Abstract
Objective
This study aimed to develop and validate a nomogram model for predicting 28-day mortality in patients with sepsis in the intensive care unit.
Methods
The health care records of 613 patients with sepsis who were hospitalized at the Affiliated Hospital of Chengde Medical University from 2022 to 2024 were retrospectively reviewed. Patients were randomly divided into training and testing sets in a 7:3 ratio. The least absolute shrinkage and selection operator regression method was used to identify potential prognostic factors for sepsis, followed by multivariate logistic regression to construct a nomogram prediction model. The predictive performance of the developed model was evaluated via receiver operating characteristic curves, decision curve analysis, and calibration curves.
Results
The predictive factors included the platelet distribution width to count ratio, mean platelet volume, N-terminal proB-type natriuretic peptide level, lactate level, respiratory tract infections, and diabetes. The area under the receiver operating characteristic curve for the nomogram model in the training set was 0.907, with sensitivity and specificity values of 0.846 and 0.831, respectively. The calibration curve demonstrated that the prediction results were consistent with the actual findings. Decision curve analysis revealed that the model showed robust performance in practical applications.
Conclusions
Platelet distribution width to count ratio, mean platelet volume, N-terminal proB-type natriuretic peptide level, lactate level, respiratory tract infection, and diabetes are closely associated with sepsis. A nomogram model based on these six variables demonstrates remarkable predictive performance and may assist clinicians in identifying high-risk patients and optimizing personalized therapy.
Introduction
Sepsis is a life-threatening multiple organ dysfunction initiated by a dysregulated host response to infection and is the leading cause of death of patients in the intensive care unit (ICU). 1 The sepsis-related mortality rate is approximately 437 deaths per 100,000 person-years, with in-hospital mortality for patients with sepsis approaching 17%. 2 Despite significant advances in scientific research and therapeutic technologies, sepsis continues to be associated with high mortality and economic burden for patients in the ICU. A previous study reported that early risk stratification and timely individualized interventions in patients with sepsis can help improve disease prognosis. 3 Several clinical scoring systems, including the Acute Physiology and Chronic Health Evaluation (APACHE) II and Sequential Organ Failure Assessment scores, are widely used to predict sepsis prognosis. 4 However, these prediction tools are complex and time-consuming for clinicians and often fail to provide an accurate estimate of outcomes in patients with sepsis.
The nomogram model has become a crucial tool to predict the prognosis of sepsis, simplifying statistical predictions into a single numerical estimate of event probability. Therefore, this study aimed to construct a reliable predictive model for poor prognosis in patients with sepsis, based on clinical data from a Chinese cohort, to comprehensively and objectively analyze the risk factors for the disease.
Materials and methods
Patients and collected variables
We retrospectively collected data from 613 eligible patients with sepsis who were hospitalized at the Affiliated Hospital of Chengde Medical University between 2022 and 2024. We divided each cohort into survival and fatality groups on the basis of prognosis. The inclusion criteria were as follows: (a) patients diagnosed with sepsis on the basis of the Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3); (b) patients aged ≥18 years; and (c) patients admitted to the ICU with complete clinical data. The exclusion criteria were as follows: (a) patients with autoimmune diseases; (b) patients with malignant tumor; (c) patients receiving glucocorticoid or immunosuppressant therapy; (d) pregnant women. A flowchart of this study is shown in Figure 1. This study was approved by the Ethics Committee of the Affiliated Hospital of Chengde Medical University (Approval number: CYFYLL2023401). The need for written informed consent was waived because of the retrospective nature of the study.

A flowchart of the study. ICU: intensive care unit.
