Abstract
Objective
To investigate the role of human leukocyte antigen (HLA-G) on proliferation, invasion, and immune escape in non-small cell lung cancer (NSCLC).
Methods
The relationship between HLA-G and overall survival (OS) of NSCLC patients was analyzed using the KMPlot database. The expression of micro (mi)R-152 or HLA-G was modulated by transfecting synthetic oligonucleotides, and the impact of the miR-152/HLA-G axis on proliferation, invasion, colony formation in soft agar, and tolerance to natural killer (NK) cell cytolysis was measured.
Results
Bioinformatics analysis showed that high HLA-G expression was correlated with poor OS in NSCLC patients. The tolerance of NSCLC cells to NK cytotoxicity was negatively correlated with HLA-G and positively correlated with miR-152 expression. Over-expressing miR-152 inhibited HLA-G expression in A549 cells and attenuated cell proliferation, migration, colony formation ability, and tolerance to NK cells. However, blocking HLA-G expression by small interfering RNA did not affect migration or colony formation, but only proliferation and tolerance to NK cells
Conclusions
miR-152/HLA-G axis plays an oncogenic role in NSCLC by affecting cell proliferation and immune escape.
Keywords
Introduction
Human leukocyte antigen (HLA)-G is one of the most important immune tolerance molecules. It was initially identified as functioning in fetal escape from maternal allorecognition, 1 but more recent research has shown it also interacts with immunoglobulin-like receptors to aid cancer cell escape from immune surveillance.2,3 Several studies have shown that HLA-G and its soluble form (sHLA-G) have prognostic value in non-small cell lung cancer (NSCLC), with the aberrant expression of HLA-G detected in the majority of NSCLC primary lesions or serum samples.4,5 Indeed, sHLA-G was reported to be an independent prognostic factor of overall survival (OS) in NSCLC, and decreased serum sHLA-G levels were associated with prolonged OS. 6 Additionally, serum sHLA-G was increased in advanced-stage NSCLC regardless of the histopathological type. 7
Recent studies have demonstrated that HLA-G is under the control of several microRNAs (miRNAs) including miR-152. For example, the delivery of miR-152 into the uterus was shown to counteract HLA-G expression and serve as a potential contraceptive therapy. 8 Additionally, hepatitis B virus infection increased tolerance to the anti-tumor immune response by down-regulating miR-152, which led to HLA-G up-regulation. 9 Several studies have found that serum miR-152 levels are associated with prognosis in NSCLC and could be used as a biomarker.10–12 However, the regulatory mechanisms of HLA-G in NSCLC are not fully understood.
Therefore, our study explored the relationship between HLA-G expression and the prognosis of NSCLC patients in a meta-analysis; on this basis, we further investigated the effect of HLA-G under the regulation of miRNA-152 on NSCLC cell proliferation, invasion, and tolerance to the cytolysis of natural killer (NK) cells. Our results showed that the miR-152/HLA-G axis plays an oncogenic role in NSCLC by affecting cell proliferation and immune escape.
Materials and Methods
Ethics and consent statements
The animal study protocol was approved by the Institutional Animal Care Committee of the Central Hospital of Wuhan (approval date: July 2020). All animal work was carried out in accordance with institutional animal care and use guidelines.
Kaplan-Meier analysis of NSCLC patients
The prognostic effect of HLA-G in NSCLC was analyzed using the public microarray database KMPlot (www.kmplot.com) with aggregate human patient data. 13 The OS probability was analyzed by Kaplan–Meier analysis with an auto-selected best cutoff and JetSet best probe. Statistical significance was analyzed using the two-tailed log-rank test incorporated in the KMPlot database. The Affymetrix ID for HLA-G is 211528.
Cell lines and culture
Three NSCLC cell lines were used in this study: A549 (human lung adenocarcinoma), L78 (human squamous lung cancer), and H460 (human large cell lung cancer). They were maintained in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS). The NK-92 cell line derived from human peripheral blood mononuclear cells was cultured with 75% α-MEM with the following supplements: 12.5% FBS, 12.5% horse serum, 0.1 mM 2-mercaptoethanol, 0.02 mM folic acid, and 200 U/mL recombinant interleukin-2 (IL-2). All cells were obtained from the American Type Culture Collection (Manassas, VA, USA), and were cultured at 37°C in a humidified incubator with 5% CO2. Basic medium and serum were purchased from Thermo Fisher Scientific (Gaithersburg, MD, USA). Recombinant IL-2 was purchased from Changchun Institute of Biological Products Co., Ltd. (Changchun, China).
