Abstract
Objective
To investigate the potential inflammatory pathways involved in the development of microscopic colitis (MC).
Methods
This prospective study analysed human intestinal tissue that was collected and classified as healthy controls (HC), microscopic colitis (MC) and ulcerative colitis (UC). An RT2 Profiler PCR Array for human inflammatory response and autoimmunity was used to evaluate the expression of 84 specific genes related to the inflammatory and autoimmunity pathways. Data were validated by means of real-time polymerase chain reaction on an independent group of MC intestinal tissue samples.
Results
This study measured the expression of inflammatory genes in HC (n = 10), in patients with MC (n = 8) and in patients with active UC (n = 10). Of the 84 genes included in the array, the expression of the C-C motif chemokine ligand 19, C-C motif chemokine ligand 21, lymphotoxin beta and complement C3 genes that are involved in the non-canonical nuclear transcription factor kappa B (NF-kB) pathway was increased by 2.96, 6.05, 5.96 and 5.93 times in MC compared with HC, respectively. These results were confirmed by real-time polymerase chain reaction.
Conclusions
The findings suggest that an impairment of the non-canonical NF-kB pathway is involved in the development of MC.
Introduction
In developed countries, microscopic colitis (MC) is one of the most common causes of chronic, watery diarrhoea. 1 Patients with MC have a normal endoscopic appearance, with occasional erythema and patchy oedema distributed along the colon.2–4 The two main histological forms of MC are lymphocytic colitis (LC) and collagenous colitis (CC). 5 The histopathological criterion of LC is a significant lymphocytic infiltration in the surface epithelium that exceeds 20 per 100 epithelial cells, while CC is defined by a subepithelial collagen band >10 µm thick with an increased inflammatory infiltrate in the lamina propria. 1 Both CC and LC show an intense infiltration of T cells in the lamina propria, with also the presence of plasma cells, eosinophils, mast cells, macrophages and neutrophils;6,7 with a normal architecture of the crypts.6,8
The heterogeneous immune profiles and disease characteristics among patients still make the pathogenesis of MC unclear.7,9–11 Currently, it is clear that MC is an immune-mediated disease in which the adaptive immune response is prominent and it shows some common features with inflammatory bowel disease (IBD). 9 Dysregulated cytokine production and signalling mechanisms by intestinal epithelial cells, lymphocytes and macrophages have been implicated in the pathogenesis of IBD. 12 Nuclear transcription factor kappa B (NF-kB) is one of the key regulatory molecules. 12 The amount of activated NF-kB correlates significantly with the severity of intestinal inflammation. 12 The involvement of NF-kB as a pivotal regulator of the immunological activation of IBD has been clearly demonstrated, however, its role in MC remains unknown.12,13
Nuclear transcription factor kappa B can be activated via two signalling pathways: the canonical pathway mediates NF-kB activation by NF-kB p65 subunit and c-Rel and the non-canonical NF-kB pathway involves p52 and transcription factor RelB; 14 with both resulting in the release of NF-kB from its inhibitors and its translocation to the cell nucleus.15–17 The canonical NF-kB pathway responds to a variety of immune receptors leading to rapid but transient activation. 14 In contrast, the activation of the non-canonical pathway is slow and persistent and induced by a subset of tumour necrosis factor (TNF) receptor superfamily members.14 Activation of NF-kB in the intestinal mucosa induces the expression of proinflammatory molecules and adhesion molecules. 12 Among these cytokines, TNF-α can enhance NF-kB activation in various cell types. 13 A well-known function of the non-canonical NF-kB pathway includes the regulation of secondary lymphoid organ development, including spleen, lymph nodes and mucosal lymphoid tissues.14,18 It is also involved in humoral immunity and germinal centre formation in secondary lymphoid organs, where B cells undergo clonal expansion after antigen-stimulation. 19 The non-canonical NF-kB pathway is indispensable for the initial activation of naïve CD4+ T cells and their differentiation to type 1 helper T (Th1) cells, Th2, Th9 and Th17 cells with helper or regulatory functions. 20
When aberrantly activated, non-canonical NF-kB signalling can promote autoimmunity and inflammation causing excessive and chronic tissue damage and inflammation. 14 Given all these previous data, the purpose of this current study was to elucidate the potential involvement of the non-canonical NF-kB pathway in MC pathogenesis.
