Abstract
Objective
The present study aimed to elucidate the underlying pathogenesis of Kawasaki disease (KD) and to identify potential biomarkers for KD.
Methods
Gene expression profiles for the GSE68004 dataset were downloaded from the Gene Expression Omnibus database. The pathways and functional annotations of differentially expressed genes (DEGs) in KD were examined by Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses using the Database for Annotation, Visualization and Integrated Discovery (DAVID) tool. Protein–protein interactions of the above-described DEGs were investigated using the Search Tool for the Retrieval of Interacting Genes (STRING).
Results
Gene Ontology analysis revealed that DEGs in KD were significantly enriched in biological processes, including the inflammatory response, innate immune response, defense response to Gram-positive bacteria, and antibacterial humoral response. In addition, 10 hub genes with high connectivity were selected from among these DEGs (ITGAM, MPO, MAPK14, SLC11A1, HIST2H2BE, ELANE, CAMP, MMP9, NTS, and HIST2H2AC).
Conclusion
The identification of several novel hub genes in KD enhances our understanding of the molecular mechanisms underlying the progression of this disease. These genes may be potential diagnostic biomarkers and/or therapeutic molecular targets in patients with KD. ITGAM inhibitors in particular may be potential targets for KD therapy.
Introduction
Kawasaki disease (KD), initially identified by the Japanese pediatric doctor Tomisaku Kawasaki, 1 is defined as an acute, multi-systemic vasculitis, particularly affecting children under 5 years old. KD is currently among the leading causes of acquired heart disease in children in developed countries. 2 It is clinically manifested by prolonged fever, cervical lymphadenopathy, polymorphous skin rashes, bilateral non-purulent conjunctivitis, peripheral extremity alterations, and diffuse mucosal inflammation. 3 Furthermore, approximately 15% to 25% of patients with untreated KD also have coronary artery lesions. With respect to therapy, high-dose and timely intravenous immunoglobulin administration, combined with aspirin during the acute phase, could significantly decrease the prevalence of CAL. 4 However, although several studies have analyzed the association between genes and KD susceptibility, the results have been inconsistent.
High-throughput platforms for gene expression analysis, such as microarrays, have been increasingly recognized as approaches with significant clinical value in areas such as molecular diagnosis, disease classification, patient stratification, prognostic prediction, identification of novel therapeutic targets, and response prediction.5,6 Microarray technology has been widely used in various gene expression profiling studies examining disease pathogenesis in the last decade, and has revealed many differentially expressed genes (DEGs) involved in various pathways, biological processes, and molecular functions. We therefore used a microarray dataset to investigate the pathogenesis of KD.
All original information in the present study was downloaded from the Gene Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/), which provides a hub for the deposition and retrieval of microarray data. We then examined the data using Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, and protein–protein interaction (PPI) network analyses.
Methods
Microarray data
Only one dataset (GSE68004) was identified following a search of the GEO database. Gene expression profiles from this dataset, which contained 76 patients with KD and 14 age-appropriate healthy controls, were downloaded from the GEO database and analyzed using the GPL10558 Illumina HumanHT-12 V4.0 expression beadchip platform. Blood samples (1±3 mL) were collected in Tempus tubes (Applied Biosystems, Foster City, CA, USA). All samples were stored at –20°C until further processing in batches. Whole blood RNA was processed and hybridized to Illumina Human HT12 V4 beadchips (47,323 probes) and scanned on the Illumina Beadstation 500 (Illumina, San Diego, CA, USA) as previously described. 7 All data were accessible online. No experiments involving humans or animals were conducted.
Data processing of DEGs
DEGs between KD and normal specimens were determined using the online analysis tool, GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/), followed by calculation of the adjusted P-value and |log fold-change (FC)|. Genes were defined as DEGs if they met the cutoff criteria of adjusted P < 0.05 and |logFC| ≥ 2.0.
