Abstract
Cerebral blood flow (CBF) can be measured with dynamic positron emission tomography (PET) of 15O-labeled water by using tracer kinetic modelling. However, for quantification of regional CBF, an arterial input function (AIF), obtained from arterial blood sampling, is required. In this work we evaluated a novel, non-invasive approach for input function prediction based on machine learning (MLIF), against AIF for CBF PET measurements in human subjects.
Twenty-five subjects underwent two 10 min dynamic 15O-water brain PET scans with continuous arterial blood sampling, before (baseline) and following acetazolamide medication. Three different image-derived time-activity curves were automatically segmented from the carotid arteries and used as input into a Gaussian process-based AIF prediction model, considering both baseline and acetazolamide scans as training data. The MLIF approach was evaluated by comparing AIF and MLIF curves, as well as whole-brain grey matter CBF values estimated by kinetic modelling derived with either AIF or MLIF.
The results showed that AIF and MLIF curves were similar and that corresponding CBF values were highly correlated and successfully differentiated before and after acetazolamide medication. In conclusion, our non-invasive MLIF method shows potential to replace the AIF obtained from blood sampling for CBF measurements using 15O-water PET and kinetic modelling.
Keywords
Introduction
Measurements of cerebral blood flow (CBF) can be used to separate pathological and healthy brain tissue as well as for functional brain research. Tracer kinetic modelling following dynamic 15O-water positron emission tomography (PET) imaging with arterial blood sampling is considered the reference standard for CBF measurements.1–6 However, arterial cannulation is an invasive, laborious and time-consuming procedure, and may, due to induction of pain and risk for complications, discourage patients and volunteers from participating in research studies. Furthermore, a useful arterial input function (AIF) curve cannot be obtained without careful cross calibration of the blood measurement detector and the PET scanner. In addition, because the blood is most commonly sampled from the radial artery, additional corrections for dispersion and delay of the tracer must be applied, to obtain the true AIF for the brain. 3 ,7–9
The use of an image-derived input function (IDIF) has been proposed as an alternative approach to overcome the challenges with the AIF. 10 In brain PET imaging, an IDIF can be measured inside a suitable intracranial blood vessel directly in the reconstructed PET images, for instance in the intracranial carotid arteries.11–13 Due to the limited spatial resolution of the PET system, and the need for short time-frames during the first pass of the bolus, image-derived methods suffer from both partial volume effects and image noise. These limitations require complex and standardized methods for partial volume correction and artery delineation, which may be difficult to achieve in practice. Recently, a few clinical studies have suggested the use of integrated PET/magnetic resonance imaging (MRI) for deriving an IDIF in brain, where the latter modality is used for artery delineation or even motion correction.14–18 However these methods are sensitive to registration errors between the modalities and require detailed knowledge of the scanner resolution. Another recent study overcame potential misregistration problems and formed a corrected IDIF by deriving total number of counts and artery volume from the two modalities separately. 19 However, this method was not yet validated with arterial blood sampling and as hybrid PET/MRI is still an emerging modality, to date, image-derived methods are rarely used in larger clinical or research studies. 10 , 20
Alternatively, a standardized, population-based AIF can be calculated as an average AIF from a group of subjects acquired with the same tracer, injection protocol and population, and scaled to the specific subject. 21 , 22 However, this method requires at least one blood sample for curve scaling while individual physiological differences and scan-dependent variations are neglected.
An approach on image data with simultaneous estimation of AIF and kinetic parameters has also been reported.23–25 This method, however, assumes a known mathematical AIF model and requires at least one late blood sample for parameter estimation. Recently, non-invasive simultaneous estimation methods were developed that obviate the need for the single late blood sample by using additional input variables from electronic health records into the models. 26 , 27 The limitation of such an approach is that a large set of clinical variables must be collected and handled for each patient. These variables may not necessarily be available in the health records for all patients and may even complicate inclusion of healthy volunteers in research studies.
In this study we use a machine learning-based approach for AIF estimation. Machine learning-methods are especially useful for function estimation and regression.
28
Briefly, one seeks to determine a function,
In our previous work, we developed and validated a machine-learning-based input function for 18F-fluorodeoxyglucose (FDG) in a mouse PET cohort. 29 In short, two learning models were evaluated that predicted an AIF from time-activity curves of up to 7 different tissue regions as input. The main limitation with our previous study was the lack of an AIF, thus the models could only be validated against a reference IDIF. However, in mouse PET scanning, the entire body of the mouse fits in the PET field-of-view, thus, time-activity curves from all organs are readily available as input data for the models. We showed that, for instance, the myocardium and liver were important for AIF prediction, because their time-activity curves closely resembled the reference IDIF. In contrast, these blood-rich organs are outside the field-of-view in clinical brain PET imaging, and thus, alternative input curves had to be derived for the MLIF model in the current study.
In the present study, we have further developed the MLIF approach for human 15O-water brain PET data and evaluated the models against an AIF obtained from continuous arterial-blood sampling. We aimed to show that an AIF could be accurately predicted by an MLIF model using multiple image-derived input data from the carotid arteries. We hypothesized that there were no significant differences in estimated CBF when using either AIF or MLIF, and investigated similarities and differences between image-derived sampling in the brain versus arterial-sampling from the arm. Further, we investigated whether the MLIF method was capable to predict a clinically relevant CBF difference between scans before and after acetazolamide medication.
Materials and methods
Subjects
Pseudonymized data from 25 subjects were retrospectively collected from a completed clinical research study at Uppsala University Hospital. The data comprised both patients with multiple sclerosis (MS) and healthy volunteers (mean age (range) in years: 40 (23–56); F:M 15:10). In this methodological study, we did not differentiate between the two groups as we considered that the subject’s health status had no impact on our evaluation of the MLIF model. Therefore, all authors were blinded for the health status of each subject and thus, no comparisons were made between healthy subjects and MS-patients. The results of the parent study will be reported elsewhere.
