Abstract
Objective
To explore the potential involvement of PANoptosis-related genes in gastric cancer susceptibility through multiomics analyses.
Methods
Summary-data–based Mendelian randomization was performed by integrating blood-derived methylation, gene expression, and protein quantitative trait loci data with genome-wide association study results. The findings were further evaluated in The Cancer Genome Atlas cohort, followed by protein–protein interaction analysis, drug prediction, and molecular docking.
Results
Summary-data–based Mendelian randomization and colocalization analyses identified several traits suggestively associated with gastric cancer risk. Genetically predicted higher expression of apoptosis and caspase activation inhibitor (AVEN) and hepatocyte growth factor (HGF) as well as higher HGF protein levels were associated with increased risk, whereas higher levels of protein phosphatase 2 regulatory subunit B beta (PPP2R2B) appeared to be protective. Multiomics integration suggested epigenetic regulation of HGF and PPP2R2B. The Cancer Genome Atlas analysis corroborated the dysregulation of these candidates, with high AVEN expression associated with poorer survival. Protein–protein interaction and drug prediction analyses highlighted functional networks and potential therapeutics, supported by molecular docking demonstrating strong HGF-binding affinities. However, these associations did not reach statistical significance in the independent validation cohort, possibly due to limited statistical power.
Conclusions
This study identified AVEN, HGF, and PPP2R2B as potential candidate genes for gastric cancer. These findings require further validation in larger cohorts.
Keywords
Introduction
Gastric cancer (GC) ranks as the fifth most common cancer worldwide and the fourth leading cause of cancer-related mortality. 1 Despite advances in therapy, the prognosis of GC remains poor, particularly due to late-stage diagnosis. Multiple risk factors contribute to the development of GC, including Helicobacter pylori infection, smoking, alcohol consumption, dietary habits, family history, and certain genetic predispositions. 2 GC is a heterogeneous disease comprising several molecular subtypes classified by The Cancer Genome Atlas (TCGA) into Epstein–Barr virus (EBV)-positive, microsatellite instability (MSI), genomically stable (GS), and chromosomal instability (CIN) subtypes, each with distinct molecular and clinical features. 3 A primary challenge in GC management is the lack of effective biomarkers for early detection. 4 Currently, surgical resection remains the mainstay of treatment for GC. However, in metastatic cases, surgery alone rarely achieves a cure, and the efficacy of chemotherapy and radiotherapy remains limited.5,6 Therefore, further evidence is urgently needed to facilitate the development of early diagnostic biomarkers and novel therapeutic targets for GC.
Resistance to cell death is a defining feature of tumor cells. To evade apoptosis, tumor cells often regulate anti-apoptotic and pro-apoptotic proteins and activate survival signaling pathways. These adaptations limit not only apoptosis7,8 but also pyroptosis9,10 and necroptosis. 11 However, targeting a single cell death pathway has shown limited efficacy in suppressing GC. 12 Recently, PANoptosis, a unique inflammatory programmed cell death pathway that integrates mechanisms of pyroptosis, apoptosis, and necroptosis, has garnered increasing attention.13,14 Emerging evidence links PANoptosis and its related genes to the pathogenesis, prognosis, and immune landscape of various cancers, including GC.15,16 Bioinformatics studies suggest that PANoptosis-related gene expression signatures can reflect tumor microenvironment characteristics, predict patient prognosis, and inform immunotherapy responsiveness in GC. However, the relationship between PANoptosis and GC susceptibility and risk remains largely unexplored. To develop new diagnostic biomarkers and therapeutic strategies, it is essential to identify PANoptosis-related genes associated with GC risk.
Mendelian randomization (MR) is a powerful approach that leverages genetic variants as instrumental variables to assess the relationship between risk factors and disease outcomes. 17 Summary-data–based Mendelian randomization (SMR) integrates genome-wide association study (GWAS) data with molecular quantitative trait locus (QTL) data, including methylation QTL (mQTL), gene expression QTL (eQTL), and protein QTL (pQTL), thereby enabling the identification of candidate target genes linked to risk variants. 18 This approach allows exploratory detection of genetic variants associated with PANoptosis-related gene expression, methylation, or protein levels that may contribute to GC risk.
Therefore, this study utilized SMR to integrate GWAS and QTL data to systematically screen for PANoptosis-related genes associated with GC risk. Furthermore, we aimed to validate the expression patterns and prognostic significance of these genes using data from TCGA. Our research provides preliminary evidence and new insights into the complex genetic landscape of GC, offering valuable clues for the future development of biomarkers and targeted therapies.