Patients and methods
Study population and sample collection
This prospective study analysed human intestinal tissue that was collected by the Endoscopy Unit staff at IRCCS Policlinico San Donato, San Donato Milanese, Italy between February 2014 and December 2014. Endoscopists provided biopsy samples of the right and left colon mucosa according to the European Guidelines on Microscopic Colitis. 1 The samples were classified as healthy controls (HC), microscopic colitis (MC) and ulcerative colitis (UC) after evaluation by a pathologist. Biopsies were immediately frozen in liquid nitrogen for further analysis. Formalin-fixed and paraffin-embedded (FFPE) samples for the verification analysis were obtained by the staff of the Pathology Unit at IRCCS Fondazione Ca' Granda Ospedale Maggiore Policlinico di Milano, Milano, Italy from their samples archive.
The research was carried out according to the Declaration of Helsinki. The local Ethics Committee of San Raffaele Hospital, Milan, Italy approved the study protocol (no. 71/INT/2014). All patients enrolled were informed about the study and they provided written informed consent before undergoing endoscopy procedures.
Analysis of genes related to inflammatory and autoimmunity pathways
An RT2 Profiler PCR Array for human inflammatory response and autoimmunity (Qiagen, Milan, Italy) was used to evaluate the expression of 84 specific genes related to the inflammatory and autoimmunity pathways. Total RNA was isolated from 10 mg of intestinal biopsy sample using the RNeasy mini kit (Qiagen) according to the manufacturer’s instructions. cDNA was synthesized using the RT2 First Strand Kit (Qiagen) after the removal of genomic DNA. Samples were analysed using the RT2 Profiler PCR Array. A total of 84 different genes were simultaneously amplified in the sample. A melting curve analysis was performed to verify that the product consisted of a single amplicon for each gene. The polymerase chain reaction (PCR) arrays were performed in 96-well plates on a StepOnePlus™ instrument (Applied Biosystems, Monza, Italy). The reaction mix was prepared from 1350 µl 2× RT 2 qPCR Master Mix (Qiagen) and 102 µl of sample cDNA. 10 µl of this mixture was applied to each well of the RT2 Profiler PCR Array. The thresholds and baselines were set according to the manufacturer’s instructions and data were analysed using software supplied by Qiagen. 21 The fold change in gene expression compared with unstimulated control was calculated using the 2−ΔΔCt method, with a more than 1.5-fold change in gene expression considered as the upregulation or downregulation of a specific gene expression. The gene pathway was reconstructed by IPA Software (Qiagen) and the Kyoto Encyclopedia of Genes and Genomes Pathways Database. 22 The experiments were performed in triplicate.
Real-time PCR analysis of a different set of FFPE sections
Total RNA was isolated from 10 FFPE intestinal tissue sections from a different group of patients with LC and CC using the FFPE RNeasy mini kit (Qiagen) according to the manufacturer’s instructions. cDNA was synthesized using the RevertAid H Minus First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, Monza, Italy). cDNA was subjected to quantitative real-time PCR using HOT FIREPol® Evagreen® qPCR Supermix (Solis BioDyne, Carlo Erba, Italy) in a final volume of 20 µl. The primer pairs designed using Eurofins Genomics design tool are listed in Table 1. 23 Ribosomal protein S14 was used as the housekeeping gene in order to standardize the amount of template for the specific primers (Eurofins Genomics Srl, Vimodrone, Italy).
List of primers used in real-time polymerase chain reaction (qRT-PCR) for validation of the data obtained using the RT2 Profiler PCR Array.