The parent study was approved by the Swedish Ethical Review Authority (reference 2014/453). All subjects signed written informed consent prior to inclusion. Since the present work was purely an image analysis methodology study using pseudonymized data, it was not covered by the Swedish or Norwegian regulations on medical research in humans and as such, no additional ethics approval was necessary.
Image acquisition
All subjects underwent two 10 min dynamic brain PET scans on either an ECAT Exact HR+ stand-alone PET scanner (Siemens, Knoxville, TN; n = 9) or a Discovery MI PET-Computed tomography (CT) scanner (GE Healthcare, Waukesha, MI; n = 16). The scans started simultaneously with an automated bolus injection of 5 MBq/kg 15O-water (15O-water in 5 ml saline at 1 ml/s followed by 35 ml saline at 2 ml/s). Each subject underwent one scan at baseline and one scan 15–30 min after intravenous administration of acetazolamide (9 mg/kg up to a maximum of 1000 mg; 5 min infusion) such that every subject was its own control. Acetazolamide medication dilates the vascular system and thereby it increases the cerebral arterial blood flow velocity.30–32 Attenuation correction was based on a 10 min transmission scan with rotating 68Ge rod sources (ECAT) or an ultra-low-dose CT scan (Discovery MI). Images were reconstructed into 26 time-frames (1 × 10, 8 × 5, 4 × 10, 2 × 15, 3 × 20, 2 × 30 and 6 × 60 s). Image reconstruction algorithms were chosen to result in a matching image resolution for the two scanners: ordered subsets expectation maximization with 6 iterations, 8 subsets and a 4 mm Hanning filter (ECAT) and 3 iterations, 16 subsets and a 5 mm Hanning filter (Discovery MI).
In addition, all subjects underwent MRI on a 3 T MRI scanner (Achieva, Philips Healthcare, Best, The Netherlands) with a 32-channel head coil. A three-dimensional T1-weighted gradient echo sequence was obtained with voxel size 1.0 × 1.0 × 1.0 mm3, repetition time = 8.2 ms and echo time = 3.7 ms.
Blood sampling
Continuous arterial blood sampling was performed during 10 min for each scan (3 ml/min) using either an ABSS V3 (Allogg, Mariefred, Sweden; subjects scanned on ECAT) or PBS-100 (Veenstra-Comecer, Joure, The Netherlands; subjects scanned on Discovery MI). A single arterial blood sample was taken through a three-way-valve on the arterial line 5 min post injection and measured in a cross-calibrated well counter for calibration of the continuous arterial blood data.
The measured blood signal, 
A mono-exponential dispersion model was assumed
8


The dispersion constant was fixed to 15 s for all subjects in this study, as suggested in the literature. 8
The dispersion-corrected AIFs were delay-corrected, as described in the ‘Image processing’ Section.
A visual assessment of the AIF curves was performed to identify abnormal AIFs due to failures in tracer administration or continuous arterial blood sampling. Three subjects were excluded after the visual assessments.
Image processing
All images were corrected for inter-frame motion using in-house written software in Matlab (Mathworks, MA, USA). A simple and objective multi-VOI thresholding method, that could capture blood information from the carotid arteries, was empirically developed. First, to remove noise close to the edge slices, a PET search-volume was defined by trimming 20 voxels in the x-y-plane periphery, and by removing 5 slices in the z-direction. The algorithm for identifying the time-frame for carotid-VOI thresholding was based on a frame-wise graph of whole-brain gray-matter total intensity. The time-frame used for VOI thresholding was the first frame where the total intensity was larger than 25% of the maximum value in this graph. Three VOIs were generated, comprising the 10, 100 and 1000 highest-intensity voxels (Figure 1a to c). The median voxel value was derived for each time-frame and VOI, resulting in three IDIF time-activity curves, named IDIF10, IDIF100 and IDIF1000 (Figure 1d), which could be interpreted as three different image-derived blood-curves with different amounts of partial volume effect.
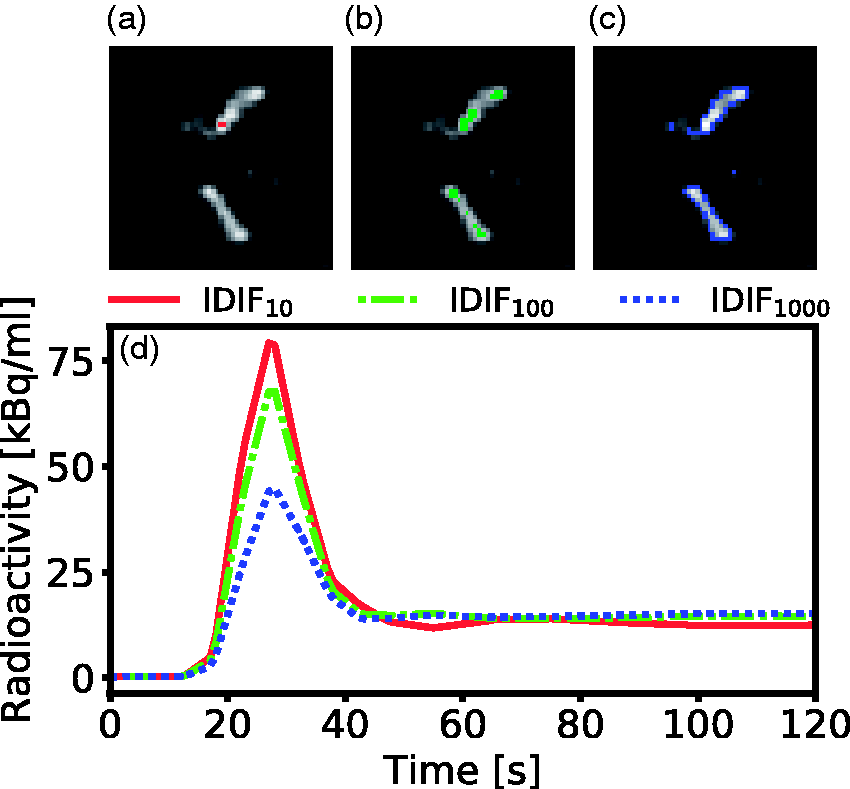
Outline of the VOI thresholding method implemented in this work. (a) to (c) shows an enlarged axial brain PET slice of the optimal time-frame for a representative subject. Highlighted are the parts of the IDIF10 (a), IDIF100 (b) and IDIF1000 (c) comprising the 10, 100 and 1000 highest intensity voxels. (d) The resulting time-activity curves during the first two minutes for IDIF10-1000. The IDIF10 captures the highest activity in the carotid artery, while the IDIF100 and IDIF1000 shows a lower activity due to a higher number of voxels included.