Materials and methods
Study design
This study was designed to explore potential associations between PANoptosis-related genes and GC risk (Figure 1). In brief, we first performed SMR and colocalization analyses on 930 PANoptosis-related genes using blood-based mQTL, eQTL, and pQTL data integrated with GC GWAS data from the FinnGen (discovery) and UKB_151 (replication) cohorts. Subsequently, SMR-identified candidate genes were subjected to a series of downstream analyses: (a) assessment of differential expression and prognostic associations in the TCGA stomach adenocarcinoma (STAD) cohort; (b) tissue-specific SMR analysis using genotype-tissue expression (GTEx) stomach tissue data; (c) construction of a protein–protein interaction (PPI) network to explore functional connections; (d) prediction of potential small-molecule drugs; and (e) molecular docking simulations to evaluate the binding affinity of predicted small molecules to key candidate proteins. Furthermore, there was no sample overlap between the exposure and outcome cohorts, ensuring independent datasets and minimizing potential bias.

Schematic workflow of the study design and analysis pipeline. A graphical overview illustrating the multistep approach of this study: (a) identification of PANoptosis-related genes; (b) integration of mQTL, eQTL, and pQTL data with GWAS data from the FinnGen and UKB-151 cohorts using SMR and colocalization analyses; and (c) validation and functional analyses, including gene expression profiling, survival analysis, protein–protein interaction network construction, drug prediction, and molecular docking simulations. GWAS: genome-wide association study; SMR: summary-data–based Mendelian randomization; mQTL: methylation quantitative trait locus; eQTL: gene expression quantitative trait locus; pQTL: protein quantitative trait locus.
All analyses were performed in October 2024. The FinnGen R10 dataset (used as the primary GWAS discovery cohort) was the most recent version available at the time (FinnGen R12 was released after completion of our analysis). The UKB-151 replication dataset has not been updated since our analysis.
Data resources
In this study, 930 PANoptosis-related genes were extracted from Yang et al. 19 for comprehensive analysis. The discovery dataset for the GC GWAS was obtained from the FinnGen database (FinnGen R10_C13_STOMACH_EXALLC), comprising 1423 cases and 314,193 controls. For validation, we used additional GC-related GWAS data from UKB_151 based on European participants, consisting of 568 cases and 391,483 controls. Detailed information is presented in Supplementary Table 1.
QTL data were primarily derived from European populations. Blood eQTL summary statistics were sourced from eQTLGen, 20 which provided gene expression data from 31,684 individuals. Blood mQTL summary statistics were obtained from a meta-analysis of two European cohorts: the Brisbane Systems Genetics Study (n = 614) and the Lothian Birth Cohorts (n = 1366). 21 Blood pQTL summary data were derived from Ferkingstad et al., 22 which included data from 35,559 participants. All summary statistics utilized in our study were obtained from previously published studies.
SMR analysis
SMR analyses were conducted using SMR software (version 1.3.1). We selected the top cis-QTL within a ±1000 kb genomic window centered on each gene, applying a p value threshold of 5.0 × 10−8 for statistical significance. Single nucleotide polymorphisms (SNPs) in linkage disequilibrium (LD) with the top-associated SNP, with an r2 value <0.9, were included. To ensure data quality, we set the maximum allowable proportion of SNPs with allele frequency differences to 0.05.
Additionally, we analyzed the association between mQTLs and eQTLs, treating mQTLs as the exposure and eQTLs as the outcome. Similarly, eQTLs were treated as the exposure and pQTLs as the outcome to identify key links between these levels of gene regulation. 23 To refine our results, we employed the multi-SNP SMR (SMR-multi) method, which incorporates multiple SNPs at QTL loci within the SMR framework. 21 Statistical significance was evaluated using pSMR and pSMR-multi values, and the heterogeneity in dependent instruments (HEIDI) test was applied to distinguish pleiotropy from linkage effects. A pHEIDI value >0.01 was considered indicative of no pleiotropy. 24 Finally, colocalization analyses of eQTLs, mQTLs, and pQTLs were conducted when pSMR values were <0.05, pSMR-multi values were <0.05, and pHEIDI values were >0.01.
Colocalization analysis
Colocalization analysis was performed using the R package ‘coloc’ (version 5.2.3) to identify shared variants between GC-related GWAS and PANoptosis-related mQTLs, eQTLs, or pQTLs. Five posterior probabilities (PPs) were computed, corresponding to exclusive hypotheses: (H0) no association with SNPs for either trait; (H1) association only with gene expression; (H2) association only with disease risk; (H3) both traits are associated but with distinct causal variants; and (H4) both traits share the same causal variant. A locus was considered to have indicative evidence of colocalization only if it satisfied a composite criterion across two analyses: (a) in an analysis with the prior probability of a shared variant (p12) set to the conventional value of 5 × 10−5, the resulting PP for a shared variant (PPH4) had to be >0.5 and (b) in a separate analysis using a more conservative prior (p12 = 1 × 10−5), the resulting PP for distinct variants (PPH3) had to be <0.5. This dual-condition approach ensures that the signal supports the evidence for a shared variant under standard assumptions and remains inconsistent with the hypothesis of distinct variants even under more stringent priors. 25 The analysis was conducted within genomic windows of ±500 kb for mQTL–GWAS pairs and ±1000 kb for both eQTL–GWAS and pQTL–GWAS pairs.26,27
Tissue-specific eQTL validation
To further investigate the relationship between the expression of the identified genes and GC in a relevant tissue context, we utilized eQTL data for stomach tissue from the GTEx project (version 8). SMR analysis was performed by integrating the GTEx stomach eQTL summary statistics with GC GWAS summary data from the FinnGen_R10_C3_STOMACH_EXALLC cohort. The same SMR criteria were applied: pSMR <0.05, pSMR-multi <0.05, and pHEIDI >0.01 to identify significant associations.
