Abstract
Targeted therapy in patients with epidermal growth factor receptor (EGFR)-mutated non-small cell lung cancer (NSCLC) often fails because of drug resistance. Here, we report a 57-year-old male patient with stage IV small cell lung cancer (SCLC) transformation during targeted therapy. Chest computerized tomography (CT), hematoxylin and eosin histological examination, immunohistochemistry, allele refractory mutation system‐based quantitative polymerase chain reaction analysis of EGFR point mutations, and next-generation sequencing were performed for diagnosis and therapeutic efficacy evaluation. A combination of chest CT, histological examination, and immunohistochemistry confirmed the initial NSCLC diagnosis. Next-generation sequencing detected only EGFR exon 19 deletion (ex19del) before treatment and later identified EGFR exon20p.T790M point mutation, EGFR amplification, myc proto-oncogene (MYC) amplification, retinoblastoma 1 (RB1) mutation, and tumor protein 53 (TP53) mutation. Histology and immunohistochemistry revealed transformation from NSCLC to SCLC during treatment, which eventually returned to NSCLC. Drug resistance to targeted therapy for patients with NSCLC frequently occurs because of EGFR exon20p.T790M point mutation, TP53 mutation, RB1 mutation, and MYC amplification. These mutations are also the major determining factors of NSCLC outcomes. Therefore, next-generation sequencing should be performed to confirm drug efficacy during targeted therapy for NSCLC.
Keywords
Introduction
Lung cancer is the most lethal cancer worldwide. It is classified into non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC), which account for 85% and 15%, respectively. 1 Lung cancer therapies include surgery, chemotherapy, radiation therapy, targeted therapy, and immunotherapy. 2 Targeted therapy inhibits the activity of epidermal growth factor receptor (EGFR) or other specific proteins known to be mutated. 3 However, most patients with NSCLC rapidly acquire drug resistance to EGFR tyrosine kinase inhibitors (TKIs). 4 The mechanisms of acquired drug resistance are complex and include the development of new gene mutations, transformation of NSCLC into SCLC, 5 and combination of NSCLC and SCLC histological subtypes at diagnosis. 6 Here, we report a patient diagnosed with EGFR-mutated NSCLC, which later transformed into SCLC. Interestingly, the SCLC eventually transformed back into NSCLC. This case indicates that transformation between NSCLC and SCLC may be another mechanism of drug resistance in targeted therapy.
Case presentation
A 57-year-old man with a >20-year smoking history was admitted to our hospital on 9 June 2018 because of a cough for 3 months. A computerized tomography (CT) scan of his chest revealed a mass in the right middle lung, right pleural effusion, and multiple distant metastases of both lungs (Figure 1). Smear analysis of the right pleural effusion and paraffin sections of the tumor mass demonstrated a few positive thyroid transcription factor 1 (TTF1) and cytokeratin 7 (CK7) cancer cells. Hematoxylin and eosin (HE) staining of the tumor combined with immunohistochemistry (IHC) confirmed the presence of lung adenocarcinoma (Figure 2a) and tumor protein 53 (TP53) mutation (Figure 2b). Allele refractory mutation system‐based quantitative polymerase chain reaction analysis of cancer cells in tumor tissues removed during surgery showed EGFR exon 19 deletion (ex19del) (Figure 1 and Table 1). The patient was administered gefitinib for more than 1 year, and his mass dramatically diminished. However, he presented with chest tightness for 2 months and was admitted to our hospital on 16 September 2019. Enhanced CT showed that the cancer cells were extensively involved in the superior vena cava, lymph nodes on the clavicle of the lower neck, and right pleura. We performed a puncture cannula to drain the pleural effusion, diuresis, albumin supplementation, and antibiotic treatment. His symptoms improved. Furthermore, we performed next-generation sequencing (NGS) of peripheral blood samples using HiSeq 4000 NGS platforms (Illumina, Geneseeq, Nanjing, China). In addition to EGFR ex19del, myc proto-oncogene (MYC) gene amplification, retinoblastoma 1 (RB1) mutation, and TP53 gene mutation were found (Table 2). HE staining of a mass from the right lung showed SCLC (Figure 2c). IHC biomarkers indicated positive TP53 mutations (Figure 2d). These results confirmed SCLC transformation from NSCLC (Figure 1). After two-cycle etoposide plus carboplatin treatment, his symptoms improved. However, enhanced CT showed growth of the preexisting lung mass. Therefore, the regimen was changed to an irinotecan + nedaplatin protocol plus icotinib for four cycles. A CT scan showed that the previous mass was significantly decreased (Figure 1). Smear and IHC analysis of a puncture biopsy showed CK7 (+++), TTF1 (++), synaptophysin (−), and CD56 (−). These cells were identified as lung adenocarcinoma. On 5 June 2020, we performed NGS of peripheral blood samples as described above. The results showed multiple gene mutations, including EGFR ex19del, EGFR exon20p.T790M (C-to-T base pair change at the second nucleotide of codon 790 in EGFR exon 20), EGFR amplification, MYC gene amplification, RB1 mutation, and TP53 gene mutation (Table 3). While waiting for the NGS results, the patient showed salivation and unresponsive symptoms. Brain magnetic resonance imaging showed multiple nodular foci of various sizes on 13 June 2020 (Figure 1). The patient did not want to continue treatment and was discharged.

Treatment and evaluation timeline for the 57-year-old male patient with advanced lung cancer reported in this case.

