Abstract
Stroke is a multifaceted disease with genetic and environmental components like diet and lifestyle. The central nervous and immune systems display complex interactions, with the peripheral immune response participating in brain injury and repair mechanisms following stroke. The bidirectional communication between the injured brain and peripheral blood presents an opportunity to investigate the molecular changes in the latter. There is substantial heterogeneity in stroke pathogenesis, pathophysiology, comorbidities, and response to treatment and outcome. This is captured and underscored by heterogeneity in the peripheral blood transcriptome. The current review highlights the role of the human peripheral blood transcriptome architecture for molecular phenotyping of different stroke etiologies and comorbidities, and for identifying underlying molecular correlates with clinically important variables and outcomes. Specific transcriptome features can potentially provide targets for clinical translation and for prioritizing genes and pathways for evaluation in experimental models. We also propose an approach to study the patient-specific transcriptional architecture and uncover the combinatorial heterogeneity in altered pathways in stroke patients that can also guide the search for treatment and prevention targets. Deciphering the molecular heterogeneity of stroke in a tissue that can be easily accessed and monitored, such as peripheral blood, may improve clinical trial success.
Rationale for studying the peripheral blood transcriptome in human stroke
Damaged brain cells after stroke release DAMPs (Damage Associated Molecular Patterns) which activate microglia and other cells to release inflammatory signals such as cytokines and chemokines. They signal to peripheral blood resulting in infiltration of leukocytes from blood to brain in a process called leukocyte extravasation.
1
Recruited neutrophils release MMPs and neutrophil extracellular traps, and together with other blood cell types also release reactive oxygen species and other neuroinflammatory molecules that damage the blood-brain barrier
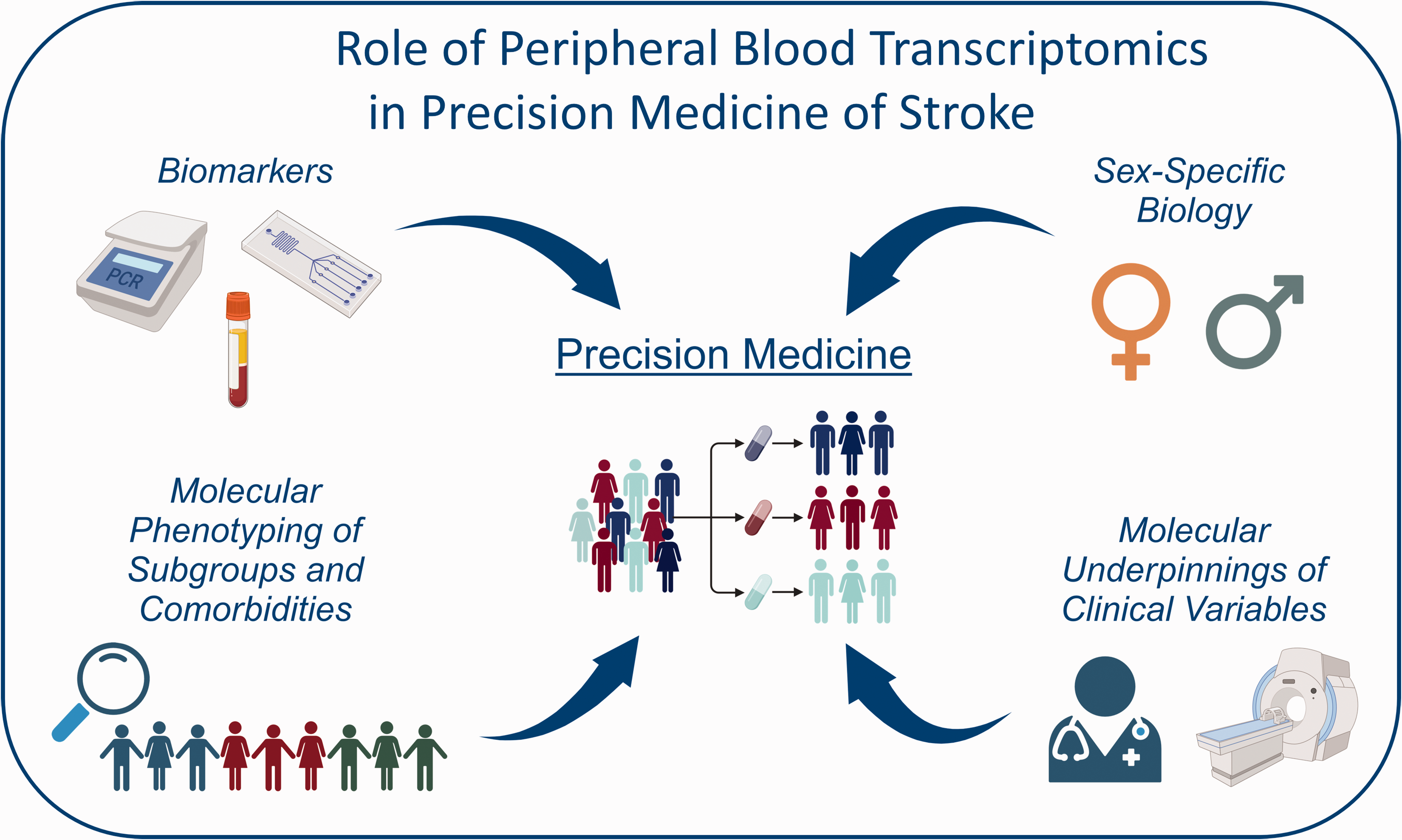
The role of peripheral blood transcriptomics in stroke precision medicine. Schematic presentation of the utility of the peripheral blood transcriptome for deciphering the sub-group heterogeneity in stroke for the advancement of precision medicine. Created with BioRender.com.
The transcriptome architecture in disease provides information on genetic, epigenetic, environmental and lifestyle influences on a condition using existing whole genome technologies that can investigate it in its entirety. Additionally, studies have revealed a significant overlap between differentially expressed genes in peripheral blood and in brain, such as for blood and brain in a middle cerebral artery occlusion nonhuman primate stroke model,
10
and in human peripheral blood
11
and perihematomal brain transcriptome
12
following intracerebral hemorrhage
Additionally, there is a need for human stroke transcriptomic studies because of the complex contribution of comorbidities to human stroke and because of difficulties in translating animal stroke studies to human stroke. For example, human and rodent peripheral blood cell composition differs significantly. 13 A mouse-human comparative peripheral blood transcriptomics study post stroke found little overlap in the affected pathways in the acute phase (9%) with more overlap in the subacute phase (47%). 14 This further underscores the need for human stroke molecular studies to help guide mechanistic animal stroke research with the overarching goal of improving translational success.
The transcriptional responses to ischemic stroke
Challenges exist in comparing omics studies, including transcriptomics studies. These challenges are due to employing different transcriptomic technologies (reviewed in 19 ) and biological samples. The biological differences may relate to whether the transcriptomics studies are performed on whole blood, cell subpopulations, such as peripheral blood mononuclear cells (PBMC), individual cell subpopulations, or single-cell/single-nucleus RNA populations. Additionally, studies may differ due to sampling different time points. Moreover, variables such as treatment, infarct size, hemorrhage and edema volumes, cause and location, vascular risk factors, age, race, and sex, if not considered in the studies, may also affect the transcriptome findings.
The purpose of the review is to underscore the utility of the peripheral blood transcriptome to investigate the heterogeneity of the peripheral blood transcriptome following stroke to molecularly phenotype causes of stroke subtypes, to uncover biological networks underlying clinically relevant features of stroke, and to decipher the effects of comorbidities including sex-specific biology. Thus, the present review highlights selected studies to illustrate what knowledge can be obtained from examining the peripheral blood transcriptome in human stroke. Due to differences in stroke populations, the evolution of molecular responses over time, cell-specific responses, and different technologies used to assess the transcriptome, direct comparison of findings among stroke transcriptome studies is challenging, as pointed out above, and not the focus of this review.
Stroke etiologies
Distinct transcriptional programs following ischemic stroke of different etiologies
IS has different etiologies including cardioembolic
Xu et al
24
investigated the peripheral blood transcriptome of CE and LVA IS in serial blood draws at ≤3 h (before thrombolytic treatment), 5 h and 24 h (post thrombolytic treatment time-points) post
SV stroke also showed specific peripheral blood transcriptomic response with pathways enriched in Monocyte and Leukocyte Activation and Recruitment, and Innate and Adaptive Immune Cell Communication biofunctions among others. 26
IS etiology-associated heterogeneity in the peripheral blood transcriptome has also been investigated at the level of the alternatively spliced transcriptome. Alternative splicing is a genetically-regulated and environmentally-influenced process by which a single gene can produce multiple alternatively spliced isoforms which have specific functions in different cells, tissues, developmental stages, and disease states. 33 It is a major process that generates vast molecular diversity by which the ∼20,000 protein-coding genes code for >250,000 RNA and protein isoforms. Dykstra-Aiello et al 29 examined the differential alternative splicing of 71 putative stroke/vascular risk factor genes expressed in peripheral blood of 245 IS and controls, and determined many had sex-specific and etiology-specific pattern of expression. This underscored the importance of considering both sex and etiology when novel treatments are developed. 29 This study also confirmed a previous pilot RNA-Seq study, 34 which showed etiology-specific alternative splicing/exon usage is involved in CE, LVA and SV IS mechanisms, and in ICH. Studying the differentially alternatively spliced genes/differential exon usage is a highly understudied area in stroke research that warrants further investigation as exon/transcript isoform resolution will likely be required to better understand stroke biology, and to develop novel treatments and biomarkers.