TCGA data analysis
To assess the expression patterns of SMR-identified candidate genes, we downloaded transcriptomic (RNA-seq) data for the TCGA-STAD cohort, including 412 primary tumor samples and 36 adjacent normal tissue samples. Differential expression analysis of the 16 candidate genes between primary tumor and solid normal tissue samples was performed using the Wilcoxon rank-sum test (unpaired, nonparametric). For survival analysis, patients were stratified into high- and low-expression groups based on the median expression level of each gene. Overall survival (OS) differences between groups were assessed using the Kaplan–Meier (KM) method, 28 with significance evaluated by the log-rank test. KM survival curves were generated using the tcga_kmplot function in the R package ‘TCGAplot’ (version 4.0.0). 29 A p value <0.05 was considered statistically significant.
PPI network analysis
To investigate the functional connections among the 16 candidate genes, a PPI network was constructed using the STRING database (version 12.0, https://version-12-0.string-db.org/). 30 An interaction score >0.4 was set as the threshold for significant interactions.
Prediction of small-molecule drugs
Based on the key PANoptosis-related genes identified through integration of the eQTL and pQTL analyses, we performed a drug prediction analysis. The Enrichr platform31–33 was utilized in conjunction with the DSigDB database, 34 which contains gene sets representing drug and small-molecule perturbations. We submitted the list of key genes to Enrichr to identify small-molecule compounds significantly associated with these genes. An adjusted p value <0.05 was used as the threshold for statistical significance.
Molecular docking simulation
To further evaluate the binding characteristics of the predicted small-molecule compounds to candidate PANoptosis-related proteins, molecular docking simulations were performed. The three-dimensional (3D) structures of selected compounds were obtained from the PubChem database, and protein crystal structures were retrieved from the Protein Data Bank (PDB, https://www.rcsb.org/). Docking was conducted using AutoDock Vina (v1.2.7, https://github.com/ccsb-scripps/AutoDock-Vina/releases) within the Dockey integrated platform (https://github.com/lmdu/dockey), 35 which also enabled PyMOL-based visualization of docking results. For each protein–ligand pair, the protein structure was preprocessed, including removal of water molecules and addition of polar hydrogen atoms, prior to docking. The docking grid was defined to encompass the entire protein structure (bounding grid box), and the number of Vina simulation runs was set to 100 to ensure robustness of the results. All other parameters were kept at their default settings. Binding affinities (kcal/mol) were recorded. Based on published criteria, 36 an affinity <−4.25 kcal/mol was considered indicative of binding activity; <−5.0 kcal/mol indicated good binding; and <−7.0 kcal/mol indicated strong docking activity.
Statistical analysis
All statistical analyses were performed using R software (version 4.3.0). The Wilcoxon rank-sum test (unpaired, nonparametric) was used for differential expression analysis. The log-rank test was applied for survival analysis within the KM framework. The R package ‘ggplot2’ (version 4.0.0) was used to generate Manhattan plots, and ‘forestplot’ (version 3.1.7) was used to create forest plots. The code for generating SMRLocusPlot and SMREffectPlot was adapted from Zhu et al. 24
Results
PANoptosis-related gene expression and GC risk
In the discovery cohort (FinnGen), we identified 12 PANoptosis-related genes (pSMR < 0.05, pSMR-multi < 0.05, and pHEIDI > 0.01) (Figure 2(a)) suggestively associated with GC risk. These genes included apoptosis and caspase activation inhibitor (AVEN), CRP, DYNLL1, HAX1, hepatocyte growth factor (HGF), PIK3CA, protein phosphatase 2 regulatory subunit B beta (PPP2R2B), RALA, RELB, SYK, TGFB2, and TIA1. Among them, nine genes (AVEN, CRP, DYNLL1, HAX1, HGF, PIK3CA, RALA, RELB, and TGFB2) demonstrated a positive association with GC risk, whereas three genes (PPP2R2B, SYK, and TIA1) exhibited a negative association. Colocalization analysis provided indicative evidence of a shared genetic signal for seven genes, including AVEN, DYNLL1, HGF (odds ratio (OR): 5.453; 95% confidence interval (CI): 1.414–21.029); PPP2R2B (OR: 0.531; 95% CI: 0.347–0.811); RELB, TGFB2, and TIA1 (PPH4 > 0.5 and PPH3 < 0.5). However, none of these eQTL associations were validated in the UKB_151 replication cohort. Complete SMR and colocalization results for PANoptosis-related eQTL–GWAS analyses are provided in Supplementary Tables 2 and 3 and Supplementary Figure 1(a).