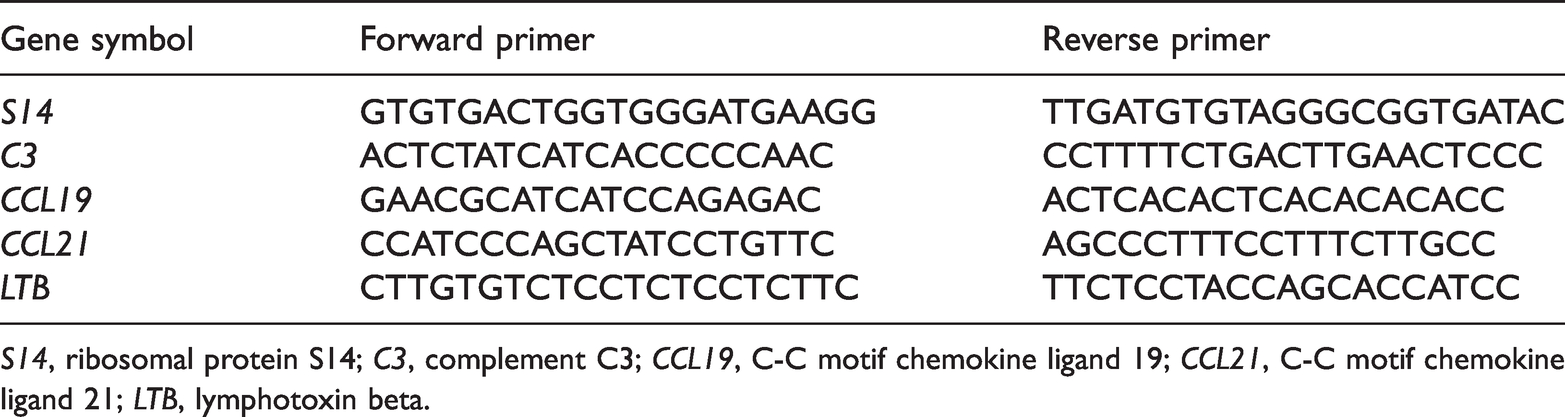
S14, ribosomal protein S14; C3, complement C3; CCL19, C-C motif chemokine ligand 19; CCL21, C-C motif chemokine ligand 21; LTB, lymphotoxin beta.
Statistical analyses
All statistical analyses were performed using GraphPad Prism 5 software (Graphpad Software Inc., San Diego, CA, USA). Data are presented as mean ± SD and compared using Mann–Whitney U-test, Wilcoxon matched-pairs signed-ranks test and Fisher’s exact test. The P-values for the RT2 Profiler PCR Array were calculated based on a Student’s t-test of the replicate 2−ΔΔCt values for each gene in the control group and treatment groups. The P-value calculation used was based on parametric, unpaired, two-sample equal variance, two-tailed distribution. P-values ≤0.05 and ≤0.001 were considered statistically significant.
Results
This study measured the expression of inflammatory genes in healthy intestinal tissue (HC; n = 10), in patients with MC (n = 8) and in patients with active UC (n = 10). The mean ± SD age of the healthy control subjects was 61 ± 15 years; and it was 71 ± 2 years for the patients with MC and 66 ± 8 years for the patients with UC. There were no significant differences between the age of the three groups. A subset of genes that were differentially regulated among the three groups were selected and four genes were identified that were exclusively modulated in MC (Table 2). The C-C motif chemokine ligand 19 (CCL19), C-C motif chemokine ligand 21 (CCL21) and lymphotoxin beta (LTB) genes are involved in the non-canonical NF-kB activation pathway and their expression was increased by 2.96, 6.05 and 5.96 times in MC compared with HC, respectively. Complement C3 (C3) was overexpressed in MC compared with HC (5.93 times), demonstrating the hypothetical involvement of a different inflammatory pathway in MC pathogenesis.
Inflammatory genes differentially regulated in patients with microscopic colitis (MC) compared with healthy control (HC) tissue.
