Abstract
Objective
We aimed to identify differentially expressed genes (DEG) in patients with inflammatory bowel disease (IBD).
Methods
RNA-seq data were obtained from the Array Express database. DEG were identified using the edgeR package. A co-expression network was constructed and key modules with the highest correlation with IBD inflammatory sites were identified for analysis. The Cytoscape MCODE plugin was used to identify key sub-modules of the protein–protein interaction (PPI) network. The genes in the sub-modules were considered hub genes, and functional enrichment analysis was performed. Furthermore, we constructed a drug–gene interaction network. Finally, we visualized the hub gene expression pattern between the colon and ileum of IBD using the ggpubr package and analyzed it using the Wilcoxon test.
Results
DEG were identified between the colon and ileum of IBD patients. Based on the co-expression network, the green module had the highest correlation with IBD inflammatory sites. In total, 379 DEG in the green module were identified for the PPI network. Nineteen hub genes were differentially expressed between the colon and ileum. The drug–gene network identified these hub genes as potential drug targets.
Conclusion
Nineteen DEG were identified between the colon and ileum of IBD patients.
Keywords
Introduction
Inflammatory bowel disease (IBD) is a chronic idiopathic disease that causes inflammation of the gastrointestinal tract, with a heterogeneous presentation.1,2 IBD has two subtypes: Crohn’s disease (CD) and ulcerative colitis (UC), which differ in presentation and treatment processes. 3 CD is a chronic granulomatous inflammation and the lesions may involve different parts of the gastrointestinal tract. 4 Lesions mainly involve the terminal ileum and adjacent colon. In contrast, UC is a chronic nonspecific colonic inflammation, 5 and UC lesions mainly involve the colonic mucosa and submucosa. Lesions may be present in the distal colon and can be retrograde to the proximal segment, even involving the whole colon and the terminal ileum. The latest epidemiological studies show that the incidence rate of IBD is increasing throughout the world. 6 The mechanism that determines the inflammatory location of IBD remains unknown.
Genetic and environmental factors are thought to be the main risk factors for IBD, contributing to abnormal activity in the intestinal immune system; however, the pathogenesis of IBD remains unclear. 7 Genome-wide association studies (GWAS) have identified common genes affecting innate defense or cellular homeostasis that are closely related to IBD.8,9 IBD patients often suffer from weight loss, bloody diarrhea, severe abdominal pain, and increased risk of cancer. 10 IBD gradually worsens over time, involves multiple recurrences, and persists for decades. 11 As a chronic, recurrent gastrointestinal disorder, IBD can have an adverse effect on mental health and quality of life, which may lead to suicidal behavior. 12 Therefore, it is important to clarify the molecular mechanisms of IBD that contribute to the development of clinical treatment decisions and reduce the burden of IBD in terms of clinical infrastructure and healthcare.
Gene expression changes affect many biological processes, including inflammation. It has been confirmed that T-helper (Th)17-related genes are differentially expressed between the inflammatory colon and ileum of patients with IBD, which suggests that gene expression patterns are regulated by different inflammatory sites. 13 Therefore, it is necessary to better understand gene expression differences between the colon and ileum of patients with IBD and determine whether this expression is regulated by different inflammatory sites.
High-throughput sequencing methods can simultaneously detect thousands of parameters for a single sample; however, systematically describing and analyzing these data and mining useful information remain challenging. Weighted gene co-expression network analysis (WGCNA) is a systematic biological tool for describing patterns related to gene expression in samples and it is widely applied in biomedical research because of its powerful analytical performance. 14 However, few studies using WGCNA have been conducted on IBD.
Our research aimed to take advantage of the Array Express public database to screen differentially expressed genes (DEG) between the inflammatory colon and ileum of patients with IBD by using gene co-expression analysis, and to determine whether intestinal region-specific gene expression is regulated by different inflammatory sites.
Materials and methods
Ethics approval and consent to participate
This study used primarily bioinformatics methods and did not require ethical review.