The above-mentioned studies have been performed in whole peripheral blood which contains different cell types with specific roles in IS. RNA-Seq studies in isolated monocytes and neutrophils from peripheral blood revealed cell type-specific- and etiology-specific gene expression
28
and time-dependent gene expression using self-organizing maps.
16
Though there was some overlap in differentially expressed genes and enriched pathways among the three IS etiologies, the majority of the differentially expressed genes and pathways were etiology-specific.
28
Most of the self-organizing map time-dependent profiles that were similar between the three IS etiologies were enriched in etiology-specific pathways, signifying the temporal evolution of the peripheral blood transcriptomes following human IS to be etiology-specific. Furthermore, there were pathways which progressed in opposite directions over time. For example, in neutrophils, genes from the toll-like receptor
Human transcriptomics studies in IS of different etiologies supplement experimental findings, which may strengthen their clinical translation. They have shown whole-blood transcriptome and cell-type specific heterogeneity in IS etiology, which is important since each cell type has specific roles in the injury and repair mechanisms following IS.1,9,36 Additionally, the etiology-specific temporal evolution may pinpoint etiology-specific treatment windows. The observed molecular heterogeneity in IS of different etiologies may have repercussions for treatment development and may require a search for novel etiology-specific therapies.
Molecular heterogeneity due to ICH etiology/location
Intracerebral hemorrhage (ICH) is caused by bleeding into the brain parenchyma. It is the second most common cause of stroke, accounting for 10–20% of all strokes, and has high short-term and long-term mortality up to 50% at 30 days.37–41 The primary ICH etiologies in the aged population are cerebral amyloid angiopathy (CAA) 42 and chronic hypertension. 37 CAA is characterized by deposition of amyloid in the media and adventitia of cortical and leptomeningeal small vessels, including arterioles and capillaries, 42 and is diagnosed based on the Boston v. 2.0 criteria. 43
To the best of our knowledge, there have been no studies of the peripheral blood transcriptome in human ICH stratified by ICH etiology in older adults. A study by Knepp et al 44 investigated the common and unique transcriptional changes in the peripheral blood transcriptome in deep ICH (usually due to chronic hypertension) and lobar ICH (usually due to CAA). The authors identified deep ICH- and lobar ICH-common pathways such as Autophagy, T Cell Receptor, Inflammasome, and Neuroinflammation Signaling. However, there was a substantial molecular heterogeneity in the transcriptome architecture in deep and lobar ICH, with Th2, Interferon, GP6, and BEX2 Signaling pathways unique to deep ICH, while Necroptosis Signaling, Protein Ubiquitination, and various RNA Processing terms were unique to lobar ICH. Additionally, amyloid processing pathways and suppression of several protein ubiquitination pathways and upregulation of ubiquitin inhibitors were also unique to the lobar ICH transcriptome. The authors suggested peripheral blood cells may participate in amyloid removal or leading to perivascular/vascular amyloid in lobar ICH due to CAA. The co-expression networks and/or the potential master regulators of the deep and lobar ICH responses also showed heterogeneity in their cell-specific landscape– with deep ICH enriched in neutrophil- and monocyte-specific genes, while lobar ICH was enriched in erythroblast- and T cell-specific genes. 44 The study also identified additional layers of heterogeneity discussed in the Heterogeneity in Common Pathways section below.
Some ICH causes in the younger population have also been investigated. Weinsheimer et al 45 studied the peripheral blood response to brain arteriovenous malformations (BAVM), an important ICH cause in young adults. The authors investigated the changes in the transcriptomes of ruptured and unruptured BAVM patients and healthy controls. 45 They found many of the ruptured-BAVM-related genes play a role in inflammatory processes, consistent with a role of inflammation in BAVM pathogenesis. 46 The findings suggested potential pathogenesis-related changes in the transcriptome for BAVM and/or BAVM rupture, which may help identify biomarkers or novel treatment targets.
Comorbidities
Stroke comorbidities, such as age, diabetes, hypertension, hyperlipidemia, smoking, and sex (discussed in a separate section below) increase stroke risk and worsen outcome (reviewed in47,48). The experimental stroke field recognizes that a critical factor for improving the clinical translation of findings is to address the presence of comorbidities in human stroke which often have not been represented sufficiently in preclinical research (reviewed in18,49). The landscape of stroke comorbidities in the peripheral blood transcriptome following human stroke is also largely unexplored. Some human stroke studies address the effect of comorbidities by including vascular risk factor-matched controls (
Age
Age is a risk factor for stroke. 18 The immune system changes with age which includes increased inflammation (inflammaging), attenuated immune response, and immunosenescence, with differences between males and females.50,51 Age-associated peripheral blood transcriptome changes following human IS have been investigated by Sykes et al, 52 where the authors identified age-associated genes in a derivation cohort (n = 94) and confirmed them in a validation cohort (n = 79) of IS patients. Sixty-nine adaptive immunity genes associated with age in IS, 49 of which were reported as age-associated in non-stroke studies as well. 52 Supporting evidence for age-associated molecular changes in stroke also came from a study by Soriano-Tárraga et al. 53 It compared stroke and control patients’ epigenetic/biological age, which is a measure of DNA methylation commonly correlated with normal aging and can be accelerated in some diseases. They found stroke patients had an older biological age than controls, indicating biological age and epigenetic modifications may be a useful stroke biomarker. Nakaoka et al. 54 analyzed gene expression of aneurysmal domes from patients with SAH and unruptured aneurysms, and superficial temporal artery samples as controls. Using hierarchical clustering, they identified two subgroups of SAH subjects with statistically different ages, demonstrating differing gene expression patterns based on patient age. These findings further underscore the molecular heterogeneity in the peripheral blood transcriptome and the importance of future studies investigating age-associated changes as they may modulate age-related stroke risk and outcome.
Atrial fibrillation
Atrial fibrillation is a major risk factor for CE IS. Many IS subjects initially deemed to have cryptogenic etiology are later discovered to have paroxysmal atrial fibrillation. A study of the peripheral blood transcriptome of IS patients uncovered additional levels of transcriptional heterogeneity based on presence or absence of atrial fibrillation in CE IS patients. 27 The study discovered specific immune responses associated with atrial fibrillation in CE IS and developed a 37-gene profile which differentiated CE due to atrial fibrillation from nonatrial fibrillation causes with >90% sensitivity and specificity. Additionally, a 40-gene profile differentiated LVA from CE with >95% sensitivity and specificity on cross-validation. When these profiles were applied to patients with cryptogenic stroke, 17% were predicted to have LVA and 41% to be CE. Of the cryptogenic strokes predicted CE, 27% were predicted to have atrial fibrillation. 27 These findings require validation in larger cohorts, as biomarkers identifying atrial fibrillation in cryptogenic strokes is of critical importance for future stroke prevention.
Smoking
Smoking is a modifiable risk factor for stroke. A whole-blood transcriptome meta-analysis of 10,233 subjects of European ancestry identified signatures of current and former smokers which included activation of platelets, immune response and apoptosis. 55 The smoking associated signature had a significant association with SNPs associated with stroke (ALDH2; CAMTA1; SH2B3; TMEM116; ERP29). 55 Cheng et al. 56 analyzed IS and control subjects stratified into current or never-smokers. Genes associated with smoking in IS patients were involved in pro-inflammatory pathways which may contribute to brain injury and worsen stroke outcomes. The authors also identified 10 genes associated with smoking which were common in the IS and control groups (GRP15, LRRN3, CLDND1, ICOS, GCNT4, VPS13A, DAP3, SNORA54, HIST1H1D, and SCARNA6). They speculated these genes might be associated with stroke risk. 56 The authors reported several of these genes have been implicated in stroke, such as ICOS which affects outcome in experimental stroke 57 and CLDND1, which is a tight-junction protein, involved in cerebrovascular disease pathogenesis of stroke‐prone spontaneously hypertensive rats. 58 Most of the smoking-associated genes in IS patients did not overlap with the ones in controls, suggesting unique interactions between smoking and ischemic stroke in peripheral blood cells. 56
Cancer
Cancer increases stroke risk and comorbid cancer in stroke is a growing area of research as roughly 13% of IS patients have a prior cancer diagnosis, 59 and 2%–10% will be diagnosed with cancer within a year post stroke. 60 A pilot study by Navi et al. 61 identified distinct peripheral blood gene expression patterns in IS patients with and without comorbid cancer, cancer alone, and VRFCs. Cancer-Stroke unique pathways included several interleukin signaling pathways (IL-1, IL-10, and IL-12), T helper-cell (Th1, Th2, Th17) activity, phagosome formation, pattern recognition receptor, TREM1 signaling, and neuroinflammation signaling pathways. 61 Additionally, major transcriptional regulators were differentially expressed between the cancer-stroke and stroke-only groups, some of which have been implicated in experimental stroke models or in stroke-associated processes. Those included CREB1 and SQSTM1. CREB1 was down-regulated in the cancer-stroke group versus the stroke-only group. CREB transcription factors modulate circuit plasticity and functional recovery after stroke; with increased CREB levels fostering stroke recovery whereas inhibiting CREB signaling hinders recovery. 62 SQSTM1 (p62) is a regulator of the hypoxia response, NF-kB and TNF signaling, 63 and was upregulated in the cancer-stroke group versus the stroke-only group. 61
A follow up study by Knepp & Navi et al 64 investigated the peripheral blood transcriptome of both protein – coding mRNA and regulatory miRNA in Cancer-Stroke patients, stroke only, cancer only, and VRFCs. They also investigated the IS-mechanism-specific transcriptome architecture of cancer-stroke patients. 64 Activation of Coagulation and Activation of Blood Platelets Biofunctions were observed in IS patients with cancer versus without cancer, supporting a hypercoagulable state in stroke with comorbid cancer. There were also differences in the complement system, growth factor signaling, and immune/inflammatory pathways. Among the differentially expressed genes, enrichment with granulocyte–specific genes was observed in both the CE and non-CE stroke response in patients with cancer compared to VRFCs, while T-cell-specific genes were associated only with CE strokes. The IS cause in cancer patients remains undetermined (cryptogenic) in ∼50% of patients, though a hypercoagulable state and CE etiology are often clinically suspected. 60 Indeed, based on a 15-gene panel, 11 of the 16 (69%) cryptogenic strokes with cancer were predicted to have CE etiology. 64 A separate 15-gene panel predicted the presence or absence of cancer in the IS subjects, which in a validation cohort classified correctly 81% of the stroke with cancer and 71% of the stroke-only subjects. 64 This suggests the possibility that peripheral blood biomarker panels could identify occult cancer in IS patients. These subgroup-specific molecular differences may indicate different treatment targets for each subgroup including the stroke subgroup with comorbid cancer.