Clinical data collection
The following clinical data of all ICU patients with sepsis enrolled in the study were collected from the medical records of the hospital: (a) sex and age; (b) common comorbidities, including hypertension, cerebrovascular disease (CVD), diabetes, coronary artery disease (CAD), and chronic obstructive pulmonary disease (COPD); (c) sites of infection, including respiratory tract, gastrointestinal tract, genitourinary tract, hepatobiliary system, skin and soft tissue, and other sites; and (d) biochemical indicators on admission, including C-reactive protein, white blood cell count, hemoglobin, platelet count (PLT), platelet distribution width (PDW), platelet distribution width to count ratio (PCR), mean platelet volume (MPV), neutrophil count (Neu), lymphocyte count (Lym), monocyte count (Mono), fibrinogen (Fib) level, albumin (Alb) level, fibrinogen to albumin ratio (FAR), D-dimer, total bilirubin (Tbil), alanine aminotransferase (ALT), aspartate aminotransferase (AST), lactate dehydrogenase (LDH), serum creatinine (Scr), blood urea nitrogen (BUN), N-terminal proB-type natriuretic peptide (NTpBNP), procalcitonin (PCT), lactate (Lac) levels, and APACHE II score.
Statistical methods
RStudio (3.5) was used to analyze all the data. Sepsis patients were randomized into training and testing sets in a 7:3 ratio. Normally distributed data were presented as mean ± SD, and Student’s t test was used to compare normally distributed data between groups. Measurement data with skewed distributions were presented as medians and quartiles, and the Mann–Whitney test was used for comparing measurement data between two groups. The chi-square test or Fisher’s exact test was used to compare count data between two groups, which were expressed as percentages. Univariate logistic regression was adopted to identify possible risk factors for the prognosis of sepsis. The least absolute shrinkage operator (LASSO) regression method was subsequently used to screen variables. The selection of the lambda value in the LASSO model was guided by the lambda.1se criterion using 10-fold cross-validation. All possible risk factors in the LASSO regression analysis were subsequently included in backward stepwise multiple logistic regression analyses to establish a predictive model. A nomogram prediction model was constructed in R on the basis of these prognostic factors and further verified with receiver operating characteristic (ROC) curve data from the testing set; the model was calibrated with calibration curve analysis and decision curve analysis (DCA).
Results
Patient characteristics
On the basis of the inclusion and exclusion criteria, 613 patients with sepsis were enrolled in the study. The patients were randomly divided into training (n = 429) and testing (n = 184) sets in a 7:3 ratio. No statistically significant differences were observed between the two groups in terms of baseline characteristics, comorbidities, or laboratory parameters (all P > 0.05). The specific baseline data of the training and testing sets are shown in Table 1.
Baseline characteristics of patients with sepsis in the training and testing sets.

Values are median (quartiles), mean ± SD, or n (%) as indicated.
Alb: albumin; ALT: alanine aminotransferase; APACHE II: Acute Physiology and Chronic Health Evaluation II; AST: aspartate aminotransferase; BUN: blood urea nitrogen; CAD: coronary artery disease; COPD: chronic obstructive pulmonary disease; CRP: C-reactive protein; CVD: cerebrovascular disease; FAR: fibrinogen to albumin ratio; Fib: fibrinogen; Lac: lactate; LDH: lactate dehydrogenase; Lym: lymphocytes; Mono: monocytes; MPV: mean platelet volume; Neu: neutrophils; NTpBNP: N-terminal proB-type natriuretic peptide; PCR: platelet distribution width to count ratio; PCT: procalcitonin; PDW: platelet distribution width; PLT: platelet; Scr: serum creatinine; Tbil: total bilirubin; WBC: white blood cell.
Predictive indicators selected from LASSO regression
LASSO regression was applied to select variables from the initial dataset of the training set. The 37 variables related to prognosis of sepsis patients were selected via LASSO regression and 10-fold cross-validation. In the lambda.1se criteria, the number of features was reduced to 13, as shown in Figure 2. The variables included PCR, MPV, Alb, Scr, BUN, FAR, D-dimer, NTpBNP, Lac, APACHE II score, respiratory tract infection, CAD, and diabetes.

Texture feature selection using binary logistic regression model with least absolute and selection operator (LASSO).
Multivariate logistic regression analysis and prediction of mortality in patients with sepsis
The 13 variables selected by the LASSO regression model were included in multivariate logistic regression analysis. The results indicated that PCR, MPV, NTpBNP, Lac, respiratory tract infection, and diabetes were independent risk factors for predicting mortality in patients with sepsis (P < 0.05), as shown in Table 2.
