrhagic area of the pharynx of animal No. 4 revealed multifocal necrosis of the mucosal epithelium. Marked ballooning degeneration and numerous intracytoplasmic eosinophilic inclusion bodies up to 15 μm in diameter were detected in the epithelial cells adjacent to the foci of necrosis (Figs. 2 and 3). This lesion was surrounded by extensive hemorrhage with a thick layer of fibrin covering the epithelium. A very mild peri- and intralesional infiltration with neutrophils was also observed. Bacterial colonies (rods) suggestive of terminal bacteremia were present in some small blood vessels. Likewise, there was multifocal necrosis of the epithelium of bronchi and bronchioles in the lung of animal No. 5 with ballooning degeneration and intracytoplasmic eosinophilic inclusion bodies similar to those detected in the pharynx of marmot No. 4 (Fig. 4). Furthermore, there were extensive pulmonary hemorrhages and numerous intravascular bacterial rods. Eosinophilic intracytoplasmic inclusions were also detected in the keratinocytes within some of the foci of hyperkeratosis observed on the skin from marmot No. 1. Affected areas of the skin also showed moderate keratinocyte vacuolization and prominent intercellular edema (spongiosis). The collagen fibrils were moderately separated by clear spaces (edema) in the superficial dermis while small numbers of lymphocytes and plasma cells were congregating in the perivascular spaces and, to a minor extent, within the remaining interstitium (Fig. 5).
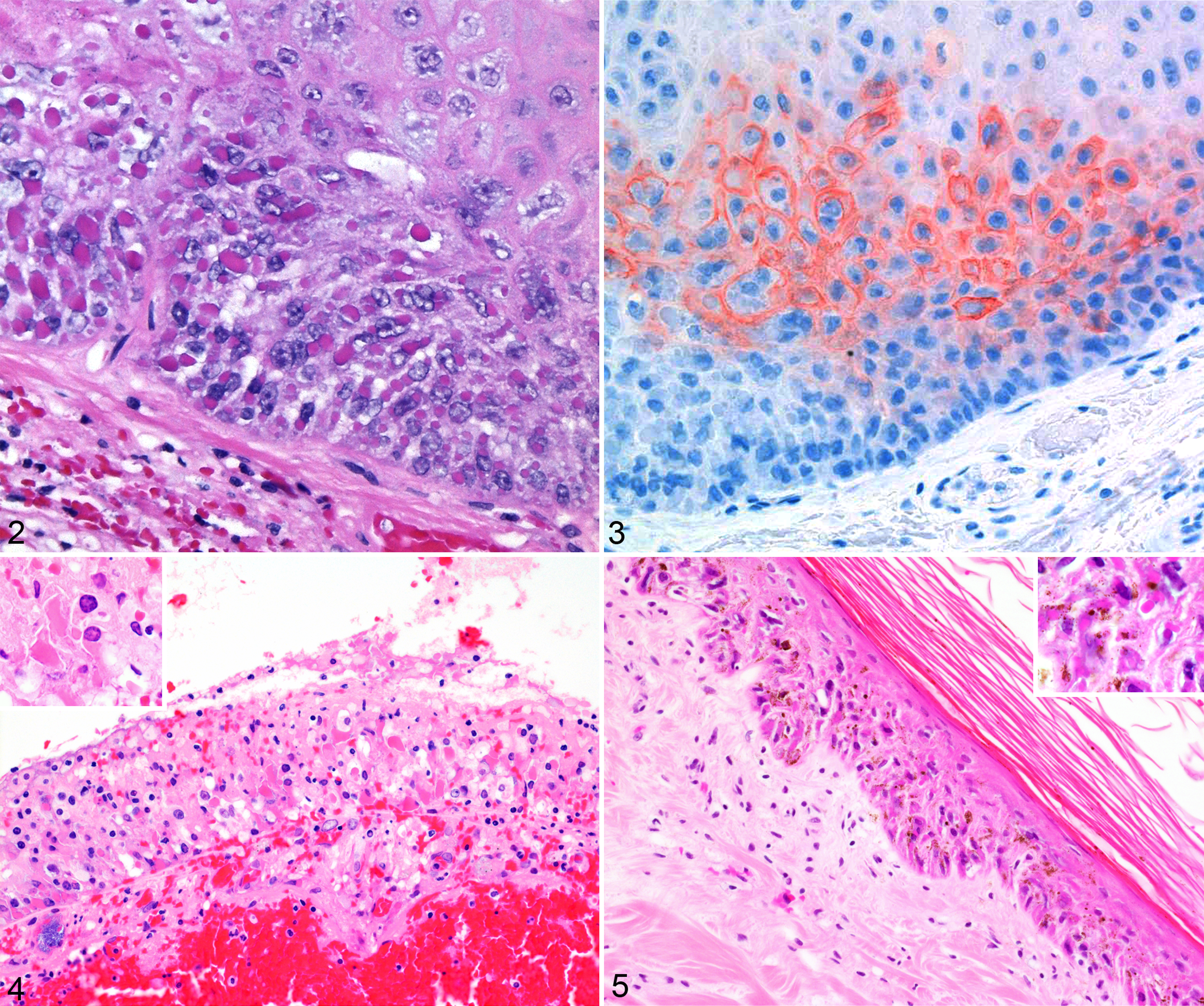
Pharynx; Asian marmot No. 4. Note the numerous prominent intracytoplasmic eosinophilic inclusions within the vacuolated and edematous epithelium of the pharynx. Hematoxylin and eosin (HE).
No microscopic lesions were detected in the remaining organs examined, except for few glial nodules in the brain of marmot No. 1. Bacteriologic examination yielded Serratia sp from the liver, kidney, and spleen of marmot No. 4 and from the lung of marmot No. 5.
CDV-Specific Immunohistochemistry