Data acquisition and preprocessing
RNA sequencing (RNA-seq) data (accession no: E-MTAB-7604) were obtained from the Array Express database (https://www.ebi.ac.uk/arrayexpress/), a public database of European Bioinformatics Institute (EBI) microarray gene expression data. 15 Total RNA-seq data contained 43 IBD samples (26 colon samples and 17 ileum samples from patients with CD and UC). We downloaded the corresponding clinical sample information. The 43 raw gene expression files were stored in corresponding text files, which were merged into an expression matrix for subsequent analysis by the R language (www.r-project.org).
Gene information for all genes with protein products was downloaded from the HUGO Gene Nomenclature Committee (HGNC; https://www.genenames.org/). 16 Based on the obtained matrix file, the expression values of all genes with protein products were extracted according to the obtained gene information, and this file was used for further analyses.
Screening of differentially expressed genes
We identified DEG between IBD patients with lesions located in the colon and IBD patients with lesion locations in the ileum (colon vs. ileum). After the raw expression data were normalized by trimmed mean of M-values (TMM) using the edgeR package, 17 we performed differential expression analysis (colon vs. ileum). The screening threshold of DEG was set to a false discovery rate (FDR) <0.01 and |logFC| >2 (where FC is the fold change), and differential expression results were visualized as volcano plots.
Functional enrichment analysis
To explore the biological functions and biological pathways involved in genes associated with IBD patients with different lesion sites, we performed Gene Ontology (GO) functional enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses through the String database (https://string-db.org/) and ClueGo+CluePedia (http://www.cytoscape.org/download.php). A
Construction of the gene co-expression network
WGCNA analysis was performed using the WGCNA package.
18
The WGCNA algorithm is commonly used to construct weighted gene co-expression networks. The data pretreatment was different from our previous data preprocessing method. The obtained expression matrix was subjected to log2(
To select the appropriate model, the different lesion sites of IBD (colon or ileum) were considered external biological parameters, and Student’s
Construction of protein–protein interaction network
Protein–protein interaction (PPI) suggests the processes by which two or more protein molecules form a protein complex through noncovalent bonds. PPI network analysis allows researchers to explore the molecular mechanisms of a disease or discover new drug targets. In the present study, we constructed a PPI network using the String database. 19 The minimum required interaction score was set to high confidence (0.7). The PPI network was visualized by using Cytoscape, 20 which was also used to calculate the connectivity of each node in the network. Nodes with a larger degree in the network have a greater role in the network.
Construction of key sub-module of PPI network
After construction of the PPI network, we used the MCODE plugin of Cytoscape to perform key sub-module analysis to obtain those sub-modules with biological significance in the PPI network. Node information of each node was calculated, including its neighbors, density, maximum k-value, the density of this k-core (core density), and the score value of the node. The score value of the node reflects the intensity of the node and its neighbors. Then, starting from the node with the largest score value, the getClusterCore() function was used to incorporate adjacent nodes that met the parameter conditions. Finally, the genes contained in the key sub-modules with MCODE score ≥5 were selected as hub genes for subsequent analysis.
Drug–gene interaction analysis
To explore whether a hub gene was therapeutically targeted or prioritized for drug development, the Drug–Gene Interaction database 2.0 (DGIdb2.0; http://www.dgidb.org/) was used to predict the obtained hub genes. 21 The parameter settings were as follows: FDA approved and the drug database was limited to DrugBank. Finally, the drug–gene interaction network was visualized through Cytoscape.
Visualization of expression of hub genes in different lesion locations
Differential expression analysis of hub genes in different lesion sites (colon or ileum) of IBD was presented as boxplots using the ggpubr package. The Wilcoxon test was performed between different lesion sites.
Validation analysis
RNA-seq data (accession no: E-MTAB-5464) from the Array Express database were used to validate the identified hub genes of IBD. The datasets included 52 IBD samples (25 CD and 27 UC). We compared the expression differences of these hub genes in the colon and ileum of IBD and in CD and UC using the Wilcoxon test.
Results
Identification of DEG between the colon and ileum of IBD
Volcano plots show the DEG (FDR <0.01 and |logFC| >2) between the colon and ileum of IBD (colon or ileum) including 86 upregulated and 293 downregulated genes (Figure 1a). To explore the biological processes and pathways involved in these DEG, functional enrichment analysis was performed using the String database. The top eight terms of GO, including biological process (BP), cellular component (CC), and molecular function (MF), KEGG, and Reactome are shown separately in Figure 1b.

