Abstract
Liver tumors from a previous National Toxicology Program study were examined using global gene expression and mutation analysis to define the mechanisms of carcinogenesis in mice exposed to oxazepam. Five hepatocellular adenomas and 5 hepatocellular carcinomas from male B6C3F1 mice exposed to 5000 ppm oxazepam and 6 histologically normal liver samples from control animals were examined. One of the major findings in the study was upregulation of the Wnt/β-catenin signaling pathway. Genes that activate β-catenin, such as Sox4, were upregulated, whereas genes that inhibit Wnt signaling, such as APC and Crebbp, were downregulated. In addition, liver tumors from oxazepam-exposed mice displayed β-catenin mutations and increased protein expression of glutamine synthetase, a downstream target in the Wnt signaling pathway. Another important finding in this study was the altered expression of oxidative stress–related genes, specifically increased expression of cytochrome p450 genes, including Cyp1a2 and Cyp2b10, and decreased expression of genes that protect against oxidative stress, such as Sod2 and Cat. Increased oxidative stress was confirmed by measuring isoprostane expression using mass spectrometry. Furthermore, global gene expression identified altered expression of genes that are associated with epigenetic mechanisms of cancer. There was decreased expression of genes that are hypermethylated in human liver cancer, including tumor suppressors APC and Pten. Oxazepam-induced tumors also exhibited decreased expression of genes involved in DNA methylation (Crebbp, Dnmt3b) and histone modification (Sirt1). These data suggest that formation of hepatocellular adenomas and carcinomas in oxazepam-exposed mice involves alteration of the Wnt signaling pathway, oxidative stress, and potential epigenetic alterations.
Hepatocellular carcinoma (HCC) is the most common primary malignancy of the liver. HCC accounts for approximately 5% of all cancers worldwide and is the third most common cause of cancer death in humans. The worldwide incidence is estimated to be between 250,000 and 1.2 million new cases each year and the mortality between 500,000 and 1 million deaths annually. 7,9,19 The cause of human HCC is multifactorial and includes exposure to environmental carcinogens, chronic viral infection with hepatitis B or hepatitis C, metabolic liver disease, industrial chemicals, and pharmacologic agents. HCC occurs more frequently in men than in women, and more than 80% of HCC cases develop in association with cirrhosis or chronic hepatitis. 6,19
Although the cause of HCC is well known, the pathogenesis remains largely unknown and the prognosis is often very poor. HCC is an aggressive tumor and is commonly at an advanced stage when detected. Cure, usually through surgery, is possible in less than 5% of patients and the median survival time following diagnosis is 6 months. Given its aggressive clinical course and poor prognosis, there is a need to determine the molecular events linked to the development and progression of HCC. 19
Several major oncogenic pathways have been identified as dysregulated in HCC, including the p53, Rb, and Wnt pathways. Mutations in the Wnt/β-catenin signaling pathway appear to be the most frequent mutational event in human HCC and have been found in 26% to 40% of HCCs. 6,7,9,19
Oxazepam is a benzodiazepine that is used therapeutically as a sedative–hypnotic and anti-anxiety medication. The benzodiazepines are one of the most commonly used classes of drugs. One of the best known benzodiazepines is diazepam (Valium). Oxazepam is also a common metabolite of many other benzodiazepines including diazepam. 4,15,22,24 All benzodiazepines share a number of effects including sedation, hypnosis, decreased anxiety, muscle relaxation, amnesia, and anticonvulsant activity. They are considered central nervous system depressants but are not general depressants. 4,22,24
A study by the National Toxicology Program (NTP) investigated the carcinogenicity of oxazepam in B6C3F1 mice. The NTP study found a significant increase in the incidence of hepatocellular adenoma (HCA) and HCC in both male and female mice. 22 H-ras mutations, which are often found in spontaneous mouse liver tumors, were not observed in the liver tumors from oxazepam-exposed mice, suggesting that the tumors may have developed by pathways independent of ras signaling. 11
Studies have demonstrated that oxazepam is a nongenotoxic carcinogen. 22,24 Exposure to nongenotoxic carcinogens leads to effects in target cells, which can indirectly lead to neoplastic transformation or enhance the development of tumors from preinitiated cells. A diversity of mechanisms exist for carcinogenic nongenotoxic chemicals, including increased production of reactive oxygen species (ROS) and nitrogen species that damage DNA, lipids, and proteins; receptor-based mechanisms; and epigenetic mechanisms such as methylation of tumor suppressor genes. 25 For example, extensive methylation is often also present in the subset of human HCCs that harbor β-catenin mutations. 20 In addition, both genotoxic and nongenotoxic hepatocarcinogens have been shown to result in rapid alterations in the rodent epigenome that result in disrupted liver homeostasis, including altered cell proliferation, apoptosis, activation state of repetitive DNA sequences, loss of genomic and chromosomal stability, and aberrant methylation of DNA. 25
Previous research has demonstrated that tumors from oxazepam-exposed mice have a high incidence of β-catenin mutations, suggesting the involvement of the Wnt signaling pathway. 10 β-catenin mutations have also been frequently reported in human HCC. 6,9 Research has suggested that β-catenin mutations in mouse liver tumors may be chemical specific and an early event in mouse hepatocellular carcinogenesis. 10 The purpose of the current study was to characterize the major genetic and epigenetic pathways involved in hepatocellular neoplasms from oxazepam-exposed mice.
