Abstract
Although still a matter of controversy, disrupted in schizophrenia protein 1 (DISC1) was suggested as a potential inhibitor of phosphodiesterase 4 (PDE4). We used Disc1 locus impairment (LI) mice to investigate the interaction between PDE4 and DISC 1 in vivo and in vitro. [11C](R)-Rolipram binding was measured by PET in LI (n = 11) and C57BL/6 wild-type (WT, n = 9) mice. [11C](R)-Rolipram total distribution volumes (VT) were calculated and corrected for plasma-free fraction (fP) measured in a separate group of LI (n = 6) and WT (n = 7) mice. PDE4 enzyme activity was measured using in vitro samples of cerebral cortices from groups of LI (n = 4), heterozygote (n = 4), and WT (n = 4) mice. Disc1 LI mice showed a 41% increase in VT (18 ± 6 vs. 13±4 mL/cm3, P = 0.04) compared to WT mice. VT/fP showed a 73% significant increase (90 ± 31 vs. 52 ± 15 mL/cm3, P = 0.004) in Disc1 LI compared to WT mice. PDE4 enzymatic activity assay confirmed in vivo findings showing significant group differences (p < 0.0001). In conclusion, PDE4 activity was increased in the absence of critical DISC1 protein isoforms both in vivo and in vitro. Additionally, [11C](R)-Rolipram PET was sensitive enough to assess altered PDE4 activity caused by PDE4–DISC1 interaction.
Keywords
Introduction
Phosphodiesterase 4 (PDE4) is part of a diverse family of PDEs responsible for the metabolic inactivation of the second messenger 3′,5′-cyclic adenosine monophosphate (cAMP). PDE4 is widely expressed throughout the brain, and its regulation is a potentially important therapy for several neuropsychiatric disorders including addiction, major depressive disorder (MDD), and other disorders that accompany mood and cognitive symptoms.1–3 PDE4 activity is regulated through phosphorylation by protein kinase A (PKA) and extracellular signal-regulated kinase (ERK).4,5
Changes in PDE4 activity can be monitored in vivo using the non-subtype selective positron emission tomography (PET) radioligand, [11C](R)-Rolipram. A previous study from our laboratory used [11C](R)-Rolipram to measure PDE4 binding in patients with MDD before and after treatment with a selective serotonin reuptake inhibitor (SSRI),1,6 and in another study, we established that PET imaging with [11C](R)-Rolipram was sensitive to the interaction between PDE4 and PKA. 7 In the latter study, PKA-mediated phosphorylation of PDE4 increased the in vivo binding of [11C](R)-Rolipram in rat brain, consistent with the predicted almost 10-fold increase in affinity of this radioligand to the phosphorylated compared to the non-phosphorylated form. 8 To our knowledge, those studies were the first to show that PET imaging is sensitive to the phosphorylation status of a protein. It should be noted that PET is currently the only tool capable of assessing the in vivo activity of PDE4, given that rapid postmortem de-phosphorylation of PDE4 after death does not allow the enzyme phosphorylation state to be reliably evaluated by in vitro binding assay. These findings suggest that PET may be a useful tool for assessing the in vivo activity of PDE4 in conjunction with its regulatory factors.
Disrupted in schizophrenia 1 (DISC1) is a scaffold protein that mediates many cellular processes, including the regulation of PDE4.9,10 Perturbation of DISC1 is known to lead to a wide range of mental manifestations, including mood symptoms.9,10 For example, sequestration of DISC1 into insoluble Huntingtin aggregates led to aberrant emotional/motivational behavior in a model for Huntington’s disease. 11 The DISC1 protein was found to interact with the regulatory domain of PDE4 and to be released upon phosphorylation of the enzyme by PKA. 12 Because this suggests a model where DISC1 binding sequesters PDE4 in its inactive state, it is thus conceivable that the absence of DISC1 would increase PDE4 activity.
The goal of this study was to determine whether PET imaging is sensitive to—and thus capable of quantifying—the predicted effects of the interaction between three proteins: PDE4, PKA, and DISC1. In order to evaluate changes in PDE4 activity in the presence or absence of DISC1 protein, we used [11C](R)-Rolipram PET to study the recently developed Disc1 locus impairment mouse model (Disc1 LI), in which a large portion of the Disc1 gene, including the section encoding for the PDE4 binding domain, is deleted.13,14 In vitro enzymatic assays were used to confirm changes in PDE4 activity observed via PET imaging.
Materials and methods
Animals
Animal parameters in the different experimental groups.