Transfection
A549 cells were seeded in 6-well plates and grown to 80% confluency. To modulate miR-152 expression levels, they were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions with 50 nM of the following: single strand miR-152 mimics (5′-TCAGTGCATGACAGAACTTGG-3′), single strand miR-152 inhibitor (5′-CCACGUUCUGUCAUGCACUGA-3′), or mock control (5′-CAGUACUUUUGUGUAGUACAA-3′). For HLA-G knockdown, cells were transfected with 10 nM of small interfering (si)RNA or a non-targeting siRNA scramble control. siRNA sequences were: sense 5′-AUGGUGGGCAGGGAAGACUGCUUCC-3′ and antisense 5′-GGAAGCAGUCUUCCCUGCCCACCAU-3′ for HLA-G knockdown; and sense 5′-UUCUCCGAACGUGUCACGUTT-3′ and antisense 5′-ACGUGACACGUUCGGAGAATT-3′ as the negative control. Oligonucleotide sequences were obtained from GenePharma (Shanghai, China).
RNA extraction and quantitative real-time (RT)-PCR
Total RNA was isolated from cells using TRIzol reagent (Invitrogen). For the detection of HLA-G, 100 ng of RNA was reverse-transcribed into cDNA using the ReverTra Ace qPCR RT kit (Takara Biomedical Technology, Beijing, China). For the detection of miR-152, small RNA was enriched using the mirVna miRNA Isolation kit (Ambion, Austin, TX, USA). HLA-G and miR-152 expression was quantified by real-time PCR using the ΔCT method 14 and SYBR green-based PCR master mix (Takara). Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and small nuclear RNA U6 were used as endogenous references. Specific primers were designed as previously reported15,16 and purchased from GenePharma, and PCR reactions were performed on an ABI 7300 system (Applied Biosystems, Foster City, CA, USA). PCR conditions were initial denaturation at 95°C for 10 minutes, then 40 cycles of 95°C for 10 s and 60°C for 45 s.
Enzyme-linked immunosorbent (ELISA) assay
Lung cancer cells were seeded in 96-well plates at a density of 1 × 104 cells/well and grown to 80% confluency. They were then cultured in serum-free medium for 48 hours and the supernatant was collected for ELISA analysis. The ELISA kit was purchased from Shanghai Enzyme-Linked Biotechnology Co., Ltd. (Shanghai, China).
Cell proliferation assay
Cells were seeded in 96-well plates at a density of 2 × 103 cells/well and growth was measured daily over the next 5 days using the MTT Cell Proliferation and Cytotoxicity Assay Kit (Beyotime Biotechnology, Shanghai, China). Briefly, 10 μL of MTT solution (5 mg/mL) was added to each well and incubated for 4 hours. Then 100 μL of formazan solution was added to each well, mixed well, and incubated for a further 4 hours. Absorbance at an optical density of 570 nm was measured with a microplate reader (Model 680, Bio-Rad, Hercules, CA, USA).
Cytotoxicity assay
Lung cancer cells were seeded in 96-well plates at a density of 5 × 104 cells/well and allowed to adhere. NK-92 cells were then added at effector:target (E:T) ratios of 10:1, 5:1, and 2.5:1. 17 Wells with lung cancer cells or NK-2 cells only were used as target controls or effector controls, respectively. To block Ig-like transcript (ILT)-2 of NK cells, which is an HLA-G receptor, 10 μg/mL of mouse anti-ILT2 (Biolegend, San Diego, CA, USA) was applied. The same volume of phosphate-buffered saline (PBS) was used as a negative control. Cells were cultured for 4 hours and the living cell density was measured using the MTT kit described above. The lysis rate was calculated as (1 – (ODE+T – ODE)/ODT) × 100%.
Transwell analysis
Cells were suspended in serum-free medium and seeded in 8 μm Millicell inserts (Millipore, Billerica, MA, USA) at a density of 3 × 104 cells/insert. The inserts were incubated in 24-well plates with complete medium in the lower chambers for 24 hours. Cells that traversed to the lower side of the insert filter were fixed with formalin and stained with 0.1% crystal violet. Five random fields (under ×400 magnification) of each well were then counted using a CX31 light microscope (Olympus, Tokyo, Japan) to determine migrated cell numbers.