Subsequently, to match the AIF sampling, all PET data were interpolated linearly to one second time framing. To correct for delay between the AIF and the PET data, the dispersion corrected AIF was shifted to provide the best overlap with the IDIF10 curve, i.e. where the dot product between the two curves was maximized.
In all following analyses, only PET and AIF data from 0–6 min were used, to minimize noise from late parts of the scans. Calibration of the AIF curves with the single arterial blood sample was only conducted for the training data, and thus no AIF or blood sample was required for the test data.
To extract time-activity curves for whole-brain grey matter, T1-weighted MRI images were co-registered to PET images and segmented using SPM8 (The Wellcome Centre for Human Neuroimaging, UCL Queen Square Institute of Neurology, London, UK). All image analysis was performed in the native PET image space.
Function prediction using Gaussian processes
Gaussian processes (GP) is a well-known, non-parametric Bayesian regression method which has been frequently used within machine learning for data-driven function estimation and regression tasks.34,35 One advantage with GP is that it predicts not only the mean function, but also its variance, thus providing an uncertainty measure of the model. 36 Furthermore, GP, is known to work well with sparse training datasets, as opposed to neural networks.37,38
In GP regression, the output 
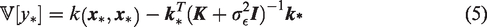
Here
Input function prediction
For input function prediction, leave-one-out cross validation was employed, which is a common validation method in machine learning with limited amounts of data. 28 In short, one sample was withdrawn from the dataset and assigned as test sample, while the remaining samples were allocated for training. The three time-activity curves (Figure 1) were used as input vectors into the GP framework, and the MLIF and variance of the test sample were predicted using equations (4) and (5), respectively. The process was repeated by assigning a new sample as test sample, until all samples had been tested once.
In all experiments, the Matérn covariance function was chosen, with
Kinetic modelling
Quantification of CBF was performed on the whole-brain grey-matter region. A single-tissue compartment model was used to generate CBF values. This model assumes that water can diffuse freely between vascular and tissue space, with activity concentrations 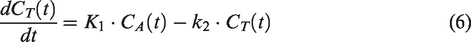
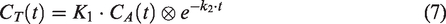
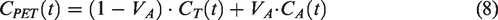
For tracers with high extraction fraction relative to the blood flow, such as 15O-water, CBF equals
Evaluation design
The aim of the current work was to investigate whether an AIF could be accurately predicted using an MLIF model in baseline and acetazolamide scans of the same subject. In a first case, a GP model named MLIF1 was trained and subsequently tested on baseline scans using leave-one-out cross validation. Similarly, another GP model, named MLIF2, was trained and subsequently tested on acetazolamide scans using leave-one-out cross validation.
In a routine setting, it is of interest to train a predictive model on normal subjects and apply the same model on a disease, or medicated group. Therefore, in a second case, an additional GP model named MLIF*1, was trained on all baseline scans, and subsequently tested on all acetazolamide scans. Here, the asterisk (*) in MLIF*1 emphasises that all subjects from the baseline scan were included in the training data for this model, as opposed to the leave-one-out training for MLIF1. Essentially, one might see MLIF*1 and MLIF1 as two models trained on the same dataset, as the difference is only one subject.
Finally, we hypothesized that MLIF*1 might be more representative for a local AIF in the brain, compared to an AIF sampled in the wrist. Therefore, in a third case, we aimed to evaluate the CBF increase from baseline to acetazolamide scans obtained by the two different methods in
The evaluation design is summarized in Table 1.
The evaluation design of the MLIF method.
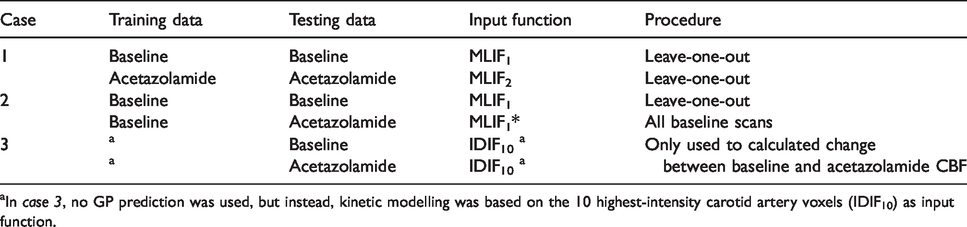
aIn
Evaluation methods
The GP predicted MLIF curves were first compared point-by-point to AIF using orthogonal regression. Subject scans with regression slopes that were more than three scaled-median absolute-deviations away from the median slope, were considered outliers, and removed from further model comparisons.
45
Time–activity curves averaged over subjects were calculated for: whole-brain grey matter, AIF, the MLIF models and IDIF10. CBF and VA were estimated for whole-brain grey matter using kinetic modelling with both AIF and the GP-predicted MLIF (
MLIF-based CBF estimates were compared with the one based on AIF by paired t-test (
The GP variance (equation (5)) was considered as a measure of prediction error. This measure was evaluated by the relationship between the magnitude of the CBF-ratio (CBFMLIF/CBFAIF) and GP variance for
Results
CBF values based on AIF (CBFAIF) and MLIF (CBFMLIF) are shown in Supplementary Table S1. The average CBFAIF and CBFMLIF were similar for both baseline and acetazolamide scans, about 0.45 and 0.60 ml·min−1·g−1, respectively. The mean CBFMLIF/CBFAIF ratio was 1.04 and 1.03 for baseline and acetazolamide, respectively. No significant differences were found between the average CBFAIF and CBFMLIF for either scan.