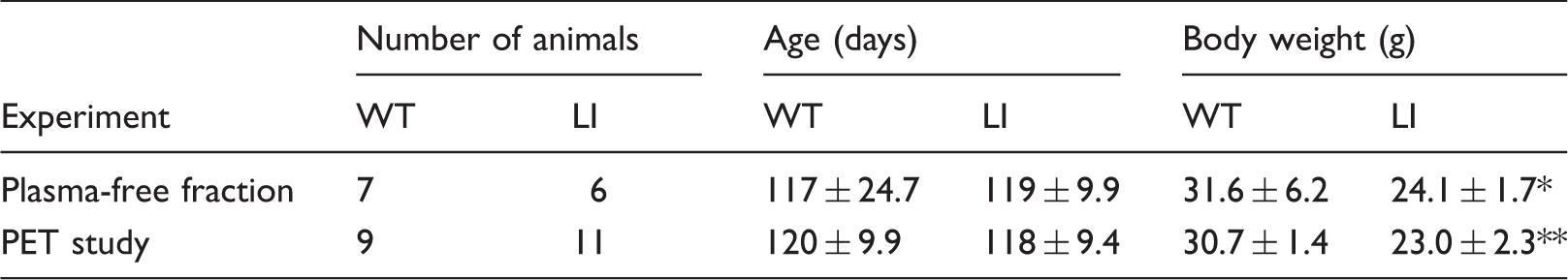

p < 0.05, **p < 0.0001.
WT: wild-type; LI: Disc1 locus impairment mouse model; PET: positron emission tomography using [11C](R)-Rolipram
Animal parameters in the PDE4 enzymatic assay.
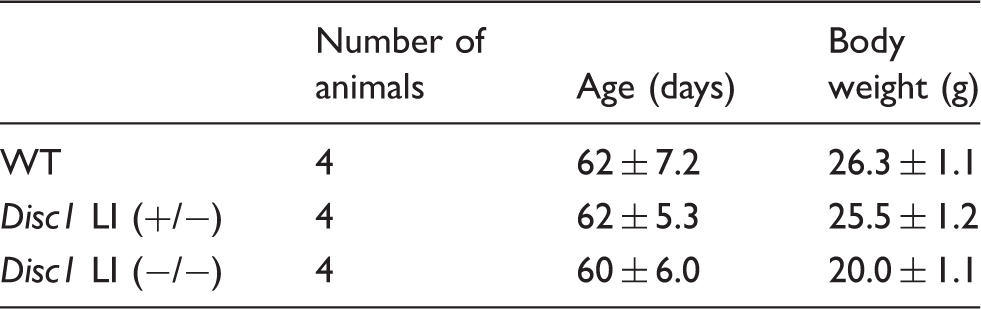

WT: wild-type; LI: Disc1 locus impairment mouse model.
Experimental design
Two separate groups of male Disc1 LI and wild-type (WT) C57BL/6 mice were used to: (1) compare [11C](R)-Rolipram binding (total distribution volume (VT)) via PET scan; and (2) measure plasma-free fraction (fP) without performing PET scans in order to normalize VT measured in goal one, above. A third group of male homozygous Disc1 LI mice (−/−), heterozygous Disc1 LI mice (+/−), and WT mice was used to compare PDE4 enzyme activity (n = 4 per group). Separate groups were used because it was not possible to perform all three of the measurements in the same animals. Mice were approximately 120 days old in all groups that underwent PET scans and fP measurement (Table 1) and approximately 60 days old in the group used for in vitro measurements of PDE4 activity (Table 2). The animals were housed in a temperature controlled (22–24℃) environment under a 12 h/12-h day/night cycle with free access to food and water. All animal experiments were conducted in accordance with the NIH Guide for the Care and Use of Laboratory Animals as well as the ARRIVE guidelines. 19 Animal protocols were approved of the institutional Animal Care and Use Committee of the site where the experiments were performed (either the National Institute of Mental Health (NIMH) in Bethesda, Maryland, USA or Kyoto University, Kyoto, Japan).
Radiochemistry
[11C](R)-Rolipram was synthesized by alkylation of the (R)-desmethyl-rolipram precursor with [11C]methyliodide as previously described. 20 The tracer was produced with a radiochemical purity of > 95%. Molar activity at the time of injection was 4618 ± 1580 GBq/µmol, which corresponds to a total injected mass dose of 78 ± 22 pmol/kg (WT animals) and 109 ± 49 pmol/kg (Disc1 LI mice).
