Abstract
Advancements in digital pathology and artificial intelligence (AI) have enormous transformative potential for nonclinical toxicologic pathology and are already changing the ways in which pathologists work. However, due to the rapid evolution of digital pathology and AI, the toxicologic pathology community would benefit from an update on these advancements, which can be used to aid drug development. Here we identify key articles published on the use of digital pathology and AI in the field and provide current regulatory statuses and guidelines. For digital pathology, we outline the requirements for equipment, validation processes, workflows, and archiving. Challenges to achieve system interoperability and to establish harmonization through Digital Imaging and Communications in Medicine compatibility are also discussed. For AI, we highlight considerations for model development, including the determination of ground truth, problems that may arise due to bias, and how the accuracy and precision of AI algorithms can be assessed. Finally, we discuss the challenges and potential for AI-assisted toxicologic pathology, picturing a future where technology and scientific expertise work hand-in-hand to improve the quality and efficiency of nonclinical drug safety evaluation. This publication is a deliverable of the European Innovative Medicines Initiative 2 Joint Undertaking, “Bigpicture.”
Keywords
Introduction
The evolution from traditional light microscopy into the age of digital pathology is incrementally transforming the field of nonclinical toxicologic pathology. Despite the benefits of digital pathology, integrating this technology into the highly regulated world of drug development has presented the pharmaceutical industry with substantial challenges. The toxicologic pathology community and regulatory bodies have started to address these challenges, but a lack of specific regulatory guidance for the use of digital pathology in the Good Laboratory Practice (GLP) environment is still perceived as a gap in regulatory frameworks by many potential users, creating uncertainty for pathologists and scientists in recent years. Nevertheless, the continued advancement and adoption of digital pathology hold promise for enhancing the efficiency, accuracy, and collaborative potential of toxicologic assessments in drug development.