Soft agar colony formation
Complete medium was mixed with 0.5% agar and added to 24-well plates to form a gel. Cells were suspended in medium with 0.3% agar and adjusted to 5 × 104 cells/mL. For each well, 500 μL of cell suspension was added and incubated for 10 days. Formed colonies were stained with MTT solution and five random fields/well were counted under microscopy.
Western blotting
Whole cell proteins were prepared by lysing cells in radioimmunoprecipitation assay buffer (Beyotime). Protein concentrations were determined by the BCA protein assay kit (Beyotime). Proteins (30 μg/sample) were fractionated by 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred onto nitrocellulose membranes (Amersham, Piscataway, NJ, USA). Membranes were blocked with 5% skim milk for 1 hour at room temperature, then incubated with mouse anti-HLA-G (dilution 1:1500; Abcam Inc., Cambridge, UK) or mouse anti-GAPDH (dilution 1:5000; Abcam) primary antibodies overnight at 4°C. The next day membranes were incubated with horseradish peroxidase-conjugated anti-mouse secondary antibody (dilution 1:5000; Abcam) at room temperature for 2 hours. Hybridized protein bands were detected with the enhanced chemiluminescence kit (Beyotime).
Dual-luciferase reporter assay
The potential binding sequence (HLA-G-WT) or truncated binding sequence (HLA-G-MUT) were cloned into the pGL3 firefly luciferase reporter vector (Ambion). HEK293 cells were seeded in 24-well plates at a density of 1 × 105 cells/well and co-transfected with 100 ng pGL-3 reporter constructs, 20 ng pRL-TK
In vivo xenograft assay
BALB/c nude mice (5-weeks-old, purchased from Shanghai SLAC Laboratory Animal Co., Ltd., Shanghai, China) were raised in a specific pathogen-free facility with free access to clean food and water. Twenty-four mice were divided into four groups (six mice/group): A549 + NK-92; A549 + NK-92 + ILT2 antibody; A549 (si-HLA-G) + NK-92; and A549 (si-HLA-G) + NK-92 + ILT2 antibody. Normal A549 cells or A549 cells transfected with si-HLA-G were subcutaneously inoculated in the back of mice (1 × 106 cells in 0.1 mL PBS). After 2 weeks, mice were administered NK-92 cells (1 × 107 cells in 0.1 mL PBS) in the tail vein, and ILT2 antibody (10 mg/kg) 18 or control PBS were administered intraperitoneally once a week. Tumor volumes were calculated once a week using the formula: volume = length × width 2 × 0.5. After 4 weeks of NK cell injection, mice were euthanized under isoflurane anesthesia.