Individual data points for MLIF and AIF from
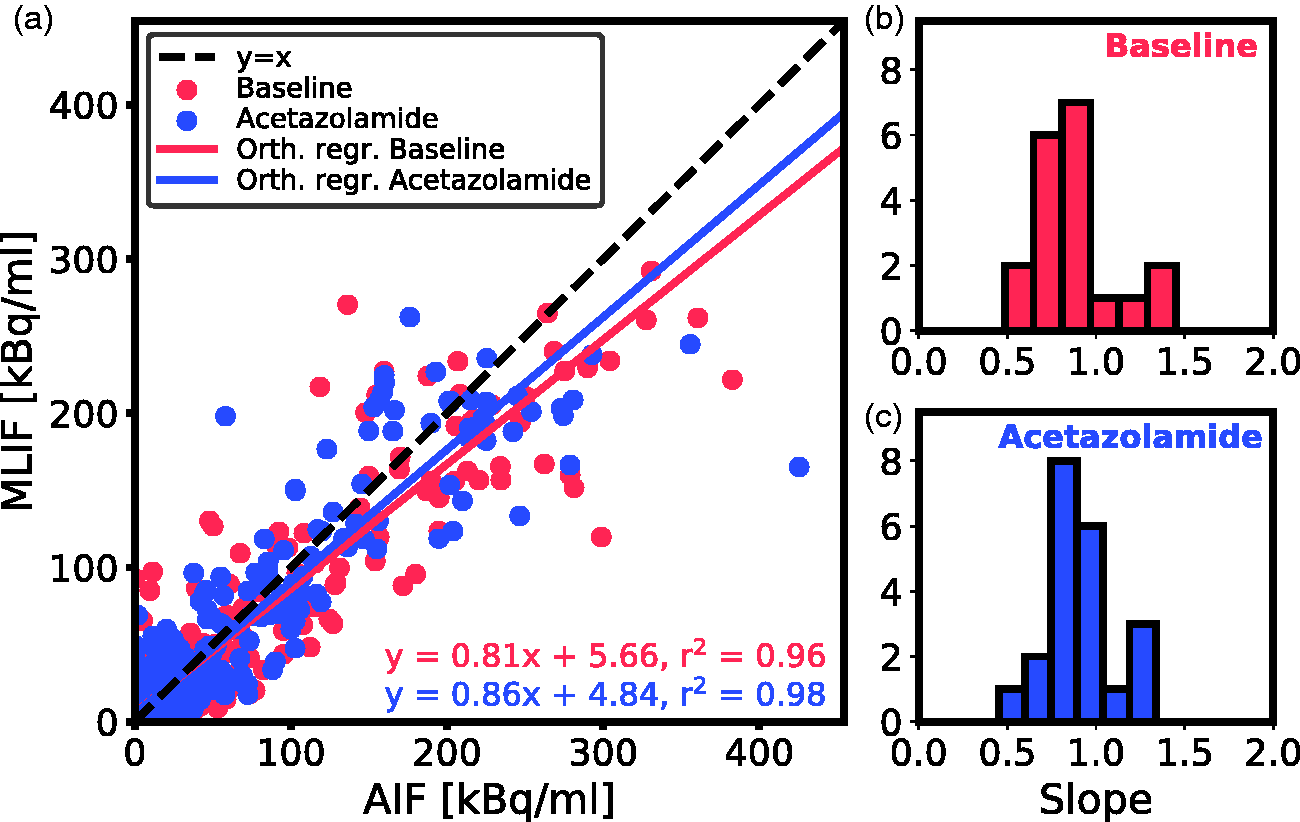
(a) Scatter plot of MLIF and AIF data points for all subjects for baseline (blue) and acetazolamide (red) scans based on
Displaying the CBF data from
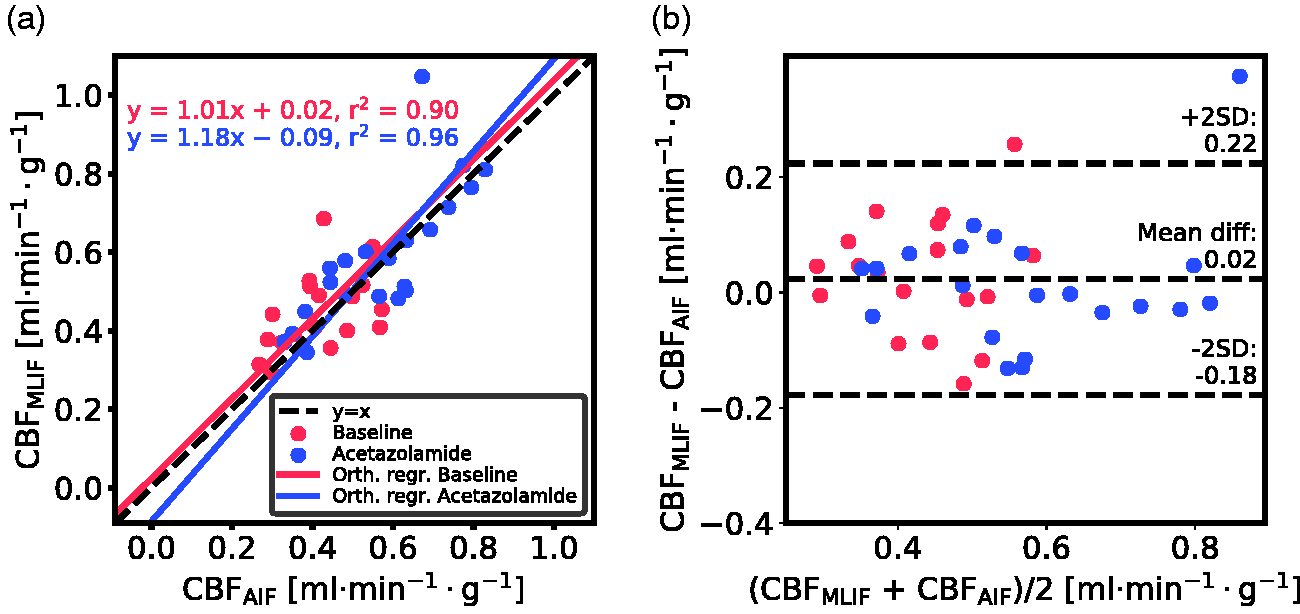
Evaluation of the GP-predicted MLIF for baseline (blue) and acetazolamide (red) scans based on
The fractional arterial blood volume, VA, from equation (8), was found to be near zero for all scans (0.001 ± 0.003).