GO and KEGG pathway analyses of DEGs
GO analysis is a widely used approach for investigating large-scale functional enrichment, in which gene function is simply categorized into cellular component (CC), molecular function (MF), and biological process (BP) terms. The KEGG database contains extensive data on genomes, drugs, chemical substances, and diseases, as well as biological pathways. GO and KEGG pathway enrichment were then analyzed using the Database for Annotation, Visualization and Integrated Discovery (DAVID) tool (https://david.ncifcrf.gov/). A value of P < 0.05 and gene count ≥ 10 indicated statistical significance.5,6 The GOCircle plot package was used to visualize the biological data from DAVID. 8
PPI network construction and hub gene identification
PPIs were established using the Search Tool for the Retrieval of Interacting Genes (STRING) database (http://string-db.org/). To assess possible PPI correlations, previously identified DEGs were mapped to the STRING database, followed by extraction of PPI pairs with a combined score >0.4. Cytoscape software (www.cytoscape.org/) was then employed to visualize the PPI network, and the Cytoscape plugin CytoHubba was used to calculate the degree of connectivity of each protein node. Specifically, nodes with a higher degree of connectivity were likely to play a more vital role in maintaining the stability of the entire network. In our study, the top 10 genes were considered as hub genes.
Results
Identification of DEGs
We identified DEGs to evaluate the gene functions and to illuminate the pathogenesis of KD. GSE68004 contained 76 KD and 14 normal samples. A total of 358 DEGs were identified from GSE68004 for subsequent bioinformatics analysis according to the criteria P < 0.05 and |logFC| ≥ 2, including 302 upregulated and 56 downregulated genes. A volcano plot of related genes is shown in Figure 1.
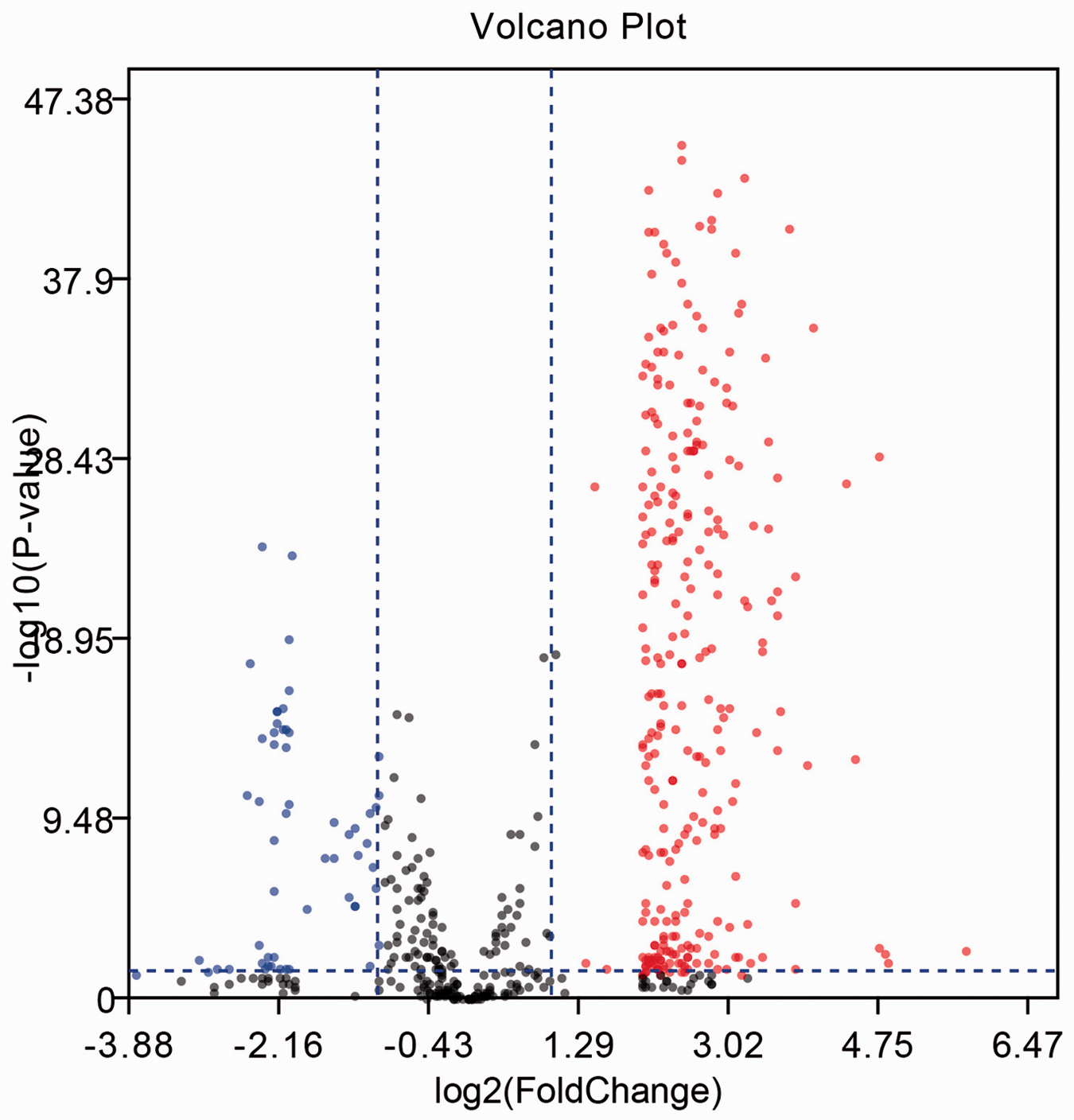
Volcano plot (red, upregulated genes; blue, downregulated genes).