Statistical analysis
Data were analyzed using GraphPad Prism 8 software (San Diego, CA, USA). Continuous values were expressed as means ± SD. Comparisons between two groups were performed by the Student’s
Results
High expression of HLA-G correlated with poor OS in NSCLC
First we explored the KMPlot database for the
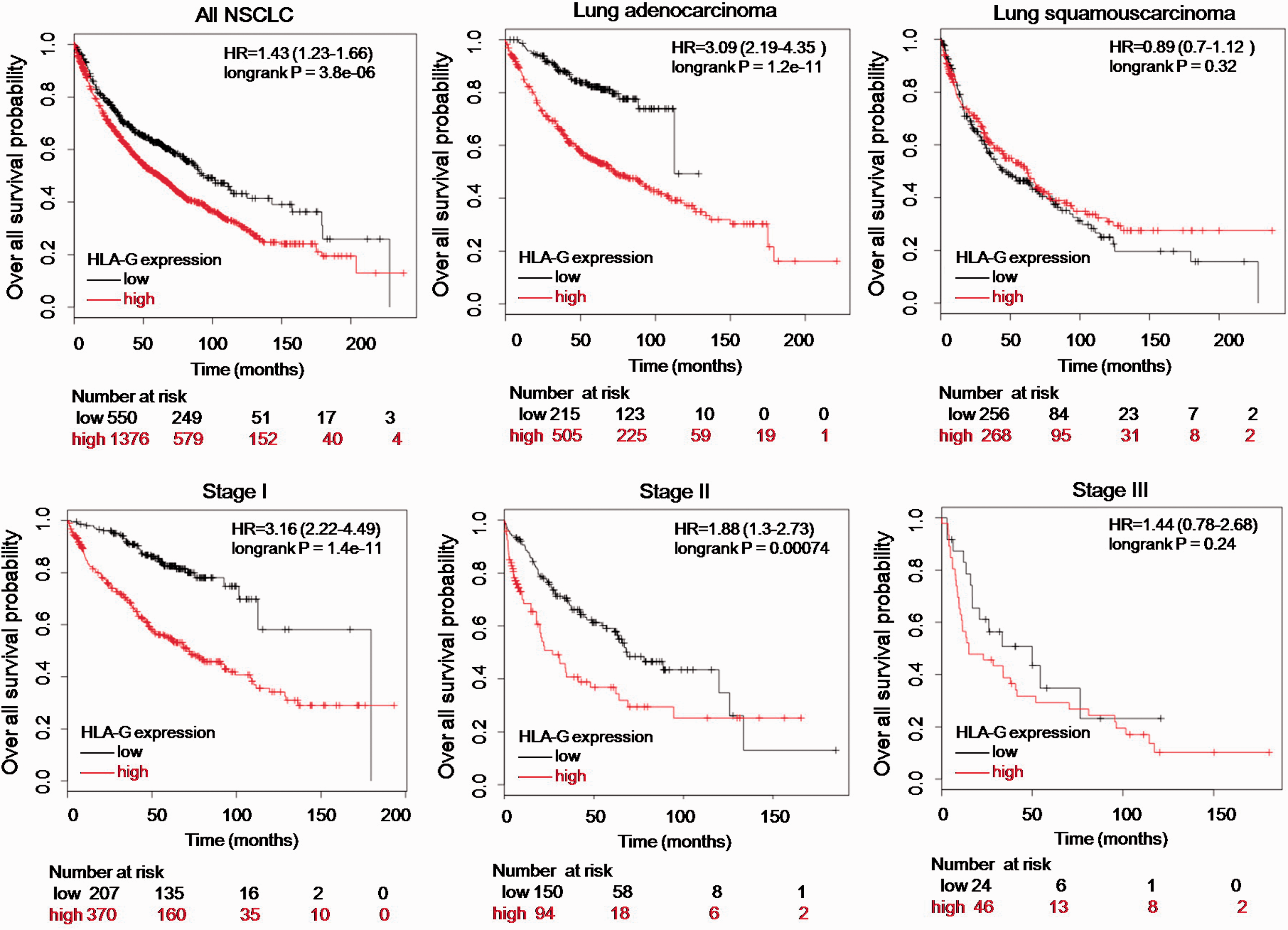
HLA-G is a prognostic marker in NSCLC. Kaplan–Meier curves were generated from the KMPlot database. An association between HLA-G and overall survival probability was shown in all patients and different histopathological subgroups.
HLA-G levels were associated with miR-152 expression and tolerance to NK cytolysis
We detected the expression of HLA-G and miR-152 in three representative NSCLC cell lines. As shown in Figure 2a, HLA-G and miR-152 were detected in all three cell lines. HLA-G mRNA expression was significantly higher in A549 cells than in the other cell types (

HLA-G is associated with miR-152 expression and tolerance to NK cytolysis. (a) Relative expression of HLA-G mRNA and miR-152 in different NSCLC cell lines. (b) Cellular HLA-G protein level and (c) sHLA-G level in three cell lines. (d) Cytolytic activity of NK-92 cells against three cell lines. *
NK cells are key effector lymphocytes in the tumor microenvironment,
19
so we next tested the tolerance of the three NSCLC cell lines to NK-92 cells. As shown in Figure 2d, the cytolysis activity of NK-92 cells against NSCLC cell lines decreased significantly with the decline of the E:T ratio (
miR-152 inhibited malignant characteristics of NSCLC cells
Previous studies showed that HLA-G expression is directly regulated by miR-152.8,9 We tested this by performing a dual-luciferase reporter assay (Figure 3a and 3b). As shown in Figure 3c–e, both cellular and soluble HLA-G levels were negatively regulated by miR-152 transfection. Overexpression of miR-152 inhibited A549 cell proliferation after 48 hours of incubation, while low expression of miR-152 significantly promoted A549 cell proliferation (Figure 4a, P < 0.001). miR-152 overexpression significantly reduced A549 cell migration through polycarbonate membranes (Figure 4b, P = 0.009) and colony formation in soft agar (Figure 4c, P < 0.001). Reducing the level of miR-152 in A549 cells was conducive to improving cell migration and colony formation.