Several articles have examined the key aspects of digital pathology in the setting of nonclinical toxicologic pathology, from quality aspects and workflow integration to regulatory issues (Table 1). In addition, to address the need for guidance and structure, two workshops were commissioned by the European Society of Toxicologic Pathology (ESTP). In the seventh ESTP International Expert Workshop, an expert panel aimed to define a recommended set of minimal requirements for regulatory acceptance, with the scope of primary slide evaluation and peer review of whole slide images (WSIs). 57 Agreed-upon recommendations included the concept of WSIs as faithful replicas of original glass slides, measures to maintain image/data integrity, optimizing user training and acceptance, and the suggestion of fit-for-purpose workflow validation. 57
Key articles on the use of digital pathology in the regulated environment of nonclinical toxicologic pathology.
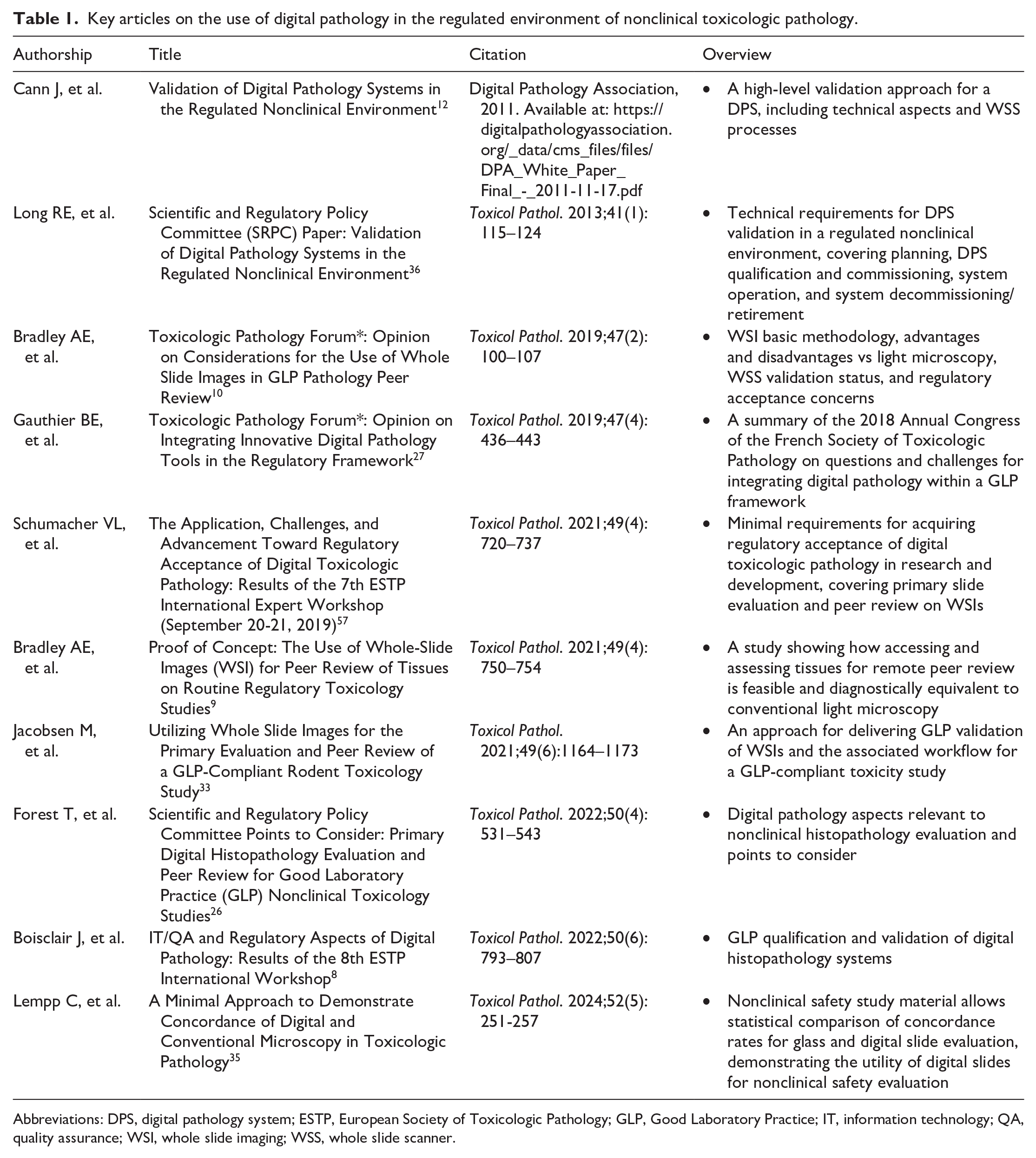
Abbreviations: DPS, digital pathology system; ESTP, European Society of Toxicologic Pathology; GLP, Good Laboratory Practice; IT, information technology; QA, quality assurance; WSI, whole slide imaging; WSS, whole slide scanner.
The eighth ESTP International Expert Workshop discussed in greater detail how to fulfill the regulatory requirements for the qualification and validation of digital histopathology evaluations in the GLP environment, which ensures that histopathology results submitted to regulatory authorities are of sufficient quality and rigor, and are verifiable.8,48,50 It was determined that the general validation, qualification, and implementation pathways for digital pathology should follow general principles applied to other GLP processes, and that close collaboration between pathologists, information technology (IT) experts, quality assurance (QA) experts, and regulatory officials are needed to ensure the success of digital pathology in GLP-compliant pre-clinical studies. 8
In computational pathology, while existing literature and regulatory guidelines predominantly address clinical applications, many of these principles are also applicable to toxicologic pathology. Based on the overall potential of artificial intelligence (AI), recognized for medicine as a whole and for pathology in particular, the implementation of AI solutions into digital toxicologic pathology needs to be considered, despite the challenges that will have to be overcome when incorporating AI into existing digital pathology workflows, especially under GLP regulations.