Manhattan plots of SMR analyses integrating QTL data with GC GWAS data from the FinnGen discovery cohort. (a) Manhattan plot showing the association between gene expression (eQTL) and GC risk; (b) Manhattan plot showing the association between DNA methylation (mQTL) and GC risk. In both panels, each point represents a PANoptosis-related gene, and the y-axis denotes the −log10 (p value) of the SMR test. The red horizontal line marks the significance threshold (p = 0.05). GC: gastric cancer; GWAS: genome-wide association study; QTL: quantitative trait; SMR: summary-data–based Mendelian randomization; eQTL: gene expression QTL; mQTL: methylation QTL.
Methylation of PANoptosis-related genes and GC risk
In the discovery cohort, we identified 83 methylation sites (corresponding to 56 genes) associated with GC risk. Among these, two methylation sites were validated in the UKB_151 replication cohort with consistent effect directions: cg15982655 (BNIP1) and cg07722995 (GRAMD4), both of which showed a positive association with GC risk (Supplementary Table 4).
Furthermore, colocalization analysis indicated that 40 sites (corresponding to 28 genes) demonstrated suggestive evidence for a shared causal variant within the colocalization region of the corresponding SNPs (Figure 2(b); PPH4 > 0.5 and PPH3 < 0.5). Among these, four sites were associated with PPP2R2B, including cg02760293, cg13983063, and cg15927927 (OR: 1.419; 95%CI: 1.044–1.929; PPH3 = 0.332, PPH4 =0.771) and cg25021259. Complete SMR analysis results are presented in Supplementary Tables 5 and 6, and colocalization results are presented in Supplementary Figure 1(b).
Protein levels of PANoptosis-related genes and GC risk
In the discovery cohort, we identified five proteins associated with GC risk. Among these, three proteins (GRP, HGF, and PK3CG) showed a positive association with GC risk, whereas two proteins (MAPK13 and SYK) demonstrated a negative association (Figure 3). Notably, colocalization analysis provided indicative evidence for three proteins, HGF (OR: 2.511; 95% CI: 1.242–5.076), SYK, and MAPK13, suggesting that their associations with GC were driven by shared causal variants within the corresponding SNP regions (PPH4 > 0.5 and PPH3 < 0.5). However, none of these pQTL associations were validated in the UKB_151 replication cohort. Complete SMR analysis results are presented in Supplementary Tables 7 and 8 and Supplementary Figure 1(c).
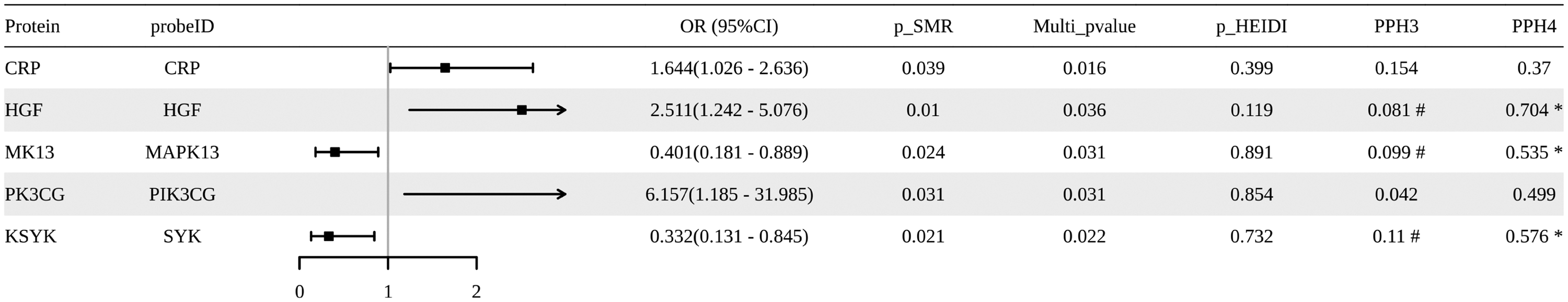
Manhattan plot of SMR analysis integrating pQTL data with GC GWAS data from the FinnGen discovery cohort. Each point represents a PANoptosis-related protein, and the y-axis denotes the −log10 (p value) of the SMR test. The red horizontal line marks the significance threshold (p = 0.05). GC: gastric cancer; GWAS: genome-wide association study; SMR: summary-data–based Mendelian randomization; pQTL: protein quantitative trait locus.