Heterogeneity in response to treatment/clinical complications
A major goal of precision medicine is developing biomarkers to predict response to treatment to identify which patients are good candidates for a particular treatment and are likely to respond well, and which patients are likely not to respond or to develop side effects or complications. Using biomarkers to predict elevated risk of complications could guide eligibility for treatment or improve allocation of care to high-risk hospitalized individuals. Discovering biomarkers for responders and non-responders for new treatments could be used to stratify patients in later phases of clinical trials and increase the chances of their success. Though some stroke clinical trials include a search for biomarkers for response to treatment or for complications, many are focused on known potential biomarkers and do not include a comprehensive omics approach. Investigators and funding agencies should consider a whole transcriptome screen, wide proteome screen, and/or multi-modal biomarker screen to identify responders in Phase I or II trials. These responder profiles could be used in Phase III trials and be extended to the clinic for screening for those most likely to respond in a precision medicine treatment approach.
Hemorrhagic transformation following tPA treatment in ischemic stroke
Recombinant tissue-plasminogen activator (rt-PA) has been the main thrombolytic IS treatment. A study found transcriptome changes in the IS peripheral blood transcriptome PRIOR to tPA administration and predicted which patients will LATER develop hemorrhagic transformation (
Delayed brain injury (DBI) following aneurysmal subarachnoid hemorrhage (aSAH)
DBI complications following SAH, such as delayed cerebral vasospasm (DCV) 67 and delayed cerebral ischemia (DCI) 68 significantly increase mortality and morbidity. 69 However, the pathophysiology of DBI is poorly understood. The search for biomarkers has been reviewed.70,71 A study by Pulcrano-Nicolas et al 72 performed miRNA-Seq in DCV+ vs DCV− in aSAH patients from the VASOGENE cohort (NCT01779713). It identified elevated miR-3177-3p levels in peripheral blood in DCV+ patients compared to DCV- patients and proposed miR-3177-3p as a potential candidate biomarker for DCV risk. 72 Pulcrano-Nicolas et al 73 in a separate study in patients from the same VASOGENE cohort analyzed the protein-coding transcriptome and identified a significant difference between the DCV+ and DCV- aSAH groups for S1PR4 gene expression (Δ(DV3-D0)). The two time-points were: admission day (D0) and 3 days before DCV+ patients experienced vasospasm, or the corresponding day for the DCV- aSAH patients (DV3). DCV- patients had lower Δ S1PR4 expression than DCV+ patients. The AUC of the prediction model integrating Δ S1PR4 expression, sex and age was 0.896. S1PR4 and its ligand SP1 are expressed mainly by platelets and play roles in arterial-associated vasoconstriction.74–76 The authors speculated that following cerebral SAH, platelets release SP1 which triggers vasoconstriction through S1PR4-mediated mechanism. 73 This human data suggests that temporal change of S1PR4 expression is associated with developing vasospasm.
Xu & Stamova et al also investigated the peripheral blood transcriptome of aSAH patients with and without vasospasm with average time post aSAH
Outcome and clinical parameters associated with outcome
Studies identifying the molecular underpinnings associated with outcome following human stroke are of utmost importance as they can help prioritize genes and pathways for follow-up in mechanistic pre-clinical studies. They can also provide the experimental stroke literature with comparable human data and with the co-expression networks which outcome-associated genes participate in. In addition, as mentioned above, transcriptomic biomarkers of outcome can aid in determining prognosis of individual patients and stratifying patients in clinical trials to improve trial success.
Ischemic stroke outcome
Peripheral blood transcriptome changes associated with severity and outcome following IS have been investigated in several studies. Barr et al
78
identified temporal gene expression changes between day 1 and 2 post
Potential predictors of outcome and prognosis in ischemic stroke from clinical studies on peripheral blood RNA.
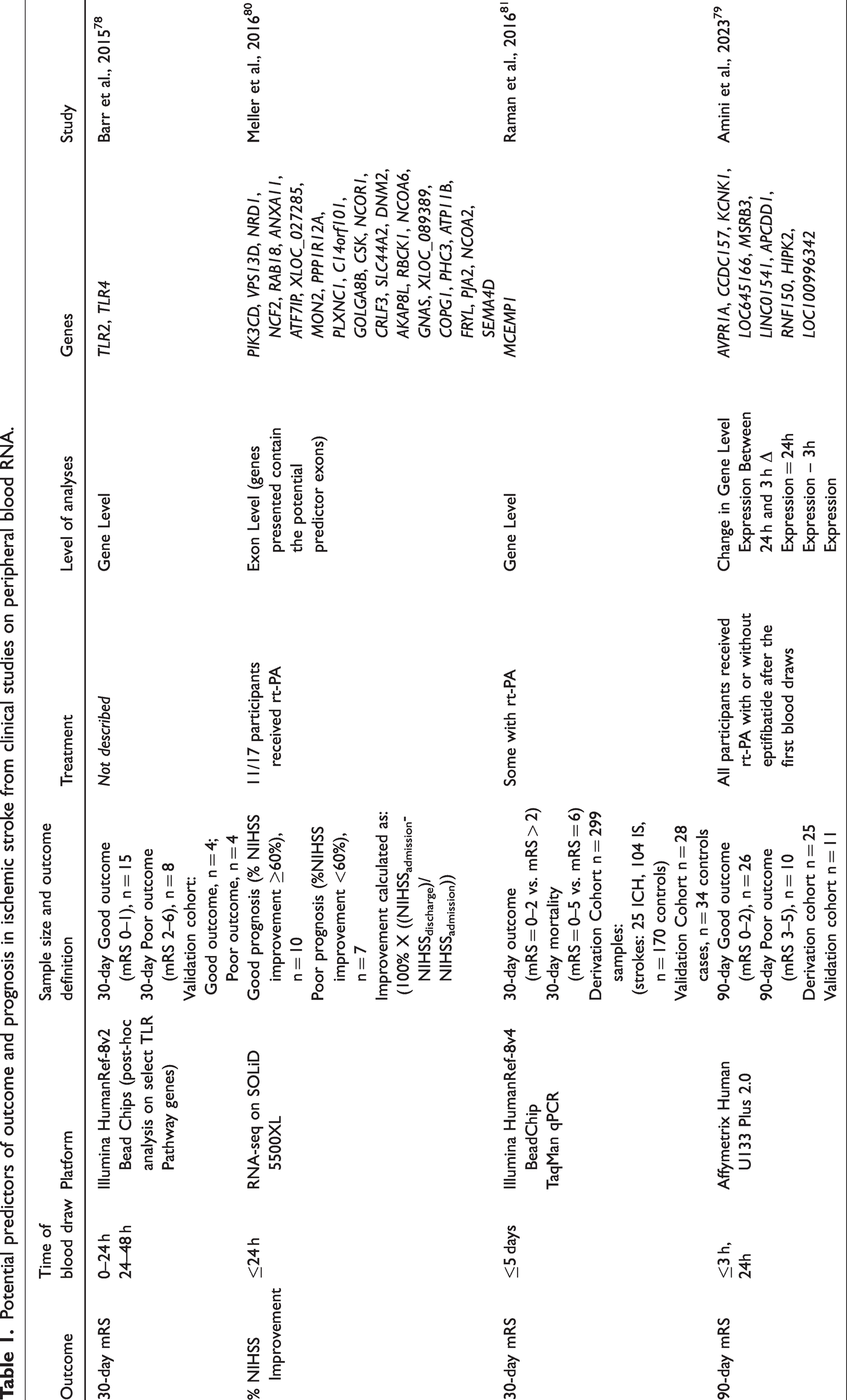
Using RNA-Seq, Meller et al 80 investigated the peripheral blood transcriptome of IS with middle cerebral artery occlusion and sex- and age-matched hypertensive controls in African Americans. They stratified the IS subjects into minor (NIHSS 0–5), moderate (6–15), and severe (>15) strokes, and identified 174 exons based on which subjects with different IS severity were separated on PCA and hierarchical clustering. Additionally, they derived a 30-exon panel that predicted the percent improvement between admission and discharge with 100% normalized correct rate (Table 1). 80 The authors studied the transcriptome changes at gene-, exon, and alternatively-spliced transcript level, and suggested exon-level resolution might have the highest biomarker potential. 80
Raman et al
81
performed peripheral blood transcriptomics from subjects from the INTERSTROKE study
82
on 299 samples in a derivation cohort (25 ICH, 104 IS, 170 controls) and a validation cohort (28 cases, 34 controls). The authors found
Amini et al 79 investigated the acute-phase peripheral blood transcriptome architecture at three time points for each IS subject and associated it with the 90-day mRS outcome. The IS subjects were from the CLEAR trial (NCT00250991) 32 described above. They were stratified based on 90-day mRS outcome (mRS = 0–2, good outcome; mRS = 3–5, poor outcome of survivors). The poor 90-day outcome group revealed many more differentially expressed genes than the good outcome group when compared to VRFCs. Poor outcomes also had significant activation in many inflammatory pathways before and after treatment, such as Acute Phase Response, HMGB1, IL-6, 17 A, STAT3, HIF1α, and NF-κB Signaling pathways. Additionally, poor outcome subjects had suppression in anti-inflammatory pathways such as LXR/RXR and PPARα/RXRα Signaling pathway (Figure 2; for details, please see Supplementary Material). Comparing the poor vs good 90-day outcome group revealed activation of inflammatory and suppression of T-cell specific pathways in the poor vs good outcome groups. The study also found up-regulation of neutrophil-specific, and down-regulation of lymphocyte-specific genes associated with poor 90-day outcome. Moreover, weighted gene co-expression network analysis (WGCNA) revealed modules of co-expressed genes associated with outcome as well as potential master regulator hub genes. A number of the outcome-associated modules contained GWAS signal, including a 24 h-module associated with ordinal 90-day mRS (unpublished data) which was significantly enriched with 15 IS risk GWAS genes, 83 and 6 IS Outcome GWAS genes84–86 (Figure 3; for details, please see Supplementary Material). These included ADAM23, which has been shown to correlate with early neurological instability, providing evidence for a role of metalloproteinases affecting cell-dell and cell-matrix interactions in influencing outcome. 84 Additionally, Amini et al derived a pilot 10-gene panel of putative biomarkers which predicted poor vs good 90-day mRS outcome with an AUC = 0.88 (Table 1).