As a visual illustration of the effect of prediction errors on estimation of CBF, the AIF and GP-predicted MLIF from the baseline scan for four representative subjects are shown in Figure 4. Figure 4(a) and (b) comprises two examples with less than 3% difference between CBFAIF and CBFMLIF, while Figure 4(c) and (d) display cases with substantial differences between both methods. Based on the CBF ratios (CBFMLIF/CBFAIF), it can be observed that an overprediction of the AIF peak (Figure 4c) results in an underestimation of CBF, while an underpredicted AIF peak (Figure 4d) ends in an overestimated CBF.
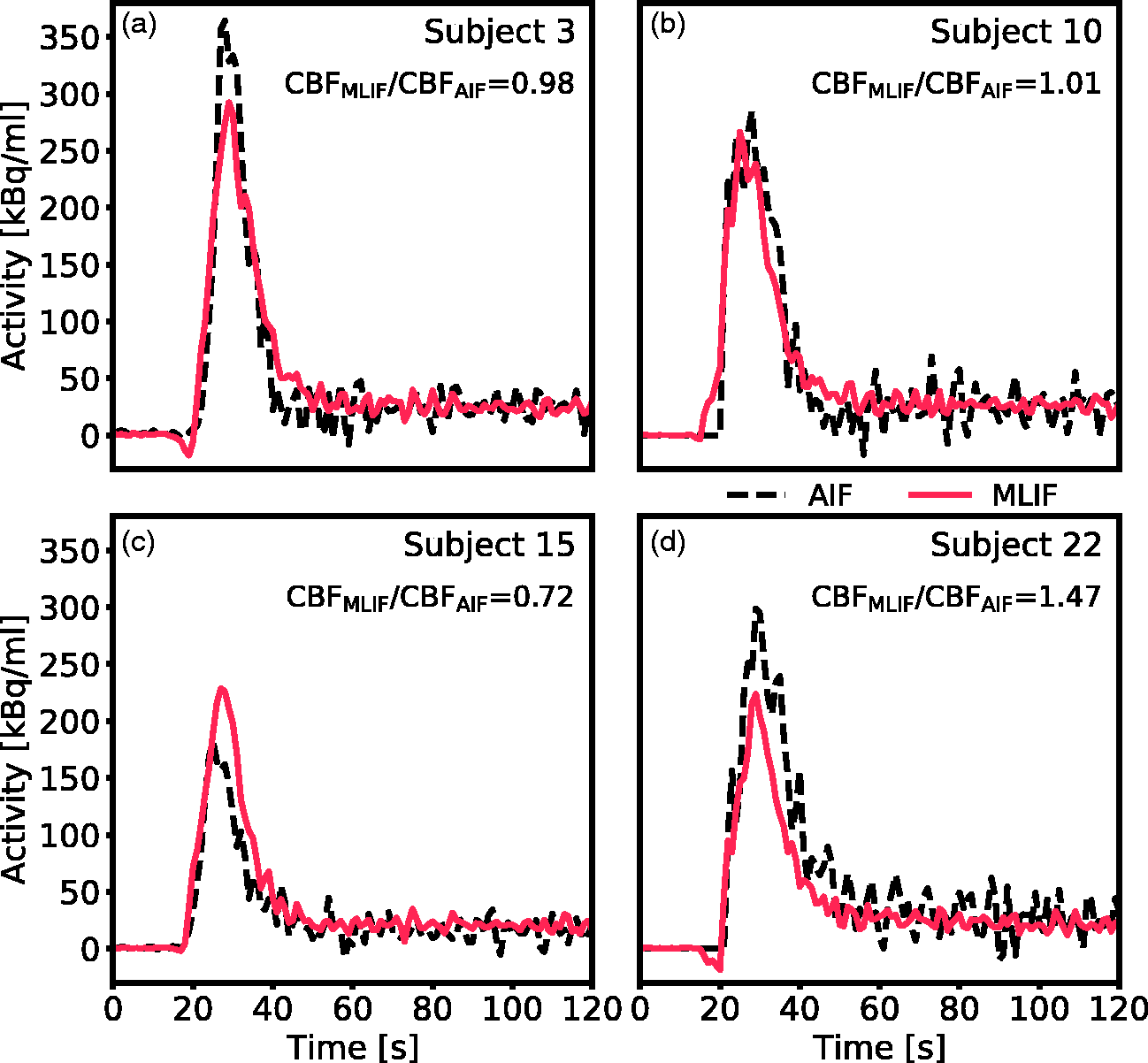
Comparison of AIF (black dashed line) and GP-predicted MLIF (red line) for the baseline scan of four representative subjects. (a and b) Two scans with less than 3% difference between CBFAIF and CBFMLIF. (c) Representative example of a scan where the MLIF overpredicts the AIF peak, resulting in an underestimation of the calculated CBF. (d) Representative example of a scan where the MLIF underpredicts the AIF peak, resulting in an overestimation of the calculated CBF.
Figure 5 displays box plots of whole-brain grey matter CBF for baseline and acetazolamide scans, when using different prediction models for MLIF (see Table 1).