As technology is a rapidly evolving field and much has been published in recent years on the advancement of digital and computational pathology, one of the aims of the European Innovative Medicines Initiative (IMI) project, Bigpicture, is to provide an update on their current state in nonclinical toxicologic assessments for drug development. Ongoing efforts to adopt and integrate these novel approaches into existing workflows are discussed herein, with a focus on regulatory requirements and recommendations, as well as challenges and evolving standards and techniques. The future perspectives of these innovative technologies and their impact on the profession are also discussed.
Digital Pathology in Toxicologic Pathology
Digital pathology, defined as an image-based environment that enables the acquisition, management, and evaluation of pathology information generated from digitized glass slides (WSIs), 27 has become more and more widely established in nonclinical toxicology in recent years, 25 at least under non-GLP conditions. Digital pathology should be able to provide pathologists with an equivalent experience to that of traditional light microscopy; however, the advantages of digital pathology over light microscopy are plentiful.10,27,30,40 The benefits that digital pathology can offer primarily include improved efficiency, increased data security, and a reduced need for travel or shipping glass slides between contract research organizations (CROs) and sponsor facilities, while simultaneously enabling remote working and virtual real-time consultations. 57 Second, access to WSIs enables the adoption of analysis techniques, such as the use of image analysis, AI, and content-based image retrieval (CBIR) systems. 57 Third, a repository of WSIs can be a valuable resource to efficiently consult for background findings, unique study designs, tissues, findings, or species. 57 Therefore, efforts are being made to validate workflows for use under GLP conditions.33,35
In the clinical environment, several digital pathology systems (DPSs) have been cleared as medical devices through the United States Food and Drug Administration (FDA) 510(k) approvals (as for the Philips IntelliSite Pathology Solution and the Aperio AT2 DX System), whereas for toxicologic pathology, an institutional fit-for-purpose validation approach is more adequate. 26 “Fit for purpose,” meaning suitable for its intended use, was originally a concept for the validation of analytical methods and signifies that, in the context of digital histopathological evaluation, the processes in place as defined in standard operating procedures (SOPs) and protocols performed by trained personnel to use the qualified DPS, allow the pathologist to reliably perform their histopathological assessment. 8 Working groups recommend that GLP DPS validation packages for toxicologic pathology demonstrate substantial equivalency or non-inferiority to traditional light microscopy in terms of sensitivity and specificity (and, at the end, in the evaluation results obtained), and that the selected slides used should support the intended use.35,57 This can be achieved by testing with representative tissue materials from nonclinical toxicology systems during installation. 57
The approaches undertaken to deliver a GLP validation of WSIs and the associated workflow for the digital primary evaluation and peer review of a GLP-compliant toxicity studies were previously described.33,35 The authors demonstrated that WSIs can effectively replace traditional glass slides for histopathological assessments and provide a robust digital alternative for nonclinical toxicologic pathology.33,35 By sharing their approaches, the authors encouraged others to adopt WSI use in a GLP-compliant manner.33,35
Status of Regulatory Guidelines
In June 2020, the Organization for Economic Co-operation and Development (OECD) issued and published the regulatory position on the use of digital pathology in a GLP environment through a Frequently Asked Questions (FAQ) document, which includes information on the use of digital pathology in regulated nonclinical toxicology studies. 46 The document emphasizes the importance of establishing the WSI as a “faithful replica of the original histology slide,” ensuring equipment and software are fit for purpose, maintaining data integrity, and warranting that the study can be reconstructed, if required. 26 In May 2023, the U.S. FDA also released a guidance document specifically addressing the use of WSIs in nonclinical toxicology studies. 67 Their guidance, presented in a short Question and Answer document, provides some basic interpretations and covers technical aspects of WSI systems, the importance of proper documentation and data integrity, and the requirements for archiving WSIs under various work scenarios in GLP studies. 67 Together, these recommendations aim to ensure that nonclinical digital pathology can be implemented in a way that maintains the high standards of scientific quality, accuracy, and data integrity that are required under GLP. The main considerations from these two guidance documents are listed in Table 2. In addition to this guidance from regulators, further recommendations derived from scientific publications are discussed below.