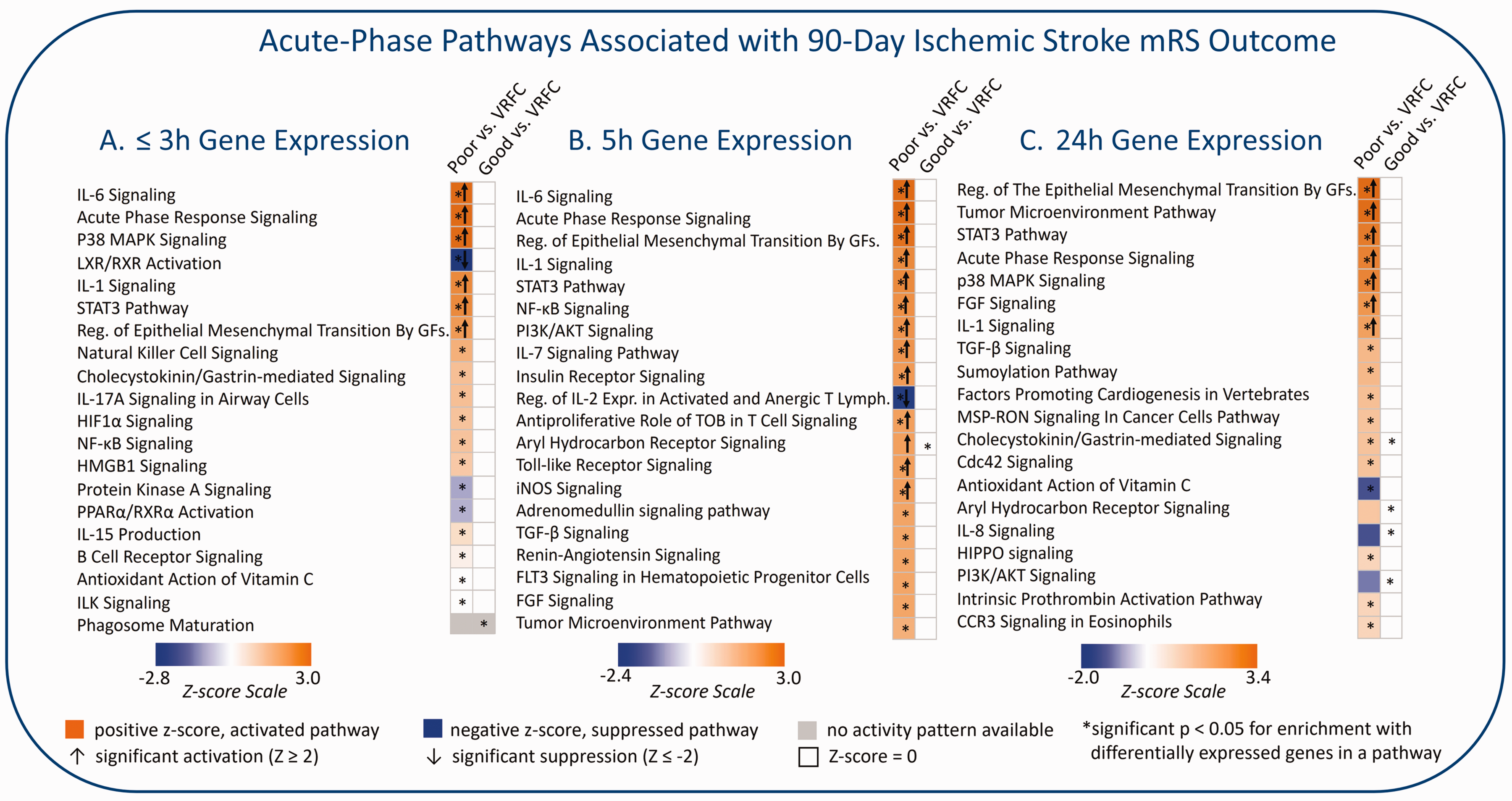
Acute-phase pathways associated with dichotomized 90-day ischemic stroke mRS outcome. Significant pathways enriched in differentially expressed genes between poor 90-day ischemic stroke outcome (mRS = 3–5, survivors) vs. vascular risk factor-matched controls (VRFC), and between good 90-day ischemic stroke outcome (mRS = 0–2) vs. VRFC. The top 20 most significant relevant pathways with activation or suppression are presented for the three time-points after stroke: (a) Gene expression at ≤3 h post
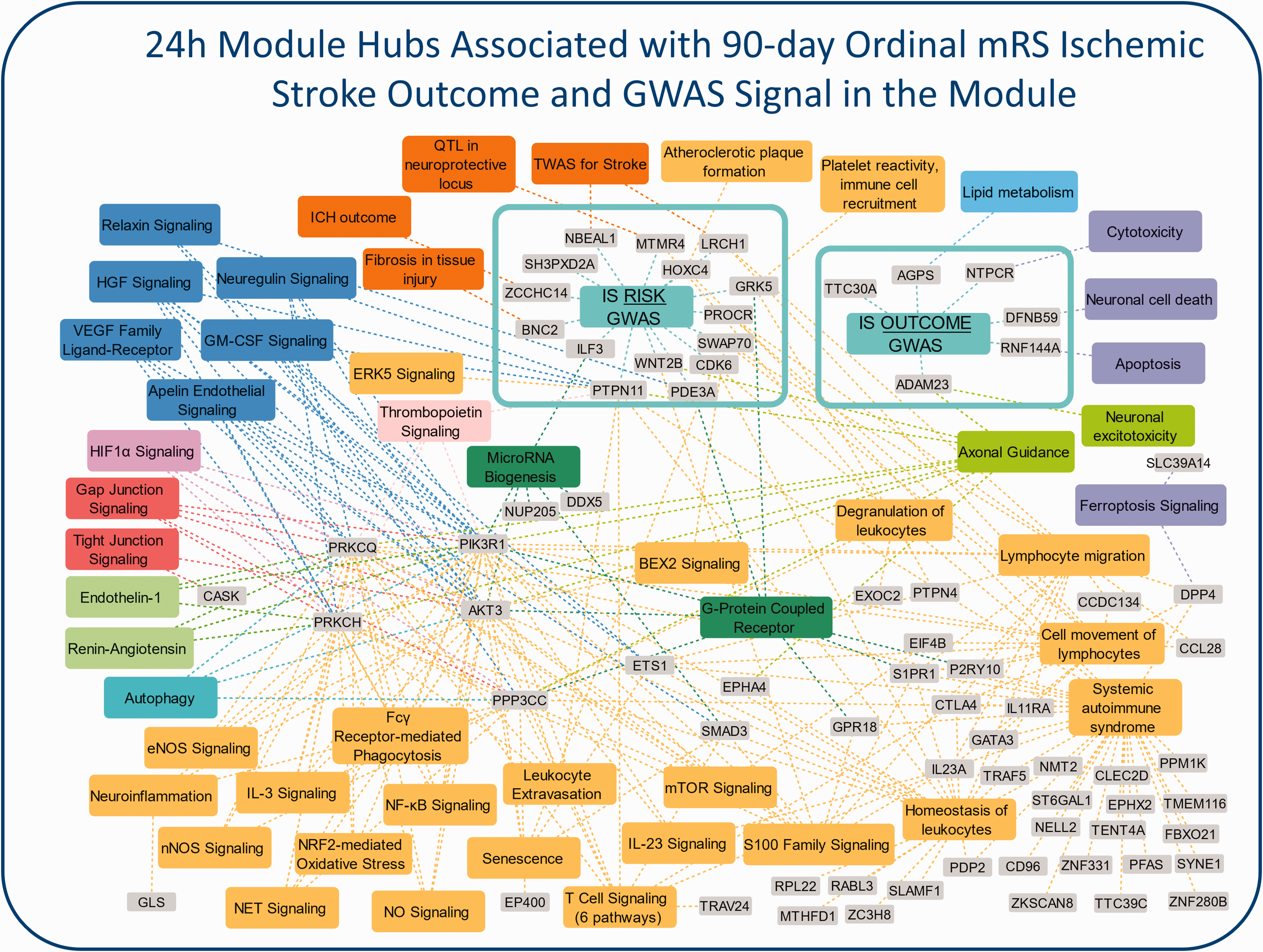
Additional analysis on Amini
Intracerebral hemorrhage outcome
Higher levels of circulating fibrinogen and HMGB1 have been associated with poorer outcomes following human ICH in a review and metanalysis by Kirby et al.