Materials and Methods
Selection of Samples
In the NTP study, 6-week-old male and female B6C3F1 mice were exposed to 0 or 5000 ppm oxazepam in feed for 104 to 105 weeks. 21 A portion of each liver tumor was fixed in 10% neutral buffered formalin, processed, embedded in paraffin, and cut into 5-μm sections for hematoxylin and eosin staining and routine histological examination. The remaining portion of each liver tumor was frozen in liquid nitrogen and stored at –80°C for molecular biology studies. The NTP histological criteria for identifying adenomas and carcinomas was adapted from the textbook Pathology of the Mouse. 16 An adenoma was characterized as a well-demarcated mass composed of cords of fairly well-differentiated hepatocytes 1 to 2 cells thick and lacking normal lobular architecture. The mass typically encompasses an area greater than 1 liver lobule and compresses the adjacent liver parenchyma. Adenomas have minimal mitotic figures and cellular atypia. A carcinoma was characterized as an expansive to infiltrative, poorly demarcated mass composed of hepatic cords greater than 3 hepatocytes in thickness. The mass displays a loss of normal lobular architecture and variable growth patterns with cells exhibiting a high mitotic rate, significant cellular atypia, and enlarged hyperchromatic nuclei. The observed variable growth patterns include trabecular, solid, and pseudoglandular. Trabecular variants are composed of thickened cords, sheets, and trabeculae greater than 3 cell layers thick, often separated by dilated vascular spaces. The solid variant is composed of sheets of neoplastic hepatocytes exhibiting nuclear pleomorphism and cellular atypia. The pseudoglandular form is characterized by a central luminal space lined by 1 to multiple layers of neoplastic hepatocytes. 16 The criteria for selecting frozen tissues for the microarray study included absence of inflammatory cell infiltration and/or necrosis in liver tumors from oxazepam-treated mice and normal liver tissues from untreated mice. Five HCAs and 5 HCCs from oxazepam-treated mice were selected for microarray analysis along with 6 histologically normal livers from untreated mice.
DNA Isolation, PCR Amplification, and Autosequencing for β-Catenin Gene
DNA was isolated and extracted from frozen normal liver and HCAs and HCCs in oxazepam-exposed animals with the DNeasy Tissue Kit (Qiagen, Valencia, CA). Amplification reactions were carried out using seminested polymerase chain reaction (PCR) using the primer sets for exon 2 (corresponding to exon 3 in humans) of the mouse β-catenin gene (BCAT-1F, 5′- TACAGGTAGCATTTTCAGTTCAC -3′; BCAT-2R, 5′- TAGCTTCCAAACACAAATGC -3′; BCAT-8R, 5′- ACATCTTCTTCCTCAGGGTTG -3′). Controls lacking DNA were run with all sets of reactions. PCR products were purified using a QIAquick Gel Extraction Kit (Qiagen, Valencia, CA). The purified PCR products were cycled with Terminal Ready Reaction Mix-Big Dye (Perkin Elmer, Foster City, CA), and the extension products were purified using the DyeEx 2.0 Spin Kit (Qiagen). The lyophilized PCR products were sequenced with an automatic sequencer (Perkin-Elmer ABI model 3100). The automated ABI DNA sequencing system detects fluorescence from different dyes that are used to identify the A, C, G, and T extension of the sequence reaction. Each sequence generates a 4-color chromatogram showing the analyzed data from which the machine determines the nucleotide sequence. An electropherogram was used for comparison between the normal control and the treated groups.
Microarray Analysis
Affymetrix microarray technology was used to examine changes in gene expression in liver tumors from oxazepam-exposed animals. RNA was collected from normal liver and liver tumors by digestion with Trizol (Invitrogen, Carlsbad, CA) and subsequent extraction with chloroform and isoamyl alcohol. RNA was further purified using the RNeasy Mini Kit (Qiagen, Valencia, CA). Quality of RNA was assessed by gel electrophoresis.
Gene expression analysis was conducted using Affymetrix Mouse Genome 430 2.0 GeneChip arrays (Mouse 430 v2, Affymetrix, Santa Clara, CA). RNA quality was determined by Bioanalyzer analysis (Agilent Technologies, Santa Clara, CA). Total RNA (1 μg) was amplified using the Affymetrix One-Cycle cDNA Synthesis protocol. For each array, 15 μg of amplified biotin cRNAs were fragmented and hybridized to the array for 16 hours at 45°C in a rotating hybridization oven using the Affymetrix Eukaryotic Target Hybridization Controls and protocol. Slides were stained with streptavidin/phycoerythrin using a double-antibody staining procedure and washed using the EukGE-WS2v5 protocol of the Affymetrix Fluidics Station FS450 for antibody amplification. Arrays were scanned with an Affymetrix Scanner 3000 and data obtained using the GeneChip Operating Software (GCOS; version 1.4.0.036). The resulting files (.dat, .cel, and .chp) were imported into the Rosetta Resolver system (version 7.0), which performed data preprocessing, normalization, and error modeling. 29
Processed microarray data were analyzed with a variety of unsupervised and supervised techniques. Principal component analysis (PCA) was performed on all samples and all probe sets to characterize the variability present in the data. To identify differentially expressed genes, a log base 10 error-weighted analysis of variance (ANOVA) with multiple test correction (Bonferroni, P < .01) was performed using Rosetta Resolver (www.rosettabio.com) to reduce the number of false positives. The ANOVA analysis yielded 1,934 significant probes. Since there were 3 groups in the data set (normal, adenoma, and carcinoma), ANOVA analysis was followed by a post hoc analysis (Tukey-Kramer, P < .01) to perform pairwise comparisons and determine which pairs were significantly different from one another.
Cluster analysis was performed to determine the intensity values (z-score) for each replicate from a treatment group. This method was used to visualize variation among the replicates within a specific treatment group and determine how the groups were different from one another. Cluster analysis places all the values on the same scale by centering the data on 0. Values greater than the baseline have a positive z-score, whereas values less than the baseline have a negative z-score.
Real-Time PCR
Real-time PCR was performed with 11 genes (Mat1a, Ctnnbip1, Zhx2, Rras2, Glul, Dhcr7, Hras1, Sirt1, Cat, Cyp1a2, and Sod2) that are known to play important roles in tumorigenesis and oxidative stress and which were significantly altered in the tumors by microarray analysis. Quantitative gene expression levels were detected using real-time PCR with the ABI PRISM 7900HT Sequence Detection System (Applied Biosystems, Foster City, CA) and TaqMan MGB probes (FAM dye labeled). Primers and probes for all genes analyzed were purchased from Applied Biosystems Assays-on-Demand Gene Expression products (Mat1a: Assay ID# Mm00522563_m1; Ctnnbip1: Assay ID# Mm01257821_m1; Zhx2: Assay ID# Mm01207642_m1; Glul: Assay ID# Mm00725701_s1; Rras2: Assay ID# Mm00491838_m1; Dhcr7: Assay ID# Mm00514571_m1; Hras1: Assay ID# Mm00476174_m1; Sirt1: Assay ID# Mm01168521_m1; Cat: Assay ID# Mm00437992_m1; Cyp1a2: Assay ID# Mm00487248_g1; Sod2: Assay ID# Mm00449726_m1). For amplification, diluted complementary DNA (cDNA) was combined with a reaction mixture containing TaqMan universal PCR Master Mix (Applied Biosystems, Catalog No. 4304437) according to the manufacturer’s instructions. Samples were analyzed in duplicate and a sample without reverse transcriptase was included in each plate to detect contamination by genomic DNA. Amplification was carried out as follows: (1) 50°C for 2 minutes (for uracil-N-glycosylase incubation); (2) 95°C for 10 minutes (denaturation); (3) 95°C for 15 seconds and 60°C for 30 seconds (denaturation and amplification) for 40 cycles. Fold increases or decreases in gene expression were determined by quantitation of cDNA from tumor samples relative to a pool of normal liver samples. The 18S RNA gene was used as the endogenous control for normalization of initial RNA levels. To determine this normalized value, 2–(ΔΔCt) values were compared between tumors and normal liver, where the changes in crossing threshold (ΔCt) = Ct Target gene – Ct18S RNA, and ΔCt = ΔCtno rmal – ΔCttumo r.