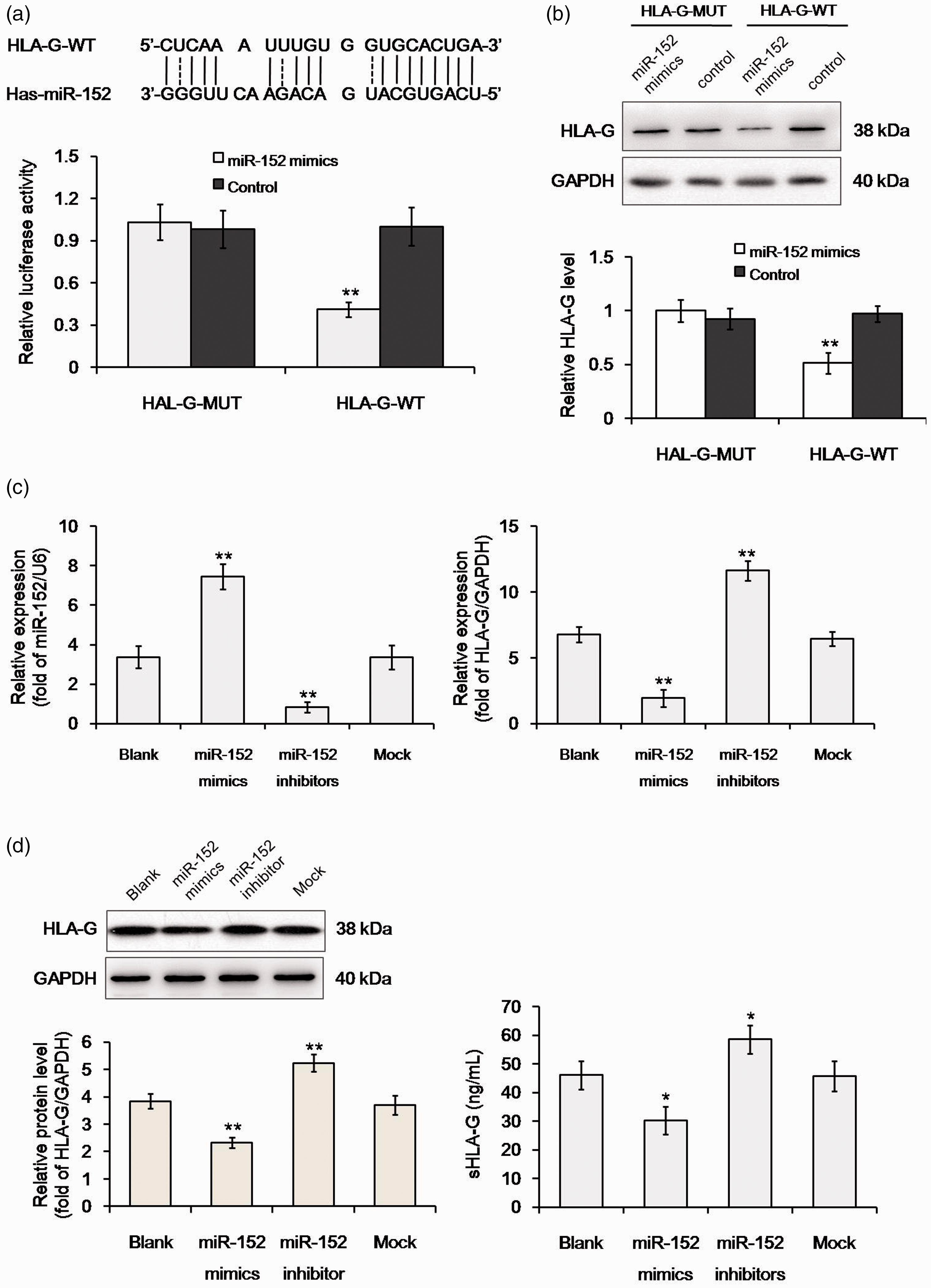
miR-152 regulates HLA-G expression in A549 cells. (a, b) Dual-luciferase assay confirming HLA-G as a direct target of miR-152. (c) Relative expression of miR-152 and HLA-G mRNA in different transfectants of A549 cells. (d) Cellular HLA-G protein level and (e) sHLA-G level in different transfectants. *

miR-152 inhibits malignant characteristics of A549 cells. (a) Effect of miR-152 on cell proliferation. (b) Effect of miR-152 on cell migration (magnification ×100, scale bar =200 μm). (c) Effect of miR-152 on colony formation. (d) Effect of miR-152 on tolerance to NK cells. *
The aberrant expression of miR-152 also influenced tolerance to NK cell cytolysis (Figure 4d). At the E:T ratio of 10, the cell lysis rate increased from 60.03% in the blank control group to 73.71% in the miR-152 mimics group (
HLA-G mediated the inhibitory effect of miR-152 on cell proliferation and tolerance to NK cytolysis
To investigate when HLA-G mediated the anti-tumor effect of miR-152, we knocked down HLA-G expression in A549 cells by siRNA technology. Figure 5 shows that both cellular and soluble HLA-G levels were significantly reduced in the si-HLA-G group (both