87
Notably,
Association with ICH and perihematomal edema (PHE) volumes
ICH and PHE volumes are key indicators of ICH outcome.
89
Investigating the molecular underpinnings of ICH and PHE volumes could delineate relevant biological processes and potential treatment targets. Durocher & Knepp et al
88
investigated the ICH peripheral blood transcriptome in the subacute phase to delineate peripheral blood genes and co-expression networks associated with ICH volume, absolute perihematomal edema (
Aneurysmal SAH (aSAH) outcome
Several studies have identified potential peripheral blood miRNA biomarkers for aSAH outcome (reviewed in 71 ). Some of the miRNA biomarkers and the pathways they regulate have also been found to be altered in experimental models of SAH, and thus may be good biomarkers for SAH and perhaps even potential treatment targets (reviewed in 91 ). It is possible that composite miRNA-mRNA panels might be even better biomarkers for aSAH outcome.
Biological sex
Sex differences exist for TLR pathways, antigen presenting cells, dendritic cells, macrophages, NK cell and neutrophils from the innate immune system, and in T cells, B cells and immunoglobulins in the adaptive immune system (reviewed in92,93). At the peripheral blood transcriptome level, extensive sex differences encompassing the sex-chromosome expressed genes and the autosome expressed genes have been reported. 94 One-third of the female-biased genes are involved in the immune system response. 94 The role of sex in the immune system aging is not well understood. A peripheral blood mononuclear cell study using ATAC-seq, RNA-seq, and flow cytometry in 172 adults revealed sex-specific differences including a greater magnitude of decline in naïve T cell function and B-cell specific loci, and a greater magnitude of increase in monocyte and cytotoxic cell functions in men than in women. 51 Sex differences in the transcriptome architecture increase after age 65, when men have higher innate and pro-inflammatory and lower adaptive activity in corresponding transcriptional pathways. 51 As age is a significant risk factor for stroke, studying sex-differences in the aging immune system in stroke is likely to be very important.
Sex differences have been reported in stroke epidemiology, pathophysiology, treatment and outcome. 95 Efforts are made to address sex differences more adequately. 96 On a molecular level, sex differences in stroke exist in the immune and coagulation systems which have been examined in both preclinical and clinical studies (reviewed in97–99). An additional layer of heterogeneity is added by the sex-specific genetic architecture of disease, and for cardiovascular disease-associated traits that are stroke comorbidities and risk factors, such as triglycerides, LDL- and HDL-cholesterol, systolic blood pressure, and lipoprotein (a). 100
Sex differences in ischemic stroke
There are sex differences in IS incidence, stroke etiology, and outcome, including the fact that males have increased stroke risk in middle age while females tend to have higher stroke risk when older. 101 Females have more cardioembolic stroke. 101 Post-stroke motor disability and mental impairment are higher in older females than older males. 102 Sex differences in treatment efficacy have also been observed. 101 Hormone-dependent and hormone-independent mechanisms are suggested to contribute to sex differences in stroke. 103 Heterogeneity in the peripheral blood IS transcriptome due to sex has been described in the protein coding and non-protein coding regulatory transcriptomes, in IS of different etiologies, and in its temporal evolution.
A much stronger immune response in the protein-coding peripheral blood transcriptome is observed in women with IS compared to men.104,105 Sex differences were studied in samples from the CLEAR trial (NCT00250991)
32
with peripheral blood draws at ≤3 h (before treatment), 5 h and 24 h post
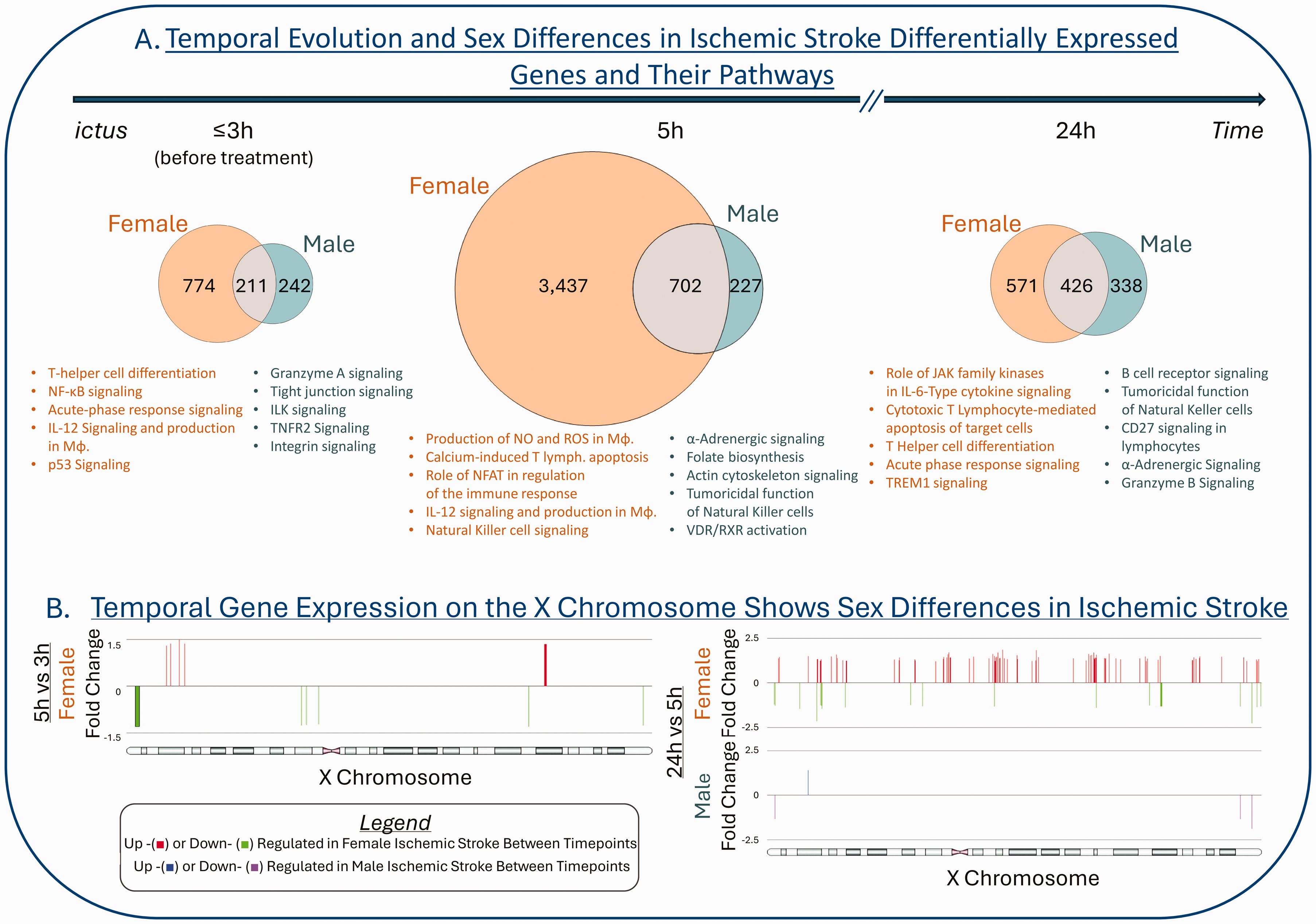
Sex Differences in the Peripheral Blood Transcriptional Response to Ischemic Stroke. (a) Proportional Venn diagrams of the numbers of the differentially expressed genes between ischemic stroke female and control female and ischemic stroke male and control male (Benjamini-Hochberg false discovery rate < 0.05, |fold change| > 1.5) for each time point (≤3 h, before treatment; 5 h and 24 h post
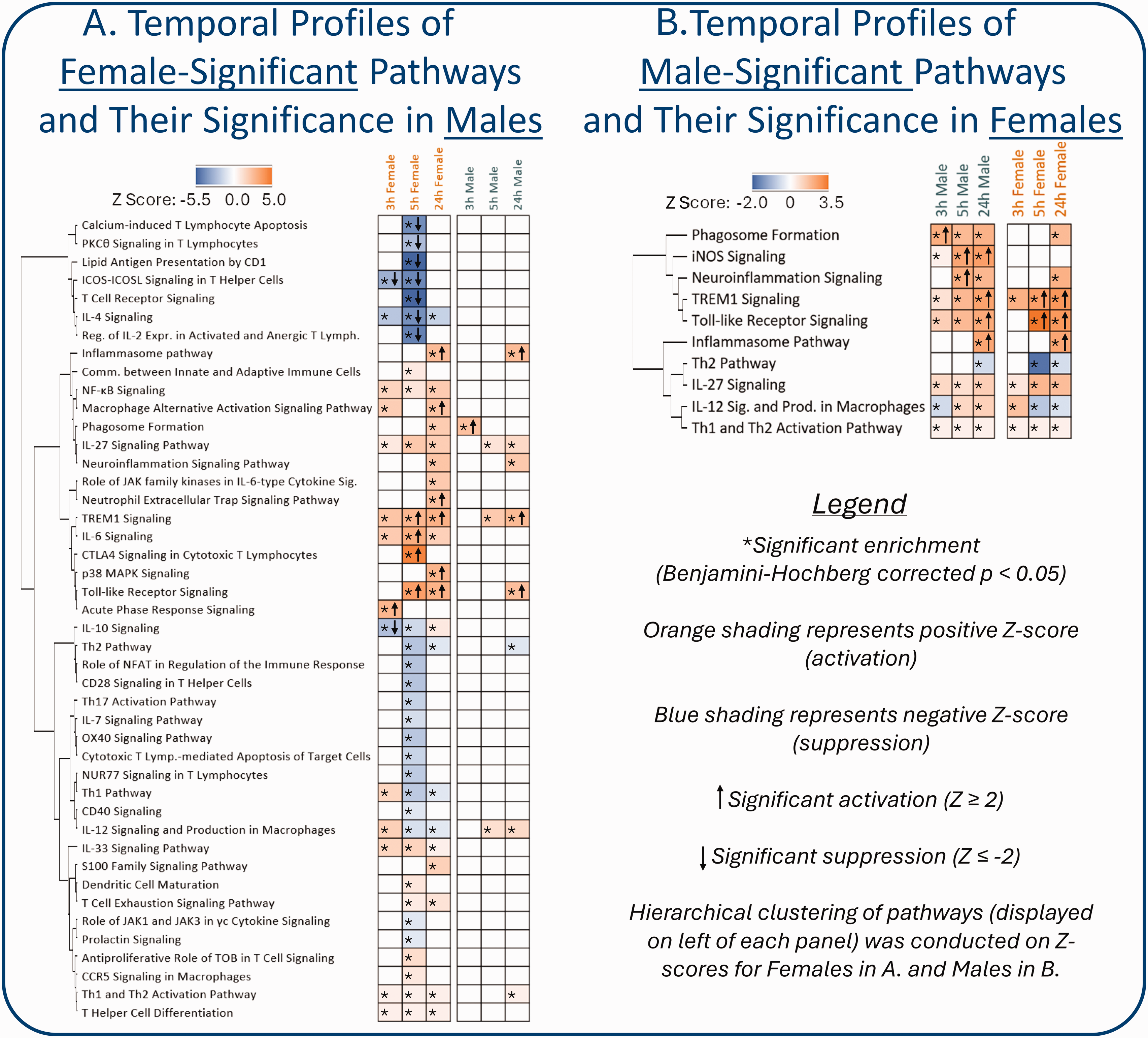

Additional analysis on Tian & Stamova
At the level of the non-coding peripheral blood transcriptome, Dykstra-Aiello et al 110 investigated the sex differences in the long-noncoding RNA (lncRNA) transcriptome in 266 whole-blood RNA samples drawn from IS patients and VRFCs. The subjects were split into derivation and validation cohorts. Notably, in this study there were more lncRNA regulated in IS males than in IS females. Only 6 lncRNA were common between the male and female IS responses, with 4 (mRNAlike lncRNA SPATA3, linc-GUSB-3, linc00671, mRNA-like lncRNA/XIST regulator) expressed in opposite directions. Additionally, IS females had significantly more down regulated lncRNAs over time than IS males. Seven lncRNA with sex-specific differential expression were near vascular risk factor/stroke risk loci. This study also showed sex-specific lncRNA expression changes in each of the three main IS etiologies: CE, LVA and SV IS etiologies. 110 Sex differences in the non-coding regulatory elements of the IS peripheral blood transcriptome are still largely unexplored and warrant further investigation.
