Abstract
Introduction
The objectives of this retrospective review were to: (a) determine the prevalence of resistant Gram-negative bacteraemia among hospitalized patients; (b) evaluate antibiotic use; (c) determine the time taken for Gram staining to final species identification.
Methods
For this retrospective study, information was extracted from patients’ electronic medical records. Eligible patients had been admitted to a 300-bed tertiary care hospital in Tucson, Arizona from October 2015 to October 2016, were over 18 years of age and had a positive blood culture for Gram-negative bacteraemia.
Results
In total, 84 patients with Gram-negative bacteraemia were identified; urinary tract infection was the most common source of infection (71%). ESBL-producing microorganisms were isolated from five (6%) patients and no MDR pathogens were identified. The, median time to Gram stain was 20.5 hours and the median time to final identification was 54.5 hours. Delayed de-escalation of broad-spectrum antibiotics (i.e., >24 hours after final culture) occurred in 25% patients with a median length of hospital stay of 118 hours (range: 56–552 hours) compared with a median length of hospital stay of 89 hours (range: 5–334 hours) in the early de-escalation group.
Conclusion
The prevalence of bacteraemia due to resistant Gram-negative microorganisms is low (6%) in this institution. However, there may be room for improvement in the antimicrobial stewardship program with regard to rapid diagnostic testing.
Keywords
Introduction
Despite the availability of effective antibiotic therapies, bloodstream infections are an important cause of morbidity and mortality. 1 Importantly, bacteraemia due to Gram-negative bacilli is associated with a poor outcome because of the emergence of antibiotic resistance to these pathogens. 2 Moreover, antimicrobial resistance is considered a growing problem worldwide. 3 For example, in Japan, the increased use of anti-pseudomonal antibiotics has been linked to an elevated risk of multidrug-resistant organisms (MDROs). 4 Therefore, in many countries, health care institutions have set up antimicrobial stewardship programs (ASP) in an attempt to optimize antimicrobial use among hospitalized patients to ensure cost-effective outcomes, avoid inappropriate use of antibiotics and decrease or avoid complications. 5
Resistance of Gram-negative bacilli to antibiotics can be caused by one or more of the following mechanisms: production of beta-lactamases, porin loss, target modification and drug efflux. 6 The production of extended-spectrum beta-lactamases (ESBL) by Gram-negative bacteria, is of growing concern because they are resistant to all β-lactam antibiotics apart from carbapenems and cephamycins. 7 ESBL-producing Enterobacteriaceae pose a significant threat not only to hospitalized patients but also to those in community settings. 8 Several risk factors have been reported to increase the risk of ESBL acquisition and include, prior exposure to beta-lactam antibiotics, length of hospital stay, residence in a long-term care facility and ≥65 years of age. 9 However, the early initiation of appropriate and effective antibiotic use within 72 hours has been linked to a decreased mortality risk due to multidrug resistant pseudomonal species.10,11
Standard techniques for microbiological identification are based on phenotypic methods, which require 48-72 hours for identification and antimicrobial susceptibility compared with rapid diagnostics tests which only need a few hours following bacterial growth. 12 For these tests, positive blood cultures are tested using molecular methods to determine causative pathogens or both causative pathogens and resistance markers. 13 The use of rapid diagnostic methods coupled with appropriate therapeutic interventions should improve the overall clinical outcome and mortality. 13
The objectives of this retrospective review were to (a) determine the prevalence of ESBL-producing Enterobacteriaceae bacteraemia and MDRO bacteraemia among hospitalized patients; (b) evaluate their antibiotic use (i.e., broad spectrum antibiotics, dose escalation/ and (c) determine the time taken for Gram staining to final species identification.
Methods
This retrospective study included all patients over 18 years of age with positive blood culture for Gram-negative bacteraemia admitted to a 300-bed tertiary care hospital in Tucson, Arizona from October 2015 to October 2016. There were no exclusions.