PET scans
To accurately measure VT without multiple arterial samples, the VT of [11C](R)-Rolipram was measured under equilibrium by a bolus-infusion protocol while obtaining a large arterial sample (0.5 mL) at the end of the scan. Throughout the experiment, mice were anesthetized using 1.5–2% isoflurane in oxygen (1 L/min). Because alterations in body temperature are known to influence radioligand kinetics, 21 the body temperature of the animals was monitored and kept constant (37℃) throughout the procedure. At the start of the PET scan, male Disc1 LI (n = 11) and WT (n = 9) mice received a bolus injection of [11C](R)-Rolipram (3.76 ± 0.75 MBq) immediately followed by a constant infusion of the radiotracer with an infusion rate of 27.7 ± 5.8 kBq/min (36 µL/h, Kbol = 135 min) using a syringe pump (Harvard PhD 2000, Harvard Apparatus, Holliston, MA, USA). Animals were scanned dynamically for 100 min. PET data were acquired in list mode, rebinned in 23 time frames (6 × 20 s, 5 × 1 min, 4 × 2 min, 3 × 5 min, 3 × 10 min, 2 × 20 min) and reconstructed using ordered subset expectation maximization (OSEM) resulting in a final pixel width of 0.866 and a slice thickness of 0.796 mm. Full-width-half-maximum of the reconstructed images was reported as slighter smaller than 2.0 mm. 22 Attenuation correction was not performed because the small head size of mice minimizes attenuation. At the end of the scan, and while constant infusion of [11C](R)-Rolipram was continued, cardiac blood was collected through cardiac puncture immediately after opening the thorax. This sample was then analyzed to quantify the concentration of intact [11C](R)-Rolipram in plasma. All Disc1 LI and WT mice except for two LI mice were scanned in sibling pairs containing one Disc1 LI and one WT mouse per scan, allowing us to control for subtle experimental conditions such as dose or length of anesthesia and obtain reliable VT values. Scans were performed using a FOCUS 120 small animal PET scanner (Siemens/Concorde Microsystems, Knoxville, TN).
Plasma analysis
Plasma [11C](R)-Rolipram levels were quantified as previously described. 20 In brief, plasma radioactivity was counted using a calibrated automatic well-type gamma-counter (model 1480 Wizard; Perkin-Elmer, Waltham, MA, USA) with an energy window set between 360 and 1800 keV (counting efficiency, 51.9%). In addition, plasma was analyzed by reversed-phase high-performance liquid chromatography on a Novapak C18 column (Waters Corp., Milford, MA, USA) within a radial compression module RCM-100, eluted with MeOH: H2O: Et3N (65: 35: 0.1 by vol) to correct for radiometabolites.
fP was measured in WT and Disc1 LI mice in a separate experiment without PET. Age-matched (three months), male Disc1 LI (n = 6) and WT mice (n = 7) were anaesthetized using isoflurane (2% in oxygen, flow: 1 mL/min). Blood was drawn from the heart of the animals through cardiac puncture immediately after opening the thorax. [11C](R)-Rolipram was then added to the plasma (550 kBq/mL of plasma) and incubated for 10 min. After incubation, plasma protein binding was measured by ultrafiltration using Centrifree membrane filters (Amicon Division, W.R. Grace and Co., Danvers, MA, USA) as previously described. 23
Quantification of [11C](R)-Rolipram binding
PMOD (version 3.4, PMOD Technologies, Zürich, Switzerland) was used to generate whole brain and heart time activity curves (TACs). To avoid selection bias, volumes of interest (VOIs) were automatically drawn based on image intensity. Because [11C](R)-Rolipram has a rather homogeneous distribution in brain, TACs were generated for whole brain rather than for individual brain regions. Heart activity was measured in PET images to estimate stability of [11C](R)-Rolipram in arterial plasma. [11C](R)-Rolipram binding in Disc1 LI and WT mice was compared by calculating VT, which was calculated as the ratio average brain radioactivity in PET images at equilibrium (60–90 min) divided by the concentration of [11C](R)-Rolipram in plasma. Due to large differences in the fP of WT and Disc1 LI mice, VT was further normalized to fP.