Patchy areas of positive staining were seen in the mucosa of the pharynx of marmot No. 4 (Fig. 3). Staining was observed both in cells containing intracytoplasmic inclusions and in those without. The pigment was mostly distributed in the cytoplasm and markedly highlighted the cell membrane, whereas the inclusions were not stained. Lack of convincing staining was observed in the lung of marmot No. 5 and in the brain or marmot No. 1.
Molecular Diagnostic
RT-PCR
CDV N gene
The partial sequence of the N gene was successfully amplified in all the marmots tested (n = 3). A 287–base pair (bp)–long amplicon was obtained from all the positive PCR reactions. Sequencing of the fragment confirmed the identity of the infectious organism as CDV.
CDV H gene: phylogenetic and glicosylation analyses
The full length of the H gene was obtained from the lung of marmot No. 5. The complete amplicon was 2199 bp long, whereas the open reading frame was 1824 bp long and encoded for 607 amino acids (JQ966309/MCDV216-L/Marmot/Ch/2011). Distance matrix analysis unambiguously clustered the novel marmot strain within the group of the Swiss CDV strains detected during a local outbreak in 2002 and the 2009–2010 nationwide (Swiss) epidemic in wildlife (Fig. 1). Maximum likelihood and maximum parsimony analyses provided similar results (data not shown).
In comparison with 6 recently sequenced hemagglutinin genes from Swiss CDV distemper strains detected during 2 CDV outbreaks in wild carnivores, the closest sequence similarities were observed between the marmot and the 44294 (dog) and W02/2253 (stone marten) strains (Fig. 1). Two nucleotide substitutions were observed between the marmot strain and 44294 and between the marmot strain and W02/2253. Three nucleotide mutations were observed between the marmot strain and W10/301 (red fox), 4 with W10/339 (red fox) and W10/845 (red fox), and 8 with W10/2733 (Eurasian lynx). The observed nucleotide mutations accounted for 2 amino acid substitutions once compared with W10/339 and W10/2733, whereas 1 amino acid substitution was observed in comparison with all the other Swiss strains. Nucleotide similarity between the marmot strain and those from Swiss wildlife ranged from 99.94% (44294 and W02/2253) to 99.56% (W10/2733), whereas similarity in the amino acid sequences ranged from 99.83% (W10/301, W10/845, W02/2253, 44294) to 99.67% (W10/2733 and W10/339).