Multivariate logistic regression analysis.

Alb: albumin; APACHE II: Acute Physiology and Chronic Health Evaluation II; BUN: blood urea nitrogen; CAD: coronary artery disease; CI: confidence interval; FAR: fibrinogen to albumin ratio; Lac: lactate; MPV: mean platelet volume; NTpBNP: N-terminal proB-type natriuretic peptide; OR: odds ratio; PCR: platelet distribution width to count ratio; Scr: serum creatinine; SE: standard error.
NTpBNP was divided by 100.
Nomogram development
Six independent variables were used to construct a regression model to predict the mortality of patients with sepsis, which is presented in the form of a visual nomogram in Figure 3. The hazard ratios of these risk factors were used to score the prognosis of sepsis in each patient. The cumulative score across all variables helped assess the risk of in-hospital mortality in patients with sepsis.
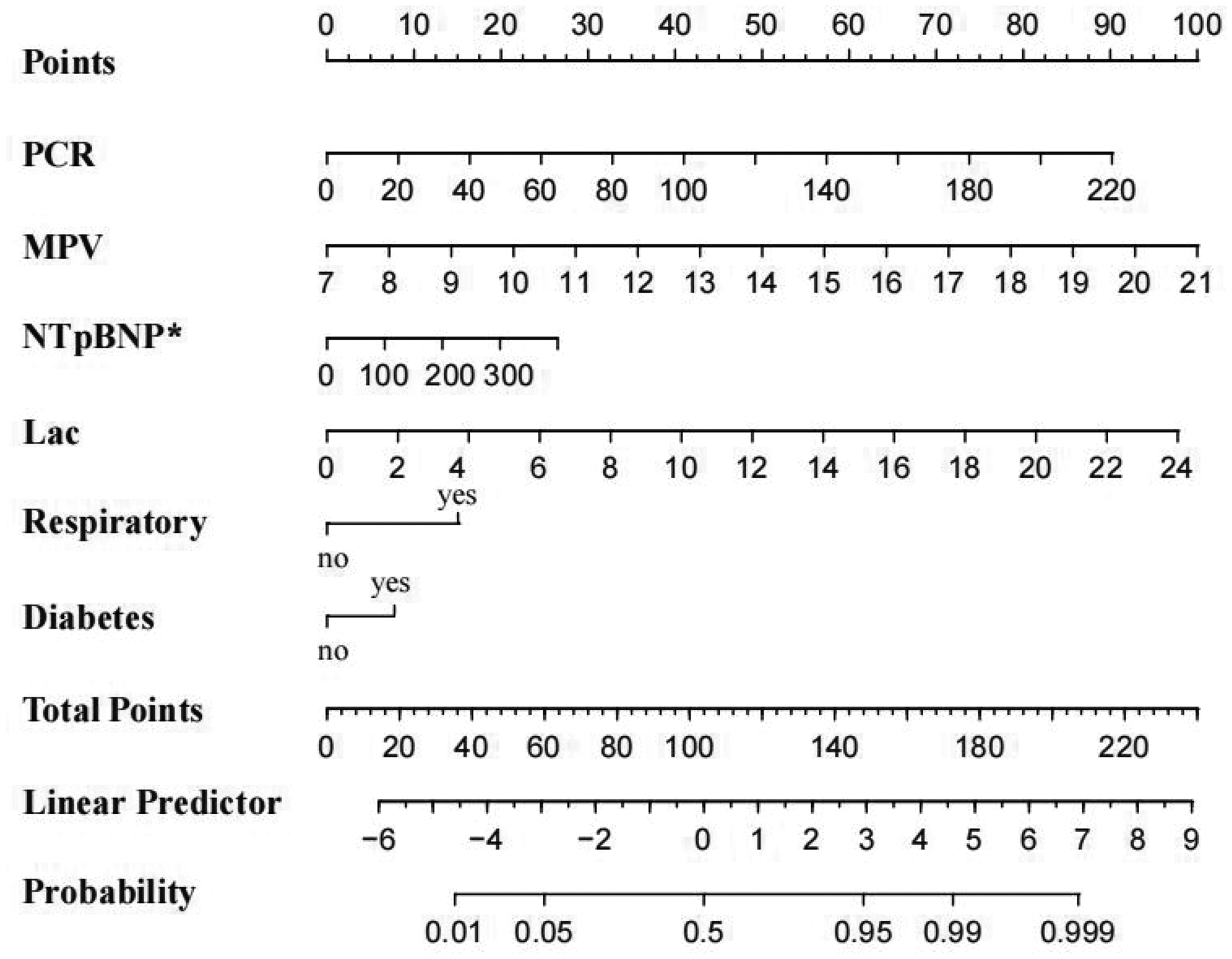
A nomogram model predicting 28-day mortality in sepsis patients. *: NTpBNP was divided by 100. Lac: lactate; MPV: mean platelet volume; NTpBNP: N-terminal proB-type natriuretic peptide; PCR: platelet distribution width to count ratio.
