Abstract
Objectives
Diabetic macular edema (DME) is a complication of diabetes mellitus that leads to diabetic retinopathy. Thus far, the role of serum exosomal microRNAs (miRNAs) in DME progression remains elusive. This study investigated serum exosomal miRNAs from patients with type 2 diabetes (T2D) and DME to identify miRNAs associated with expression of vascular endothelial growth factor (VEGF), a pivotal component in DME progression; it also evaluated the diagnostic values of these miRNAs for DME.
Methods
Serum was collected from patients with T2D who did (n = 20) and did not have DME (n = 24). Exosomes were isolated from serum and subjected to real-time polymerase chain reaction, western blotting, luciferase reporter, and miRNA profiling analyses.
Results
VEGF was significantly upregulated in ARPE-19 cells treated with exosomes from patients with T2D and DME, compared with exosomes from patients with T2D alone. Among the top 10 downregulated miRNAs identified during exosomal miRNA profiling, miR-377-3p inhibited the expression of VEGF. Luciferase reporter assays confirmed that miR-377-3p could directly regulate VEGF expression. Receiver operating characteristic analysis identified serum exosomal miR-377-3p as a potential biomarker for DME.
Conclusion
Serum exosomal miR-377-3p inhibits VEGF expression to suppress retinal pigment epithelium proliferation and offers a diagnostic biomarker for DME.
Keywords
Introduction
The most common sight-threatening complication of diabetes is diabetic macular edema (DME), which causes diabetic retinopathy and vision loss through macular thickening due to increased levels of extracellular fluid derived from hyperpermeable retinal capillaries.1,2 Approximately 20% of patients with type 1 diabetes and 25% of patients with type 2 diabetes (T2D) developed DME within 10 years after the initial diagnosis.1,3,4 In China, the population with diabetes comprises 9.7% of people aged >20 years. 5 Thus, there is an urgent need to understand the mechanism by which diabetes causes DME and identify potential biomarkers for DME, thereby providing insights for clinical treatment.
Exosomal microRNAs (miRNAs) have been identified as biomarkers in several diseases, such as hepatocellular carcinoma and osteosarcoma.6,7 However, the involvement of exosomal miRNAs in DME is unclear. Exosomes contain various components, such as lipids, proteins, and nucleic acids (e.g., mRNAs, miRNAs, and non-coding RNAs), which are mainly located in the exosomal lumen. 8 Exosomes must interact with target cells to mediate their intended biological functions; for example, exosomal miRNAs degrade target proteins by directly silencing target mRNAs. 9 Many cell types can release exosomes; these cells include immunocytes and retinal pigment epithelium (RPE). Under conditions of oxidative stress, vascular endothelial growth factor (VEGF)-overexpressing exosomes released from RPE can strongly promote angiogenesis in endothelial cells. 10 Furthermore, exosomes isolated from human umbilical cord mesenchymal stem cells can freely enter breast cancer cells to downregulate the expression of VEGF, thus inhibiting angiogenesis and associated tumorigenesis, 11 which demonstrates that exosomes can regulate VEGF expression. VEGF is mainly produced by RPE and plays a pivotal role in retinopathy; 12 accordingly, VEGF is a critical target for the treatment of DME, but its regulation remains largely uncharacterized.
In this study, we investigated the serum exosomal miRNA profiles in patients with T2D, with and without DME, to identify miRNAs associated with VEGF expression. We also evaluated their potential value in the diagnosis of DME.
Materials and methods
Serum collection
Ten-milliliter samples of peripheral blood were collected from patients with T2D, with and without DME, using tubes containing coagulant (KJ1002Z, Kang-Jia Co. Ltd., Yuhuan, China). Only patients not receiving treatment for any condition were included in the study. All patient samples were obtained with written informed consent from participating patients. All experiments were conducted in accordance with the Helsinki Declaration. This study protocol was approved by the Ethical Committee of Shenzhen People’s Hospital (Ref: LL-KY-2019397).
Exosome extraction and quantification
Exosome extraction was conducted as previously described. 13 Briefly, serum was obtained by centrifugation of blood samples at 900 × g for 10 minutes, then transferred to RNase-free tubes (Eppendorf, Hamburg, Germany) and stored at −80°C until use. Exosome isolation from serum was performed using ExoQuick Exosome Precipitation Solution (EXOQ5A-1, System Biosciences, Palo Alto, CA, USA), in accordance with the manufacturer's protocol. Specifically, serum samples were thawed and centrifuged at 3000 × g for 15 minutes to remove cell debris. Subsequently, 250 µL serum and 63 µL ExoQuick reagent were added to RNase-free tubes and samples were incubated at 4°C for 12 hours. Finally, exosomes were collected by centrifugation at 1500 × g for 30 minutes. Exosome Capture & Quantification Kits (NBP2-49785, Novus Biologicals, Centennial, CO, USA) were used for exosome quantification, in accordance with the manufacturer’s instructions. The particle size and concentration were measured using a Nanoparticle Tracking Analysis 300 device (Malvern Instruments, Malvern, UK), in accordance with the manufacturer’s instructions.