GC/MS Analysis of Isoprostane Expression
Six-week-old male B6C3F1 mice were administered either 0 or 2500 ppm oxazepam ad libitum in the diet for 6 months. Liver samples were harvested and sample extraction and derivatization were performed as previously described. 23 Gas chromatography/negative ion chemical ionization/mass spectrometry (GC/NICI/MS) was performed on a Hewlett-Packard model 5890 GC coupled to a VG Trio 1 mass spectrometer (VG/Fisons, VG Masslab, Manchester, UK). An on-column injection of 3–5 μL was performed onto a 5-m length of deactivated fused silica of ID 0.53 mm, which was coupled to the analytical column. The chromatographic column was an SPB-5 (Sigma-Aldrich, St. Louis, MO) at a length of 25 m, ID of 0.32 mm, and film thickness of 0.25 μm. The GC oven was held at 250°C for 1 minute, ramped to 300°C at a rate of 10°C/min, and held for 15 minutes. Methane was used as the chemical ionization gas. Selected ion monitoring of m/z 569.5 and m/z 573.5 was used for detection of isoprostane and the d4 internal standard. Comparison of the means for isoprostane levels between control and oxazepam-exposed animals was performed using the Student’s t-test. Statistical significance was set at P < .05.
Immunohistochemical Staining
Immunohistochemical staining was performed on selected formalin-fixed paraffin-embedded sections of normal liver from untreated controls and HCAs and HCCs from oxazepam-exposed animals. After deparaffinization, tissue sections were incubated with prediluted normal serum for 20 minutes at room temperature (Jackson ImmunoResearch Laboratories Inc., West Grove, PA). Endogenous peroxidase activity was quenched in 0.6% H2O2 in 60% methanol for 30 minutes, and antigen retrieval was performed using a decloaking chamber (0.1 M citrate buffer pH 6.0). After antigen retrieval, endogenous biotin was masked by incubation with Vector Avidin/Biotin Block (Vector Laboratories, Burlingame, CA). Sections were incubated with the primary antibody at the following dilutions: rabbit anti-glutamine synthetase; 1:10,000 (ab49873; Abcam, Cambridge, MA) and mouse anti-Dnmt3a; 1:25 (IMG-268A; Imgenex, San Diego, CA) for 60 minutes at room temperature. Normal serum was applied to the negative slides for an equivalent incubation period. After washing, sections were incubated with horseradish peroxidase (HRP)–conjugated IgG for 30 minutes at room temperature. Immunoreactivity was detected with 3′ diaminobenzidine tetrahydrochloride (DakoCytomation Liquid DAB Substrate Chromogen System; DakoCytomation, Carpentaria, CA). Sections were counterstained with hematoxylin, dehydrated in graded ethanol, and coverslipped.
Results
Microarray Analysis: ANOVA and Post Hoc Tukey-Kramer
Using genome-wide Affymetrix microarrays, changes in gene expression were examined between HCAs and HCCs from oxazepam-exposed mice and normal liver from control mice. ANOVA was used to identify differentially expressed genes. The ANOVA analysis yielded 1,934 significantly altered probes. ANOVA and post hoc analysis identified 1,513 probe sets that were differentially expressed between normal liver and HCA and 1,976 probe sets that differed between normal liver and HCC. Additionally, 269 probe sets were differentially expressed between HCAs and HCCs (Figure 1 ; Supplemental Table 1). These probe sets represent genes involved in cellular processes such as growth and proliferation, adhesion, motility, metastasis, apoptosis, DNA replication, transcription, translation, and oxidation reduction and in signaling pathways such as the mitogen activated protein kinase (MAPK) pathway and the Wnt pathway.
