Abstract
Objective
Alterations in the structure and function of intervertebral discs by multifaceted chronic processes can result in intervertebral disc degeneration (IDD). The mechanisms involved in IDD are still unknown.
Methods
We investigated the possible mechanisms underlying IDD using a bioinformatics analysis of publicly available microarray expression datasets and built a circular RNA–microRNA–mRNA (circRNA–miRNA–mRNA) network based on the results. Datasets GSE67566 and GSE116726 were downloaded from the Gene Expression Omnibus (GEO) and analyzed using the limma package in R. The CircInteractome database was used to detect miRNAs related to circRNA, and TargetScan, miRDB, and miRTarBase were used to predict target mRNAs. Key target genes were annotated using Gene Ontology terms.
Results
The circRNA hsa-circ-0040039 was found to have the top log fold-change score. Analysis using Metascape showed that the associated genes were enriched mainly in the cell cycle. The Cytoscape plugin MCODE predicted that two members of the RAS oncogene family—RAB1A and RAB1B—and multiple coagulation factor deficiency (MCFD2) may play key roles in IDD.
Conclusion
Our results suggested that hsa-circ-0040039 and the related network may be potential biomarkers for IDD.
Keywords
Introduction
Intervertebral disc degeneration (IDD) is an orthopedic disease that affects 97% of individuals 50 years or older worldwide.1–3 IDD affects the stability of the lumbar region, leading to disc herniation and cervical spondylosis.4–6 Numerous factors have been associated with IDD, including nutrition, genetics, aging, and mechanical loading. 7 The pathological basis of IDD; that is, excessive apoptosis, leads to changes in cell composition and a decrease in active cells. 8 Predominant lower back pain as a result of IDD is difficult to reverse because of changes inside the cells and the cell matrix. 9 Hence, it is imperative to find new methods for IDD treatment. 10 IDD causes annulus fibrosus of endplate cartilage at the top and bottom of the vertebral disc and affects the nucleus pulposus at the center of the disc. A routine spine surgery technique, spine fusion, is widely used to treat cervical spine instability, intervertebral disc injury, spinal deformity, and lumbar spine degeneration. 11 The advent of new technologies in molecular biology such as gene therapy may help to make the treatment of IDD more efficient and effective. 12
RNA molecules that are covalently closed and have no 5′ cap structure or 3′ polyadenylated tail are called circular RNAs (circRNAs). 13 The rapid growth of RNA sequencing and bioinformatics has led to the discovery of a large number of circRNA structures with known functions. Four types of circRNAs have been characterized: exon circRNAs, intron circRNAs, exon–intron circRNAs, and intergenic circRNAs. 14 These circRNAs play crucial roles in many pathological and biological processes, including apoptosis, cell cycle, metastasis, invasion and migration, and cell proliferation. 15 CircRNAs act as microRNA (miRNA) sponges that mediate downstream gene expression. 16 Hence, circRNAs continuously provide a cushion to highly fluctuating miRNA effects, and circRNA–miRNA–mRNA networks are activated by overall gene expression that is directed to a specific disease process by the altered expression of circRNAs. 17 Various types of circRNAs, such as circSEMA4B, VMA21, and circRNA-104670, have been shown to play crucial roles in IDD. Several diseases have been linked to the critical function of the circRNA–miRNA–mRNA axis.18–20
Bioinformatics analysis and DNA sequencing methods have produced large amounts of data and led to the discovery of genomic mutations related to cancer. 21 Critical roles that circRNAs, miRNAs, and mRNAs play in IDD also have been revealed.22,23 Accurate predictions of drug therapy mechanisms together with rapid screening of drug targets have become more realistic with the development of bioinformatics tools.
In this study, we used the microarray expression profile datasets GSE67566 and GSE116726 from the Gene Expression Omnibus (GEO) to obtain circRNAs, miRNAs, and mRNAs related to IDD. Gene ontology (GO) enrichment analysis was applied to the differentially expressed genes (DEGs). A novel circRNA–miRNA–mRNA network that may play an important role in disc degeneration was constructed. Hub genes from among the common DEGs were detected by constructing a protein–protein interaction (PPI) network.
Material and methods
Ethical statement
Ethical approval for this study was deemed unnecessary because only data from bioinformatics databases were used.