Cell culture and treatment
ARPE-19 cells (Cobioer Biosciences, Nanjing, China) were cultured in Dulbecco's modified Eagle's medium/Nutrient mixture F12 (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 5 mM HEPES buffer, 7.5% NaHCO3, 10% fetal bovine serum, and 1% penicillin/streptomycin; cells were maintained at 37°C and 5% CO2.
To investigate the impacts of exosomes from different patient groups on VEGF expression, ARPE-19 cells were treated with 50 μg of exosomes. During repetitive exosome treatment, ARPE-19 cells were treated three times at 24-hour intervals. After treatments, ARPE-19 cells were harvested for real-time polymerase chain reaction and western blotting analysis.
To investigate the impacts of miRNAs on RPE proliferation, ARPE-19 cells (transfected as described in the next section) were seeded in 96-well plates (1 × 104 cells/well) and incubated for 0, 1, 2, 3, 4, and 5 days at 37°C. Subsequently, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT; 5 mg/mL) was added at the indicated growth intervals and the cells were incubated for an additional 4-hour interval. Dimethyl sulfoxide (100 µL) was then added to each well. Finally, the absorbance was measured at 490 nm using a standard microplate reader.
Transfection with miR-377-3p and VEGF expression plasmids
ARPE-19 cells were seeded in six-well plates at 4 × 105 cells/well. When cells reached approximately 60% confluency, they were transfected with pCMV-miR-377-3p plasmid or pCMV negative control plasmid (OriGene, Rockville, MD, USA) using Lipofectamine 2000 (Thermo Fisher Scientific), in accordance with the manufacturer’s instructions. Stable clones were obtained via puromycin selection. For proliferation assays, cells were transiently transfected with VEGF plasmid (sc-29520-SH; Santa Cruz Biotechnology, Dallas, TX, USA) using Lipofectamine 2000.
Transmission electron microscopy
Isolated exosomes were re-suspended in 1 mL of phosphate-buffered saline; 3 μL of sample was then placed onto a glow-discharged copper grid and covered with carbon film. After incubation for 30 s at room temperature, samples were stained with 3% uranyl acetate. Transmission electron microscopy observations were conducted using a Tecnai T120 microscope (Thermo Fisher Scientific). Whole exosomes in the images were measured to determine their size.
RNA extraction and real-time polymerase chain reaction
Total RNA was extracted using an RNA extraction kit (Sigma-Aldrich, St. Louis, MO, USA). cDNA was synthesized from extracted RNA using a High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA). Quantitative real-time polymerase chain reaction analyses were performed using cDNA and SYBR green dye (Applied Biosystems). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the internal control and the 2−ΔΔCt method was used for analysis. Primers were as follows: VEGF Forward:
Western blotting analysis
Total protein was extracted from ARPE-19 cells by CytoBuster™ Protein Extraction Reagent (Novagen, Madison, WI, USA), then separated by 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis. Proteins in gels were transferred onto polyvinylidene difluoride membranes (Millipore, Burlington, MA, USA). Membranes were blocked with 1% bovine serum albumin, then incubated with primary antibody (described below) overnight at 4°C. Membranes were then incubated with horseradish peroxidase-conjugated anti-rabbit IgG secondary antibody (Abcam, Cambridge, MA, USA) at room temperature for 1 hour. Enhanced Chemiluminescence Reagent (Bio-Rad Laboratories, Hercules, CA, USA) was used to visualize immunoreactive bands. Primary antibodies were as follows: rabbit anti-VEGF (sc-7269; Santa Cruz Biotechnology), rabbit anti-GAPDH (5174; Cell Signaling Technology, Danvers, MA, USA), rabbit anti-CD9 (8457; Cell Signaling Technology), and rabbit anti-CD63 (sc-15363; Santa Cruz Biotechnology). Protein levels were quantified by ImageJ software (National Institutes of Health, Bethesda, MD, USA).