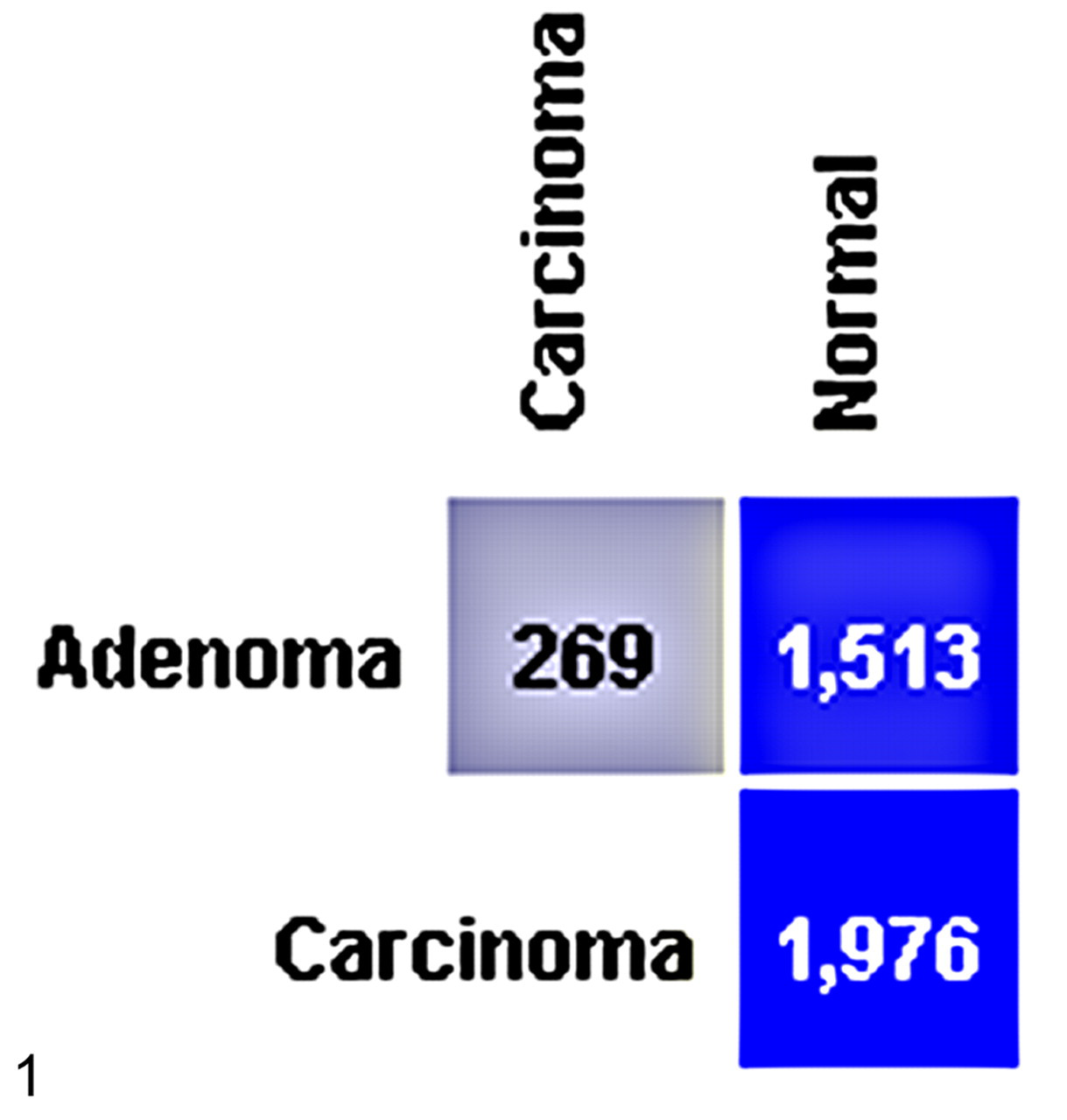
ANOVA was used to test for differential gene expression across the 3 groups (control, adenoma, and carcinoma) and was followed by post hoc analysis. ANOVA and post hoc Tukey-Kramer at P < .01 identified the differential expression of 1,513 probe sets between normal liver and hepatocellular adenoma, 1,976 probe sets between normal liver and carcinoma, and 269 probe sets between adenoma and carcinoma.
Summary of β-Catenin Mutations in Mouse Hepatocellular Neoplasms Following Oxazepam Exposure
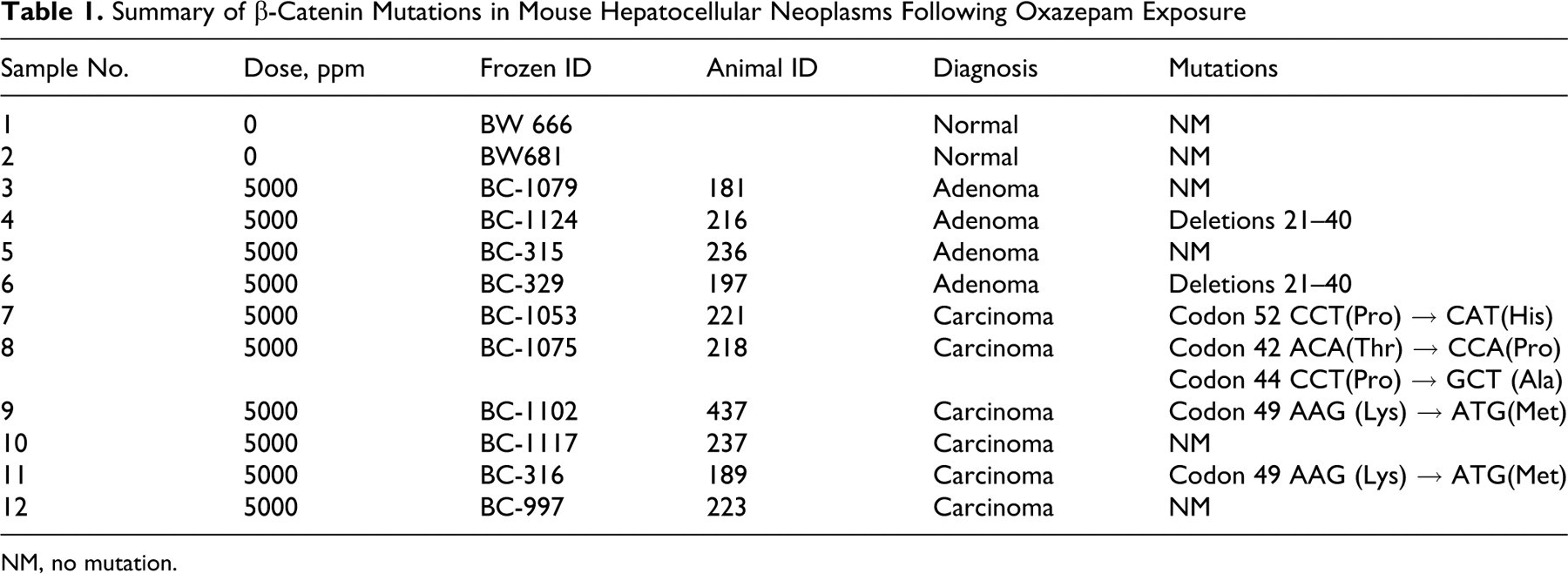
NM, no mutation.
A limitation of the current study is the lack of inclusion of normal adjacent liver tissue from oxazepam-treated mice. The inclusion of these samples would have allowed further studies examining the effects of chronic oxazepam exposure.
Wnt Signaling
Previous research has demonstrated an increased incidence of β-catenin mutations in liver tumors from oxazepam-exposed mice and suggested a role for the Wnt signaling pathway in oxazepam-induced liver tumorigenesis. 10 Consistent with the previous results, we found that 6 of the 10 liver tumors (60%) in oxazepam-exposed mice had mutations in β-catenin (Table 1 and Figure 2).