Sex differences in ICH
Sex differences in ICH exist in early PHE expansion which is greater in male ICH patients. 111 Male ICH patients also have higher rates of hematoma expansion, 90-day mortality rates, 112 and pneumonia risk. 113 Though females have lower ICH incidence than men, they have a higher 1-year mortality possibly related to older age at ICH onset. 114
Though little is known about the sex differences in the ICH peripheral blood transcriptome in humans, a small pilot investigation of Deep and Lobar ICH did show transcriptome sex differences in ICH. 44 The study found many more peripheral blood differentially expressed genes in ICH females compared to ICH males. 44 The male ICH response was associated with DNA Methylation and Transcriptional Repression and Apelin Liver Signaling Pathways in deep ICH, and with Protein Binding in Lobar ICH. The female ICH response, on the other hand, was enriched in many immune and coagulation pathways. For example, several T cell pathways were predicted to be significantly suppressed. TLR Signaling and Neuroinflammation Signaling were predicted significantly activated in female ICH, and NF-κB showed a trend towards activation in both lobar and deep female ICH.
The finding of a greater immune response in the coding peripheral blood transcriptome in female ICH compared to male ICH is similar to the sex differences observed in IS discussed above. It is possible males have a greater response to stroke in other layers of the blood transcriptome such as the non-protein coding transcriptome as was observed in IS above.
The human peripheral blood transcriptome findings, the epidemiological sex differences, as well as the experimental stroke literature on sex differences in stroke97,99 all highlight that it is essential to consider sex in both clinical and pre-clinical studies to improve clinical translation. Understanding sex differences is critical to developing sex-specific medicine in stroke, such as sex-specific treatment targets, prevention strategies, and diagnostic, prognostic, predictive, and pharmacotranscriptomic biomarkers.
Stroke pathophysiology and diagnostic biomarkers
The peripheral blood transcriptome architecture following stroke has been examined in several studies (reviewed in2,30,50,115). Since the focus of this review is heterogeneity in stroke subgroups, these studies will not be reviewed here. In addition, diagnostic biomarker development, such as for distinguishing ischemic stroke from ICH and SAH, 116 IS etiology and predicting the stroke mechanism of cryptogenic IS 30 and diagnosing IS117,118 are discussed in other reviews and not repeated here. However, given the transcriptional heterogeneity documented in this review, it seems likely that separate biomarker panels might be needed to more accurately predict/diagnose molecular subgroups for different sexes with different comorbidities.
Transcriptional heterogeneity in common pathways
Though common pathways have been identified in different phenotypic subgroups, caution must be exercised since different genes may be modulated in the common pathways. For example, even though there were common pathways in the peripheral blood transcriptome of ICH patients with hemorrhage in Deep ICH and Lobar ICH as compared to controls, 44 we found many of the affected genes in the common pathways were ICH-location-specific. For instance, the NF-κB signaling pathway was a common pathway over-represented with differentially expressed genes in lobar ICH and deep ICH. However, the genes were mainly location-specific (Figure 6(a), based on data from 44 ; for details, please see Supplementary Material). Genes from the NF-kB pathway are potential treatment targets, 40 but any target should consider ICH location. We also found heterogeneity in the differentially expressed genes in common biofunctions associated with the peripheral immune response to ischemic stroke in males and females. For example, deeper investigation of data from Tian & Stamova et al 105 (described above; for details, please see Supplementary Material), revealed that the Cellular Infiltration of Granulocytes biofunction was significantly over-represented with differentially expressed genes between female IS and female control, as well as between male IS and male control at the ≤3 h timepoint (before thrombolytic treatment). However, many of the genes were sex- specific (Figure 6(b); for details, please see Supplementary Material). Out of the 33 genes in females and the 16 genes in males participating in the Cellular Infiltration of Granulocytes biofunction, only 8 overlapped (Figure 6(b)). The largest fraction of the peripheral blood granulocytes are neutrophils. They are among the first cells to infiltrate the injured brain post ischemic stroke. Neutrophils are a major source of matrix metalloproteinases (MMPs) which disrupt the BBB and exacerbate the brain damage. 7 Inflammatory cell infiltration/function has been considered a major therapeutic target in ischemic stroke. 7 Thus, consideration for sex differences in this biological process and/or in its temporal evolution needs to be considered. Overall, molecular heterogeneity in stroke subgroups can have significant repercussions when treatment and prevention strategies are developed and may warrant search for subgroup-specific treatments.