Luciferase reporter assay
VEGF 5ʹ-untranslated region fragments of VEGF, containing the wild-type binding site of miR-377-3p or a corresponding mutant fragment, were cloned into a luciferase reporter vector (GeneCopoeia, Rockville, MD, USA). ARPE-19 cells were seeded in 24-well plates and co-transfected with wild-type or mutant VEGF 5ʹ-untranslated region vector and miR-377-3p mimic or miR-negative control. After cells had been incubated for 48 hours, they were collected for the measurement of luciferase activity using the Luc-Pair™ Luciferase Assay Kit (GeneCopoeia).
miRNA microarray analysis
To investigate the miRNA expression profile, microarrays were performed as previously described. 14 In brief, 100 ng of total RNA was used to construct miRNA libraries, which were then investigated with the Human microRNA Microarray Kit (version 3; Agilent Technologies, Santa Clara, CA, USA), in accordance with the manufacturer’s instructions. Microarray data were processed for standardization, quality evaluation, and statistical analysis. The 75% quartile method was used for normalization. Analysis of miRNAs that regulate VEGF expression was performed using Targetscan Human 7.2 (www.targetscan.org/vert_72/).
Statistical analysis
Differences in miRNA expression levels between sample groups were determined using the Mann–Whitney U test. Comparisons of patient characteristics and gene expression levels (real-time polymerase chain reaction and western blotting analyses) were performed using Student’s t-test. Differences in cell viability between two groups were determined by repeated-measures analysis of variance. Data are presented as means ± standard deviations. GraphPad Prism, version 5 (GraphPad Software Inc., San Diego, CA, USA) was used for data analysis. Differences with P < 0.05 were considered statistically significant.
Results
Exosomes from patients with T2D downregulate VEGF expression in ARPE-19 cells
Exosomes were isolated from patients with T2D who did (n = 20) and did not have DME (n = 24). Clinical characteristics of all included patients are shown in Table 1; only central subfield thickness significantly differed between groups (P = 0.002). Exosomes isolated from different patients were subjected to nanoparticle tracking analysis, which revealed an acceptable size range (Figure 1a). CD9 and CD63 are markers of exosomes. 15 Using western blotting, we confirmed expression of both CD9 and CD63 in exosomes randomly picked from six patients (Figure 1b). Transmission electron microscopy revealed that serum exosomes exhibited a typical saucer-like spheroid morphology (Figure 1c).
Characteristics of patients with T2D alone and patients with T2D and DME.

All values are shown as mean ± standard deviation or n (%).
BMI, body mass index; DME, diabetic macular edema; HbA1c, glycated hemoglobin; T2D, type 2 diabetes.

Serum exosomes isolated from patients with T2D and DME serum led to upregulation of VEGF expression, compared with exosomes from patients with T2D alone. (a) Nanoparticle tracking analysis showed that the size of exosomes was within a normal range. (b) Transmission electron microscopy revealed that exosomes exhibited a saucer-like morphology. Scale bar = 100 nm. (c) Western blotting results demonstrated that exosomal markers CD9 and CD63 were expressed in exosomes isolated from patient serum. (d) ARPE-19 cells treated with exosomes isolated from patients with T2D and DME showed greater VEGF expression, compared with cells treated with exosomes isolated from patients with T2D alone (left graph, mRNA expression; middle panel and right graph, protein expression). **P < 0.01.
To determine the effects of exosomes on VEGF expression, ARPE-19 cells were treated with exosomes isolated from patients with T2D and DME or patients with T2D alone. Real-time polymerase chain reaction and western blotting analyses showed that VEGF expression was significantly greater in exosomes isolated from the serum of patients with T2D and DME than in exosomes isolated from the serum of patients with T2D alone (both analyses, P < 0.01; Figure 1d). These results implied that exosomes from patients with T2D and DME could not inhibit VEGF expression.
miR-377-3p is downregulated in patients with DME
miRNAs play a pivotal role in regulating protein expression. 16 To determine the miRNA expression profiles in patients with DME, we isolated serum exosomes from 10 patients with T2D alone and nine patients with T2D and DME; the characteristics of these randomly selected patients are shown in Table 2. miRNA microarray analyses showed substantially altered miRNA expression levels between patients with T2D alone and those with T2D and DME (Figure 2a). Because of the pivotal role of VEGF in DME progression, 12 we searched for miRNAs that regulate VEGF expression from among the top 10 miRNAs that were significantly downregulated (Figure 2b); only miR-377-3p was predicted to directly regulate VEGF expression, based on analysis using Targetscan Human 7.2 (www.targetscan.org/vert_72/). Further analysis by real-time polymerase chain reaction revealed that miR-377-3p expression was significantly lower in exosomes from patients with T2D and DME than in exosomes from patients with T2D alone (P = 0.014; Figure 2c).