Knockdown of HLA-G expression in A549 cells. (a) Relative expression of miR-152 and HLA-G mRNA in different transfectants. (b) Cellular HLA-G protein level and (c) sHLA-G level in different transfectants. **
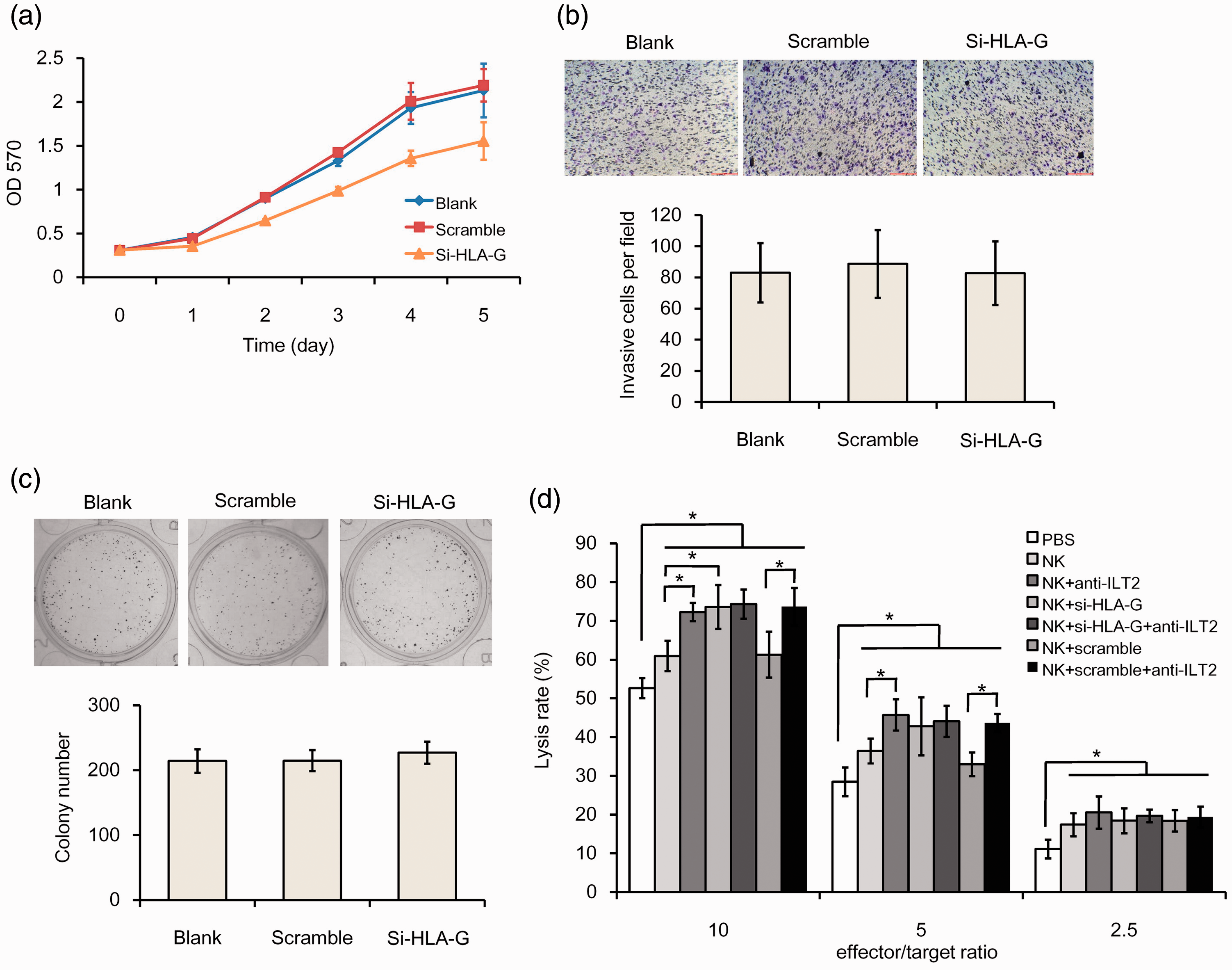
HLA-G affects proliferation and tolerance to NK cells
The cytotoxicity of NK cells depends on the binding of leukocyte Ig-like receptor 1 (LIT2) to the HLA-G ligand on the target cell surface. 20 We further blocked ILT2 of NK-92 cells by adding an ILT2 antibody to the culture medium and repeating the cytotoxicity assay. Compared with the PBS blank control, co-culture with NK cells significantly increased the lysis rate (Figure 6d, P < 0.05 at all E:T ratios). Interference of HLA-G expression significantly increased the cytotoxicity of NK cells at E:T=10. For cells with normal HLA-G expression, anti-ILT2 treatment also increased the lysis rate. However, the lysis rate was not further increased by anti-ILT2 when the expression of HLA-G was blocked. This suggested that the main role of HLA-G is not to affect the migration or colony formation of cancer cells, but to influence proliferation and immune escape.