PDE4 enzymatic activity assay
The enzymatic activity of PDE4 in brain was evaluated by measuring fluorescence polarization (FP) via the IMAP™ FP Phosphodiesterase Evaluation Assay Kit (Molecular Devices, Sunnyvale, CA, USA) according to the manufacturer’s instructions. To ensure that anesthetics did not interfere with experimental outcomes, mice were humanely sacrificed using a rapid decapitation method without anesthesia. The heads of the mice were immediately immersed in liquid nitrogen for 6 s to cool down. The prefrontal cortical area of the brain (∼50 mg/mouse, four mice/genotype) was then rapidly dissected on an ice-cold surface within 30 s and frozen in liquid nitrogen before protein extraction. These freshly frozen tissues were sonicated (10 s, 10 times) in ∼800 µL of ice-cold KHEM homogenization buffer (50 mM potassium chloride, 50 mM HEPES, 10 mM EGTA, 1.92 mM magnesium chloride) supplemented with protease inhibitor cocktail (Roche, Basel, Switserland) and 1 mM dithiothreitol. These homogenates were then centrifuged (13,000 g, 10 min at 4℃) and the resulting supernatants were used for assays. Protein concentrations in the lysates were measured using a Pierce BCA protein assay (Thermo Fisher Scientific, Waltham, MA). The lysate (1 µL, 0.2 mg/mL) was mixed with 19 µL of the assay buffer containing the fluorescein-labeled cAMP in the presence or absence of rolipram (final 10 µM) and incubated for 45 min at room temperature; 60 µl of the binding buffer was then added and this mixture was incubated for 20 min. The FP values were then measured using the M5 plate reader equipped with the excitation (485 nm) and emission (525 nm) filters (Molecular Devices). The experiments were done in triplicate, and PDE4 activity was calculated by subtracting the FP values obtained in the presence of rolipram from the values without, normalized by protein concentration.
Statistical analysis
All data are reported as mean ± SD. Conventional statistics were carried out using Graphpad Prism 5. An unpaired t-test was performed to compare different parameters in WT and Disc1 LI mice after performing the D’Agostino and Pearson omnibus test to check for normality. A one-way Kruskal–Wallis test was used to check for the influence of genotype on PDE4 enzymatic activity. Differences with 95% probability level were considered significant.
Results
Evaluation of the bolus infusion protocol