Subgroup characteristics of randomly selected patients with T2D alone and patients with T2D and DME who were included in the microarray analysis.

All values are shown as mean ± standard deviation or n (%).
BMI, body mass index; DME, diabetic macular edema; HbA1c, glycated hemoglobin; T2D, type 2 diabetes.

miR-377-3p was downregulated in exosomes isolated from patients with T2D and DME. (a) miRNA microarray analysis revealed differentially altered miRNA expression levels between exosomes isolated from patients with T2D alone and exosomes isolated from patients with T2D and DME. (b) Top 10 downregulated miRNAs in exosomes isolated from patients with T2D and DME, compared with patients with T2D alone, based on p values. (c) TargetscanHuman 7.2 predicted that miR-377-3p regulated VEGF expression; real-time polymerase chain reaction analysis confirmed the downregulation of miR-377-3p in exosomes isolated from patients with T2D and DME, compared with patients with T2D alone. *P < 0.05
miR-377-3p negatively regulates VEGF expression
To confirm the relationship between miR-377-3p and VEGF, we conducted luciferase reporter assays. Transfection with a miR-377-3p mimic reduced luciferase activity associated with the wild-type VEGF promoter (P < 0.01), while transfection with a miR-377-3p mimic did not alter the luciferase activity associated with the mutant VEGF promoter in ARPE-19 cells (Figure 3a); these results implied a direct interaction between miR-377-3p and VEGF mRNA. Consistent with this hypothesis, overexpression of miR-377-3p led to reduced VEGF expression at both mRNA and protein levels (P < 0.01; Figure 3b). These results indicated that miR-377-3p negatively regulates VEGF expression.

miR-377-3p directly regulated VEGF expression in ARPE-19 cells. (a) Luciferase activity was significantly reduced in cells with WT VEGF promoter sequence, compared with NC; luciferase activity was not significantly affected by transfection with Mut VEGF promoter sequence. (b) Real-time polymerase chain reaction and western blotting analyses of ARPE-19 cells transfected with miR-377-3p or control miRNA revealed that miR-377-3p significantly reduced VEGF expression (left graph, mRNA expression; middle panel and right graph, protein expression). **P < 0.01.
miR-377-3p–VEGF signaling regulates proliferation of ARPE-19 cells
VEGF is an important factor involved in regulating angiogenesis and vascular permeability. In a high glucose environment, VEGF overexpression leads to RPE proliferation, while promoting angiogenesis and vascular permeability, thereby causing DME. 12 Here, we investigated the impact of miR-377-3p–VEGF signaling on RPE proliferation. Analysis using MTT revealed that miR-377-3p overexpression resulted in significant reduction of RPE proliferation (P < 0.01; Figure 4a,b). In contrast, VEGF restoration in miR-377-3p-overexpressing ARPE-19 cells resulted in significant enhancement of RPE proliferation (P < 0.05; Figure 4a,b). These observations suggested that miR-377-3p–VEGF signaling regulates RPE proliferation.
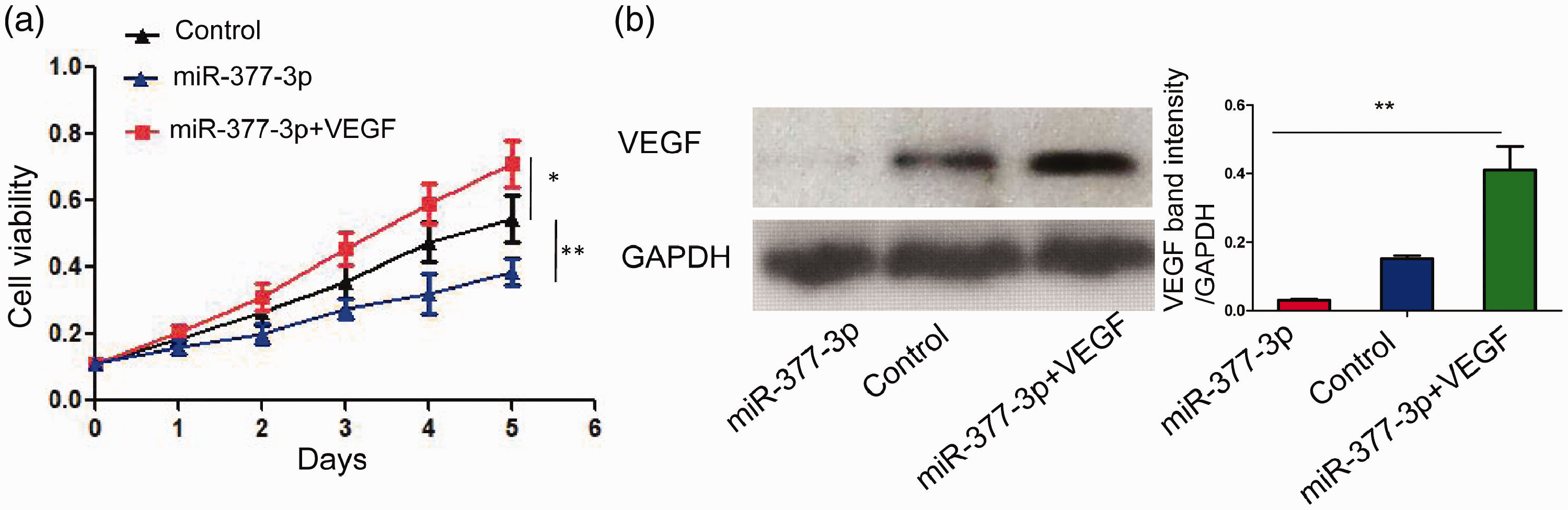
Reactivation of VEGF resulted in increased ARPE-19 cell proliferation. Cell viability assay analysis (a) showed that overexpression of miR-377-3p resulted in significantly reduced cell proliferation, compared with control cells. However, overexpression of VEGF in miR-377-3p-overexpressing cells (b) led to significantly enhanced cell proliferation (a). *P < 0.05, **P < 0.01.
Serum exosomal miR-377-3p offers a potential diagnostic biomarker for DME