Software analysis revealed the presence of 7 potential N-glycosylation sites (ectodomain) in the H protein overlapping with those observed in the strains detected during the 2002 and 2009–2010 CDV outbreaks in Switzerland and in those from recent CDV outbreaks in Italy and Germany. 10,21,23,31
OPXV poxDNApol and poxH genes
The PCR amplicon obtained for poxDNApol was 178 bp long and was detected in all the tissues available. The PCR amplicon obtained for poxH was consistent with expected size (133 bp long) and was detected in the tissues from all available marmots (n = 3).
Sequencing of the entire poxDNApol amplicon and partial sequencing of the poxH PCR product confirmed the identity of the infectious organism as an OPXV. The closest match for both sequences (using the “BLAST” software; http://blast.ncbi.nlm.nih.gov/Blast.cgi) was a cowpoxvirus.
Discussion
Canine distemper virus is an organism with a remarkably large host spectrum among both domestic and wild carnivores. Occasionally, it has also been reported in groups of noncarnivores, but it has never been detected in a rodent until now. A similarly wide host spectrum characterizes also OPXVs, which are known to be able to infect humans and nonhuman primates, large numbers of mammalian species (including rodents and lagomorphs), birds, and reptiles. 9,16,19
The source of the OPXV, presumptively a cowpoxvirus, is considered to be wild rodents,14 which are most likely to have had easy access to the marmots' enclosure. Differently, the source of the CDV infection for these marmots is not clear. Since the marmots were kept in captivity, it is relatively unlikely that canidae or animals of comparable size would have had access to them. However, there was a suspicion of wild ermines entering in this zoo, which, as mustelidae, are known to be susceptible to CDV infection. 24 Their small body size might have allowed them to penetrate into the enclosure of the marmots. However, no lesions consistent with bites wounds inflicted intra vitam were detected despite full skinning of the animals. Nevertheless, the partial skin and muscle loss over the ribcage of marmot No. 1 and from the abdominal area of marmot No. 7 (without hemorrhages) indicates that carnivores (or other scavengers, including small rodents) had access to the marmots, albeit after their death.
The marmots that were fatally infected with both CDV and OPXV were found dead in hibernation during a routine check, and no prior clinical signs were noticed. The timing of the infection could not be conclusively determined. The nature of the lesions would suggest a very short interval between the infection and the time of death, consistent with a recent infection followed by an acute to hyperacute course of the disease. This is as would be expected in accidental hosts for CDV such as Asian marmots but also in rodents with a disseminated form of OPXV infection. Evidence suggests that OPXVs might express a significant degree of virulence in individuals with impaired immune response. 9 CDV might have contributed to weaken the host’s immune response, thus facilitating the OPXV infection with the development of the systemic form of the disease. OPXV might have further impaired the host’s immune defenses, given its large array of immune-evading gene armory. 1 In addition, the high virulence of CDV and OPXV might have well been enhanced by the “dormant” immune system of the hibernating marmots. This hypothesis is supported by multiple examples of indirectly increased pathogen virulence during hibernation spanning from reptiles to mammals. 25,29 Testudinid herpesvirus, for example, can induce significant disease in tortoises either during hibernation or in close proximity to it (early spring and late fall). 25 Similarly, hibernating bats have been shown to be particularly susceptible to Geomyces destructans, the agent of the white nose syndrome, 12,29 and it has been suggested that the quiescent status of their immune system might play a role in the pathogenesis of the disease. 29 In addition, it is known that in hibernating mammals, the immune system undergoes multiple adaptations, which may lead to a reduction of the circulating lymphocytes and of the T-lymphocyte response, all extremely relevant in terms of antiviral response. 6 Within this scenario, it would appear that the low winter temperature could have represented a relevant predisposing condition, which might have significantly reduced the host’s chances to respond to the CDV and OPXV infection. Presumptive increased exposure to CDV, given the recent nationwide epidemic and bona fide enhanced virulence of the virus, 24 might have been significant additional factors promoting development of the disease. The immune-suppressive nature of CDV combined with the highly efficient immune evasion of OPXV and the dormant immune system of the hibernating animals might have led to the “perfect storm” resulting in a hyperacute death of the marmots.
The most common macroscopic lesion observed in the affected marmots was edema in multiple regions of the body. Interestingly, edema is also one of the most common lesions observed in pox-infected animals. 9 Differently, the classic cutaneous rash could not be detected.