Integration of mQTL and eQTL data for GC risk assessment
Based on key findings from mQTL and eQTL data, we integrated these datasets using SMR to investigate methylation-driven regulation of critical genes. The mQTL–eQTL SMR analysis identified causal associations between two methylation sites and gene expression. Specifically, methylation at cg18944653 was negatively associated with HGF expression (β = −0.174), and methylation at cg15927927 was negatively associated with PPP2R2B expression (β = −0.658). Detailed results of this SMR analysis are provided in Table 1 and Supplementary Table 9.
Summary of SMR analyses for mQTL–eQTL and eQTL–pQTL associations.

Exposure ID (CpG site for mQTL, Ensembl ID for eQTL).
HEIDI: heterogeneity in dependent instruments; SMR: summary-data–based Mendelian randomization; SE: standard error; mQTL: methylation quantitative trait locus; eQTL: gene expression quantitative trait locus; pQTL: protein quantitative trait locus; SMR-multi: multi–single nucleotide polymorphism SMR.
Integration of eQTL and pQTL data for GC risk assessment
Furthermore, integration of eQTL and pQTL data revealed a significant association between HGF gene expression and its protein levels (β = 1.840). Detailed results of this SMR analysis are presented in Table 1 and Supplementary Table 10.
Integrating evidence from multiomics levels
Finally, we integrated the key findings from the SMR analyses. Supplementary Figure 2 presents a Manhattan plot showing the chromosomal distribution of mQTL, eQTL, and pQTL signals for HGF. Integration of blood-derived eQTL, mQTL, and pQTL data indicated that HGF and PPP2R2B had multiple lines of indicative evidence supporting their involvement in GC (Table 1).
For HGF, SMR analysis demonstrated that genetically predicted higher gene expression (OR: 5.453; 95% CI: 1.414–21.029) and lower methylation at site cg18944653 (OR: 0.736; 95% CI: 0.556–0.973) were associated with an increased risk of GC (Figure 2). Integration of mQTL and eQTL data further indicated that HGF expression is negatively regulated by methylation at cg18944653 (Supplementary Table 9). At the protein level, genetically predicted higher HGF protein abundance was also associated with increased GC risk (OR: 2.511; 95% CI: 1.242–5.076; Figure 3). This finding was consistent with our eQTL–pQTL SMR analysis, which showed that higher HGF gene expression was associated with higher HGF protein levels. Further integration of eQTL and pQTL data confirmed concordance between HGF gene expression and protein abundance (Supplementary Table 10). Taken together, these results suggest a model in which lower methylation at cg18944653 may upregulate HGF expression and subsequently increase protein abundance, thereby potentially elevating GC risk. LocusZoom plots were used to illustrate the distribution of significant signals for HGF (Supplementary Figure 3), and SMR effect plots were generated to demonstrate the impact of these signals on GC risk (Supplementary Figure 3).
Additionally, for PPP2R2B, SMR analysis showed that genetically predicted higher gene expression was associated with a decreased risk of GC (OR: 0.531; 95% CI: 0.347–0.811), whereas higher methylation at site cg15927927 was associated with an increased risk (OR: 1.419; 95% CI: 1.044–1.929) (Figure 2). The mQTL–eQTL SMR analysis supported this regulatory relationship, demonstrating that higher methylation at cg15927927 was associated with lower PPP2R2B expression (Supplementary Table 9). These findings suggest a model in which increased methylation at this site may downregulate PPP2R2B expression, thereby potentially increasing GC risk. However, further validation of the relationship between PPP2R2B protein abundance and GC risk was not feasible due to the lack of available pQTL data for the encoded protein.
Clinical validation and functional network analysis of SMR-identified genes
Our SMR analyses of eQTLs and pQTLs in the discovery cohort identified 12 and 5 potentially associated genes, respectively. After removing the overlapping gene (HGF), a total of 16 unique candidate genes were selected for downstream analyses. We analyzed the expression of all 16 candidate genes in the TCGA-STAD cohort to provide supportive evidence for our SMR findings. Notably, 11 of the 16 genes showed differential expression between tumor and normal tissues in a direction consistent with their SMR-inferred causal effects on GC risk (Figure 4(a)). For example, SMR-identified risk genes such as AVEN, HGF, CRP, DYNLL1, HAX1, PIK3CA, RALA, RELB, TGFB2, and TIA1 were significantly upregulated in tumor tissues, whereas the SMR-identified protective gene PPP2R2B was significantly downregulated.