Developing a bolus-infusion protocol allows the precise and simple calculation of VT values from plasma and brain concentrations, provided that equilibrium is reached. TACs extracted from whole brain in Disc1 LI and WT mice showed stable radioactivity levels starting from 50 min post injection (p.i.) of [11C](R)-Rolipram (Figure 1). Due to expression of PDE4 in the myocardium,
24
[11C](R)-rolipram is expected to bind specifically to myocardium,
25
suggesting that changes caused by the absence of DISC1 can also be expected in the heart. Indeed, [11C](R)-Rolipram uptake in the heart of Disc1 LI mice was higher compared to that in WT animals (Figure 1). TACs from the heart were thus examined as an indirect measure of the stability of plasma activity rather than as a measure of plasma activity itself. TAC stability was evaluated by calculating the percent change/hour starting from 50 min p.i. (Table 3). Both brain and heart showed minimal changes in [11C](R)-Rolipram concentrations after 50 min of scanning. In addition, TAC stability in heart and brain did not differ significantly between WT and Disc1 LI mice.
Time activity curves of whole brain (a) and heart (b) derived from wild-type (O) and Disc1 locus-impairment mice (▴). Mice were scanned after a bolus-infusion protocol using [11C](R)-Rolipram that resulted in stable steady state concentrations in both brain and heart. Time activity curves are presented as the normalized concentration, which is the radioactive concentration normalized for body weight and infusion rate. Data are presented as average of wild-type (n = 9) and Disc1 LI mice (n = 11) with error bars representing the standard deviation. WT: wild-type; LI: Disc1 locus impairment mouse model. Stability of the time activity curves after bolus injection with [11C](R)-Rolipram followed by constant infusion. WT: wild-type; LI: Disc1 locus impairment mouse model.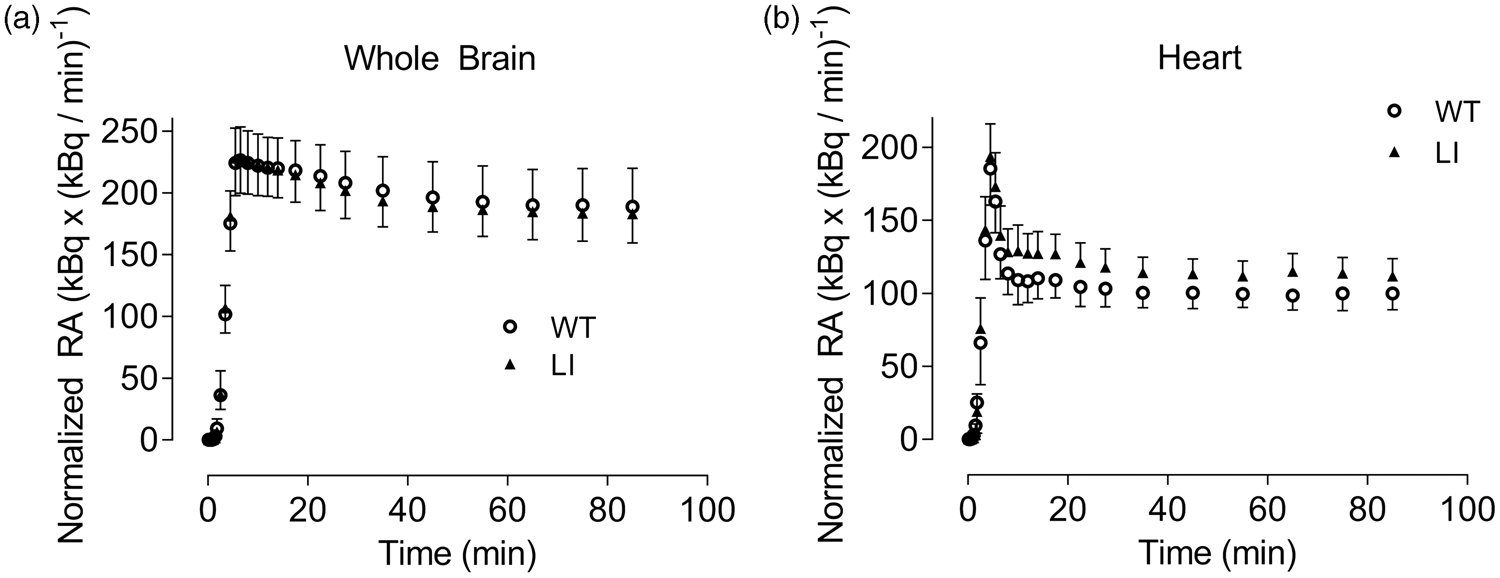
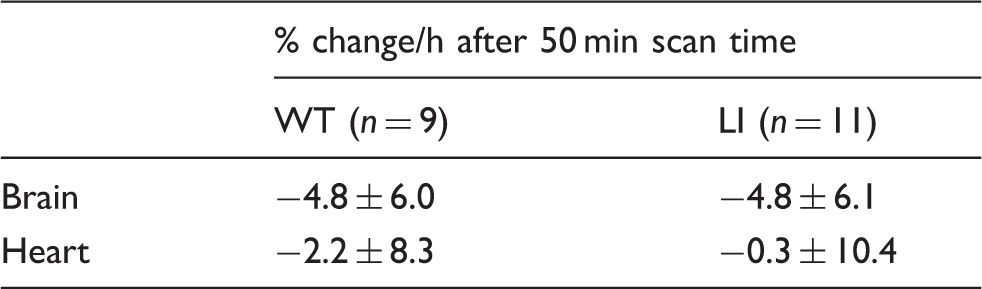
[11C](R)-Rolipram imaging in Disc1 LI and WT mice
VT values were calculated by dividing the equilibrium brain concentration with the radiometabolite corrected plasma concentration of [11C](R)-Rolipram (average fraction of intact [11C](R)-Rolipram in plasma = 12.3 ± 4.5% in WT and 9.9 ± 2.7% in LI mice, not significantly different). The calculated VT values in Disc1 LI mice were 41% higher (VT = 18.3 ± 6.4) than in WT mice (VT = 13.0 ± 3.6; p < 0.05; Figure 2a). SD/mean of VT was about 30%, indicating that the measurement was precise and did not display the variability caused by measurement errors.