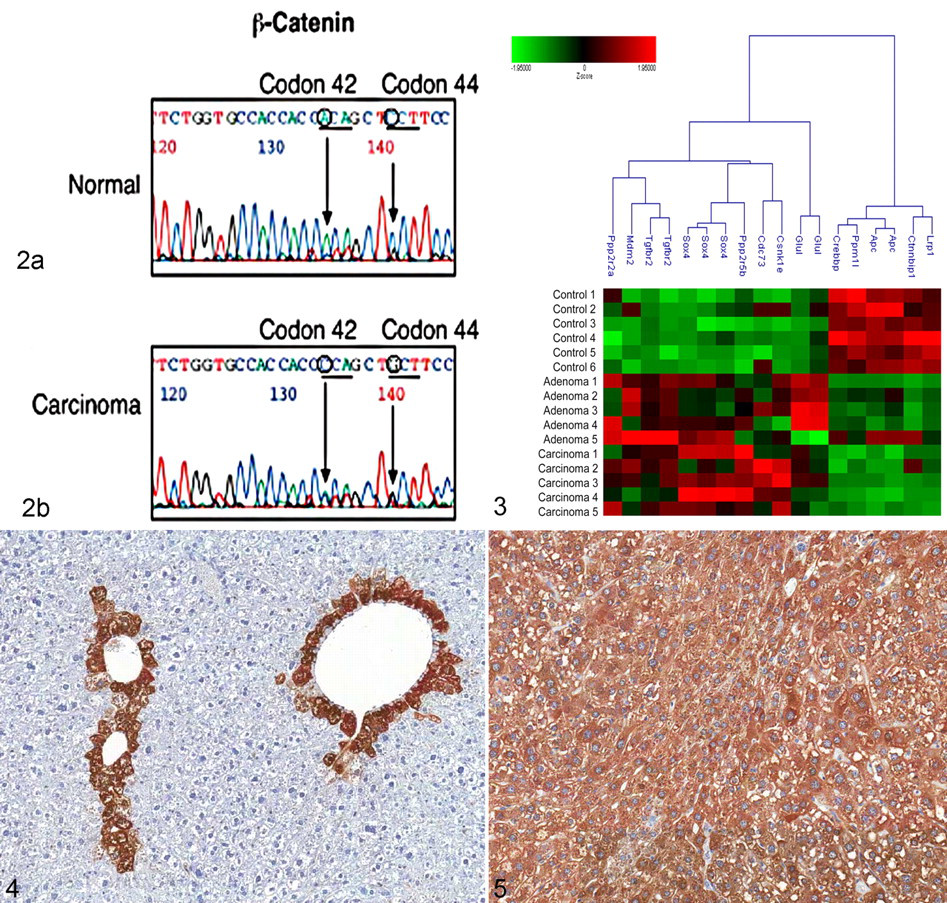
Autosequencing of exon 2 of β-catenin in liver tumors from oxazepam-exposed mice. (a) DNA sequence from normal frozen liver (sample No. 1) and (b) DNA sequence from oxazepam-induced HCC (sample No. 8). The mutations in codon 42 ACA → CCA and codon 44 CCT → GCT are indicated.
Ingenuity pathway analysis was used to identify Wnt signaling genes that were altered in the microarray analysis. Figure 3 is a cluster analysis of these genes. There was a clear difference in the expression of Wnt signaling pathway genes between normal liver and oxazepam-induced liver tumors. There was an increased expression of genes that activate β-catenin (Sox4) and decreased expression of genes that inhibit Wnt signaling (Lrp1, APC, and Crebbp).
To identify downstream signaling of Wnt, we examined the expression of glutamine synthetase, a downstream target in the Wnt signaling pathway. Glutamine synthetase is strongly upregulated in mouse liver tumors that contain mutations in β-catenin and has been used as a marker for their identification. 1,5 By immunohistochemistry, normal liver displays a centrilobular staining pattern for glutamine synthetase whereas tumors display a homogeneous staining pattern (sample No. 4) (Figures 4 and 5). An increased expression of glutamine synthetase was observed in 10 of the 10 tumors (100%), with no obvious difference between the staining in adenomas and carcinomas (data not shown). We also used immunohistochemical staining to examine the expression of β-catenin protein in the tumors. Five of the 5 adenomas (100%) and 4 of the 5 carcinomas (80%) from oxazepam-exposed mice displayed membrane and cytoplasmic localization of β-catenin, whereas normal liver contained only membrane localization (data not shown). Nuclear localization of β-catenin was not detected in any of the samples. Nine of the 10 samples (90%) displayed both increased expression of glutamine synthetase and cytoplasmic and membrane localization of β-catenin.
Oxidative Stress
It has been suggested that oxidative stress is involved in the formation of liver neoplasms. 17,26 Isoprostanes are prostaglandin-like compounds that are formed in vivo from the free radical–catalyzed peroxidation of essential fatty acids. They are accurate markers of oxidative stress–induced lipid peroxidation in both humans and animals. 18 Using mass spectrometry, we determined that mice exposed to oxazepam for 6 months showed a 1.5-fold increase (P = .026) in isoprostane expression in their livers compared with control mice (Figure 6), consistent with increased oxidative stress.