The microscopic lesion that is considered the hallmark of CDV infection consists of the presence of intracytoplasmic and/or intranuclear eosinophilic inclusions, whereas infections with OPXVs are associated solely with intracytoplasmic eosinophilic or basophilic inclusions (type I and type II inclusions, respectively). Intracytoplasmic eosinophilic inclusions were seen in the epithelial lesions in the pharynx of marmot No. 4, the airways of the lung of marmot No. 5, and the skin of marmot No. 1. The main lesions were observed in the upper respiratory tract and in the lung, where CDV- and OPXV-associated lesions may be observed in the earlier stage of the disease following the infection through aerosols. The absence of significant lesions and of detectable CDV viral antigen in the brain and other tissues is also consistent with an acute disease. The CDV-specific IHC revealed the presence of chromogenic substrate limited to the pharynx epithelium. The positive stain was confined to the cell membranes and cytoplasm of several but not all infected cells, and no inclusions could be stained. Possible explanations for these results could be poor antigen retrieval, degeneration of the viral protein because of prolonged fixation of the tissue for up to 10 days, or some undetermined significant mutation that might have occurred at the binding site of the monoclonal antibody that was used for this study (D110: an anti–N MAb). Alternatively, the inclusions might have been composed mainly of poxviral antigenic material, whereas the CDV antigen might have not been predominant in these cellular structures.
The Serratia spp infection detected in 2 animals is most likely a secondary opportunistic infection and might be a sequel of an immune suppression due to the distemper virus infection and/or occurring during hibernation. 6,18
Phylogenetic, sequencing, and glycosylation analyses strongly support the wildlife origin of the marmot viral strain. This finding would be consistent with the hypothesized contact between the captive marmots and local wild carnivores (Fig. 1). Although the phylogenetic tree shows a fox (W10/845) and a dog (44294) strain to be the most closely related to that of the marmot, the low bootstrap value does not rule out the possibility that other Swiss strains might share a similar relatedness, as suggested by the sequence analysis. This presumptive spillover is not an isolated case, since it is known that CDV infection of animals in the permanent collections of zoological gardens can be primed by free-ranging local wildlife. 20 Nevertheless, this is generally regarded to be the case only in well-known CDV susceptible species such as the Zoo-housed wild carnivores. Our results indicate that the spectrum of species at risk might be wider. In fact, according to some of the recent reports of CDV infections in “nonconventional” species, 27,33,40 and in light of the present outbreak in marmots, it appears that host specificity in distemper infection might be a relatively loose term. No phylogenetic inference was performed on the OPXV detected in the marmot tissues since the limited length of the sequences available and their very conserved nature would have not provided enough data for an unambiguous resolution of the different OPXVs that might have been considered. Nevertheless, the close resemblance with several strains of cowpox virus might suggest that the latter could be the actual OPXV species involved in the outbreak. Cowpox viruses are considered endemic in Europe, 9 with rodents being their presumptive vectors. 1 Therefore, it is more than likely that wild mice and/or rats might have been the actual source of the virus. Although an OPXV infection in marmots is less surprising than that caused by CDV, this represents the first documented poxviral infection in this animal species.
In light of these results, preventive CDV vaccination protocols even in nonconventional hosts along with the adoption of novel strategies aiming to dissuade or prevent contact between animals in the collections with local wildlife, including small rodent populations, are warranted. Finally, our results emphasize once more the necessity of considering the possibility that multiple agents might be responsible for a single disease outbreak and that screening for multiple compatible agents, despite apparently initial unambiguous but partial clinical, pathological, and laboratory findings might provide very significant insights.
Footnotes
Acknowledgements
We thank Dr Marie Pierre Ryser-Degiorgis for her very precious “networking” support, Miss Maja Stodte for her much appreciated help with the editing of the article, Mr Pashk Selitaj for performing the immunohistochemistry technique, and all the personnel at FIWI, ITPA (Vetsuisse Faculty, University of Bern, CH), and Galli Valerio (Lausanne, CH) for their continuous help. The manuscript has been prepared according to the Uniform Requirements format.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