Main considerations from OECD and FDA guidance documents regarding WSI use in GLP nonclinical toxicity studies.46,67

Abbreviations: CFR, code of federal regulations; DPS, digital pathology system; FDA, U.S. Food And Drug Administration; GLP, good laboratory practice; OECD, Organization for economic co-operation and development; WSI, whole slide imaging.
GLP Status of WSI
WSIs and their metadata, although digitally replicated from a physical specimen, do not meet existing definitions of specimens or raw data. 26 The current GLP status of WSI for nonclinical toxicologic pathology is that use of WSIs in lieu of glass slides is acceptable, providing it can be proven that WSIs are a faithful representation of the original glass slides, including all elements needed for histopathological examination or pathology peer review.46,67 It is important to note that a “faithful replica,” a term mentioned in the OECD FAQ document, is not defined with specific GLP requirements. A WSI, considered a faithful replica, is created from a specimen (i.e., a histology slide) and contains all necessary information (i.e., the labeling, markings, artifacts, and defects), whereas a true copy is a version of a record verified to contain the same information as the original record. 26
The current regulatory guidance from the U.S. FDA foresees that WSIs and associated metadata need to be archived if they contribute to the creation of raw data or study reconstruction in GLP studies. 13 This is the case for digital primary study evaluation and for retrospective peer review. Also, under certain circumstances of contemporaneous peer review, WSI retention would be required if WSIs are the basis for resolution of differences of opinion between pathologists, if there is an issue resolution process documented in the peer review memo, or if there is a peer review in which histopathology findings are only evident in the WSIs. 26 Although not explicitly required by the OECD, currently, in the scenario of primary slide evaluation on glass slides and contemporaneous digital peer review, archiving of WSIs can only be avoided if any discrepancy between pathologists is resolved using original tissue sections. 26 This is not a practical solution. In the future, with growing confidence in the equivalency of glass and digital slides, and the ongoing dialog with regulatory agencies, this requirement should possibly be reviewed if certain requirements are met in terms of “faithful replication” or “concordance.” Where there are disagreements between pathologists, the toxicologic pathology community considers that it can be left to the scientific judgment of the two pathologists involved, to base their agreement on the WSIs without the necessity for archiving. This flexibility would open the possibility for many institutions to use digital peer review without the need to establish a GLP-compliant digital archive solution. The concept of WSIs being equivalent to glass slides is discussed further in Box 1.
Concept of equivalency/concordance studies.

Requirements for IT Equipment
A core requirement for the successful implementation of digital pathology is the availability of high-resolution slide scanners and monitors, as well as computers that fulfill specific hardware requirements. 8 Scanners must be capable of capturing images at multiple magnifications with high fidelity, preserving the quality of the original glass slides. 10 Calibrated, high-resolution, large-screen monitors should ensure there is consistent color representation to allow for accurate image interpretation;26,57 however, pathologists should be aware that ambient lighting and reflections can also affect display performance. 10 The colors displayed on any viewing device should be vivid, with good white balance capabilities, and the display should be free from glare. 10
Color calibration procedures for both scanners and monitors must be described in relevant SOPs. 57 A process to ensure consistent color reproduction across WSI scanners and enhance color homogeneity in WSIs has recently been described. 15 The process includes two modules: (1) assessing scanner color reproducibility and (2) applying color correction to minimize deviation/variation. 15 Color variability can arise from inconsistencies in slide preparation, scanner hardware, and display devices. 15 Techniques reported to address this issue included stain normalization, internal color calibration, and external monitor calibration. 15 While some pathologists consider color standardization less critical for digital primary reads and peer review due to the adaptability of toxicologic pathologists to different stain profiles, it is more essential for image analysis purposes. 8 Therefore, routine scanner calibration and maintaining minimum standards for monitor quality and calibration are recommended. 8
As digital pathology can generate a large amount of data, associated IT infrastructure must be able to support high-speed data transfer and ensure storage systems can accommodate large collections of image files. 57 GLP-compliant archiving is further discussed below, but cloud-based storage solutions are increasingly being used to support multi-site collaborations; however, these systems must comply with GLP data integrity requirements, including those for traceability and secure data access, and their use as a GLP archive is still a matter of debate.27,57 Protecting the integrity and confidentiality of nonclinical study data are also critical and IT infrastructure supporting digital pathology must comply with cybersecurity standards to prevent unauthorized access, data breaches, or loss of data. 8
Validation Requirements for Equipment and Workflow