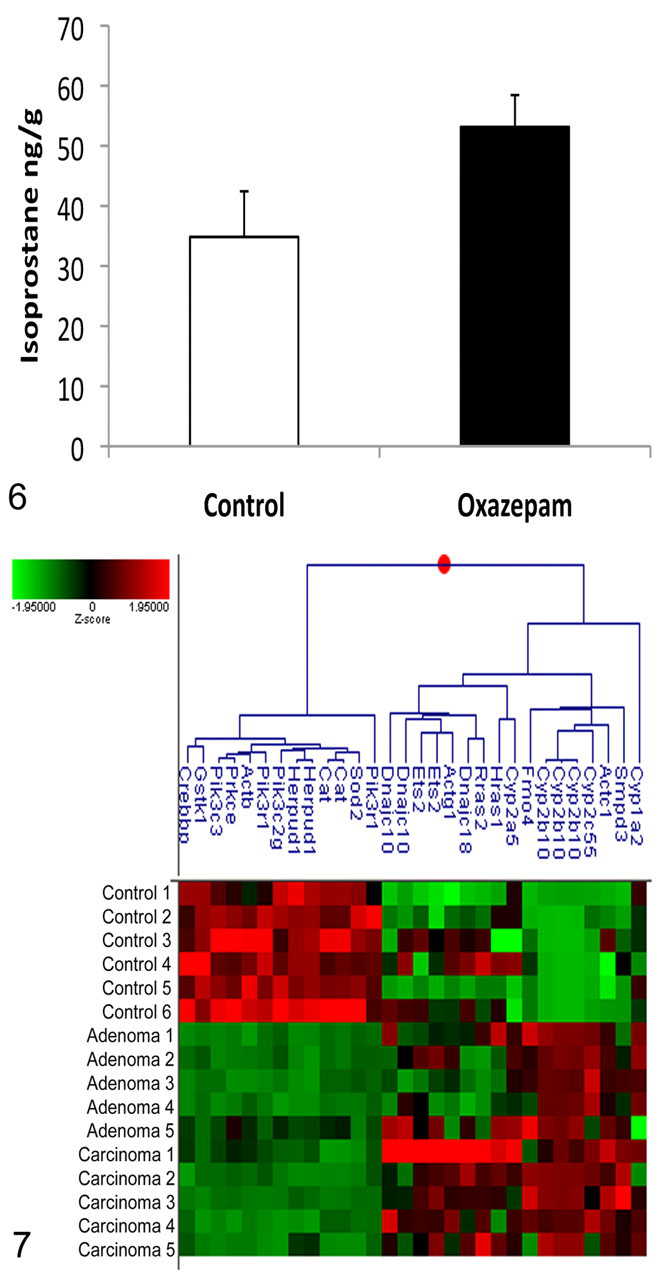
Expression of the oxidative stress marker, isoprostane, in the livers of B6C3F1 mice following 6 months of oxazepam exposure (n = 3). Oxazepam-exposed mice displayed a 1.5-fold increase in isoprostane as measured by mass spectrometry (P = .026).
Ingenuity pathway analysis was used to identify other genes related to oxidative stress. Cluster analysis of the identified genes showed a clear difference in expression between normal liver and liver tumors from oxazepam-exposed mice (Figure 7). The tumors had increased expression of cytochrome p450 genes including Cyp1a2, Cyp2c55, Cyp2b10, and Cyp2a5, whose induction has been associated with oxidative stress, and decreased expression of genes that protect against oxidative stress including Sod2, Cat, and Gstk1.
Epigenetic Mechanisms
Since oxazepam is a nongenotoxic chemical and epigenetic mechanisms including methylation are known to be important in the formation of liver tumors, 25 we examined liver tumors from oxazepam-exposed mice for epigenetic changes important in human liver cancer. Potentially methylated genes were identified using MeInfoText and cluster analysis of the identified genes was performed (Figure 8). We found decreased expression of tumor suppressors such as APC and Pten, consistent with hypermethylation, and increased expression of the oncogene H-ras, consistent with hypomethylation.
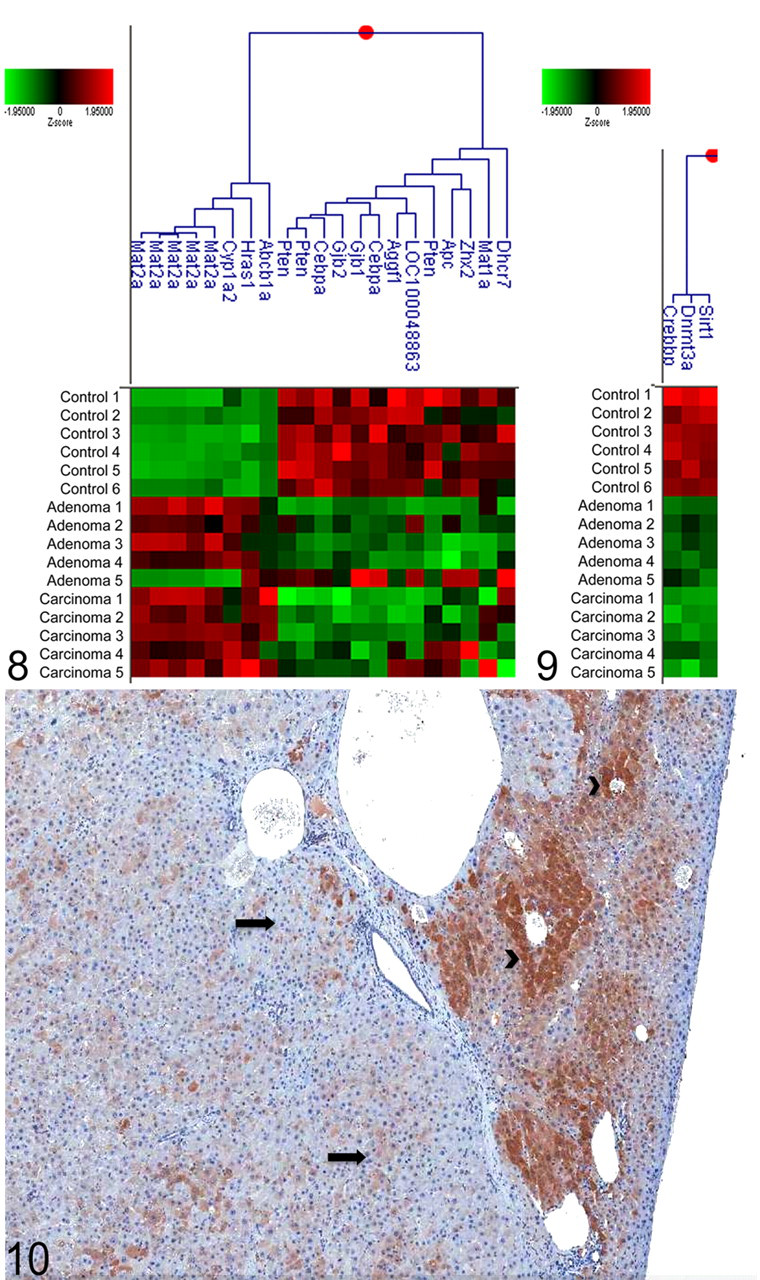
Cluster analysis depicting the log base 10 z score of genes that are regulated by methylation identified as significantly altered (P < .01) in hepatocellular adenomas and carcinomas from oxazepam-exposed mice.