Differential expression analysis and functional enrichment analysis. (a) Differential expression analysis (colon vs. ileum). Red indicates upregulated genes in the colon compared with the ileum of IBD and blue indicates downregulated genes in the colon compared with the ileum of IBD. (b) Functional enrichment analysis showing the top eight enrichment terms of differentially expressed genes, including GO (BP, CC, and MF), KEGG, and Reactome. IBD, inflammatory bowel disease; GO, Gene Ontology; BB, biological process; CC, cellular component; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes; DE, differential expression; FDR, false discovery rate.
Construction of the gene co-expression network
First, we checked the sample for outliers, which were found and removed from all samples (Figure 2a). Then, the appropriate power value was screened. When the soft threshold power value was equal to 14, scale independence reached 0.8 and mean connectivity was higher than zero (Figure 2b). Therefore, the soft threshold power value was set to 14 for subsequent analysis. As shown in Figure 2c, 11 co-expression modules were identified according to the soft threshold power value, in which the gray module represented a gene that was not assigned to any module. Different modules are represented by different colors. The eigengene adjacency heatmap was used to identify correlations between different modules (Figure 2d). Modules that were divided into one branch may have similar functions. Furthermore, we explored module–trait relationships. As shown in Figure 2d, the green module had the highest correlation (r = 0.7,

The gene co-expression network analysis. (a) Sample clustering to detect outliers. (b) Analysis of different soft-thresholding powers on the scale independence of co-expression modules. (c) Hierarchical clustering analysis showing each co-expression module; different modules are represented by different colors. (d) Eigengene adjacency heatmap (left) and module–trait relationships (right). The color change from blue (0) to red (0) in the heat map represents weak to strong connectivity of key genes in different modules. Module–trait relationships show the correlation between different modules and different lesion sites of IBD (colon or ileum). Each unit contains the corresponding correlation and
Identification of meaningful module with different inflammatory sites of IBD (colon or ileum)
To further identify meaningful modules with different lesion sites of IBD (colon or ileum), we verified modules in the gene co-expression network. As shown in Figure 3a, genes in the green module had the highest correlation with lymphovascular invasion. Moreover, the highest gene significance was found in the green module. Therefore, the green module was identified as meaningful for different inflammatory sites of IBD (colon or ileum). Membership in the green module was significantly related to gene significance for different lesion sites of IBD in Figure 3b (r = 0.71,

Identification of meaningful modules with different inflammatory lesion sites of IBD (colon or ileum) and functional enrichment analysis. (a) The correlation between different modules with lymphovascular invasion (top), and gene significance in different modules (bottom). (b) Module membership vs. gene significance in green module. (c) GO function enrichment and KEGG pathway enrichment analysis of genes in the green module. GO, Gene Ontology; BB, biological process; CC, cellular component; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes.
Identification of hub genes in the green module
To obtain DEG between the colon and ileum of IBD, the 379 DEG were intersected with 879 genes in the green module (Figure 4a), and 200 genes were obtained for PPI using the String database. In the PPI network, there were 133 nodes and 237 relationship pairs (Figure 4b). The different sizes of the nodes represented the degree of the node in the PPI network. The larger the circle, the greater the degree of the node in the network, indicating that the node was more important. After obtaining key genes from the PPI network, the MCODE plug-in was used to perform further key sub-module analysis. A sub-module with a MCODE score ≥5 was considered a key sub-module, and three key sub-modules were obtained. Sub-module 1 had a total of eight nodes, with an MCODE score of 8; sub-module 2 had a total of six nodes, with an MCODE score of 6; and sub-module 3 had a total of five nodes, with an MCODE score of 5 (Figure 4c). The 19 genes in the three key sub-modules were differentially expressed between the colon and ileum of IBD.

Identification of hub genes in the meaningful module. (a) Venn diagram showed the common genes between differentially expressed genes and genes in the green module. (b) PPI network constructed based on the common genes using the string database. (c) Analysis of key sub-modules in the PPI network using MCODE. DEG, differentially expressed genes; WGCNA, weighted gene co-expression network analysis; PPI, protein–protein interaction.