The use of WSI in GLP toxicity studies requires validation of the entire digital pathology workflow, from glass slide generation/labeling/scanning to viewing at the pathologist’s workstation through archiving. DPSs involve multiple hardware and software interfaces, and so there is potential for unnoticed critical failures that may produce a nonobvious loss of image quality. 57 To address this, all equipment used in this workflow must be qualified and staff must be adequately trained. 27 Risk-based validation efforts should consider the entire “pixel pathway” from image acquisition to display as a single system, acknowledging that, with time, some components such as monitors, may need to be swapped/upgraded without a requirement for revalidation if performance remains fit for purpose.8,57 Components of this pathway may vary according to hardware/software elements, the number of sites and their geographic locations, as well as the business need and intended use cases. 8 If the entire pathway occurs at a single institution (either single- or multi-site), the qualification and validation are the sole responsibility of the institution. 8 On the contrary, if the pathway results from a collaboration between institutions (i.e., between a CRO and a Sponsor), communication must be optimal as each partner is accountable for their respective activities, and this accountability must be clearly outlined in their respective validation documentation, written procedures, and in the study documentation.8,33 In addition, in multisite scenarios across international borders, one must be aware of possibly notable differences between national GLP regulations if pathologists are generating data in remote locations. 32
The validation of GLP-compliant DPSs should ensure scanned images accurately represent the original glass slides, with no omission of tissue areas and no loss of quality or data integrity throughout the workflow.3,33,57 DPSs are subject to the same requirements as any other GLP-computerized system, meaning they must meet AlCOA standards of being Attributable, Legible, Contemporaneous, Original, and Accurate. 56 As such, security measures ensuring image and metadata integrity must be validated. Validation processes should demonstrate that there is limited and authorized access to systems, WSIs, and metadata, and that roles have been established to define the level of access pathologists and other study personnel have to study material. 36 Electronic records should be retained securely and the ability to expediently retrieve the records must be in place. 36 In addition, raw images must remain unmodifiable. 36 If any alterations are required, these should not change the raw image but should instead be layered over it; these changes should be tracked via audit trails employing user-independent computer-generated time stamps. 36 In contrast to this, simple annotations like pins may be considered as pathologist notes and do not need to be validated nor retained. Another consideration to address during the planning of validation is the security of the WSIs and metadata when transmitting to external users. 36
An example outline for the validation of a scanner and image acquisition software for nonclinical toxicologic pathology has been previously published. 57 The plan states that test cases should be selected from relevant study types, and user acceptance criteria should be established to ensure digital images faithfully replicate the original glass slides, findings are visible, and annotations/labels are correct. 57 Additional outcomes of the validation plan are to ensure the fulfillment of user acceptance criteria, the purpose of the validation, stating in which cases the scanner will be used and to ensure responsibilities for troubleshooting are assigned. 57
Following a successful validation, the use of WSIs on GLP toxicity studies should be detailed in study plans and SOPs, and must cover quality control (QC), metadata, image inclusion/exclusion criteria, and chain of custody, referencing any relevant SOPs for the image management system (IMS) and WSI archiving. 26
GLP-Compliant Archiving of WSI and Metadata
To enable digital slide evaluation of GLP studies, dedicated storage must be provided, which requires the use of qualified servers. 8 The associated database must also be subject to minimum requirements (including, but not limited to, defined roles, user account management, and audit trails, etc.). In addition, the current regulatory view on use cases necessitating the archiving of WSIs and metadata is that archiving is required for a primary read (generating raw data) or for a retrospective peer review, just as glass slides and corresponding blocks should be archived appropriately.27,49 In situations where WSIs should be archived, the retention period should be consistent with applicable data retention policies and national regulations. 26
GLP-compliant archiving of WSIs presents unique challenges, particularly regarding long-term data preservation. WSI files must be stored securely, with appropriate audit trails, and GLP compliance requires that archived files should remain accessible and unaltered as they move through the system, between institutions, and over time. 57 One of the key considerations for archiving is how to manage technology obsolescence and ensure that archived WSI file formats can still be accessed decades into the future.9,33 This is a particular concern for the archiving of scanner-proprietary file formats. In addition, if “Software as a Service” is used in the digital pathology workflow, it must be ensured that WSIs are still readable in their native format, even if the service ends (exit strategy). The aspect of system interoperability is further discussed in Box 2.