To verify whether HLA-G affected the tolerance of tumor cells to NK cells
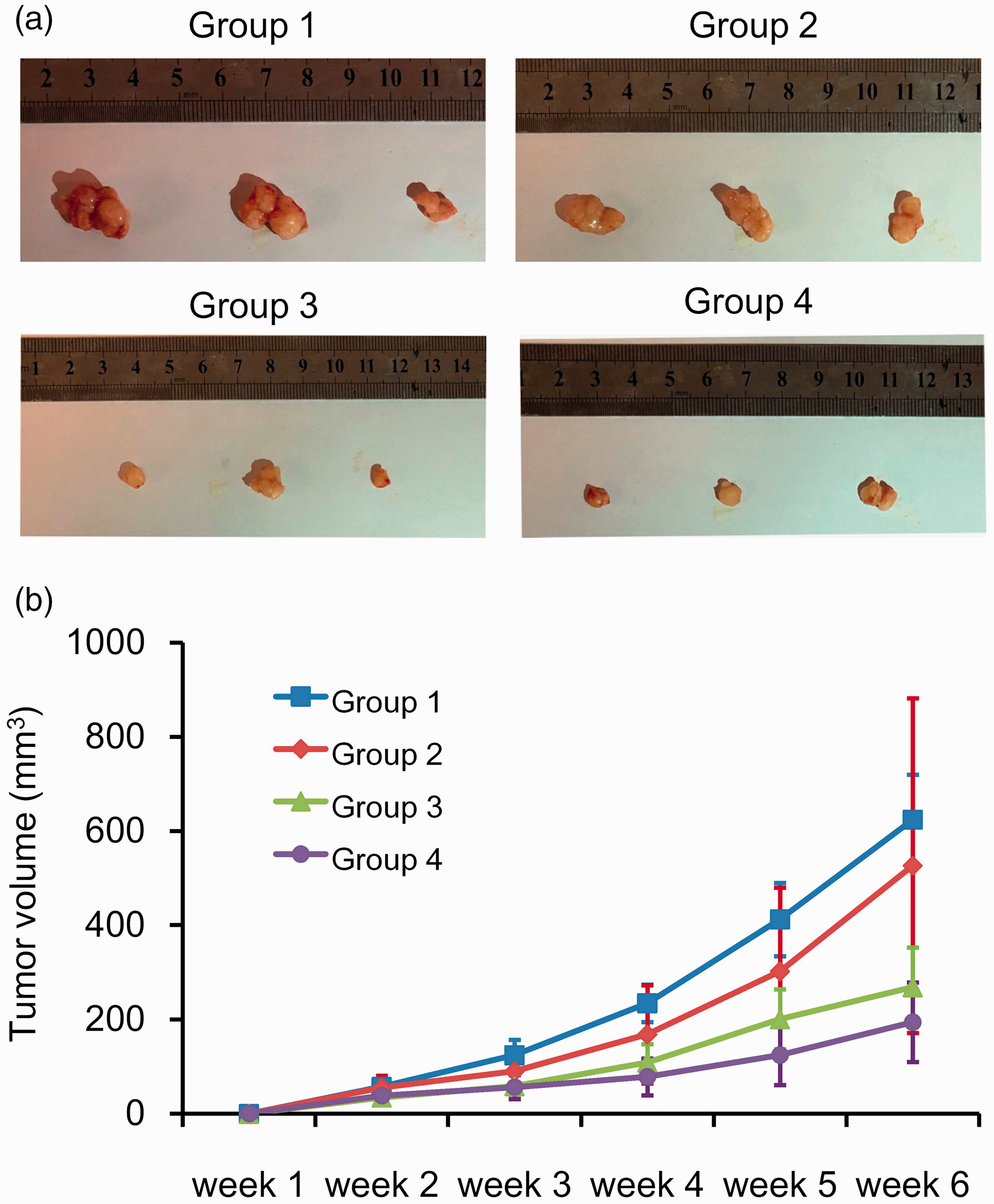
HLA-G affects proliferation and tolerance to NK cells
Discussion
In this study, we explored the KMPlot database with integrated genome-wide microarray data to show that HLA-G could be used as a prognostic marker for OS in NSCLC, especially in adenocarcinoma and stage I/II NSCLC. Further cellular experiments showed that high HLA-G expression, under the regulation of miR-152, was beneficial for tumor cells to proliferate and to escape NK cytolysis. A mouse xenograft assay further confirmed that HLA-G expression was associated with the growth and immune escape of A549 cells
Yie et al. 4 previously showed that HLA-G expression in NSCLC tissues was correlated with both survival time and disease stage. Similarly, a study by Lin et al. found that HLA-G expression in NSCLC lesions was associated with disease stage, but not with histological type, and that plasma sHLA-G rather than lesion HLA-G expression was correlated with OS. 5 Consistent with this, Amor et al. 7 reported an association between sHLA-G with disease stage and survival time, while Schutt et al. 6 found that HLA-G was exclusively elevated in NSCLC, especially in squamous NSCLC, but not in small-cell lung cancer. Taken together, these results suggest that HLA-G, especially sHLA-G, is associated with disease stage but the relationship between HLA-G expression and histological type is less clear. This may be related to the small sample size of a single study, albeit one with the strong evidence of immunohistochemistry and quantitative RT-PCR. The KMPlot database can merge multiple studies to achieve a larger sample size, but the relative evidence strength is only moderate because the data derive from microarrays. Our cellular assays showed that HLA-G expression in adenocarcinoma A549 cells was higher than in squamous carcinoma L78 cells and large cell carcinoma H460 cells, supporting the notion that HLA-G expression varies according to NSCLC histological type.
HLA-G is a non-classical human leukocyte antigen class I molecule which inhibits the activities of T lymphocytes, NK cells, and antigen-presenting cells.21–24 In this study, we found that when HLA-G expression was down-regulated in A549 cells by transfecting miR-152 mimics or siRNA, more A549 cells could be recognized and lysed by NK cells. A similar association between HLA-G expression and the NK cell-killing effect was reported in other cancers such as melanoma,