GO term enrichment analysis
The functions of the significant DEGs were analyzed using DAVID software. The enriched GO terms were categorized into BP, CC, and MF terms. BP analysis demonstrated that the DEGs were significantly enriched in innate immune response, defense response to Gram-positive bacteria, inflammatory response, and antibacterial humoral response. Regarding CC, DEGs were significantly enriched in terms of the extracellular region, plasma membrane, extracellular space, and extracellular exosome (Table 1). GO enrichment terms for the upregulated and downregulated genes were illustrated as a GOCircle plot (Figure 2). KEGG pathway analysis indicated that the majority of the DEGs in KD were enriched in systemic lupus erythematosus (Table 1).
Significantly enriched GO terms and KEGG pathways of DEGs.

GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; DEG, differentially expressed gene; BP, biological process; CC, cellular component.

GOCircle plot (red, upregulated genes; blue, downregulated genes). Outer ring displays scatterplots of expression levels for genes in each term.
PPI network construction and hub gene identification
We further aimed to examine the functional correlations among the hub genes using STRING (a database of known and predicted protein interactions). The nodes in the PPI network represented the genes, and the edges between the nodes represented the interactions between the genes. The number of edges linked to a given gene was defined as the degree of connectivity of that gene, and only experimentally validated interactions (edges) were used in this analysis. A gene with a high degree of connectivity was deemed as a hub gene with an essential biological function. The PPI network was subsequently visualized using Cytoscape software, and the connectivity degree in the PPI network was assessed to identify the top 10 genes (Figure 3 and Table 2). Integrin αM (ITGAM) and myeloperoxidase (MPO) showed the greatest degrees of connectivity (both 22), followed by mitogen-activated protein kinase 14 (MAPK14) and solute carrier family 11 (SLC11A1) (both 19), histone cluster 2 (HIST2H2BE) and elastase, neutrophil expressed (ELANE) (both 16), cathelicidin antimicrobial peptide (CAMP; 15), matrix metallopeptidase 9 (MMP9; 14), neurotensin (NTS; 13), and histone cluster 2 (HIST2H2AC; 12).

Top 10 hub genes with high degrees of connectivity.
Top 10 hub genes with high degrees of connectivity.

Discussion
KD is an acute, self-limited vasculitis that specifically affects children under 5 years of age. Infectious factors, genetic vulnerability, and autoimmunity have all been proposed as probable causes of KD. 9 However, despite extensive investigations, the pathogenesis of KD remains unclear. Previous studies have aimed to identify the genetic background of KD and to provide clues to its etiology and pathogenesis. 3 However, no previous studies have examined the internal correlations among significantly associated genes in relation to the probable pathogenic process of KD using bioinformatics analysis.