Functional enrichment analysis of hub genes and drug–gene network construction
To explore the biological functions and biological pathways involved in the 19 hub genes, functional enrichment analysis was performed. Figure 5a shows the top eight terms of GO, including BP, CC, and MF, KEGG, and Reactome using the String database. Pathway enrichment analysis of the 19 hub genes was performed using ClueGo+CluePedia. The 19 hub genes were mainly enriched in protein digestion and absorption, renin-angiotensin, vitamin digestion and absorption, cholesterol metabolism, and fat digestion and absorption (Figure 5b). To explore whether the 19 hub genes could be therapeutically targeted or prioritized for drug development, drug–gene relationships were predicted using the DGIdb2.0 database (Figure 5c). Among 19 hub genes, it was predicted that

Function enrichment analysis of hub genes and drug-gene network construction. (a) The top eight terms of GO including BP, CC, and MF, KEGG, and Reactome using the String database. (b) Pathway enrichment analysis using ClueGo+CluePedia. (c) The construction of drug–gene network. GO, Gene Ontology; BB, biological process; CC, cellular component; MF, molecular function; KEGG, Kyoto Encyclopedia of Genes and Genomes.
Expression of hub genes between the colon and ileum of IBD
We analyzed the expression of hub genes between the colon and ileum of IBD using the ggpubr package. We found that expression of

Boxplots showing the differential expression of hub genes between the colon and ileum of IBD. IBD, inflammatory bowel disease. The box shows the interquartile range, the line indicates the median, the whiskers indicate the minimum and maximum values, and symbols indicate the expression levels of genes.

Validation of the differential expression of hub genes between the colon and ileum of IBD in an independent dataset. IBD, inflammatory bowel disease. The box shows the interquartile range, the line indicates the median, the whiskers indicate the minimum and maximum values, and symbols indicate the expression levels of genes.
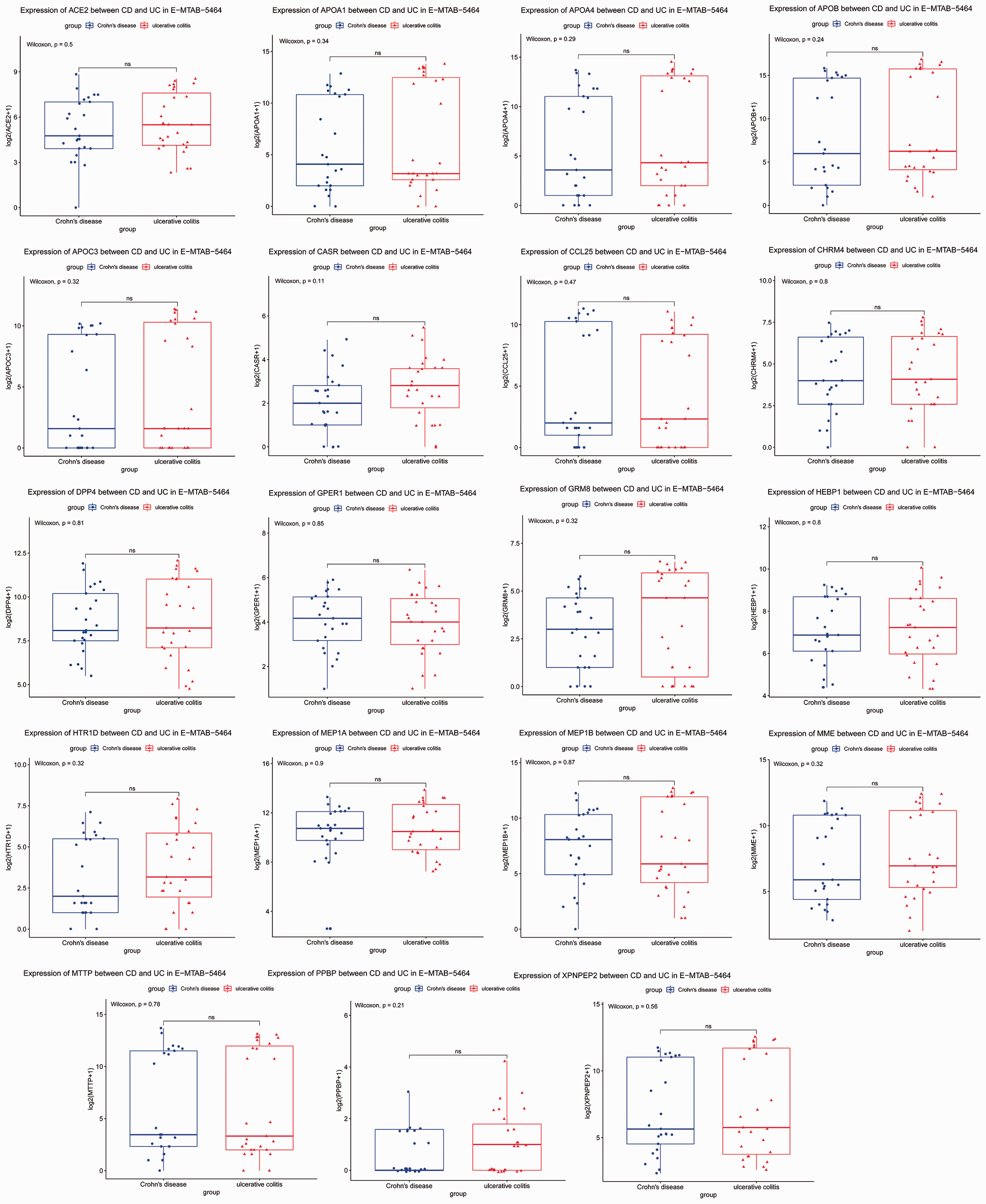
The differential expression of hub genes between samples from Crohn’s disease and ulcerative colitis. CD, Crohn’s disease; UC, ulcerative colitis. The box shows the interquartile range, the line indicates the median, the whiskers indicate the minimum and maximum values, and symbols indicate the expression levels of genes.
Discussion