Discrimination and calibration
The nomogram based on data in the training set presented an area under the ROC curve (AUC) of 0.907 (95% confidence interval (CI): 0.877–0.937) for the prediction of 28-day mortality. Similarly, the AUC was 0.889 (95% CI: 0.834–0.944) for the prediction of 28-day mortality in the testing cohort, as shown in Figure 4.

The area under the receiver operating characteristic curves. AUC: area under the curve; CI: confidence interval.
The nomogram demonstrated good agreement with actual 28-day mortality for both cohorts, as shown by the calibration curves in Figure 5, despite slight deviations in the logistic calibration curve and nonparametric curve from the ideal line. The Hosmer–Lemeshow test indicated no significant deviation from a perfect fit, with P-values of 0.151 in the training set and 0.714 in the testing set.
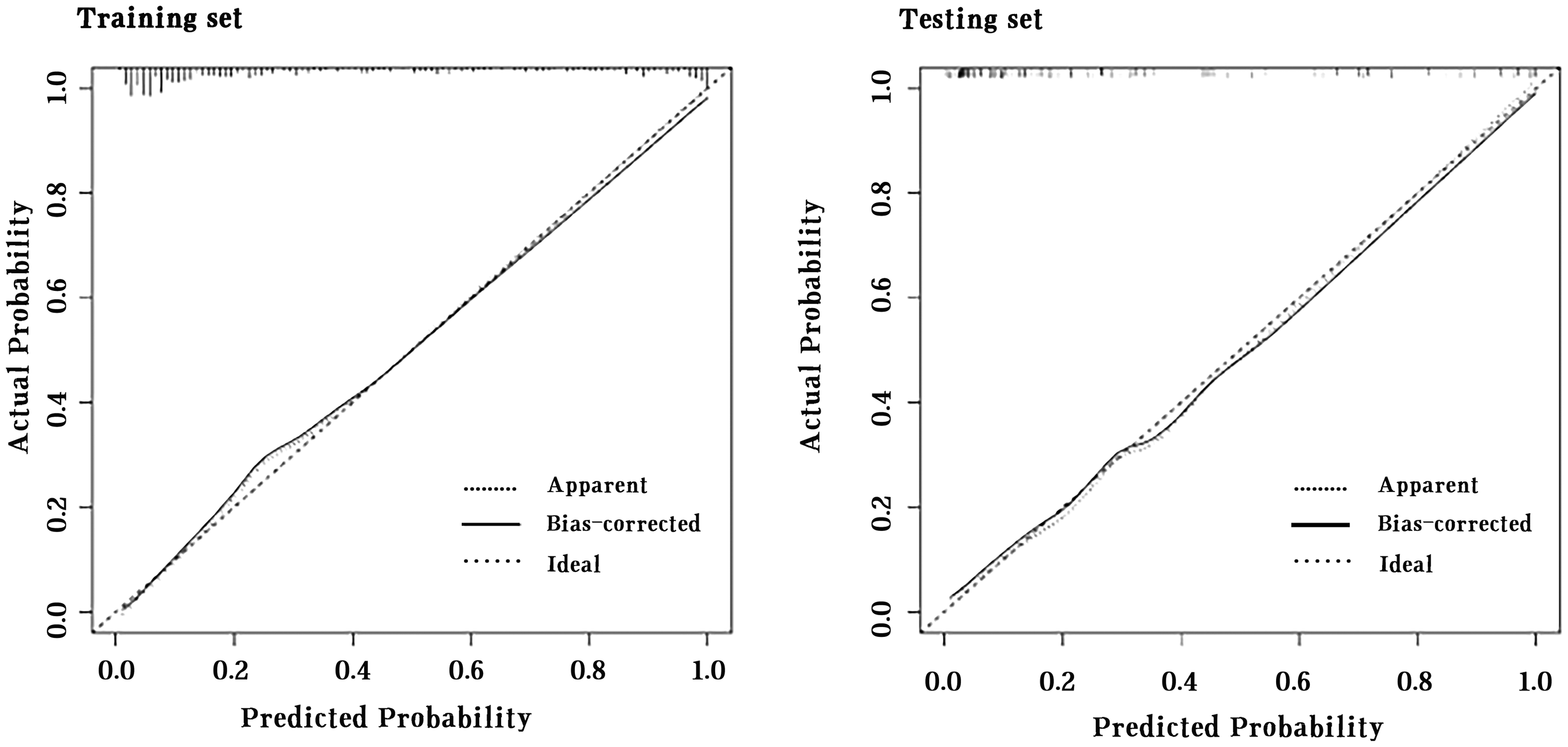
Calibration plot of predicted and observed probabilities of 28-day mortality.
Clinical utility
DCA was applied to evaluate the clinical utility of the predictive model and scoring methods in the training and testing sets, where the X-axis measures the threshold probability and the Y-axis measures the net benefit, as shown in Figure 6. The horizontal axis represents a scenario in which all samples are negative (Treat-None) and have zero net benefit. The oblique line represents a scenario in which all samples are positive (Treat-All). The range formed between the DCA curve and the horizontal and oblique lines represents the clinical net benefit interval. These findings indicate that there is a net clinical benefit from the use of a predictive model for the prognosis of sepsis.

Decision curve analysis to assess the benefit of clinical intervention based on the predictive nomogram.