In this research, we performed gene expression and PPI analyses based on accessible datasets to determine possible core genes related to KD. DEGs between KD and normal specimens were screened according to gene expression profiling information from GEO. KEGG pathway analysis indicated that most DEGs were enriched in systemic lupus erythematosus, and may thus be immune response-associated genes. In total, 302 upregulated and 56 downregulated DEGs were identified, which were related to the BP terms defense response to Gram-positive bacteria, antibacterial humoral response, innate immune response, and inflammatory response. Infection has long been postulated as the main cause of KD, and superantigens related to infection are considered to be the most crucial factors involved in its pathogenesis. 10 Previous studies reported that bacterial agents may be related to the onset of KD, and Staphylococcus aureus and Streptococcus pyogenes have been isolated from patients with KD and considered to have a possible association with the pathogenesis of this disease.11,12 We also constructed a PPI network to investigate the interrelationships among the DEGs, which identified ITGAM, MPO, MAPK14, SLC11A1, HIST2H2BE, ELANE, CAMP, MMP9, NTS, and HIST2H2AC as hub genes in KD.
The current bioinformatics approach identified immune response-associated genes as being involved in KD. Integrins, comprising α and β subunits, are a family of receptors for extracellular matrix (ECM) and cell surface ligands that participate in cell migration and ECM attachment. Bound integrins can transmit and receive intracellular signals, subsequently modulating endothelial cell migration, angiogenesis, cell survival, and connecting components of the ECM, as well as cellular proliferation, motility, and adhesion.10–12 Integrins are currently used as therapeutic targets in inflammatory disorders such as Crohn’s disease and multiple sclerosis. 13 Natalizumab (Tysabri) is an anti-α4 integrin monoclonal antibody approved by the US Food and Drug Administration for the treatment of multiple sclerosis. 14 ITGAM (also known as CD11b) is located at chromosome 16p11.2 and encodes an α-chain subunit of a leukocyte-specific integrin, which regulates leukocyte activation, adhesion, and migration from the blood stream and is important in the phagocytosis of complement-coated particles. 15 A meta-analysis of case-control studies demonstrated that the ITGAM rs1143679 polymorphism was significantly associated with an increased risk of systemic lupus erythematosus. 16 Furthermore, a recent study showed that protein expression levels of ITGAM were upregulated in KD patients. 21 In KD coronary artery lesions, ITGAM might enhance subacute/chronic vasculitis, resulting in the transition of smooth muscle cells to myofibroblasts and their subsequent proliferation. 17 ITGAM was also reported to be upregulated in the peripheral blood of KD patients who were refractory to initial therapy. 18 In this regard, ITGAM-knockout mice subjected to endothelial denudation and arterial stretching showed reduced intimal thickening and cell proliferation, as well as reduced leukocyte accumulation compared with control mice. 19 Another murine model of vascular injury demonstrated decreased inflammatory cell infiltration and reduced long-term intimal thickening in the presence of anti-integrin αM antibodies. 20 Overall, overexpression of ITGAM may thus be a unfavorable prognostic factor in patients with KD. However, further studies are needed to explore the value of ITGAM inhibitors in the treatment of KD.
In addition to ITGAM, we detected another nine hub genes associated with KD. MPO is a highly oxidant enzyme that produces reactive oxygen species, and is considered to be a critical biomarker of vascular inflammation. 22 Oxidative stress can disturb the balance between reactive oxygen species and antioxidants, further destroying proteins, lipids, and nucleic acids. Increased levels of MPO have been observed in patients with acute KD. 21 Studies of MPO polymorphisms have mainly concentrated on the -463G>A polymorphism (GenBank ID: rs2333227), and a recent case-control study suggested that the G allele of this polymorphism may be a possible genetic risk factor for KD.22–24 Additionally, SLC11A1 can modulate the interactions between macrophages and interferon-γ derived from bacterial lipopolysaccharide and/or natural killer cells or T cells. 24 A previous study demonstrated that allele 1 of the 5' promoter (GT)n repeat in the SLC11A1 gene was related to KD. 25 MMP9 has been implicated in various pathological situations, including tumor metastasis, KD, inflammation, and atherosclerosis. 26 However, there is currently limited information on the relationships between the above-mentioned core genes and KD, and further studies are warranted to investigate these associations.
The current study had some limitations. Notably, the sample size was relatively small, and further larger studies are needed to confirm these results.
In summary, we investigated KD DEGs in the GSE68004 dataset by systematic bioinformatics analyses. We identified 10 hub genes with potentially important roles in KD progression, which could also act as possible biomarkers for KD. However, further experiments should be performed to validate the functions of these identified genes in KD.
Footnotes
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