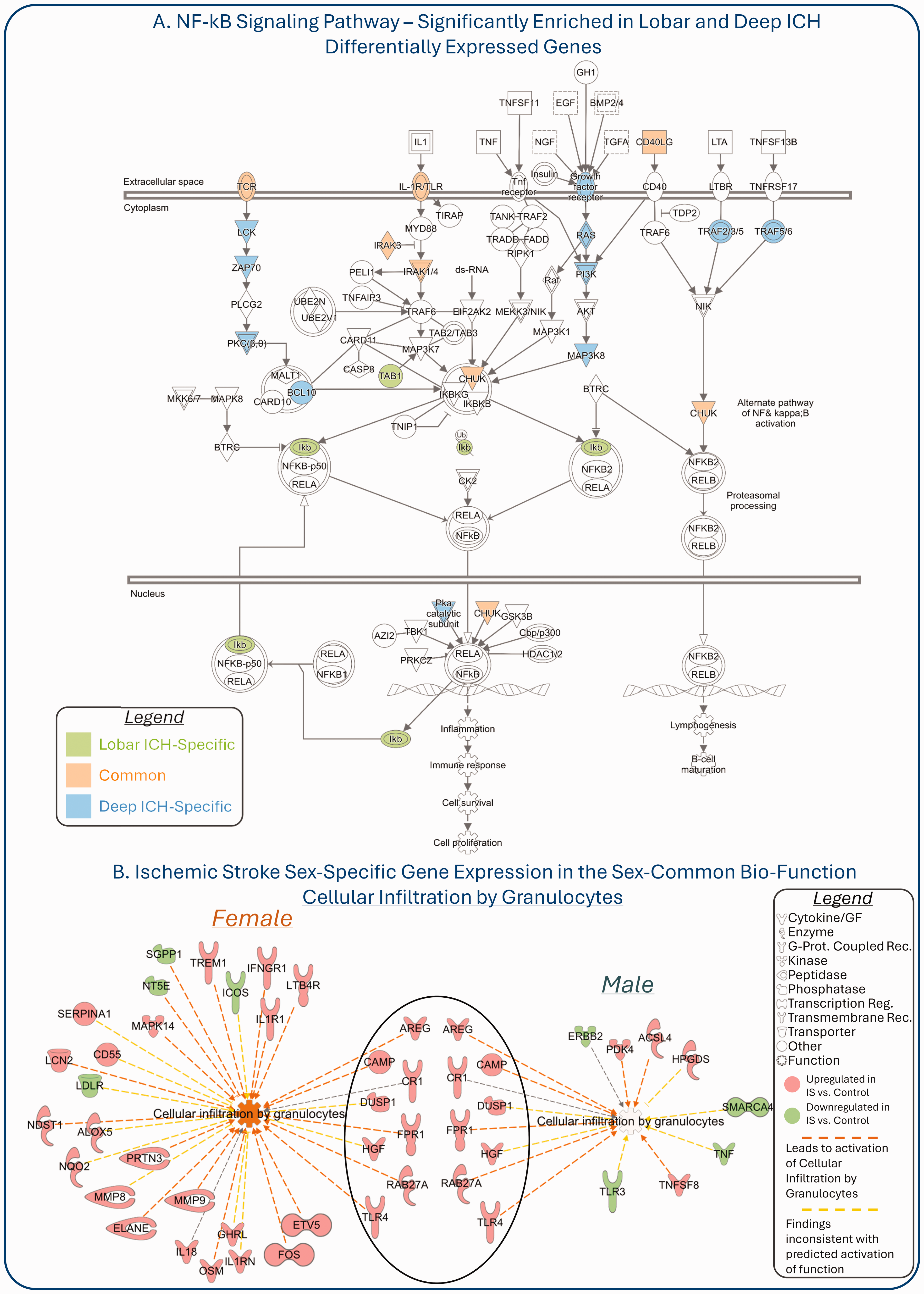
Subgroup-specific heterogeneity in common pathways. (a) Based on data from Knepp et al, Brain Hemorrhages, 2022 (PMID: 36936603). ICH location-specific differentially expressed genes (DEGs) in the ICH location-common NF-κB Signaling Pathway. DEGs between Lobar ICH and Control are highlighted in green; DEGs between Deep ICH and Control are highlighted in blue; DEG in common – in orange. (b)
Combinatorial molecular heterogeneity of affected pathways in the human ICH peripheral blood transcriptome: Patient-Specific architecture
As noted throughout this review, significant and pervasive molecular heterogeneity in stroke subgroups exists. To molecularly phenotype this heterogeneity at an even more refined level, we applied an approach we developed previously 119 to delineate the combination of affected pathways at an individual patient-level resolution. To adapt this approach for use in stroke, we re-analyzed the peripheral blood transcriptome of 66 participants (33 ICH, 33 VRFC) we previously utilized to define the inflammatory, regulatory and autophagy gene co-expression modules associated with ICH, and the differentially expressed genes between ICH and VRFC. 11 We originally identified 1,225 differentially expressed genes (FDR p < 0.05, FC>|1.2|) between the ICH and VRFC transcriptomes. 11 The differentially expressed genes were over-represented in 116 canonical pathways (Benjamini-Hochberg -corrected p < 0.05). 11 To evaluate the subject-level heterogeneity here, we first decreased the number of differentially expressed genes by increasing the stringency (FDR p < 0.05, FC > |2|), which resulted in a total of 105 differentially expressed genes. Ingenuity Pathway Analysis (IPA®) revealed over-representation in 29 significant pathways (Benjamini-Hochberg p < 0.05) (Figure 7). Many inflammatory pathways such as Neutrophil Degranulation, Pyroptosis Signaling (an inflammatory cell death pathway), Inflammasome pathway, and IL-6 Signaling were predicted significantly activated or with a trend towards activation in ICH vs VRFC. Some anti-inflammatory pathways, such as IL-10, LXR/RXR signaling, and PPAR/RXR signaling were predicted to be suppressed in ICH vs VRFC.
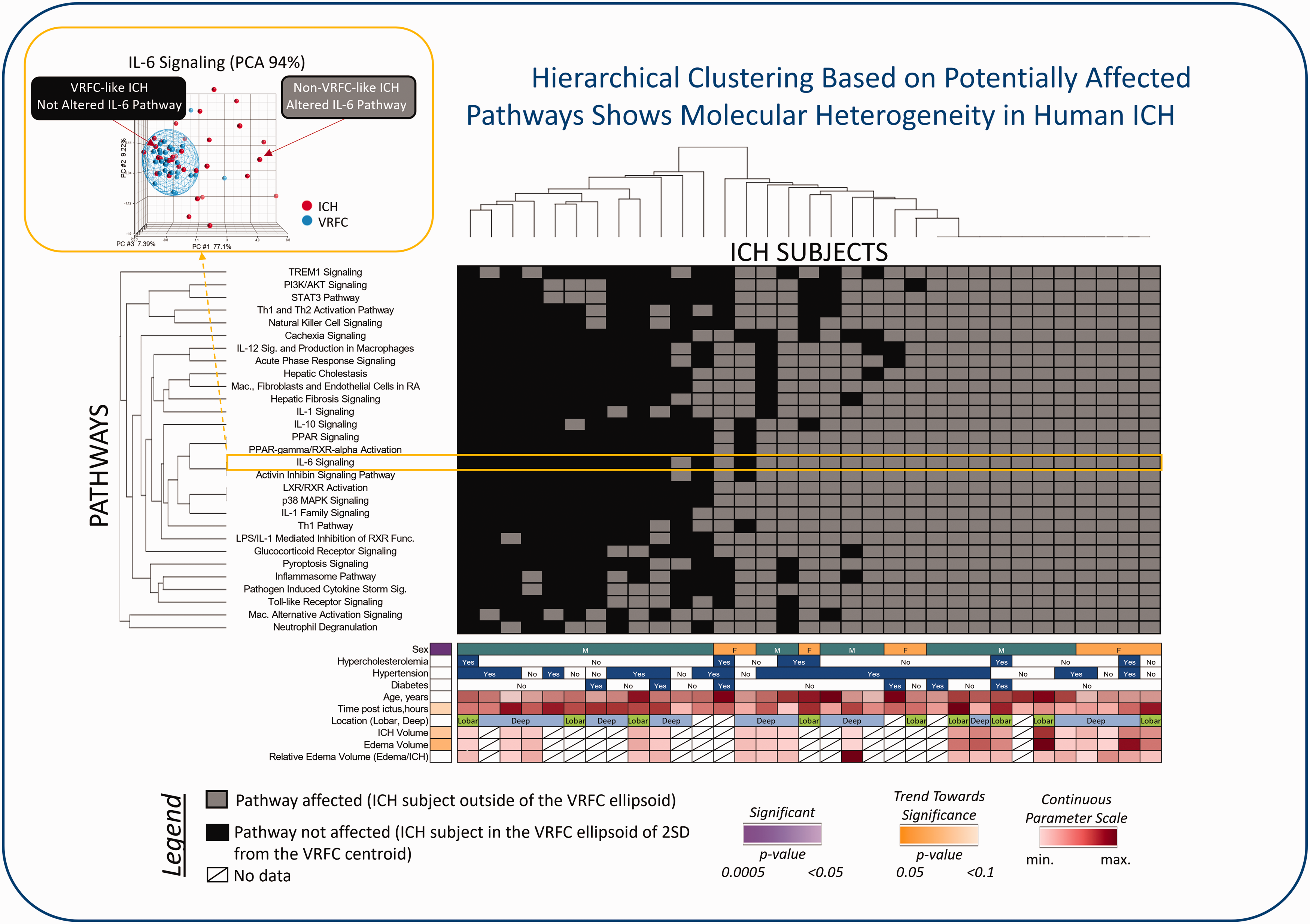
Hierarchical clustering of ICH subjects based on their affected pathways. On the y-axis – pathways significantly enriched with differentially expressed genes in peripheral blood between ICH and vascular risk factor-matched control (VRFC) subjects. On the x-axis – ICH patients. We scored an ICH subject as “VRFC-like ICH” (represented by a black box in the heatmap) if the ICH subject fell within 2 Standard Deviations (SD) of the VRFC centroid, suggesting this ICH subject did not have this particular pathway altered. Alternatively, we scored an ICH subject as “non-VRFC-like ICH” (represented by a grey box in the heatmap) if this ICH subject fell outside the 2SD of the VRFC centroid. PCA of the IL-6 signaling pathway based on the 5 DEGs in this pathway is presented in the upper left as an example. In this PCA, the blue mesh (VRFC’s ellipsoid) is drawn at 2SD of the VRFC centroid. Arrows point to an example of an ICH subject for which the IL-6 signaling pathway is not altered (the ICH subject falls within the blue ellipsoid; VRFC-like) and to an example of an ICH subject for which the IL-6 signaling pathway is altered (the ICH subject falls outside of the VRFC ellipsoid; Non-VRFC-like). Clinical and demographic characteristics of each ICH patient are represented on the bottom of the heatmap, as well as the significance of the association between these parameters and the number of altered pathways. For example, Sex was found significant, indicating that the average number of altered pathways in Females was different from the average number of altered pathways in Males. For details, please see Supplementary Material.