Systematic validation and functional analyses of SMR-identified candidate genes. (a) Box plots showing differential expression of 16 candidate genes between normal adjacent tissues (normal, n = 36; red) and gastric cancer tissues (Gastric cancer, n = 412; blue) in the TCGA-STAD cohort. Statistical significance Continued.was assessed using the Wilcoxon rank-sum test (unpaired and nonparametric). Significance levels are indicated as follows: ****p < 0.0001; ***p < 0.001; **p < 0.01; *p < 0.05; ns, not significant. (b) Kaplan–Meier survival curves for overall survival (months) of patients with gastric cancer from the TCGA-STAD cohort, stratified by high and low AVEN expression based on the median expression level. The p value was calculated using the log-rank test. (c) Protein–protein interaction network of the 16 candidate genes constructed using the STRING database (version 12.0). Nodes represent proteins, with filled nodes indicating that the 3D structure is known or predicted. Edges (connecting lines) represent functional and physical protein associations. Edge color denotes the type of interaction evidence: light blue and purple lines represent known interactions (from curated databases and experimental determination, respectively); yellow and black lines represent associations derived from textmining and coexpression, respectively; and dark blue lines indicate gene co-occurrence. Line thickness is proportional to the confidence score, reflecting the strength of the association. The network illustrates the potential functional relationships among the candidate proteins. AVEN: apoptosis and caspase activation inhibitor; 3D: three-dimensional; SMR: summary-data–based Mendelian randomization; TCGA-STAD: The Cancer Genome Atlas–stomach adenocarcinoma.
Further survival analysis revealed that, among the 16 candidate genes, high AVEN expression was significantly associated with poorer OS (p = 0.031; Figure 4(b)), consistent with its role as a risk factor identified by both SMR and differential expression analyses. The remaining candidate genes identified by eQTL/pQTL-SMR did not show significant associations with OS in the TCGA-STAD cohort (Supplementary Figure 4 and Supplementary Figure 5).
To explore the functional relationships among these genes, we constructed a PPI network. The network revealed multiple interactions among the 16 candidate proteins, suggesting that they function within a coordinated network (Figure 4(c)). Notably, PIK3CA emerged as a potential hub gene with the highest number of connections, interacting with other key risk genes such as HGF and RALA. This highlights the potential importance of the PI3K signaling pathway in mediating the effects of this PANoptosis-related gene network on GC.
Tissue-specific validation of eQTLs in stomach tissue
Subsequently, we conducted tissue-specific SMR analysis using GTEx stomach eQTL data. However, none of the 16 candidate genes demonstrated a significant association that replicated the direction of effect observed in our primary blood-based SMR analysis (Supplementary Table 11).
Prediction of small-molecule drugs
To explore the therapeutic potential of the 16 SMR-identified candidate genes, we performed a drug prediction analysis. Several small-molecule compounds were identified as significantly targeting this gene set (adjusted p value < 0.05; Supplementary Table 12). Notably, resveratrol, the most significantly enriched compound, was predicted to target nine of the candidate genes, including the key oncogene HGF. Another compound, genistein, was predicted to target both HGF and the putative tumor suppressor PPP2R2B. Intriguingly, decitabine, a DNA methyltransferase inhibitor currently used in cancer therapy, was also identified as a significant hit, targeting seven genes within the network, including PPP2R2B.
Molecular docking validation of HGF with candidate drugs
Given the absence of an available PDB structure for PPP2R2B, molecular docking analysis was performed exclusively for the core protein HGF and its predicted candidate drugs. Specifically, resveratrol (PubChem CID: 445154) and genistein (PubChem CID: 5280961) were selected for in silico docking validation, as both compounds were highlighted in our enrichment analysis and have been reported to target HGF.
The crystal structure of HGF (PDB ID: 7B3Q) was used as the receptor. Both resveratrol and genistein exhibited strong predicted binding affinities to HGF, with docking energies of −8.287 kcal/mol and −9.566 kcal/mol, respectively (Table 2). Notably, both values are substantially lower than −7.0 kcal/mol, a commonly accepted threshold for strong binding activity in molecular docking studies, indicating a high likelihood of stable and specific interactions between these compounds and the HGF protein.
Molecular docking results of HGF with candidate drugs.

HGF: hepatocyte growth factor; PDB: Protein Data Bank.
The detailed binding poses of resveratrol and genistein with HGF are shown in Figure 5. These visualizations illustrate close interactions between the ligands and key residues within the HGF-binding pocket, supporting the potential of these molecules as effective HGF inhibitors.

Molecular docking models of candidate drugs with the HGF protein. (a) Predicted binding mode of resveratrol (PubChem CID: 445154) with HGF (PDB ID: 7B3Q); (b) predicted binding mode of genistein (PubChem CID: 5280961) with HGF (PDB ID: 7B3Q). For each compound, the left panel shows the overall docking pose within the protein, whereas the right panel presents a close-up view of interactions between the ligand and key amino acid residues of HGF. Hydrogen bonds and hydrophobic interactions are indicated as appropriate. Binding affinities (kcal/mol) are reported in the main text. HGF: hepatocyte growth factor; PDB: Protein Data Bank.