System interoperability.

Both on-site and cloud-based solutions can be employed for WSI archiving, each with unique advantages and challenges. The choice between data storage on premises or in the cloud for archiving should be based on suitable risk assessments, and the following regulatory guidelines should be considered: OECD documents #17 50 (including Supplement 1) 47 and #22, 51 and the U.S. FDA guidance for industry on the use of WSI in nonclinical toxicology studies. 67 On-site storage solutions are more controlled but require significant investment in infrastructure and maintenance. 57 Systems that use cloud-based solutions offer scalability and remote access, including read-only access. However, they may face challenges if stored in unknown or geographically dispersed locations, as regulatory inspections often require on-site visits under general GLP requirements so that protection measures can be inspected.8,57 To avoid such problems, some countries only allow archiving of GLP data in GLP-certified facilities, which is often not the case for data centers. For cloud-based solutions in GLP environments, irrespective of whether they are internally managed or outsourced to an external cloud service provider, it is critical that appropriate knowledge, awareness, and oversight, and control of the system remain with the test facility. Transparency and awareness of the responsibilities of all involved parties is crucial and this is outlined in the OECD document #17 Supplement 1. 47 For the implementation of a cloud-based solution, the following key elements are instrumental for its GLP compliance: a detailed risk assessment, thorough cloud service provider assessment(s), clearly defined service level agreements, and the validation of the computerized systems hosted in cloud-based services. 47 Therefore, cloud-based solutions should be carefully considered and based on an open dialogue with national GLP authorities. 8
Any IT infrastructure supporting long-term WSI storage should guarantee data is maintained over time and allow data access/retrieval in the event of regulatory inspection. 8 The physical address of data centers hosting WSIs and their metadata may be required by some inspecting regulatory agencies, and secured access and GLP compliance, including disaster recovery plan, should also be specified in the service-level agreement when WSI storage/archival is taken care of by a third party. 8
An increasing number of software vendors are turning to cloud-based storage solutions, suggesting regulators and QA should consider virtual inspections to verify data center compliance. 57 Without this, the progress of digital pathology could be hindered, necessitating the invention and implementation of alternative technical solutions. 57 Archiving systems must, therefore, balance accessibility with the need to preserve data in an unaltered state over time, and this challenge is exacerbated by the growing size of WSI files. 26
Although debates are still ongoing regarding which file format should be archived, the original file from the scanner or the copy assessed by the pathologist, a risk-based decision is warranted.50,51 Other outstanding questions include: 5 Will cloud-based archiving be accepted universally by GLP authorities? What requirements must be fulfilled for archives to serve as read-only database simultaneously? Will the increasing confidence in the digital pathology workflow lead to more flexibility in terms of GLP retention period of WSIs if certain requirements are met in terms of “faithful replication” or “concordance,” since the original glass slides will always allow reconstruction of the study?