Microarray expression data
We selected two GEO microarray datasets (GSE67566 and GSE116726) to screen for differentially expressed circRNAs (DECs), differentially expressed miRNAs (DEMs), and DEGs in IDD tissues compared with normal tissues.24–26 The Agilent-070156 Human miRNA (GPL20712) platform was used for GSE116726, and the Agilent-069978 Arraystar Human CircRNA microarray V1 (GPL19978) platform (Agilent, Santa Clara, CA, USA) was used for GSE67566. The two datasets had eight control (normal) samples and eight IDD samples; GSE116726 had three control and three IDD samples, and GSE67566 had five control and five IDD samples (Table 1).
Details of the GSE67566 and GSE116726 microarray expression profile datasets.

IDD, intervertebral disc degeneration.
Data analysis to detect DEGs in the IDD tissues
The Bioconductor affy package (https://www.bioconductor.org/packages/release/bioc/html/affy.html) was used to analyze the annotation files for log2 conversion, standardization, and background correction of the raw data, and the limma package in R (https://www.R-project.org/) was used to screen the DEGs in the IDD tissues. P-values <0.05 and |logFC| >1 (where FC is fold change) were considered significant.
Prediction of miRNA target genes
The top highly upregulated circRNAs were used for further investigation. To predict potential interactions between the miRNAs and circRNAs, we used the CircInteractome database (https://circinteractome.nia.nih.gov/); 27 to predict the target genes of the identified miRNAs, we used starBase v3.020 (http://starbase.sysu.edu.cn/). Target genes were identified by the miRNA target prediction programs TargetScan (http://www.targetscan.org), miRDB (http://mirdb.org/), and miRTarBase (http://mirtarbase.mbc.nctu.edu.tw/php/index.php).28–31
Venn plot
We used the Venn diagram website (http://bioinformatics.psb.ugent.be/webtools/Venn/) to calculate and draw a diagram of overlapping genes. The result of target miRNA prediction was based on top upregulated DEC prediction and DEM from GSE116726. The overlapping genes from TargetScan, miRDB, and miRTarBase were considered miRNA target genes.
Gene Ontology and PPI analysis
The GO enrichment analysis was conducted using Metascape. 32 GO enrichment analysis has been applied extensively to gene products or to specific gene annotations. 33 To better understand the relationships among the enriched terms, we constructed a network plot by connecting terms that had similarities >0.3 to form edges. Twenty clusters were formed and the terms with the lowest P-values were selected, with a cap of 250 terms and no more than 15 terms per cluster. Cytoscape was used to visualize the network. 34 The PPI analysis for proteins encoded by each listed gene was conducted using the OmniPath (https://omnipathdb.org/), InWeb_IM (https://inbio-discover.intomics.com/map.html#search), and BioGrid (https://wiki.thebiogrid.org/doku.php/statistics/) databases. The subset of proteins that formed physical interactions with at least one other protein was contained within the resultant network. We used the molecular complex detection (MCODE) plugin in Cytoscape to identify the components of the densely connected network.35–37
Results
Identification of DEGs in IDD tissues
After processing the expression profile data and standardizing the gene expression, we screened the DEGs in the GSE67566 and GSE116726 datasets using bioinformatics tools (Table 1). A total of 970 DEMs were detected in GSE116726 and 105 DECs were detected in GSE67566 using the limma package. Volcano plots of the DEGs from each dataset are shown in Figure 1A and B.

Differentially expressed genes (DEGs) and microRNAs (miRNAs) in intervertebral disc degeneration (IDD) tissues. (a, b) Volcano plots of DEGs between IDD and normal tissues in the (A) GSE67566 and (b) GSE116726 datasets. Red dots indicate significantly upregulated genes in IDD; green dots indicate significantly downregulated genes in IDD; black dots indicate genes that are not differentially expressed. P < 0.05 and |log2 fold change| >1 were considered significant. (c) Venn diagram of the 970 differentially expressed miRNAs detected from GSE116726 and their overlap with miRNAs from CircInteractome. (d) Venn diagram of the miRNA target genes predicted by TargetScan, miRDB, and miRTarBase.
Circ-0040039 functions as a sponge of several miRNAs
The ability of circRNAs to sponge miRNAs and inhibit their activities has been shown in numerous studies.38–41 Analysis using the CircInteractome database (https://circinteractome.nia.nih.gov) identified two possible binding site types between hsa-circ-0040039 and its predicted target miRNAs. Twenty-three target miRNAs were identified; hsa-miR-370, hsa-miR-545, and hsa-miR-665 had both types of binding sites, 16 had one type of binding site and 4 had the other type of binding site (Table 2).