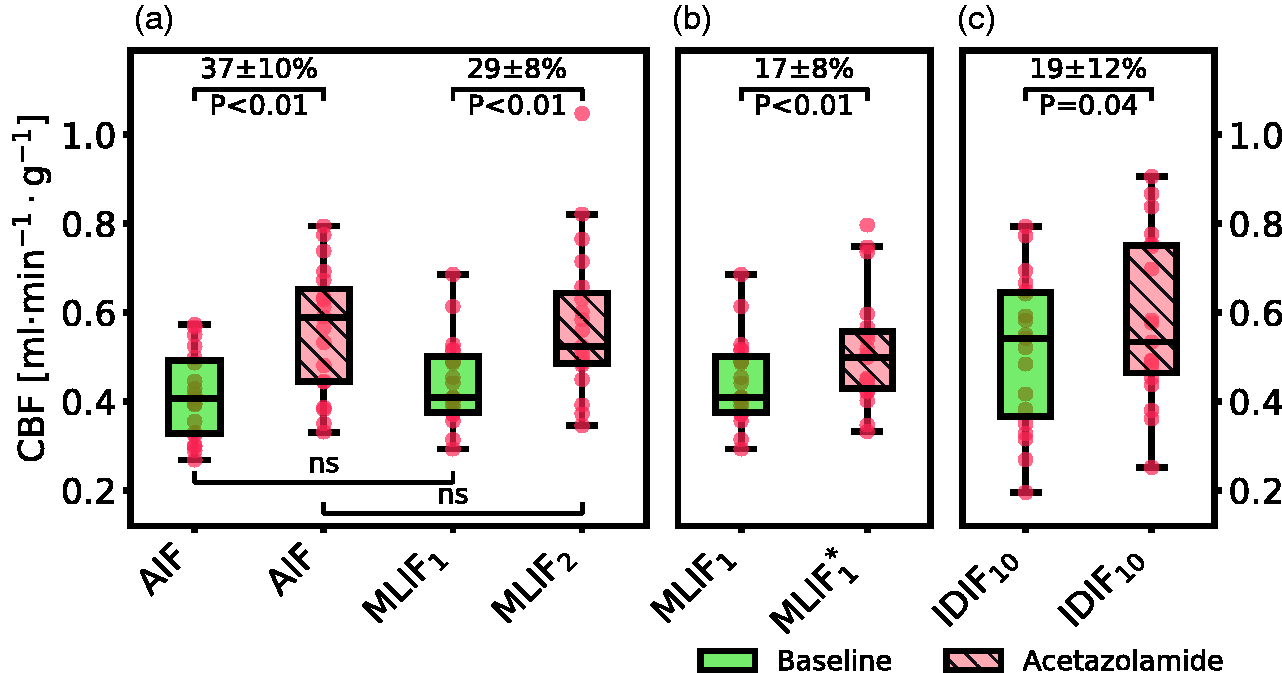
Box plot of estimated whole-brain grey matter CBF for baseline and acetazolamide scans, when using different input functions for
The GP variance was evaluated as prediction error measure in
In
In
In a final case, the relative CBF increase from baseline to acetazolamide scans was investigated by using the IDIF10 as input function for each scan. A significant CBF increase of 19% (P = 0.04) was found after acetazolamide medication (Figure 5c), which was comparable to the change observed in
As CBF is based on underlying time-activity curves, we proceeded to investigate the differences in CBF after acetazolamide medication in the first two cases by inspecting the mean time-activity curves across subjects in Figure 6. We observed that the local brain input function (IDIF10) (Figure 6e) showed a shape-dependence on acetazolamide medication which was not reflected in the AIF measured in the wrist (Figure 6b). In the AIF, the baseline and acetazolamide curves were similar, while for IDIF10, the acetazolamide curve had a larger area-under-curve compared to baseline. Also, for IDIF10, there was a slight shift in the mean time–activity curves between baseline and acetazolamide, which was not visible for the AIF. Thus, the AIF measured in the wrist does not reflect physiological changes due to acetazolamide, which are apparent in the local brain input (IDIF10).
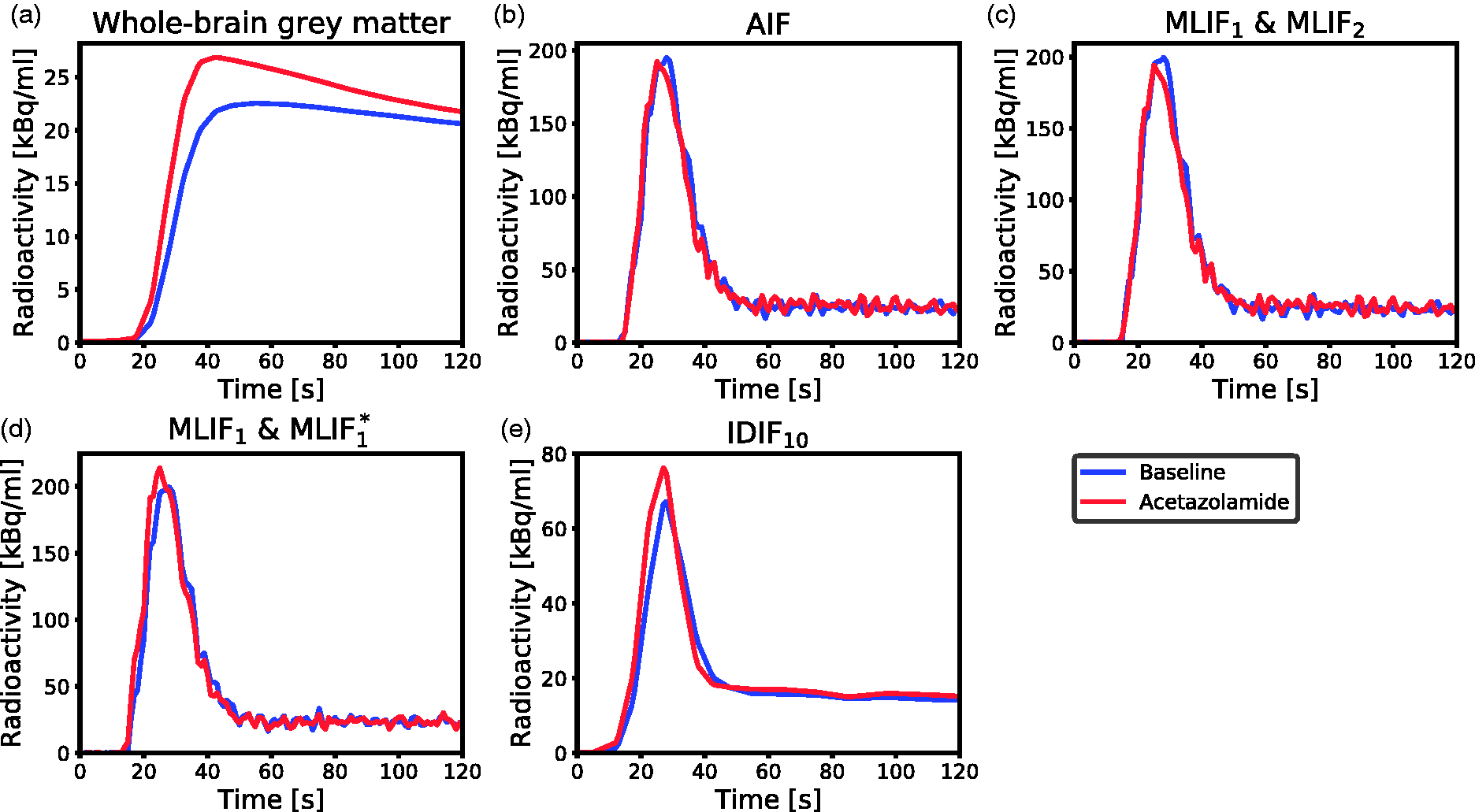
Mean time-activity curves across subjects for baseline (blue) and acetazolamide (red) scans during the first 2 min of PET scanning. (a) Measured radioactivity uptake in whole-brain grey matter. (b) AIF. (c) MLIF1 model for baseline, and MLIF2 for acetazolamide scans. (d) MLIF1 model for baseline, and MLIF*1 for acetazolamide scans. (e) IDIF10 input function based on the 10 highest voxels in the carotid arteries.