Discussion
Sepsis poses an escalating threat to public health in China, making early personalized intervention and treatment essential. Our study found that PCR, MPV, NTpBNP level, Lac level, respiratory tract infections, and a history of diabetes were independent predictors of 28-day mortality in patients with sepsis in China. These predictors were used to develop a nomogram for quickly and visually assessing the prognostic risk in patients with sepsis.
Our study provides evidence that platelet-related parameters, including PCR and MPV, are significantly associated with increased 28-day mortality in patients with sepsis in the ICU. PCR combines two platelet indices, PDW and PLT, which are readily available through routine blood tests. Platelet dysfunction is recognized as a risk factor for sepsis, influencing prognosis through the platelet–endothelial cell coagulation cascade, which can lead to sequential organ dysfunction. 5 An analysis using the Medical Information Mart for Intensive Care (MIMIC-III) database indicated that PLT was independently associated with outcomes and was included in a predictive model to help clinicians assess the overall survival of patients with sepsis. 6 Platelet indices include PDW, which is closely associated with platelet aggregation and thrombus formation, and MPV, a marker of platelet size. An elevated MPV indicates increased platelet turnover by the bone marrow in response to stress. 7 At present, several research efforts have explored the use of PCR and MPV as promising methods for the early detection of sepsis and for evaluating patient prognosis.8,9 Our study revealed that the use of PCR and MPV can independently increase 28-day mortality in patients with sepsis.