Comparison of VT and VT/fP in wild type (O) and Disc1 locus-impairment mice (▴). Distribution volume (VT; a) was calculated as the ratio of average brain radioactivity in PET images (60–90 min) divided by the concentration of [11C](R)-Rolipram in plasma. Distribution volume corrected for plasma-free fraction (VT/fP; b) was calculated as the ratio of average brain radioactivity in PET images (60–90 min) divided by the concentration of free [11C](R)-Rolipram in plasma. Both VT and VT/fP were significantly higher in Disc1 LI (n = 11) compared to WT mice (n = 9). *p < 0.05, **p < 0.005; unpaired t-test.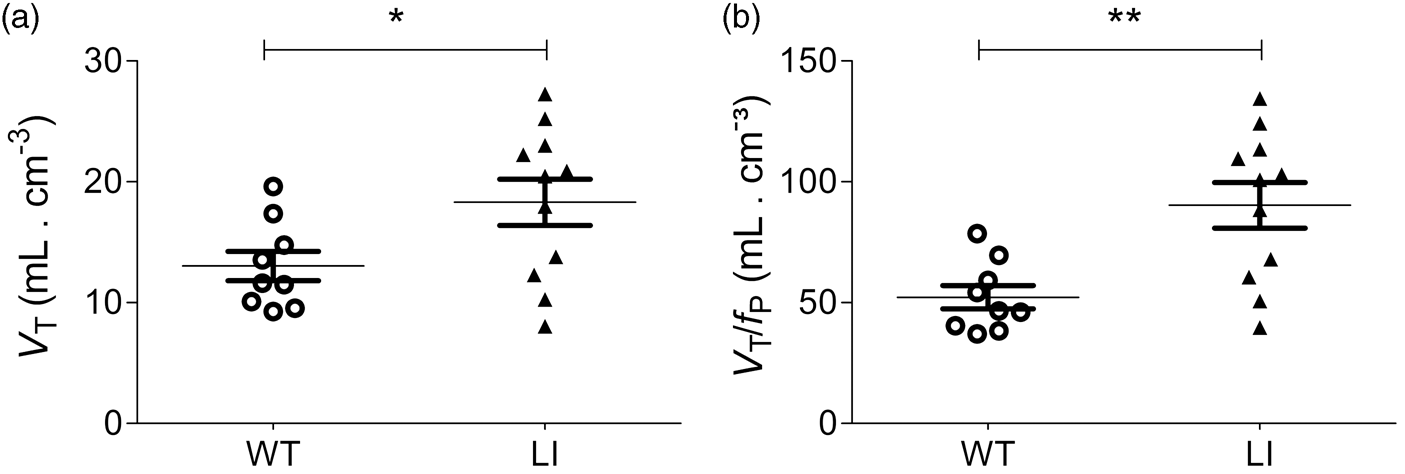
Another subset of animals was used to compare fP values in WT and Disc1 LI mice because we suspected that the significant difference in body weight (Table 1) might be associated with differences in fP; notably, only free [11C](R)-Rolipram enters the brain. fP levels did, in fact, differ significantly between WT (fP = 24.9 ± 1.9) and Disc1 LI mice (fP = 20.3 ± 2.6) (p < 0.005, Supplementary Figure 1). This suggests that VT/fP is more appropriate than VT for comparing [11C](R)-Rolipram binding between the groups. Disc1 LI mice showed a 73% significant increase (p < 0.005) in VT / fP (VT/fP = 90.3 ± 31.3) compared to WT mice (VT/fP = 52.2 ± 14.5.3; Figure 2(b)). Parametric VT/fP parametrical images showed a remarkably higher uptake in Disc1 LI mice compared to WT mice (Figure 3)
Parametric VT/fP images in wild type and Disc1 locus-impairment mice. These are representative coronal and transversal images showing VT/fP of [11C](R)-Rolipram in Disc1 locus-impairment mice and wild-type mice image intensity in each voxel is radioactivity normalized to free [11C](R)-Rolipram levels in plasma (i.e., VT/fp). The arrows indicate the location of the brain in the image. Note the higher uptake of [11C](R)-Rolipram in the brain of Disc1 locus-impairment.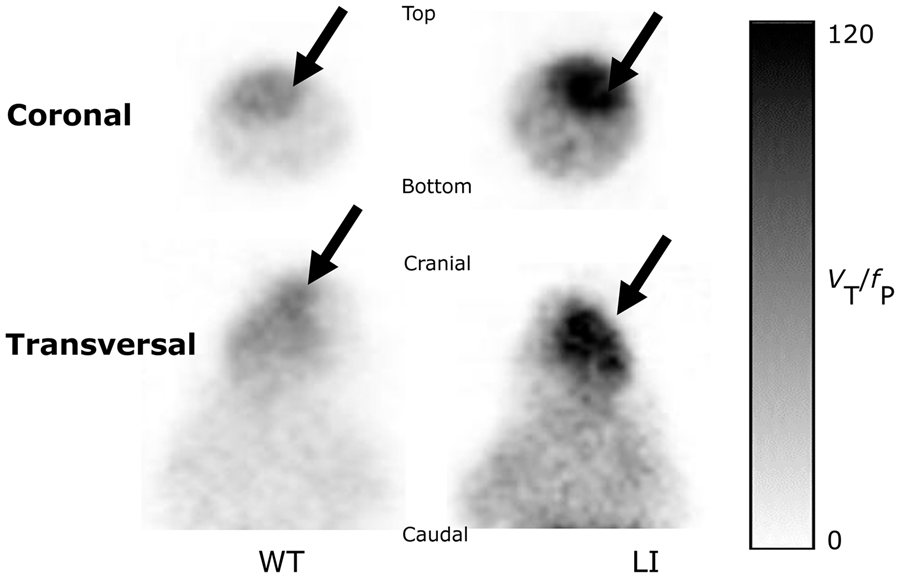
Because of the small body size of mice, the mass dose of [11C](R)-Rolipram could have affected VT/fP. To confirm that this did not occur, we examined the relationship between VT/fP and body weight or molar concentration of [11C](R)-Rolipram calculated from brain activity measured in PET images and specific radiosynthesis activity. If mass dose had had an effect, we would have expected lower body weight or higher concentrations of [11C](R)-Rolipram to cause lower VT/fP. No correlation between VT/fP and body weight or rolipram concentration was observed in Disc1 LI or WT mice (Supplementary Figure 2), indicating that mass dose did not affect VT/fP.