25
choriocarcinoma,
26
leukemia,
27
ovarian cancer,28,29 and hepatocellular carcinoma.
30
It was also found that the NK cytolysis of leukemia cells was dependent on their HLA-G expression level.
27
ILT2/HLA-G recognition is a dominant step during NK activation.
20
In this study we found that blocking ILT2 reduced NK cytolysis of A549 cells
Here we found that HLAL-G was controlled by miR-152 in NSCLC. Several previous studies reported that serum miR-152 had good biomarker potential for NSCLC screening and recurrence prediction in resectable NSCLC.10,11 Our work suggested that HLA-G mediated the regulation of miR-152 on A549 cell proliferation and tolerance to NK cytolysis. However, miR-152 also affected A549 cell migration and colony formation, which could not be explained in relation to HLA-G. Recent research revealed several other targets of miR-152 in NSCLC including fibroblast growth factor 2, which was down-regulated following miR-152 over-expression and suppressed migration and invasion in several NSCLC cell lines. 33 Additionally, miR-152 targeted neuropilin-1 to mediate A549 cell invasiveness, 34 and ADAM metallopeptidase domain 17 to suppress proliferation, colony formation, migration, and invasion. 35
The present study has some limitations. First, the regulatory relationship between miR-152 and HLA-G was only investigated in cellular experiments, not in clinical samples. This is because we did not accumulate enough clinical samples, nor did we conduct a sufficient follow-up at the beginning of this study. However, the clinical value of HLA-G and miR-152 is expected to be better evaluated in future work. Second, other immune cells, such as B cells, also express ILT2 on their surface. Therefore, future studies should block the expression of ILT2 in NK cells rather than using an anti-ILT2 antibody in the xenograft assay.
In conclusion, this study showed that HLA-G could serve as a prognostic marker in NSCLC. Molecular experiments suggested that HLA-G could be regulated by miR-152 and modulate the immune escape of A549 cells. These findings provide support for the use of HLA-G as a therapeutic target in NSCLC and a candidate immune therapy checkpoint.