In addition, by cluster analysis, we examined the expression of DNA methyltransferases (Dnmts) and histone modifiers (Figure 9). Decreased expression of the DNA methyltransferase Dnmt3a and decreased expression of 2 genes involved in histone modification, Sirt1 and Crebbp, were detected in both adenomas and carcinomas from oxazepam-exposed mice. By immunohistochemistry, there was a marked decrease in the expression of Dnmt3a in the oxazepam-induced adenomas and carcinomas compared with the normal liver (sample No. 8) (Figure 10), confirming that the decreased RNA expression translated to a decrease in protein expression.
Validation of Microarray Analysis (Real-Time PCR)
To validate the microarray results, the expression of a subset of genes was examined by TaqMan qPCR. These genes were identified by microarray as dysregulated in tumors from oxazepam-exposed mice. The expression of Mat1a, Ctnnbip1, Zhx2, Rras2, Glul, Dhcr7, Hras1, Sirt1, Cat, Cyp1a2, and Sod2 as determined by real-time PCR correlated well with the microarray results (Figure 11).
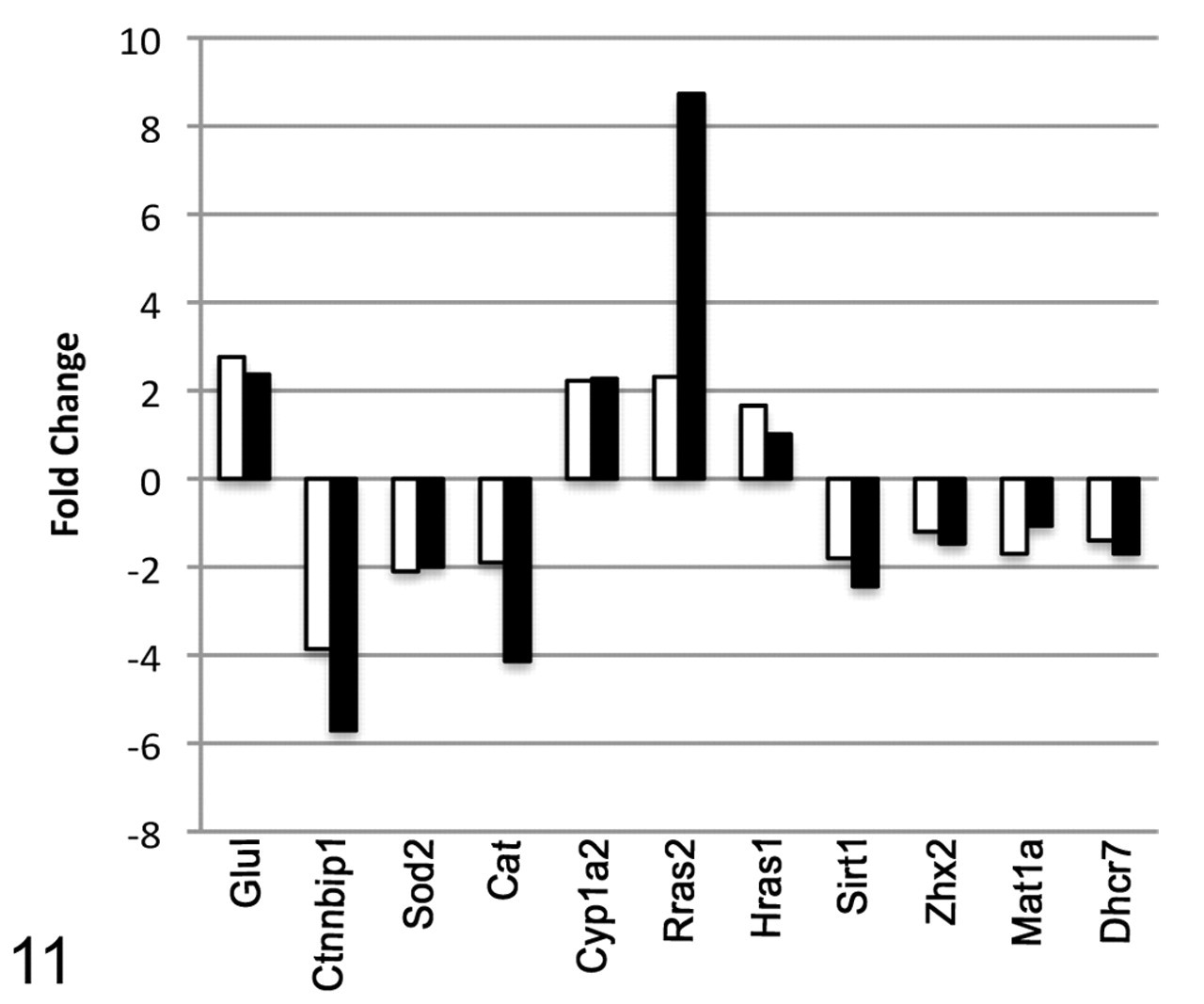
Validation of microarray data using real-time PCR analysis. The expression of Mat1a, Ctnnbip1, Zhx2, Rras2, Glul, Dhcr7, Hras1, Sirt1, Cat, Cyp1a2, and Sod2 was examined in carcinomas from oxazepam-exposed mice by real-time PCR and compared with the results of the microarray analysis. White bars = microarray; black bars = real time PCR.
Discussion
The current study demonstrated that the formation of HCAs and HCCs in oxazepam-exposed mice occurs through mechanisms involving alteration of the Wnt signaling pathway, through oxidative stress, and through epigenetic changes. Importantly, the genes and signaling pathways identified in this study play important roles in human liver carcinogenesis.
The altered expression of genes that are involved in or regulated by the Wnt signaling pathway was identified in liver tumors from oxazepam-exposed mice compared with normal liver using microarray analysis. For example, the Wnt signaling antagonists APC and Crebbp were decreased in adenomas and carcinomas compared with normal liver. Also, the expression of genes known to act as agonists of Wnt signaling was increased in HCC. Csnk1e, which can act as both a positive and a negative regulator of Wnt signaling, was increased in oxazepam-induced HCC along with the transcription factor Sox4, which can interact directly with β-catenin or TCF/LEF family members to enhance Wnt signaling. Five of the 13 Wnt signaling genes (38%) demonstrated significant differential gene expression in adenomas compared with carcinomas, with expression being higher in the carcinomas. The increased gene expression in the carcinomas suggests that progression to a malignant phenotype involves additional alterations leading to severe dysregulation of the Wnt signaling pathway.