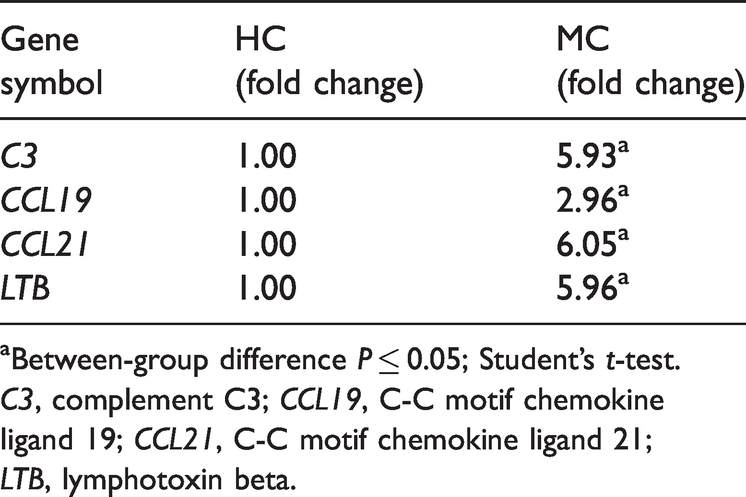
aBetween-group difference P ≤ 0.05; Student’s t-test.
C3, complement C3; CCL19, C-C motif chemokine ligand 19; CCL21, C-C motif chemokine ligand 21; LTB, lymphotoxin beta.
In order to further validate the RT2 Profiler PCR Array data, real-time PCR analysis was performed on a different set of FFPE intestinal tissue from patients with MC (LC, n = 2; CC, n = 3). The results confirmed the RT2 Profiler PCR Array findings (Figure 1). All four genes were upregulated in MC compared with HC; CCL19, CCL21 and LTB gene expression was increased by 1.79, 4.75 and 5.66 times compared with HC, respectively. The C3 gene was overexpressed 4.01 times in MC compared with HC.

Real-time polymerase chain reaction (qRT-PCR) gene expression results for formalin-fixed and paraffin-embedded samples from a different group of patients with microscopic colitis (MC) compared with health control (HC) tissues undertaken to verify the results for the four genes identified by the RT2 Profiler PCR Array. Data presented as mean ± SD; *P ≤ 0.05; Student’s t-test. CCL19, C-C motif chemokine ligand 19; CCL21, C-C motif chemokine ligand 21; C3, complement C3; LTB, lymphotoxin beta.
Discussion
Microscopic colitis comprises immune-mediated disorders affecting mostly the colonic mucosa in adults and elderly patients presenting with chronic non-bloody diarrhoea. 4 The two main forms of MC are CC and LC. 4 They share clinical presentation and many histopathological characteristics. 9 Recently, scientific research has highlighted some potential new hypotheses on the pathogenesis of MC. 9 Among these new hypotheses, a genetic predisposition and the dysregulation of the adaptive immune responses appear to have important roles.9,24–27 More specifically, MC appears to be mediated by Th1/Th17 responses and cytotoxic T cells.24–27
These current data suggest that NF-kB is one of the key regulators of this immunological condition. In particular, the analysis of the gene expression profile demonstrates that the non-canonical signalling pathway of NF-kB is dysregulated in MC. The non-canonical NF-kB pathway is responsible for the development of immune cells and is required for the maturation and function of thymus epithelial cells (TECs), which are essential for T cell development in the thymus.28–30 The T cells become effector cells in the adaptive immune response and are also involved in autoimmune diseases.28–30 This pathway is also involved in the development of secondary1,14,18,31 and tertiary lymphoid organs;32–34 and in chronic inflammatory diseases.14,35 The non-canonical NF-kB pathway mainly functions in T cell development in the thymus; and in the generation and maintenance of effector and memory T-cells in the periphery.14,17 It was reported that a deficiency of the non-canonical NF-kB inducers lymphotoxin alpha (LTA), LTB, their receptors, and other components of this pathway (Figure 2), results in defects of medullary thymic epithelial cell (mTEC) development, leading to disturbed self-tolerance.28,36–38 Additionally, lymphotoxin beta receptor (LTβR) signalling regulates the function of mTECs and in vivo experiments have demonstrated that LTA/LTB deficient mice show disorganized medullary region and impaired communication between developing thymocytes and mTECs, causing defects in T cell selection and autoimmunity. 39 Further studies in LTβR deficient mice demonstrated that impaired CCL21 expression in mTECs compromises the interaction between the developing thymocytes and mTECs.(36,38) Mature naïve T cells emigrate from the thymus to the periphery, where they are activated by antigens presented on antigen presenting cells and differentiate into effector and memory T cells.14,17 Activated CD4+ T cells differentiate into distinct effector subsets including Th1, Th2, Th9, Th17, follicular helper T cells and regulatory T cells.20,40 The non-canonical NF-kB pathway is necessary for Th17 differentiation and these T cells are involved in antimicrobial immunity at mucosal barriers, stimulating epithelial cell production of antimicrobial proteins and recruiting neutrophils. 