Discussion
Tracer kinetic modelling of dynamic PET data requires accurate knowledge of an AIF, conventionally measured through arterial blood sampling. Our aim was to investigate whether an AIF could be predicted as accurately by an MLIF model using solely image-derived input data from the carotid arteries.
AIF and predicted MLIF curves were found to be similar, with no significant difference between whole-brain grey matter CBFAIF and CBFMLIF estimates. Furthermore, the correlation between the CBF estimates was r2>0.9 and the mean differences were close to zero. The MLIF model was also able to accurately predict an increased CBF after acetazolamide medication, when trained on post-acetazolamide data. Altogether the results indicate that the MLIF method has potential as an alternative AIF for generation of CBF values using 15O-water PET and kinetic modelling, which in clinical practice implies evading of arterial cannulation.
Initially, we evaluated an MLIF model trained on pooled baseline and acetazolamide data. However, this resulted in inferior generalization to new samples compared to when training was done separately for baseline and acetazolamide scans (data not shown). The reason for this difference between the approaches may be because the input IDIFs under the two conditions vary in amplitude (Figure 6e), while the AIF (Figure 6b) does not. Therefore, in a first case, input functions of baseline scans were predicted using an MLIF model trained on baseline data (MLIF1), and, similarly, input functions of acetazolamide scans were predicted using an MLIF model trained on acetazolamide data (MLIF2). There was a strong linear relationship between the data points from AIF and predicted MLIF curves, although the peak values were systematically underestimated (slope <1 in Figure 2). In the time-activity curves, the number of high data values, acquired during the first pass peak, is low compared to the number of low values, acquired during the rest of the scan. Also, the standard deviation around the peaks was observed to be larger than at the tails (data not shown). We speculate that this imbalance affects the GP models and results in the systematically underestimated peak values. Previous work has shown that when data is limited, the GP model may underestimate the mean function. 47
For both baseline and acetazolamide, the mean time-activity curves for AIF (Figure 6b), and MLIF1, MLIF2 (Figure 6c) appeared similar in shape. Following kinetic modelling, the average CBF values obtained using an AIF are in line with previously published work before48,49 and after administration of acetazolamide.
48
No significant differences were found between AIF-based and MLIF-based CBF estimates in whole-brain grey matter for neither baseline nor acetazolamide scans (Figure 5a). Also, a slope close to unity, an r2>0.9 and low bias between CBFAIF and CBFMLIF estimates pointed towards an acceptable agreement of both methods (Figure 3a). Across both baseline and acetazolamide scans, differences between CBFAIF and CBFMLIF estimates were relatively small (Figure 3b), although several subjects had relative CBF errors of > 20% for the baseline scans while there was a somewhat lower spread for the acetazolamide scans. Our hypothesis is that a GP model trained on acetazolamide data (MLIF2) generalize better to new samples, compared to a model trained on baseline data (MLIF1). We speculate that the reason for this is that a larger range of CBF values is a result from a larger range of input values (IDIF10-1000), for acetazolamide, compared to baseline scans. This may have resulted in MLIF2 being valid for a wider range of unseen samples, compared to MLIF1, as mentioned and illustrated in Supplementary material. Nevertheless, the CBF increase between baseline and acetazolamide scans was highly correlated to that of the AIF in
In a second case, we trained an MLIF model on all baseline scans and then applied that model on the acetazolamide scans (MLIF*1). The idea was to resemble a situation where the baseline scans reflected a database comprising healthy subject with a normal CBF whereas the acetazolamide scans reflected clinical data from patients with an altered CBF. Even in this scenario significant differences were found between the whole-brain grey matter CBF from the baseline and acetazolamide scan. However, the CBF increase between baseline and acetazolamide scans in this case displayed small between-subject variation (data not shown) and, maybe because of that, low correlation to the AIF CBF changes. Furthermore, unexpectedly, the difference in CBF was only 17% (Figure 5b) compared to 29% in the previous case (Figure 5a).
We found this difference between baseline and acetazolamide response striking. The local brain input function (IDIF10) (Figure 6e) showed a shape-dependence on acetazolamide which was not reflected in the AIF measured in the wrist (Figure 6b). Consequently, CBF calculated with a local brain input would by necessity be different from AIF-based CBF. This effect remained also for the MLIF1 and MLIF1* models (Figure 6d) and could possibly explain the observed differences in CBF change.