Predicted microRNA (miRNA) targets of hsa-circ-0040039.

Binding sites: hsa-miR-370, hsa-miR-545, and hsa-miR-665 had both types of binding sites, 16 had one type of binding site and 4 had the other type of binding site.
Venn plots
Among the 970 DEMs that were detected from GSE116726, five overlapped with miRNAs from the CircInteractome database, as shown in the Venn plot in Figure 1C. The five overlapping DEMs were hsa-miR-648, hsa-miR-638, hsa-miR-502-5p, hsa-miR-574-5p, and hsa-miR-662. Among the predicted miRNA target genes, 46 genes were predicted by all three prediction programs (TargetScan, miRDB, and miRTarBase; Figure 1D).
Gene Ontology and PPI
To investigate the biological functions of the DEGs associated with IDD, GO enrichment analysis was performed using Metascape. The results indicated that the target genes were significantly enriched in vesicle targeting, negative regulation of cell cycle G2/M phase transition, cell junction assembly, negative regulation of cell cycle, negative regulation of cellular component organization, synapse assembly, and skeletal system morphogenesis (Figure 2). The PPI analysis showed that RAB1A, RAB1B, and MCFD2 play important roles in IDD (Figure 3).
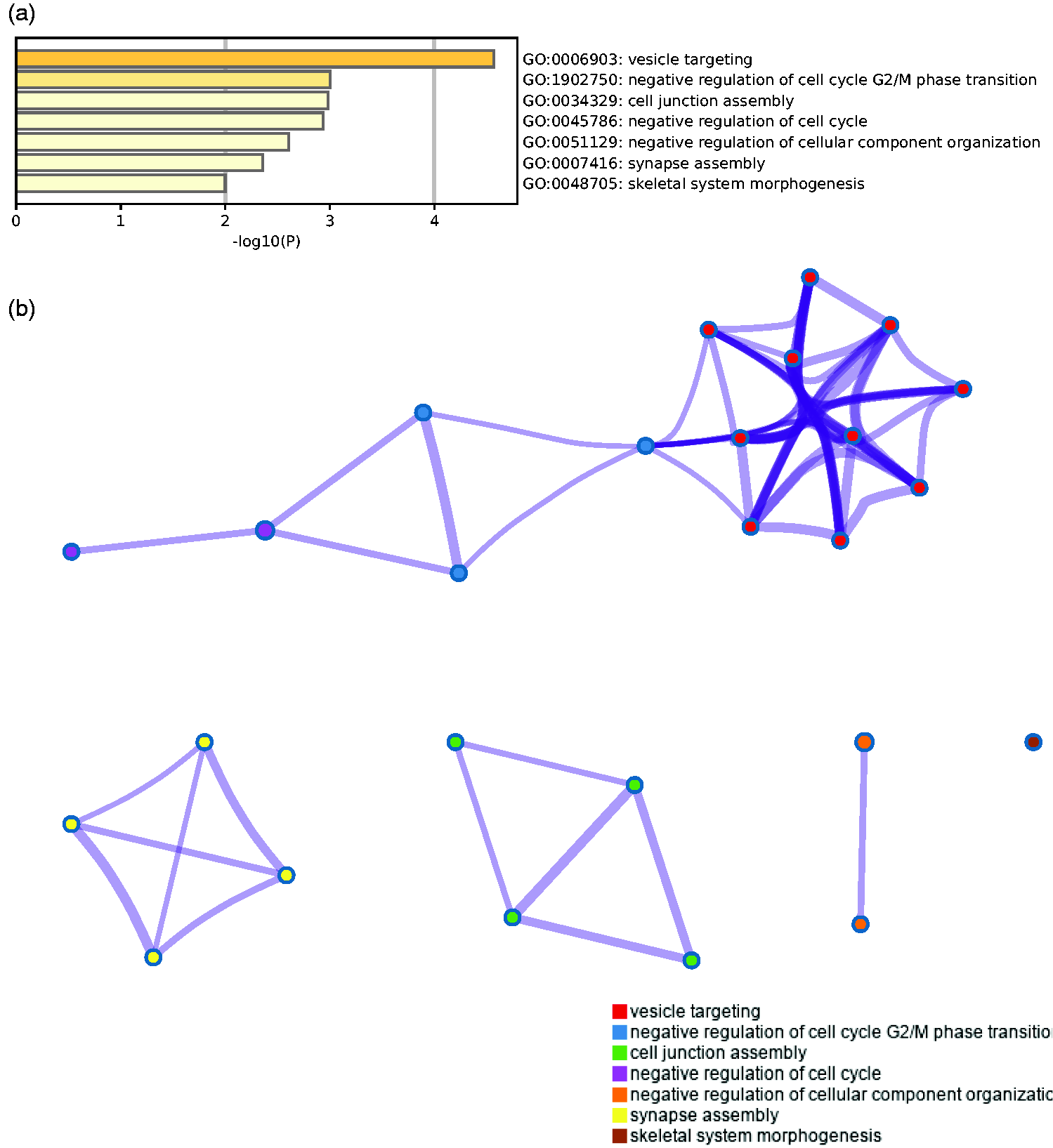
Gene Ontology (GO) enrichment analysis of the differentially expressed target genes associated with intervertebral disc degeneration tissues. (a) Heatmap of enriched GO terms assigned to the target genes. (b) Network of genes with enriched GO terms.