24 Interleukin 17A induces claudin production in intestinal epithelial cells, strengthens tight junctions and induces mucin production in the gut. 26 However, when the Th17 response is inappropriate it can be a trigger in the pathogenesis of several immune and inflammatory diseases, such as rheumatoid arthritis, psoriasis, Sjögren syndrome, IBD, multiple sclerosis and systemic lupus erythemathosus.26,27 The expression of the C3 gene in MC mucosa can cause the enhancement of the local innate immune response and also amplify the cascade linking innate and adaptive immune responses.41,42 The complement system plays an important role as an effector of innate immunity.41,42 It is regarded as the pattern-recognition receptors that recognize pathogen-associated molecular patterns. 43 The complement system is one of the regulatory systems of T cell functions such as the direct opsonization of non-self antigens by antigen-presenting cells, the modulation of cytokine release and also stimulates the differentiation of regulatory T cells.41,43

The non-canonical nuclear transcription factor kappa B (NF-kB) pathway selectively responds to a subset of tumour necrosis factor receptor (TNFR) superfamily members such as lymphotoxin alpha (LTA) and lymphotoxin beta (LTB) that activate the kinase NF-kB-inducing kinase (NIK). NIK phosphorylates and activates IKKα, which in turn phosphorylates p100, initiating the degradation of the C-terminal IkB-like structure of p100, leading to the generation of p52. The nuclear translocation of p52 and RelB activate the transcription of the C-C motif chemokine ligand 19 (CCL19) and C-C motif chemokine ligand 21 (CCL21) genes. TRAF2, TNF receptor associated factor 2; TRAF3, TNF receptor associated factor 3; LT-βR, lymphotoxin β receptor; cIAP1/2, cellular inhibitor of apoptosis 1 and 2; IKKα, IkB kinase α. The colour version of this figure is available at: http://imr.sagepub.com.
This study had several limitations. First, the number of patients included in the study was small. Despite this, it was possible to demonstrate the important role of the non-canonical NF-kB pathway in the pathogenesis of MC. Future research will further explore this pathogenic mechanism and analyse patients with CC and LC separately in order to determine if they share some characteristics in the activation of this specific pathway.
In conclusion, this current study suggests a role for the non-canonical NF-kB pathway in the development of MC. Given the role of the non-canonical NF-kB pathway in Th17 differentiation and the involvement of the C3 gene in T cell function, it is possible to hypothesize a role for these processes in the pathogenesis of MC. A clearer understanding of the mechanisms involved in the development of MC could lead to the identification of biomarkers for the early, non-invasive diagnosis of MC and provide new molecular targets for therapy.
Footnotes
Author contributions
Conceptualization: Laura Francesca Pisani, Gianeugenio Tontini; Investigation: Laura Francesca Pisani; Resources: Laura Francesca Pisani, Gianeugenio Tontini, Giorgio Alberto Croci; Supervision: Luca Pastorelli; Writing – original draft, Laura Francesca Pisani; Writing – review & editing, Luca Pastorelli, Maurizio Vecchi and Gianeugenio Tontini.
Acknowledgements
Authors would like to thank Beatrice Frassini (ISIS Zenale e Butinone, Treviglio, Italy) for her graphical contribution.
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and publication of this article: This research was partially supported by Ricerca Corrente funding from the Italian Ministry of Health to IRCCS Policlinico San Donato and by the Research Grant ‘LETIZIA CASTELLI SCHUBERT’.