Discussion
This study systematically explored potential associations between PANoptosis-related genes and GC risk using an SMR approach, followed by supportive analyses in the TCGA cohort. Our SMR screening identified several promising candidate genes, and subsequent TCGA analyses demonstrated that the expression patterns of 11 of these genes were consistent with their genetically predicted effects. Among these, AVEN, HGF, and PPP2R2B emerged as particularly compelling candidates for further investigation. For AVEN, our findings suggest a potential link between its SMR-inferred risk effect and its clinical relevance as a prognostic marker. For the well-characterized oncogene HGF, our results propose a hypothetical model of epigenetic regulation mediated by DNA methylation at cg18944653. In addition, PPP2R2B was highlighted as a potential tumor suppressor, a hypothesis supported by both SMR evidence and its consistent downregulation in tumor tissues.
A key finding of our study is the identification of AVEN as a promising candidate risk factor and prognostic marker in GC. AVEN is a well-established anti-apoptotic protein that functions by inhibiting apoptosome formation and activating the ATM-dependent DNA damage response.37,38 Its role as an oncoprotein has been demonstrated in several malignancies, particularly hematopoietic neoplasms and lung adenocarcinoma, where its overexpression is consistently associated with increased malignancy and poor prognosis.39,40 However, despite its known anti-apoptotic function, the role of AVEN in the initiation and progression of GC has remained largely unexplored. Our study helps address this gap. First, our blood-based eQTL-SMR analysis provided novel genetic evidence supporting a risk-increasing effect of elevated AVEN expression on GC. This finding was further corroborated by analyses in the TCGA cohort, which demonstrated that AVEN was significantly upregulated in GC tumors and that high AVEN expression was an independent predictor of poorer OS. These results are consistent with a recent integrative proteomic study that also identified AVEN as a highly expressed molecule with prognostic significance in GC. 41 Collectively, our findings support a potential role for AVEN as both a therapeutic target and a prognostic biomarker in GC.
HGF is a multifunctional cytokine that promotes cell proliferation, migration, and survival through activation of the c-Met receptor and its downstream signaling pathways.42–44 Consistent with its established pro-tumorigenic role in GC,45,46 our SMR analysis provided supportive evidence for a risk-increasing effect associated with higher HGF expression and protein abundance. Specifically, lower methylation levels at the CpG site cg18944653 were associated with both increased HGF expression and elevated GC risk. These findings suggest that hypomethylation at this locus may represent an upstream regulatory event contributing to HGF overexpression during gastric tumorigenesis. In the context of PANoptosis, HGF has been reported to suppress multiple forms of programmed cell death, including apoptosis,47–50 pyroptosis, 51 and necroptosis. 52 Therefore, the epigenetically driven upregulation of HGF identified in our study may contribute to GC development by fostering a cellular environment resistant to PANoptotic cell death. However, because the underlying eQTL signal was not replicated in the UKB_151 cohort and colocalization evidence was moderate, this proposed mechanistic model should be considered hypothesis-generating and therefore warrants further validation.
PPP2R2B emerged as another crucial gene, with high methylation levels at the cg15927927 site potentially suppressing its expression and increasing GC risk, suggesting that PPP2R2B may function as a protective factor in GC. PPP2R2B encodes a widely expressed serine/threonine phosphatase that is involved in numerous essential cellular processes. Although no studies have directly linked PPP2R2B to GC or PANoptosis, its downregulation has been reported in several other cancers, including breast cancer, 53 pancreatic cancer, 54 and bladder cancer. 55 Studies have demonstrated that suppression of PPP2R2B results in the activation of the mechanistic target of rapamycin (mTOR) signaling pathway and promotes pancreatic ductal adenocarcinoma growth. 54 In addition, studies suggest that methylation of PPP2R2B is critically associated with survival and prognosis in patients with gliomas. 56 Furthermore, PPP2R2B has been reported to induce neuronal apoptosis by facilitating mitochondrial division. 57 These findings suggest that PPP2R2B may play a similar role in GC by influencing cell death pathways, consistent with the conclusions of our study. Further investigation is required to fully elucidate the specific role of PPP2R2B in GC and PANoptosis.
To explore potential therapeutic strategies, we predicted small-molecule drugs targeting the 16 key PANoptosis-related genes identified through eQTL and pQTL analyses. For instance, resveratrol was predicted to interact broadly with this gene set, including HGF, whereas genistein was associated with both HGF and PPP2R2B. Molecular docking analyses demonstrated strong binding affinities between HGF and both resveratrol (−8.287 kcal/mol) and genistein (−9.566 kcal/mol), with values below the commonly accepted threshold for robust ligand–receptor interactions. These results provide supportive structural evidence for our drug predictions and offer a preliminary mechanistic basis for targeting HGF in GC. However, docking analysis could not be performed for PPP2R2B because of the lack of an available crystal structure, a limitation that is common in computational drug discovery. The strong binding observed for HGF underscores the therapeutic promise of these compounds for future clinical translation and highlights the feasibility of structure-based drug discovery in this context.