To determine the diagnostic value of serum exosomal miR-322-3p, receiver operating characteristic curve analysis was performed to distinguish patients with T2D and DME from patients with T2D; the area under the curve was 0.7781 (95% confidence interval = 0.6383–0.9180, P = 0.001; Figure 5). These results indicated that serum exosomal miR-377-3p offers a potential diagnostic biomarker for DME.

Receiver operating characteristic curve analysis demonstrated that miR-377-3p was a diagnostic marker for DME (AUC = 0.7781).
Discussion
DME is a common complication of diabetes, which leads to substantial vision problems. 1 The main cause of DME is the onset of macular thickening. 17 VEGF overexpression plays a pivotal role in DME progression, but its mechanism and diagnostic role have remained elusive. Serum exosomal miRNAs have been identified as biomarkers of several diseases, although their value in DME has been unclear. In this study, we isolated serum exosomes from patients with T2D and DME, as well as patients with T2D alone. To determine the effects of exosomes from these patients on VEGF expression, we cocultured ARPE-19 cells with each type of exosomes. Importantly, VEGF expression was significantly upregulated in ARPE-19 cells cocultured with exosomes from patients with T2D and DME. We then compared miRNA profiles between exosomes from patients with T2D and exosomes from patients with T2D and DME, which revealed substantial differences. Analysis of the top 10 miRNAs downregulated in exosomes isolated from patients with T2D and DME revealed that miR-377-3p was likely to regulate VEGF expression; this prediction was confirmed by in vitro experiments, which showed that miR-377-3p could directly bind to VEGF and negatively regulate its expression. Proliferation assay analysis showed that miR-377-3p reduced RPE proliferation by inhibition of VEGF.
Previous studies have shown that miR-377-3p has an inhibitory effect on the development of other diseases;18,19 consistent with the prior findings, our data suggest that miR-377-3p may broadly prevent disease progression. Additionally, we revealed the potential value of exosomal miR-377-3p as a diagnostic biomarker in DME, based on the findings of receiver operating characteristic analysis.
A notable limitation of our study was that it only included patients with T2D (with and without DME); thus, the role of miR-377-3p in the progression from type 1 diabetes to DME requires further investigation. Another limitation was that serum exosomal miR-377-3p could not be fully validated as a diagnostic biomarker for DME; moreover, the isolation of serum exosomes is difficult. Future investigations should focus on the clinical value of miR-377-3p in larger numbers of patients and methods to simplify the isolation of serum exosomes. In summary, our study demonstrated that miR-377-3p can negatively regulate VEGF expression; moreover, miR-377-3p offers a diagnostic biomarker for DME.
Footnotes
Author contributions
HC, TD, MY, TM, HY, LJ, and XL carried out the studies, participated in collecting data, conducted cell experiments, and drafted the manuscript. HC and TD performed miRNA microarray analysis. MY analyzed microarray data. TM and HY isolated exosomes from serum. All authors read and approved the final manuscript.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This study was funded by the Basic Research Project of Shenzhen Municipal Health and Family Planning System Research (No. SZFZ20170).