Patient data were collected from the hospital’s electronic medical records system (Cerner®). Demographic characteristics, source of infection, length of hospitalization, isolated microorganisms, time of sample collection, time of Gram stain, time of final culture sensitivities and susceptibilities, results of final sensitivities and susceptibilities, first empiric antibiotics used, timing of antibiotic administration, time on broad-spectrum antibiotics and time to de-escalate narrow-spectrum antibiotics were extracted from the database.
The following definitions were used:
time to Gram stain = time between sample collection and first Gram stain results time to final culture sensitivities and susceptibilities = time between sample collection and final culture sensitivities and susceptibilities time to de-escalate = time to de-escalate the antibiotics to narrower spectrum after final culture sensitivities and susceptibilities. delayed de-escalation = >24 hours after final culture sensitivities and susceptibilities broad spectrum antibiotics = antibiotics with pseudomonal coverage or two injectable antibiotics with different spectra of activity first antibiotics use = any antibiotic used after sample collection but before final culture sensitivities and susceptibilities
Patients with ESBL-producing Enterobacteriaceae bacteraemia and MDRO bacteraemia had their characteristics (i.e., initial and direct antibiotic use and disposition) identified. MDRO was defined as non-susceptibility to at least one agent in three or more antimicrobial categories. 14 Patients who had antibiotics de-escalated in less than 24 hours were compared with those who had delayed de-escalation (i.e., >24 hours) in terms of resistant pathogens and length of hospital stay.
This was a quality improvement project as part of an antimicrobial stewardship rotation and was exempt from Institutional Review Board (IRB) approval.
Statistical analyses
Data were analysed using IBM SPSS software (version 25.0 for Windows®; IBM Corp, Armonk, NY, USA). Normally distributed continuous variables were reported as means with standard deviation (SD). Non-normally distributed continuous variables were reported as medians with interquartile range (IQR) and Mann-Whitney U test was used to compare data where appropriate. A P-value <0.05 was considered to indicate statistical significance.
Results
Eighty-four patients with Gram-negative bacteraemia were identified. Their demographics and baseline characteristics are shown in Table 1. Mean (SD) age of the cohort was 72 (15.9) years and most patients were older than 60 years (80%) and female (69%) In total, 17% patients had recently been hospitalised and 12% were in long-term care facilities. Approximately a quarter of the cohort had recently received antibiotics most of which were broad spectrum. The urinary tract (71.4%), followed by gastrointestinal tract (17.8%) were the most common sources of infection (Table 2).
Demographics and baseline characteristics.

Data are presented as mean ± SD or n (%).
*Antibiotics with pseudomonal coverage or two injectable antibiotics with different spectrums of activity.
Sources of infection.
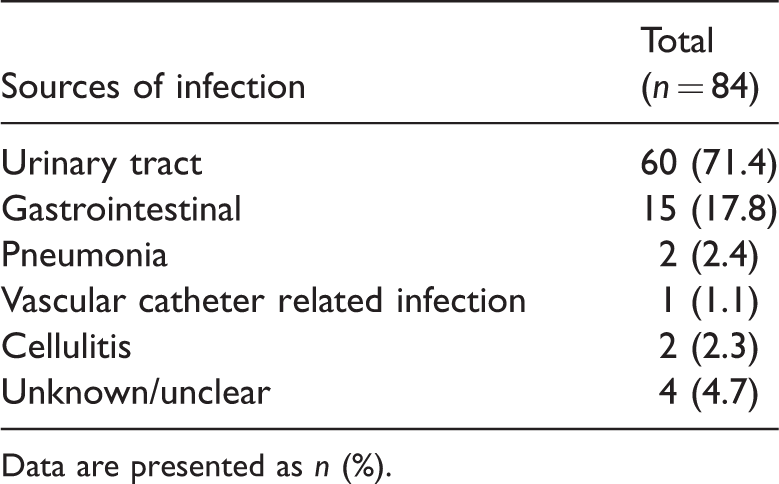

Data are presented as n (%).