Protein–protein interaction of three proteins—two members of the RAS oncogene family (RAB1A, RAB1B) and multiple coagulation factor deficiency (MCFD2)—associated with intervertebral disc degeneration.
Discussion
IDD is considered a global threat to human health because of the significant associated healthcare costs and because it is a gateway to other disc-related diseases. 42 IDD is caused by a variety of traumatic, mental, and nutritional factors. 43 As well as the known behavioral and environmental factors, genetic factors have now been found to be associated with risk of IDD.44–47 Traditional treatments for disc degeneration include surgery and preventive treatments 10 . The recent development of biological techniques to repair IDD has provided new avenues of research and potential new treatments for IDD. It has been suggested that Wnt signaling may offer insight into targets for pharmacologic or nonpharmacologic therapeutics for IDD. 48 More biomarkers and related signaling pathways must be identified to further improve the treatment of IDD.
In this study, we downloaded microarray expression data from GSE67566 and GSE116726 and used it to detect differences between IDD and normal tissues. The identified DECs and DEMs were used to create a circRNA–miRNA–mRNA network. GO enrichment analysis showed that the mRNAs in the network were significantly enriched in vesicle targeting, negative regulation of cell cycle G2/M phase transition, cell junction assembly, negative regulation of cell cycle, negative regulation of cellular component organization, synapse assembly, and skeletal system morphogenesis. IDD has been shown to be closely associated with apoptosis of nucleus pulposus cells. 49 The pathology of IDD is influenced by decreases in the composition and synthesis of the extracellular matrix because of a reduction in the activity of the intervertebral disc cells due to increased cell apoptosis. 50 The PPI showed that RAB1A, RAB1B, and MCFD2 played important roles in disc degeneration. RAB1A and RAB1B are members of the RAS oncogene family and are associated with neoplasms, infections, and kidney diseases.51–54 MCFD2, RAB1B, and RAB1A were enriched mainly in vesicle targeting, which is associated with phenotype-keeping of intervertebral disc chondroid cells and enhanced proliferation of notochord cells. 55 Further studies are necessary to elucidate the actions of the circRNA–miRNA–mRNA network in IDD to understand their potential relevance to IDD.
In summary, we identified a novel circRNA–miRNA–mRNA network related to the progression and initiation of IDD through a comprehensive bioinformatics analysis of two GEO microarray datasets of IDD. The identified hub genes could have potential as biomarkers in the treatment, prognosis, and diagnosis of IDD. Their crucial role in the pathogenesis of IDD was confirmed further through a series of detailed analyses. However, the specific role of hsa-circ-0040039 in the pathogenesis of IDD needs to be further studied. Overall, our results shed light on the potential integration of multiple biomarkers for clinical practice in the diagnosis and treatment of IDD.
Supplemental Material
sj-pdf-1-imr-10.1177_0300060520960983 - Supplemental material for A novel circRNA–miRNA–mRNA network reveals hsa-circ-0040039 as a biomarker for intervertebral disc degeneration
Supplemental material, sj-pdf-1-imr-10.1177_0300060520960983 for A novel circRNA–miRNA–mRNA network reveals hsa-circ-0040039 as a biomarker for intervertebral disc degeneration by Jianhua Tang, Chenlin Zhang, Shengru Wang and Jianfeng Chen in Journal of International Medical Research
Footnotes
Acknowledgment
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Scientific Research Projects of Wuxi Health Planning Commission (grant number Z201703Z201703); and the Training Program of Innovation and Entrepreneurship for College Students in Jiangsu (grant numbers 201913993001Y, 201913993021H).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