The pathogenesis of IBD is extremely complex, involving multiple pathways and cytokines.22,23 The disease is unpredictable and incurable. 24 Therefore, understanding the mechanism underlying IBD would be helpful in finding the best treatment options as early as possible, which would improve long-term treatment results and quality of life in patients with IBD.25,26 Gene expression patterns are altered in different inflammatory sites of IBD. Therefore, identification of DEG between the colon and ileum of IBD may help us understand the pathogenesis of IBD.
Traditional biological research is based on a single gene or protein, so it can only explain the local part of a biological system, and it is difficult to comprehensively explore the whole system. In contrast, WGCNA is an algorithm for mining module information from high-throughput data. In this method, a module is defined as a group of genes with similar expression profiles. If some genes have similar expression patterns within a physiological process or in different tissues, there is reason to believe that these genes are functionally related. 27 The clustering criteria of WGCNA have biological implications, such as the use of geometric distances between data. 28 Therefore, the results obtained using this method have higher credibility and are more conducive to identifying gene functions and internal associations from the whole biological function. In the present study, after constructing the co-expression network, we identified the green module as meaningful, because it had the highest correlation with different lesion sites of IBD. To identify DEG between the colon and ileum of IBD, we took the intersection of DEG and genes in the green module. Among these, we identified complex relationships according to the PPI network. After obtaining key genes from the PPI network, 19 hub genes were identified using the MCODE plug-in.
In this study, we focused on hub genes associated with lesion sites of IBD. Previous studies have found that mutations in
Conclusion
In the present study, we identified 19 DEG of IBD, which are regulated by different inflammatory sites. These genes participate in biological processes of IBD and could become potential drug targets. These 19 DEG might be important in the development of IBD.
Footnotes
Authors' contributions
Yong Xie conceived and designed the study; Yuting Zhang conducted most of the experiments and data analysis and wrote the manuscript; Bo Shen and Liya Zhuge participated in collecting data and helped to draft the manuscript. All authors reviewed and approved the manuscript.
Availability of data and material
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was funded by National Key Research and Development Grogram of China (2016YFC1302201), the National Natural Science Foundation of China (81760105) and Special Funds of Graduate Student Innovation Project in Nanchang University (CX2016315).