In total, 83 microorganisms were isolated from the 84 patients. Multidrug resistant carbapenemase-producing Providencia stuartii was initially thought to be isolated from one patient but this was found to be negative after using a modified Hodge Test to perform phenotypic testing for carbapenemase activity. Escherichia coli was the most commonly isolated microorganism (56/83, 67.4%), followed by Klebsiella pneumoniae (13/83, 15.6%), Enterobacter species (5/83, 6%) and Pseudomonas aeruginosa (2/83, 2.4%) (Figure 1). No MDROs were identified but five (6%) ESBL-producing pathogens were identified (four E. coli and one K. pneumoniae).

Isolated microorganisms (n = 83).
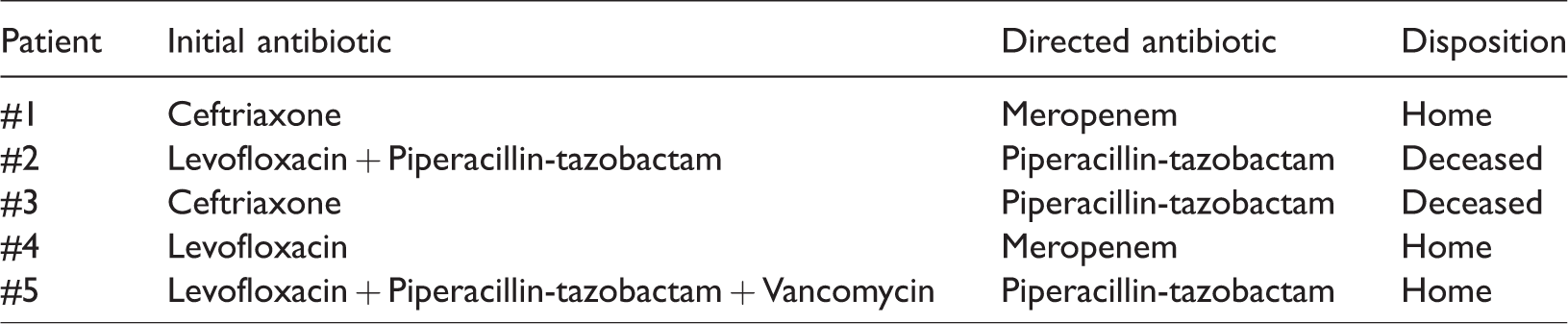
All ESBL-infected patients were older than 60 years. Some details of these patients are shown in Table 3. Of the five patients, one had a recent hospitalization, three had a history of broad-spectrum antibiotic exposure and one was a long-term care facility resident. Four ESBL-infected patients had experienced delayed first effective antibiotic initiation; for three the source of infection was the urinary tract and for one patient it was unknown. Two were discharged home, one was discharged to a skilled nurse facility and one died.
Extended-spectrum beta-lactamases (ESBL)-infected patients (n = 5) and their characteristics.
Across the entire group, the median time to Gram stain was 20.5 hours (interquartile range 12.31 hours) and the median time to final culture sensitivities and susceptibilities was 54.5 hours (interquartile range 19 hours) (Figure 2). Therefore, the median time from gram stain to final culture was 29.4 hours (interquartile range 27.4 hours)
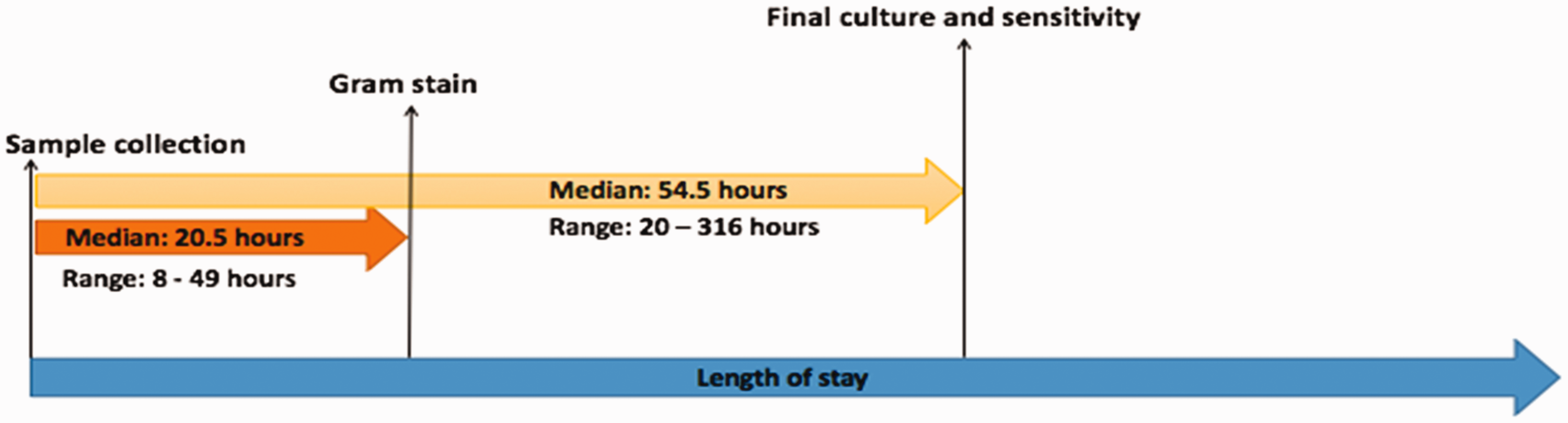
Time to Gram stain and final sensitivities.