HE and IHC staining of the tumor. a, staining of the primary NSCLC tumor; b, anti-TP53 mutant antibody staining of the primary NSCLC tumor; c, HE staining of SCLC; d, anti-TP53 mutant antibody staining of the SCLC tumor. All pictures were taken at a 200-fold magnification using a light microscope
EGFR mutations before treatment.

EGFR, epidermal growth factor receptor; WT, wild type.
Next-generation sequencing of a mass in the right lung.

EGFR, epidermal growth factor receptor; MYC, myc proto-oncogene; RB1, retinoblastoma 1; TP53, tumor protein 53.
Next-generation sequencing of peripheral blood.
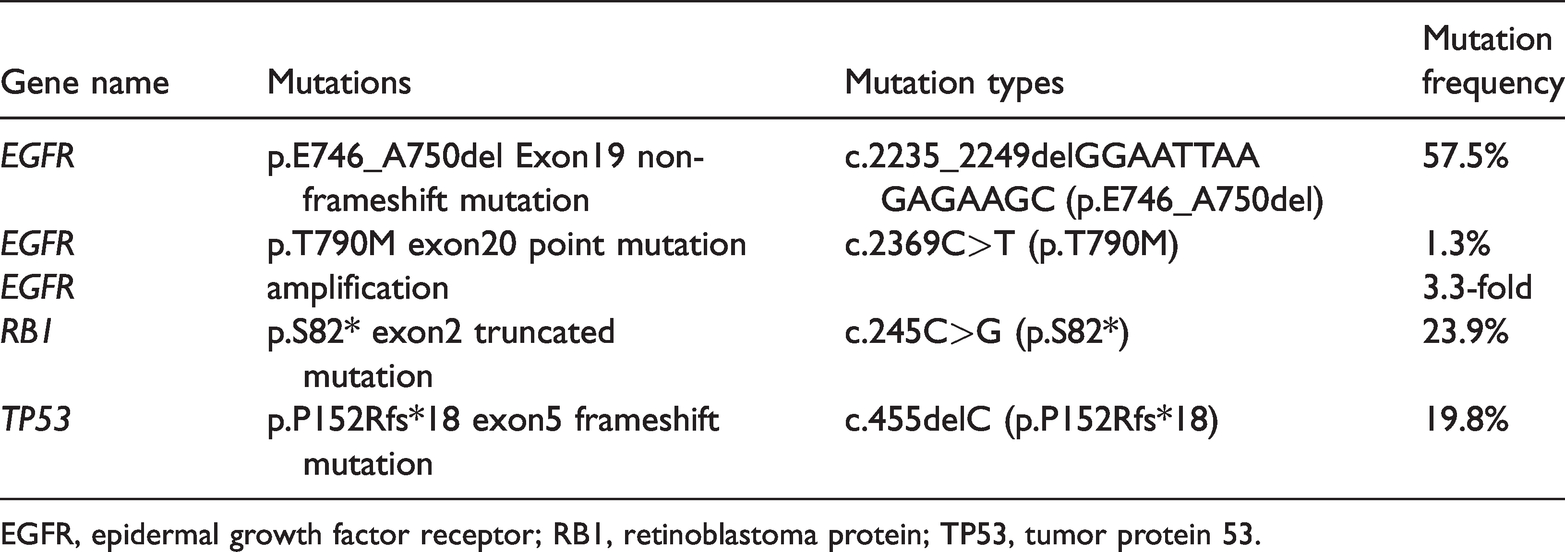
EGFR, epidermal growth factor receptor; RB1, retinoblastoma protein; TP53, tumor protein 53.
This report was reviewed by the Ethical Committees of the Cancer Hospital of Shantou University Medical College, and the need for approval was waived because this article is a case study. Written informed consent was obtained from this patient. The reporting of this study conforms to CARE guidelines. 7
Discussion
Here, we report a case in which NSCLC transformed into SCLC after TKI treatment and eventually returned to NSCLC. Several studies have revealed that the most frequent mechanisms underlying resistance to EGFR TKIs were the occurrence of EGFR gene mutations and SCLC transformation.1,3,8 The patient in our case had EGFR ex19del before his initial treatment. However, EGFR exon20p.T790M point mutation, EGFR amplification, MYC amplification, RB1 exon 2 truncated mutations, and TP53 exon 5 frameshift mutations were detected after gefitinib treatment. EGFR exon20p.T790M point mutation results from a C-to-T base pair change in codon 790 of exon 20 in the EGFR gene, which gives rise to an amino acid substitution from threonine to methionine. 9 This mutation is the most common resistance mechanism in patients treated with EFGR TKIs and increases the affinity of the receptor for ATP. 10 To date, the molecular mechanisms reported to contribute to SCLC transformation included EGFR mutation in patients with NSCLC and the combination of NSCLC and SCLC at the initial diagnosis.1,11 The concept of NSCLC and SCLC coexistence is supported by the idea that NSCLC and SCLC share stem cells of a common origin. In our case, we did not find SCLC cells at the initial diagnosis. However, we cannot exclude that this may be due to limited tissue collection.
NGS sequencing in patients with SCLC has revealed a high incidence of TP53 and RB1 mutations. 12 In addition, MYC amplification has been reported in patients with SCLC. 13 These data indicated that the loss of suppressor genes and oncogene activation are major drivers of SCLC transformation. Our data were consistent with these findings. In addition, after the development of resistance to EGFR TKI therapy, we detected EGFR exon20p.T790M point mutations. This mutation may drive the transformation of SCLC back into NSCLC, but future experiments are needed to support this claim.