We then determined which of the 29 pathways were dysregulated/altered in each of the 33 ICH subjects. To do this unique analysis, we performed PCA mapping based on the differentially expressed genes present in each of the 29 significant pathways. In each of these 29 PCAs, we scored an ICH subject as “VRFC-like ICH” if the ICH subject fell within 2 Standard Deviations (SD) of the VRFC centroid, suggesting this ICH subject did not have this particular pathway altered. Alternatively, we scored an ICH subject as “non-VRFC-like ICH” if this ICH subject fell outside the 2SD of the VRFC centroid. PCA of the IL-6 signaling pathway based on the 5 differentially expressed genes in this pathway (CEBPB, IL18RAP, IL1R, IL1RAP, MAPK14) is presented in Figure 7 (for details, please see Supplementary Material), upper left as an example. In this PCA, the blue mesh (VRFC’s ellipsoid) is drawn at 2SD of the VRFC centroid. Arrows point to an example of an ICH subject for which the IL-6 signaling pathway is not altered (the ICH subject falls within the blue ellipsoid; VRFC-like ICH) and to an example of an ICH subject for which the IL-6 signaling pathway is altered (the ICH subject falls outside of the VRFC ellipsoid; Non-VRFC-like ICH). We scored each of the 33 ICH subjects as either having a particular pathway not altered, represented by a black rectangle in the hierarchical clustering dendrogram in Figure 7, or as having a particular pathway altered, represented by a gray rectangle in Figure 7. This procedure was performed for each of the 29 significantly over-represented canonical pathways. Hierarchical clustering along the 33 ICH subjects and the 29 canonical pathways’ alteration status showed varying combinations of pathways were affected in each of the ICH subjects (Figure 7). Eleven out of the 33 ICH subjects had all 29 pathways altered. The remaining 22 ICH subjects showed different combinations of altered pathways, revealing what we refer to here as combinatorial pathway heterogeneity.
Females had significantly more altered pathways than males (p = 4.8E-4). A trend toward significant negative correlation between the number of altered pathways and time post
Additionally, we investigated the 29 pathways individually by comparing the ICH subjects that had a particular pathway altered
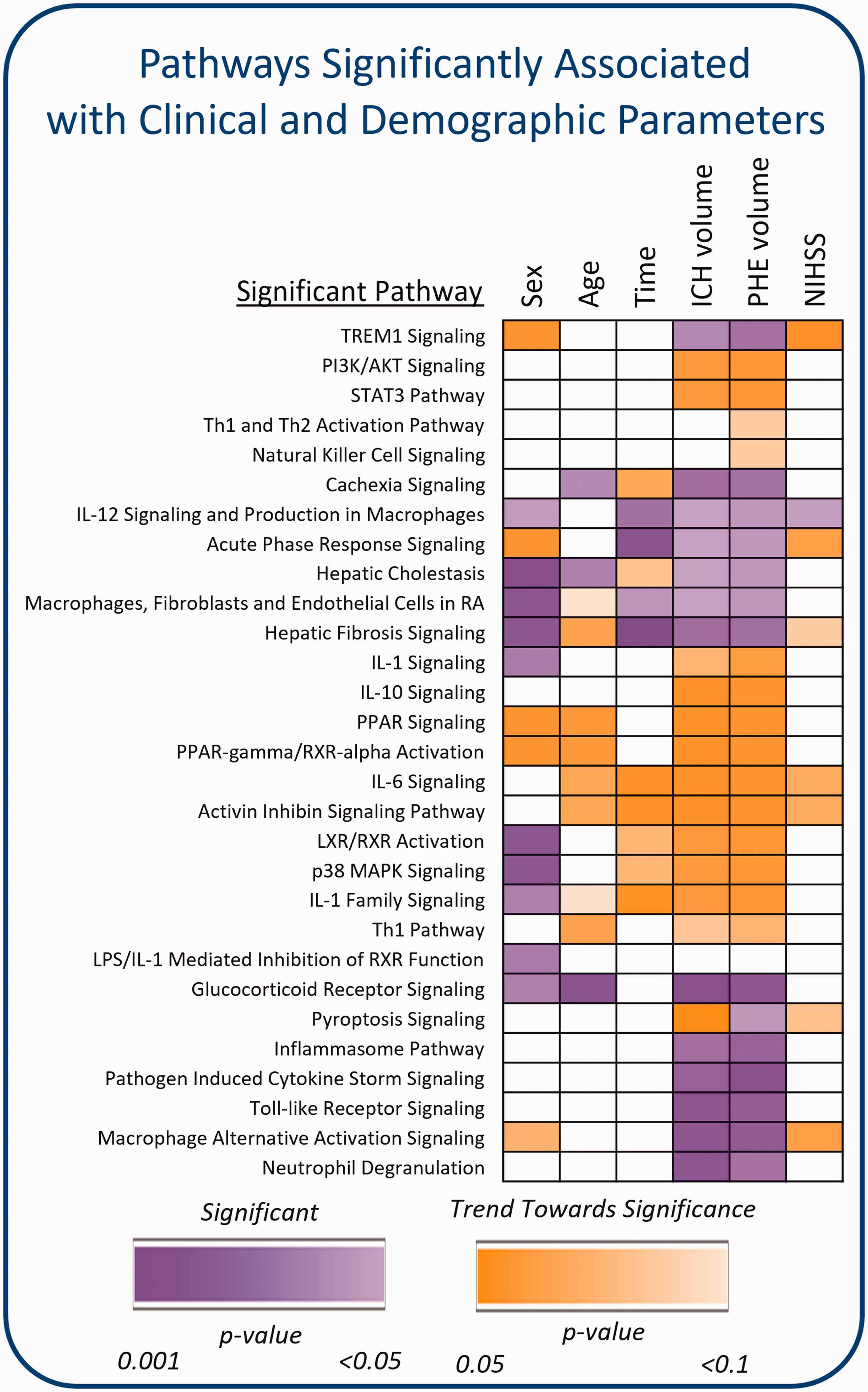
Association between significant pathways from Figure 7 and clinical and demographic parameters. The heatmap represents differences between the ICH subjects with a particular pathway altered (non-VRFC-like ICH)
We propose that this method can identify the combinatorial heterogeneity in ICH altered pathways, based on ICH-patient level pathway subgroups. The findings further our understanding of the ICH molecular heterogeneity at an individual-patient level. The proposed approach can be applied to other phenotypic levels, such as proteome, metabolome, epigenome, and imaging data. It strongly underscores potential repercussions for choosing treatment targets and may provide consideration in identifying candidate patients in clinical trials targeting a particular pathway. For example, 64% of the subjects (21 out of the 33 ICH subjects) in Figure 7 showed potential alterations in the PPARγ/RXRα Activation pathway. PPARγ agonists have been shown to improve experimental ICH outcomes. 120 Thus, identifying which ICH subjects have alterations in this pathway may help pinpoint potential candidates in clinical trials targeting this pathway. The approach may contribute to the overarching goal of advancement towards precision medicine in stroke. Combining this approach with recently developed framework for automatically identifying responsive subgroups regarding treatment effects from randomized clinical trial data 121 may further help advance personalized stroke treatments.
Conclusions
The peripheral immune and coagulation responses contribute to injury and repair mechanisms in the injured brain following stroke. This is underlined by molecular changes at many biological levels including the transcriptome. Transcriptomic studies in peripheral blood following human stroke have revealed considerable heterogeneity in the transcriptome architecture in subgroups related to etiology, comorbidities, biological sex, outcome, response to treatment, temporal evolution, cell-specific responses, and even common pathways. Additionally, we show that subject-level combinatorial heterogeneity exists with different stroke subjects having different combinations of pathways potentially altered. Differences in the individual’s response to stroke reveal molecular subgroups and emphasize the need for precision medicine in future treatments and therapeutics. Knowledge of the peripheral transcriptome architecture in human stroke is evolving, and new technologies, such as single-cell RNA sequencing (reviewed by Shi, Chen, & Zhang et al in this special issue) and long-read RNA sequencing (for directly examining the transcript-level resolution) will continue refining it. Additional studies deciphering deeper differences in the human stroke peripheral blood transcriptome, such as ones due to RNA modifications and RNA editing in stroke subgroups, as well as long-distance communication between the brain and peripheral blood via extracellular vesicles, will help decipher stroke phenotypic subgroups. The molecular heterogeneity may have significant repercussions for the development of treatment targets and prevention strategies, and sub-group specific approaches may be needed to advance towards personalized medicine.
Supplemental Material
sj-pdf-1-jcb-10.1177_0271678X251322598 - Supplemental material for Molecular heterogeneity in human stroke – What can we learn from the peripheral blood transcriptome?
Supplemental material, sj-pdf-1-jcb-10.1177_0271678X251322598 for Molecular heterogeneity in human stroke – What can we learn from the peripheral blood transcriptome? by Boryana Stamova, Bodie Knepp and Fernando Rodriguez in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by R01NS127976 and R33NS119346 from the NIH National Institute of Neurological Disorders and Stroke (NINDS) to B.S.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: B.S. holds a patent on gene expression signatures for predicting stroke and its causes.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