PDE4 enzymatic activity assay
As an alternative measure of PDE4 enzymatic activity in the brains of Disc1 LI mice, tissue homogenates from the prefrontal cortical area of the brains were tested for their ability to break down fluorescently labeled cAMP in the presence or absence of rolipram using the IMAP PDE assay system (Molecular Devices). The assay is based on the high affinity binding of the phosphate newly generated by PDE4 to the metal immobilized on nanoparticles. A significant effect of genotype was detected across the three groups (p = 0.0073) Homozygous and heterozygous mice displayed ∼50% and ∼25%, respectively, higher levels of PDE4 activity than WT mice (Figure 4). These data were in good agreement with the data obtained from [11C](R)-Rolipram imaging.
Enzymatic activity assays in wild-type mice as well as homozygous and heterozygous Disc1 locus-impairment mice. The effect of genotype across the three groups was significant (p = 0.0073) based on Kruskal–Wallis test, n = 4 per genotype group. WT: wild-type; LI: Disc1 locus impairment mouse model.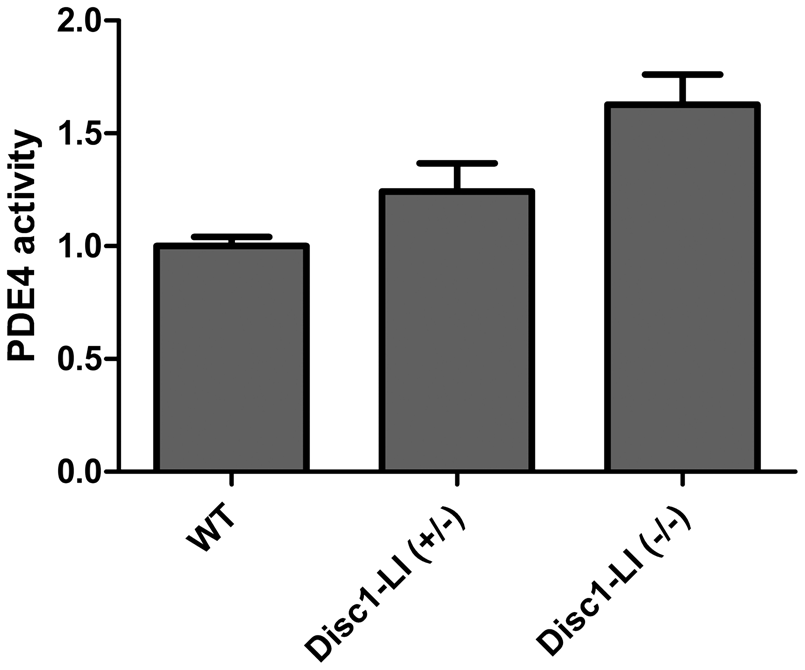
Discussion
This study demonstrated the feasibility of using [11C](R)-Rolipram PET imaging to measure protein-protein interactions between DISC1 and PDE4. We previously reported that PET imaging with [11C](R)-Rolipram was sensitive to phosphorylation of PDE4 by PKA. 7 Building on this work, the present study quantified the effects of the interaction between PDE4, PKA, and DISC1 in a Disc1 LI mouse model. Disc1 LI mice had significantly higher levels of [11C](R)-Rolipram binding than WT mice. The results of PET matched those of the enzyme assay, which detected greater enzyme activity in the LI mice. To our knowledge, this study is the first to demonstrate an interaction between PDE4 and DISC1 protein in vivo.
A major advantage to using PET rather than in vitro techniques is that PET allows us to estimate changes in PDE4 enzyme activity in vivo. Enzymatic activity of PDE4 is regulated through phosphorylation by PKA. 5 Therefore, rapid dephosphorylation of PDE4 after death is a limiting factor for ex vivo measurement of PDE4 enzymatic activity. The fact that the in vivo VT of [11C](R)-Rolipram correlates with PDE4 activity 7 allows PDE4 activity and phosphorylation status to be quantified unbiased by postmortem dephosphorylation. In the present study, in vitro enzymatic assays showed altered PDE4 activity in both homozygous and heterozygous Disc1 LI mice. However, this PET imaging studies only quantified differences between homozygous Disc1 LI and WT mice to focus limited resources on the condition where the effect was expected to be highest. Although the changes in VT observed in this study did not allow us to differentiate between changes in expression (Bmax) or activity (KD), the changes do reflect increased PDE4 enzymatic capacity in the Disc1 LI model.