In this report, using logistic regression analyses, we showed that high baseline Lac and NT-proBNP levels were correlated strongly with increased 28-day mortality in patients with sepsis in the ICU. Blood Lac, a common clinical indicator of sepsis progression, is believed to result from tissue hypoxia in septic patients. Various previous studies have indicated that high levels of Lac show a strong correlation with elevated mortality in those affected by sepsis.10,11 NT-proBNP is derived from an inactive fragment of the neurohormone brain natriuretic peptide, which is produced in the cardiac atria under normal conditions and is predominantly cleared by the muscles, liver, and kidney.12,13 NT-proBNP levels are significantly elevated in response to severe myocardial injury and are strongly correlated with systemic inflammation. 14 However, only a few studies have used NT-proBNP level alone as a prognostic indicator in septic patients. NT-proBNP level has been combined with other clinical markers to predict mortality in septic patients, further demonstrating its significance in determining sepsis prognosis.15,16
The clinical presentation of sepsis is heterogeneous, and its prognosis largely depends on the primary site of infection and comorbidities. The respiratory tract is the most common site of infection (as in pneumonia) and is strongly associated with the highest mortality in patients with sepsis. 17 A retrospective cohort study by Leligdowicz et al. 18 revealed that the site of infection was strongly associated with hospital mortality, even after adjusting for confounders such as the timing and appropriateness of antimicrobial therapy. The emergency department sepsis (MEDS) score, developed by Shapiro et al.,19,20 includes infection site (as in pneumonia as well as skin and soft tissue infections) and can predict both 28-day and 1-year mortality in patients with sepsis. Although the impact of diabetes on sepsis-related mortality remains controversial, diabetic patients represent the largest group affected by postsepsis complications and increased mortality. 21 Two large retrospective cohort studies demonstrated that diabetic patients experienced higher mortality from infectious diseases than nondiabetic patients.22,23 These studies further underscore the significance of respiratory tract infections and diabetes in determining the prognosis of sepsis.
Recently, nomograms have been widely used as a novel predictive tool in survival analysis of patients with sepsis. In this study, six independent risk factors—including PCR, MPV, NT-proBNP level, Lac level, respiratory tract infection, and diabetes—were integrated to establish a nomogram model based on the results of multivariate logistic analysis. These six factors are clinical characteristics and biochemical indicators collected during initial routine examinations of patients with sepsis, with results that are readily available. Our nomogram model demonstrates strong predictive performance and accuracy in predicting 28-day sepsis-related mortality, with satisfactory calibration curves and relatively high AUC values in both the training and testing sets. Furthermore, the model shows broad clinical applicability in predicting 28-day sepsis-related mortality, as demonstrated by DCA in both cohorts. In addition, it may be more convenient for medical staff and decrease the mortality of sepsis in the ICU.
This report has several limitations. First, the investigation is based on a retrospective cohort study with a limited sample size, which may restrict the generalizability of our findings. Validating the practicality and application of this nomogram model requires comprehensive studies across multiple centers with substantial sample sizes. Second, we only validated our data internally, and it is essential that future studies conduct additional validation to confirm the strength and efficacy of the nomogram. Finally, the duration of follow-up in our study was insufficient.
Conclusion
We developed a nomogram model based on six common clinical variables—including PCR, MPV, NTpBNP level, Lac level, respiratory tract infection, and diabetes—to predict 28-day mortality in sepsis patients. The training and testing cohorts showed that our model demonstrated strong predictive performance and clinical utility, as assessed by the ROC curve and DCA. Thus, the developed model may provide greater practicality and convenience for clinicians in predicting prognosis and formulating therapeutic strategies for patients with sepsis.
Footnotes
Acknowledgments
The authors have no acknowledgments to disclose.
Author contributions
All authors contributed to data analysis related to sepsis patients, were involved in drafting or revision of the manuscript, approved the final version for publication, and accepted the responsibility for all aspects of the work.
Data availability
The data investigated in this research were approved by the Ethics Committee of the Affiliated Hospital of Chengde Medical University. Further inquiries can be directed to the corresponding author.
Declaration of conflicting interest
The authors report no conflicts of interest in this study.
Funding
The study was supported by the Medical Science Research Project plan of Hebei Provence (Approval number: 20241528).