In addition to noting altered expression of Wnt signaling genes, we detected mutations in β-catenin. Sequencing of β-catenin exon 2 revealed mutations in 6 of the 10 liver tumors (60%) from oxazepam-exposed mice. No clear mutation signature was detected in liver tumors from oxazepam-exposed mice, which is in consistent with that of nongenotoxic chemicals. 10 We also found increased protein expression of a marker of β-catenin activation, glutamine sythetase, in 10 of the 10 adenomas and carcinomas (100%) as well as cytoplasmic expression of β-catenin in 9 of the 10 tumors (90%). The mutations in β-catenin and the altered expression of glutamine synthetase were detected equally in adenomas and carcinomas, suggesting that alterations in β-catenin may occur early in liver carcinogenesis.
Approximately 40% of human HCCs have mutations in β-catenin. 6,9 Since β-catenin mutations are important in the pathogenesis of human HCC and since chemically induced tumors in mice such as those induced by oxazepam have mutations similar to those observed in humans, 10 the gene expression profile of oxazepam-induced HCC was further compared with human HCC with β-catenin mutation to determine whether the pathways altered are similar between mouse and human and whether HCC in oxazepam-exposed mice is a good model of human HCC. 27 Thirty-three percent of upregulated genes in human HCC with β-catenin mutation were also upregulated in HCC in oxazepam-exposed mice (Supplemental Table 2). These included genes involved in cell signaling, cell growth, and proliferation (GLUL, MME, FABP4, CSRP2, and ESM1). Conversely, 47% of downregulated genes in human HCC with β-catenin mutation were also downregulated in HCC from oxazepam-exposed mice, including the complement factors C9 and CFHR1, the signaling molecule IGF-1, and genes involved in metabolic processes (SLC3A1, ARG1, and NNMT) 27 (Supplemental Table 2). This indicates that similar pathways are involved in HCC resulting from β-catenin mutation in mice and humans.
Another pathway that was determined to be important in liver tumors from oxazepam-exposed mice was the oxidative stress pathway. Oxidative stress entails an excessive production of ROS and/or a reduction in antioxidant defenses. Oxidative stress can affect a number of cellular processes involved in tumor development including cell proliferation, senescence, inflammation, and metastasis. Unregulated or prolonged ROS production has been linked to mutation as well as modification of gene expression through both genetic and epigenetic mechanisms. 13,26 We found increased expression of isoprostane, a marker of oxidative stress that is associated with increased lipid peroxidation, in oxazepam-exposed mice compared with controls.
Our gene expression analysis also supports oxidative stress as a mechanism important in liver tumors from oxazepam-exposed mice. The oxazepam-induced adenomas and carcinomas had strong upregulation of the cytochrome P450 genes, Cyp2a5, Cyp2b10, Cyp2c55, and Cyp1a2. Upregulation of cytochrome p450s increases the production of ROS as a byproduct of microsomal oxidation 30 and uncoupling of the P450 catalytic cycle. Metabolism of phenobarbital by cytochrome P450 enzymes is known to result in uncoupling of the P450 catalytic cycle and the release of superoxide anion. 17 Studies have suggested that oxazepam is a phenobarbital-type inducer of cytochrome P450 enzymes in the mouse liver and that the early liver responses in oxazepam-exposed mice are similar to those observed in phenobarbital-exposed mice. 8,14,15 This suggests that the increased oxidative stress observed in oxazepam-exposed mice may be due to uncoupling of the P450 catalytic cycle during its metabolism. Furthermore, there was increased expression of genes induced by oxidative stress (Ets2 and Smpd3) and decreased expression of other genes that protect against oxidative stress (Gstk1, Cat, and Sod2). The microarray analysis suggests that oxidative stress is an important mechanism in the formation of adenomas and carcinomas in oxazepam-exposed animals. Four of the 21 oxidative stress genes (19%) identified in the cluster analysis were significantly different between adenomas and carcinomas, with expression higher in the carcinomas.
In addition to genetic alterations, epigenetic alterations, most commonly aberrant methylation of CpG dinucleotide sequences in the promotor regions of cancer-related genes, are a common feature of HCCs. 2,3,25 Altered DNA methylation has previously been found in the livers of mice exposed to nongenotoxic hepatocarcinogens. 2,3,25 Consistent with this, genes known to be regulated by methylation in human liver cancer were also altered in liver tumors from oxazepam-exposed mice. In oxazepam-exposed mice, there was increased expression of genes known to be hypomethylated in human HCC (Mat2a, Cyp1a2, and the oncogene Hras1) and decreased expression of tumor suppressors (Pten and APC) and other genes known to be hypermethylated in human HCC (Cebpa, Gjb2, Gjb1, Aggf1, Zhx2, Mat1a, and Dhcr7). The expression of potentially methylated genes was similar between adenomas and carcinomas.
Our results further demonstrated decreased expression of a gene involved in the regulation of epigenetic mechanisms, Sirt1. Sirt1 is a histone deacetylase that has been found to be a negative regulator of β-catenin activity 12 and has been shown to play a role in human HCC. 28 Human HCC exhibits reduced Sirt1 protein and messenger RNA (mRNA) expression levels compared with normal liver. The observation that Sirt1 levels are reduced in many human cancers provides strong evidence that SIRT1 may function as a tumor suppressor. 28 Tumors from oxazepam-exposed animals demonstrated reduced mRNA expression of Sirt1, suggesting a similar signaling mechanism in human and mouse HCC.
Our studies demonstrate that animal models of liver cancer can be useful in providing clues to both genetic and epigenetic factors that may contribute to the development of human disease. We were also able to show that the gene expression profile of oxazepam-induced liver tumors was linked to the Wnt signaling pathway and increased oxidative stress. The significance of these findings is that many of the genes with altered expression in the mouse tumor model represent genes and signaling pathways that may play a role in human liver cancer.
Footnotes
Acknowledgements
We thank Theodora Devereux, Carol Parker, and LeRae Graham for their help with isoprostane measurement.
The authors declared that they had no conflicts of interests with respect to their authorship or the publication of this article.
This research was supported in part by the Intramural Research Program of the National Institute of Environmental Health Sciences/National Institutes of Health.