The current finding that PDE4 activity is regulated by DISC1 is consistent with our recent observation that loss of function of DISC1 increases PDE4 activity. 11 It should be noted that a previous study found that a Disc1 mutant model had lower levels of PDE4 activity in the brain 26 ; however, the Disc1 LI mice used in the current study demonstrated the opposite effect. The discrepancy between these studies could be explained by the retention of a shorter isoform of the DISC1 protein present in the Disc1 mutant model, 26 whereas Disc1 LI mice lack the domain encompassing exon 1 through exon 3, which includes the PDE4 binding domain. Interestingly, this N-terminal portion of the DISC1 protein appears to be expressed transiently due to premature termination of the Disc1 coding region. 26 The fact that this domain includes the PDE4 binding domain 11 suggests that this DISC1 isoform dominantly inhibits PDE4 activity. Indeed, DISC1 has been reported to have multiple isoforms,15,16 and the activity of such isoforms remains largely unstudied. A relevant example from the literature is that of brain-derived neurotrophic factor (BDNF), whose transcription is regulated by nine individual promoters; disrupting the different transcripts results in mouse models expressing different molecular, cellular, and behavioral phenotypes. 27
The [11C](R)-Rolipram PET probe used in the current study could potentially be used for detailed evaluation of PDE4 activity in multiple Disc1 mutant mouse models. As noted earlier, [ 11 C](R)-Rolipram was previously used to successfully measure PDE4 binding in patients with MDD before and after treatment with an SSRI. Unmedicated patients experiencing a major depressive episode had globally reduced binding to PDE4, consistent with the low cAMP signaling thought to be associated with this disorder and consistent with decreased activation of PDE4 by PKA. 6 Two months of treatment with an SSRI increased PDE4 binding in MDD patients. 1 Interestingly, the involvement of DISC1 protein in regulating PDE4 enzymatic activity found in the present study further implicates altered PDE4 enzymatic activity in several other neuropsychiatric disorders in which DISC1 is dysregulated, including Huntington’s disease. [11C](R)-Rolipram PET could therefore be an additional tool to investigate this correlation.
In summary, the present study demonstrated a clear increase in PDE4 activity in the absence of most critical DISC1 protein isoforms both in vivo and ex vivo. These data warrant further investigation into the potential biological activity of DISC1 isoforms. Since we showed that [11C](R)-Rolipram is sensitive to the interaction between PDE4 and DISC1, [11C](R)-Rolipram PET imaging could help the complex role of DISC1 and PDE4 in diseases.
Supplemental Material
Supplemental Material for [11C](R)-Rolipram positron emission tomography detects DISC1 inhibition of phosphodiesterase type 4 in live Disc1 locus-impaired mice
Supplemental Material for [11C](R)-Rolipram positron emission tomography detects DISC1 inhibition of phosphodiesterase type 4 in live Disc1 locus-impaired mice in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Intramural Research Program of the National Institute Mental Health, NIH: projects ZIAMH002795 and ZIAMH002793, and by grants from the DOD/CDMRP (W81XWH-11-1-0269) (to T. Tomoda), the CTF-DDI (to T. Tomoda), the NIH (DA040127, MH092443, MH-094268, and MH105660) (to A. Sawa), the Brain and Behavior Research Foundation (to A. Sawa), the Stanley Foundation (to A. Sawa), and the S-R/RUSK Foundations (to A. Sawa).
Acknowledgements
The authors would like to thank Ioline Henter (NIMH) for providing invaluable editorial assistance.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Authors’ contributions
Maarten Ooms, Tetsuya Tsujikawa, Talakad G. Lohith, Sanché N. Mabins, Sami S. Zoghbi, Yasuyuki Kimura and Sanjay Telu substantially contributed to acquisition, analysis, and interpretation of the PET imaging data.
Akiko Sumitomo and Toshifumi Tomoda substantially contributed to acquisition, analysis, and interpretation of the PDE4 enzymatic assay data.
Hanna Jaaro-Peled, Victor W. Pike, Robert B. Innis, and Akira Sawa substantially contributed to the conception and design of the study.
Masahiro Fujita substantially contributed to the conception and design of the study as well as to the acquisition, analysis, and interpretation of data.
All authors contributed significantly to draft the article and revised it critically for important intellectual content.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