In order to quantify the differences in CBF after acetazolamide medication in the first two cases, we attempted using a carotid artery region (IDIF10) as input function for both baseline and acetazolamide scans (
We suggest that the above discussed difference in input function curve shape might be explained in part by effects on the vascular system after acetazolamide medication. Acetazolamide dilates the vascular system and increases the cerebral blood flow velocity,30–32 which explains the increased mean time–activity curve in whole-brain grey matter after acetazolamide medication (Figure 6a). A lower back-pressure due to dilated vasculature together with increased blood velocity30,31 could possibly also explain the observed effect on MLIF*1 (Figure 6d) and on the IDIF (Figure 6e). Figure 6e also indicated that the 15O-water tracer arrived earlier to the brain after acetazolamide medication compared to the baseline scan, resulting in a slight shift of the mean time–activity curves. This observation might also support that the differences in the IDIF10 peaks between scans were partly due to an enhanced cerebral blood-flow velocity in the acetazolamide scan. An additional contributing effect to the increased amplitude of the IDIF after acetazolamide may be an increased spill-over from tissue due to increased brain uptake.
In summary, when training and testing on the same scans, similar CBF estimates for whole-brain grey matter are obtained when using AIF and MLIF (
A prerequisite for the MLIF approach is that representative training data have been collected for the specific tracer and imaging system, including both images and blood AIFs. Once an MLIF model has been trained, it offers several advantages, compared to various other image-derived and population-based methods.11–13,20–25 A trained MLIF model is a non-invasive method describing both the shape and the amplitude of an AIF, without any need for calibration blood samples. The MLIF models represents a learned transformation, that directly maps the image-derived input data, to a ready-to-use AIF, by inherently correcting for partial volume effect, with no predefined assumption of the model function. Furthermore, the input data required by the MLIF approach consists of only three carotid artery regions which can be objectively and automatically segmented in the PET images. This makes the MLIF approach simple and convenient to use, without the need for MRI-based artery segmentation.14–19
One limitation of this study is that the GP models were trained on at most 22 samples (MLIF*1), which might have resulted in an inferior generalization of the model to new samples which were dissimilar from the training data. To investigate the robustness of the MLIF model to unseen test data, new IDIF curves were created by scaling the existing input IDIF time–activity curves for each subject during the leave-one-out testing (Supplementary material). When the input data was scaled, the MLIF model was stable for input ranges encountered in the training data. For higher and lower scale factors, the model performance was gradually degraded (Supplementary Figure S2A). The drop of performance of machine learning models outside the range of training data, so called
In a clinical setting where an existing MLIF model is applied to a patient, it is important that the resulting CBF is reliable. We have reported four outliers based on abnormal regression slopes between the AIF and MLIF data points. However, we cannot know if it was the AIF or the MLIF curves that were abnormal. The corresponding CBF values for these outlier scans all indicate that the AIF was abnormally high as indicated by CBFMLIF/CBFAIF ≪ 1 (Table S1). This suggests that the outlier was caused by the AIF and not by the MLIF model. Future research should investigate methods for quality control of predicted MLIF curves and CBF values from a trained model.
Another limitation of the study is the combination of healthy and MS patients in the data set, as well as data from two different scanners. This increased the heterogeneity in the input data to the MLIF models, and in combination with the effects of acetazolamide, this might have affected the results. However, the limited number of subjects does not allow to study these effects further in detail. Further, the evaluation of our MLIF model was mainly based on differences in whole-brain gray matter CBF. We expect similar results to be obtained for CBF in smaller brain regions, but this aspect should be investigated in future studies. Also, the test-retest variability of the MLIF method should be investigated and compared to that of the blood AIF, 53 as well as the evaluation of the model sensitivity in relation to aspects such as injected dose, time-framing, reconstruction settings and different type of scanners. Finally, the generalizability of the MLIF method to other diseases should be investigated. For example, it is known that carotid stenosis alters the temporal shape of the AIF, 54 which might have implications for MLIF models trained on baseline scans and applied to patients with pathological arterial vasculature.
In this study, different MLIF models were evaluated with 15O-labeled water. In our previous research 29 a machine learning approach was also feasible for AIF prediction using FDG, although not yet evaluated in clinical data. Thus, we suggest that the method can be adopted to other tracers by merely training similar MLIF models. With proper validation, it may also be conceivable that tracers requiring metabolite correction of the AIF can be included in the prediction model. As for all data-driven models, the accuracy of the MLIF approach for a particular PET application will depend on the quality and quantity of the available training data. Nevertheless, MLIF opens for simplified and non-invasive input function measurements, and thereby potentially eliminating the need for extensive arterial blood sampling in future PET studies.
In conclusion, we demonstrated that our non-invasive MLIF prediction method may be a viable alternative for CBF measurements using 15O-water PET and kinetic modelling, which in clinical practice implies evading of arterial cannulation. The MLIF method successfully differentiated CBF values before and after acetazolamide medication.
Supplemental Material
sj-pdf-1-jcb-10.1177_0271678X21991393 - Supplemental material for Cerebral blood flow measurements with 15O-water PET using a non-invasive machine-learning-derived arterial input function
Supplemental material, sj-pdf-1-jcb-10.1177_0271678X21991393 for Cerebral blood flow measurements with 15O-water PET using a non-invasive machine-learning-derived arterial input function by Samuel Kuttner, Kristoffer Knutsen Wickstrøm, Mark Lubberink, Andreas Tolf, Joachim Burman, Rune Sundset, Robert Jenssen, Lieuwe Appel and Jan Axelsson in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Northern Norway Regional Health Authority [grant number HNF1349-17]. The parent study was supported by BiogenIdec Sweden AB and The Swedish Foundation for MS Research.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ contributions
All authors contributed substantially to the scientific process leading to this manuscript. JA, KKW, ML, RJ, RS and SK contributed in conception and design. AT, JB, LA and ML were responsible for acquisition of clinical PET data (parent study). KKW, RJ and SK implemented the machine learning models. JA and SK analysed the data. JA, LA, ML and SK interpreted the results and drafted the manuscript. All authors reviewed and approved the final manuscript.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