Our study builds upon and advances prior research by integrating multiomics QTL data to systematically investigate the potential causal roles of PANoptosis-related genes in GC. Although previous studies have implicated some of these genes in cancer biology, few have applied an integrative SMR framework that combines blood-based QTL, GWAS, methylation, and proteomic data, together with validation in clinical cohorts. By adopting this approach, our research provides novel hypothesis-generating evidence and identifies new candidate genes for future functional and translational studies. Overall, these findings suggest that genetic dysregulation of PANoptosis-related components is an important contributor to GC susceptibility, highlighting how interactions among multiple programmed cell death pathways shape gastric tumorigenesis and therapeutic resistance.
However, our study has several limitations. First, the tissue-specific SMR analysis relied on eQTL data derived from GTEx normal gastric tissue. In contrast, the pathological processes underlying GC occur within the tumor microenvironment, where gene regulation may differ substantially from that in normal tissue. In addition, the relatively small sample size of GTEx gastric tissue may limit the statistical power to detect true associations. Consequently, the absence of replication in GTEx does not exclude the possibility of tissue- or context-specific regulatory effects in GC. Future studies incorporating larger sample sizes and tumor-derived eQTL data are required to address this limitation more comprehensively. Second, this study is primarily based on bioinformatics analyses of publicly available datasets and lacks in vitro, in vivo, or clinical validation of the identified targets. Therefore, the findings should be regarded as hypothesis-generating and interpreted with caution, as computational results alone may not be fully reliable without experimental confirmation. Future work should focus on validating the candidate genes and predicted drug–target interactions through cellular, animal, and clinical studies. Third, most of the QTL associations identified in this study were not replicated in the UKB_151 cohort, which may be attributable to limited sample size and reduced statistical power. Moreover, blood-based QTL signals may not fully reflect the regulatory landscape present within tumor tissues. Fourth, for several key signals, particularly HGF and PPP2R2B, the strength of colocalization evidence was moderate (PPH4: approximately 0.6–0.74). Although this suggests the possibility of a shared causal variant, it does not preclude the influence of LD or other confounding factors. Accordingly, these results should be interpreted cautiously, and further fine-mapping and functional validation studies are warranted. Finally, the molecular docking analyses conducted in this study provide supportive but indirect evidence for predicted drug–target interactions. These computational findings require subsequent experimental validation to confirm their biological relevance, therapeutic efficacy, and pharmacodynamic properties.
Conclusion
Our multiomics SMR and TCGA analyses generated testable hypotheses regarding the roles of several PANoptosis-related genes in GC risk. Our results provide a consistent body of evidence supporting AVEN as both a risk factor and a prognostic marker, warranting further validation. In addition, our study offers novel insights into the epigenetic regulation of HGF and the potential tumor-suppressive role of PPP2R2B. These findings underscore the importance of PANoptosis in GC pathogenesis and identify promising candidate genes for future development as diagnostic biomarkers and therapeutic targets.
Supplemental Material
sj-jpg-1-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-jpg-1-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Supplemental Material
sj-pdf-2-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-pdf-2-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Supplemental Material
sj-jpg-3-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-jpg-3-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Supplemental Material
sj-jpg-4-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-jpg-4-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Supplemental Material
sj-jpg-5-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-jpg-5-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Supplemental Material
sj-xlsx-6-imr-10.1177_03000605261430163 - Supplemental material for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk
Supplemental material, sj-xlsx-6-imr-10.1177_03000605261430163 for A multiomics Mendelian randomization study on PANoptosis-related genes and gastric cancer risk by Yunliang Wang, Yujian Xia, Lin Weng, Jin Wang, Wenzi Meng, Jiahui Zhu, Xin Zhao and Xinguo Zhu in Journal of International Medical Research
Footnotes
Acknowledgment
The authors would like to acknowledge the use of Gemini (Google) for language polishing and grammatical refinement during the preparation of this manuscript. The final interpretation of the data and the overall scientific content remain the sole responsibility of all authors.
Authors’ contributions
Yunliang Wang and Yujian Xia carried out the studies, participated in collecting data, and drafted the manuscript. Lin Weng, Jin Wang, Wenzi Meng and Jiahui Zhu performed the statistical analysis and participated in its design. Xin Zhao and Xinguo Zhu participated in acquisition, analysis, or interpretation of data and draft the manuscript. All authors read and approved the final manuscript.
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Consent for publication
Not applicable.
Declaration of conflicting interests
The authors have no relevant financial or nonfinancial interests to disclose.
Ethics approval and consent to participate
Not applicable.
Funding
This research was supported by grants from the National Science Foundation of China (NSFC; no. 82073180); Science and Education Promoting Health for Young Scholar, Science and Technology Project of Suzhou City (no. KJXW2021086); and Research of Medical Innovation and Application, Science and Technology Development Project of Suzhou City (no. SKY2023109).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