According to our pre-defined definition, antibiotics with antipseudomonal activity were used in 48/84 (57.1%) patients. This included one or more of the following: piperacillin-tazobactam, levofloxacin, meropenem, and aztreonam. Of the two patients who had P. aeruginosa bacteraemia, one received meropenem and vancomycin and was then de-escalated to meropenem following final blood culture sensitivities. The other patient received piperacillin-tazobactam upon admission, which continued throughout the hospital stay. Both patients had acquired their infections in the community; for one patient the source of infection was cellulitis and for the other patient it was a urinary tract infection. In our opinion, vancomycin had been used inappropriately in 10/84 (11.9%) patients with urinary source bacteraemia.
Delayed de-escalation of broad-spectrum antibiotics (i.e., >24 hours after final culture) occurred in 12/48 (25%) patients who had a median length of hospital stay of 118 hours (range: 56–552 hours). Resistant pathogens were not identified in any of the delayed de-escalated group. By contrast, 33/48 (75%) of patients had their broad-spectrum antibiotics de-escalated appropriately (i.e., within 24 hours of final culture). The median length of hospital stay for these patients was 89 hours range: 5–334 hours. The difference in hospital length of stay between delayed and early de-escalation groups was statistically significant (P 0.02).
Discussion
The prevalence of ESBL-producing Enterobacteriaceae differs among patients and geographic regions. 15 Moreover, rates of ESBL-producing bacilli in K pneumoniae isolates tend to be higher than those in E coli. 16 For example, data from a global surveillance database showed the rate of ESBL production was highest among the K. pneumoniae isolates collected in Latin America, followed by Asia/Pacific Rim, Europe, and North America (44.0%, 22.4%, 13.3%, and 7.5%, respectively). 15 By contrast, data from a global study of the in vitro susceptibilities of E. coli strains isolated from individuals with urinary tract infections at 88 international hospitals, found that overall, 17.9% of isolates were ESBL producers with multidrug resistance. The highest rates were seen in Asia/ Pacific (27.7%), while the lowest rates were seen in North America (7.4%). 17 In addition, a retrospective study of data collected from five hospitals in the Netherlands during the period 2008 to 2010, found that of the 232 consecutive patients with ESBL bacteraemia due to E coli, and K. pneumoniae the ESBL producing prevalence among blood culture isolates of these bacteria were 6.6% and 8.7%, respectively. 18 Therefore, our findings of 6% bacteraemia caused by resistant Gram-negative microorganisms in our institution is reassuring and encourages us to continue with the adherence to national and local guidelines. In our study, ESBL-producing pathogens were identified in four E. coli and one K. pneumoniae isolates. These results are not surprising since urinary tract infections were common in this cohort and E. coli is the most important uropathogen. 17 In addition, while the U.S. Centers for Disease Control and Prevention (CDC) rated the level of carbapenem-resistant Enterobacteriaceae (CRE) as urgent 1 and data from a nationwide US Surveillance Network identified that CRE accounted for 4% of bloodstream infections, 19 no CRE was identified at our institution.
Timely administration of appropriate antibiotics is essential for decreasing hospital mortality in patients with blood infections. For example, one study found that with each hour delay of effective antimicrobial therapy, there was a 7.6% decrease in survival in patients with sepsis. 20 Moreover, delaying appropriate antibiotic use can increase healthcare costs, length of stay and antimicrobial resistance.21,22 In this present study, we identified only 4/84 patients with delayed initiation of effective antibiotics, and all had ESBL-producing pathogens. All of these patients had ESBL bacteraemia and only one patient died.
For traditional methods of microbial identification, the length of time taken for the detection of the pathogen is an important consideration. Most laboratories take 48–72 hours to complete the antimicrobial susceptibility testing.12,23 This study showed that in our institution the median time from sample collection to final culture sensitivities and susceptibilities was 54.5 hours and from gram stain to final culture the median time was 29.4 hours. In addition, 75% of patients had their broad-spectrum antibiotics de-escalated appropriately (i.e., within 24 hours of final culture) and had a significantly shorter hospital stay than patients who had late de-escalation of antimicrobials. Therefore, similar to the findings of other studies, 24 our ASP is efficient and has led to decreased length of hospital stay. However, the introduction of rapid diagnostic testing at our institution may result in faster de-escalation times and avoid any unnecessary use of broad-spectrum antibiotics. 13 Nevertheless, data from a prospective, randomized, controlled trial over two years involving 1489 patients, found that although rapid bacterial identification and susceptibility testing led to earlier changes and a significant reduction in antibiotic use, it did not reduce mortality. 23 Similar findings were reported in another study that compared an ASP coupled with polymerase chain reaction (PCR)-based blood culture identification with ASP and conventional organism identification or conventional organism identification (control) alone. 25 Although microorganism identification was significantly shorter for the PCR-blood culture identification group, there was no difference in mortality. However, significantly shorter times to de-escalate were observed in the PCR-blood culture identification group compared with the other groups. While results from a retrospective study involving 232 patients with ESBL bloodstream infections found that inappropriate antimicrobial agent within the first 24 hours was not associated with increased risk of mortality, 18 early identification of resistant microorganisms is imperative and more studies are needed, especially in patients with deep-seated infections. 26
Our current study has some limitations. For instance, it was a retrospective single-centre study using a small sample size and so the findings may not be robust compared with data from other facilities that have higher rates of MDROs. Additionally, we did not assess the severity of disease nor did we include a control group. We were also unable to assess potential complications from the use of broad spectrum or multiple antibiotics. We defined delayed de-escalation as >24 hours after final sensitivities and susceptibilities rather than the time from the gram stain results and we did not evaluate whether doses were optimized based on minimum inhibitory concentration (MIC) results for specific microorganisms and antibiotics. Finally, we did not evaluate whether the treating physician or the ASP pharmacist performed the de-escalation.
Selecting the most appropriate and cost effective rapid diagnostic test is challenging given the wide range of commercially available diagnostic tools, some of which require labour-intensive clinical laboratory work with high complexity. 12 In addition, practitioner awareness of the usefulness of rapid diagnostic testing may be limited. For example, a study found that only 60% of practicing physicians adjusted their patients’ therapy based on rapid-blood-culture-identification results, and therefore, de-escalation of antimicrobial therapy was not optimal. 27 Although we observed a low prevalence of resistant Gram-negative pathogens, the median time from Gram stain to final culture was 29.4 hours and so the introduction of rapid diagnostics tests may be of beneficial value in reducing any unnecessary use of broad-spectrum antibiotics. While we did not examine the cost of therapy, the use of rapid diagnostic testing may have an impact on the overall cost, as well as decrease resistance rates. 25 A prospective study is required to determine the utility and possible cost reduction involved in the implementation of rapid diagnostic testing coupled with ASP in a large patient sample.
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Footnotes
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Acknowledgements
The authors would like to thank the University of Arizona Libraries Business Office for providing financial support. Abdulaziz Almulhim would like to thank the Deanship of Scientific Research for continuous support.