Lessons Learned and Outlook
The success of implementing a GLP-compliant, end-to-end digital pathology workflow hinges heavily on the collaborative efforts between the main stakeholders. These stakeholders include:
End users, including pathologists, laboratory technicians, and archivists.
IT colleagues, who outline and ensure adherence to a validated software development life cycle.
QA colleagues, who provide compliance oversight according to available regulatory guidance and assist in drafting the necessary documents that demonstrate all components were validated according to their intended use.
The rapid pace of technological advancement in digital pathology poses a substantial challenge for regulatory compliance and validation efforts. The pathologists and IT teams must continually re-validate through change control processes to account for new software updates, scanner models, and image management platforms. 33 Once digital pathology is better established, the integration of AI tools, which are discussed below, may further augment pathologists’ capabilities.8,27,33
AI in Toxicologic Pathology
In September 2021, the first U.S. FDA market authorization for an AI system in pathology was granted to Paige Prostate Detect, a software algorithm intended to assist pathologists in the detection of foci suspicious for cancer in WSIs from prostate needle biopsies. 11 Classified as a Class II medical device under the generic name “software algorithm device to assist users in digital pathology,” special controls were established for Paige Prostate by the U.S. FDA, including extensive requirements regarding design verification and validation, across three key areas: 66
A detailed description of the device software, including its algorithm and its development, together with a description of any datasets used to train, tune, or test the software algorithm.
Analytical studies to demonstrate acceptable analytical device performance, including detailed documentation.
Validation studies with clinical specimens and detailed documentation.
This first market authorization of an AI tool in diagnostic pathology can be seen as a door-opener for strategies to obtain regulatory acceptance of AI tools for routine clinical work. 54
Regulatory Status and Guidance
Current regulatory status of AI in nonclinical toxicologic pathology
Despite the U.S. FDA’s approval of the Paige Prostate Detect system, the limited regulatory guidance for a validation concept of AI is still the greatest hurdle for the successful development and implementation of AI in the lifecycle of medicines, especially in the field of toxicologic pathology under GLP. 53 Machine learning (ML) techniques integrated into image analysis software, such as random forest and neural networks, enable the segmentation, classification, and quantification of histological structures. However, these approaches often rely heavily on manually engineered features and exhibit low reusability. Subject matter experts typically tailor the software for specific studies, analyze the data, and frequently need to redesign the system for subsequent studies. Examples of qualification methods of AI-enhanced image analysis software applied to toxicologic pathology have been previously described. 73 To the authors’ knowledge, more advanced AI tools that can eliminate the necessity for time-consuming manual feature engineering have not yet been validated or widely deployed in the field of toxicologic pathology. While AI tools capable of WSI QC, slide triaging, and/or quantifying specific common lesions in rats’ organs are emerging, they still require validation. Nevertheless, since the end of the last decade, a number of publications from clinical and toxicologic pathologists, as well as opinion pieces and white papers resulting from initiatives of professional societies, have started to outline basic principles for AI tool generation and application. Publications covering key aspects regarding the use of AI in toxicologic pathology are listed in Table 3.
Key articles on the use of AI in nonclinical toxicologic pathology.

Abbreviations: AI, artificial intelligence; DL, deep learning; ML, machine learning.
In November 2019, Abels and other experts of the Digital Pathology Association published a white paper on best practices and recommendations for computational pathology, with a focus on its application to WSI. 1 As a general principle for training any AI algorithm, it is recommended to split annotated data into “training” and “validation” datasets and to compare results with a test/‘ground truth’ dataset, established by subject matter experts (Figure 1). Likewise, in the field of toxicologic pathology, the importance of strictly separating training, validation, and test datasets has been emphasized. 2 In this regard, the “gold-standard paradox” 4 arises from the dilemma that the gold standard is histopathological results from a human pathologist, whereas the algorithm data may, in fact, be more reproducible than human assessment; however, this is still matter of debate. 1
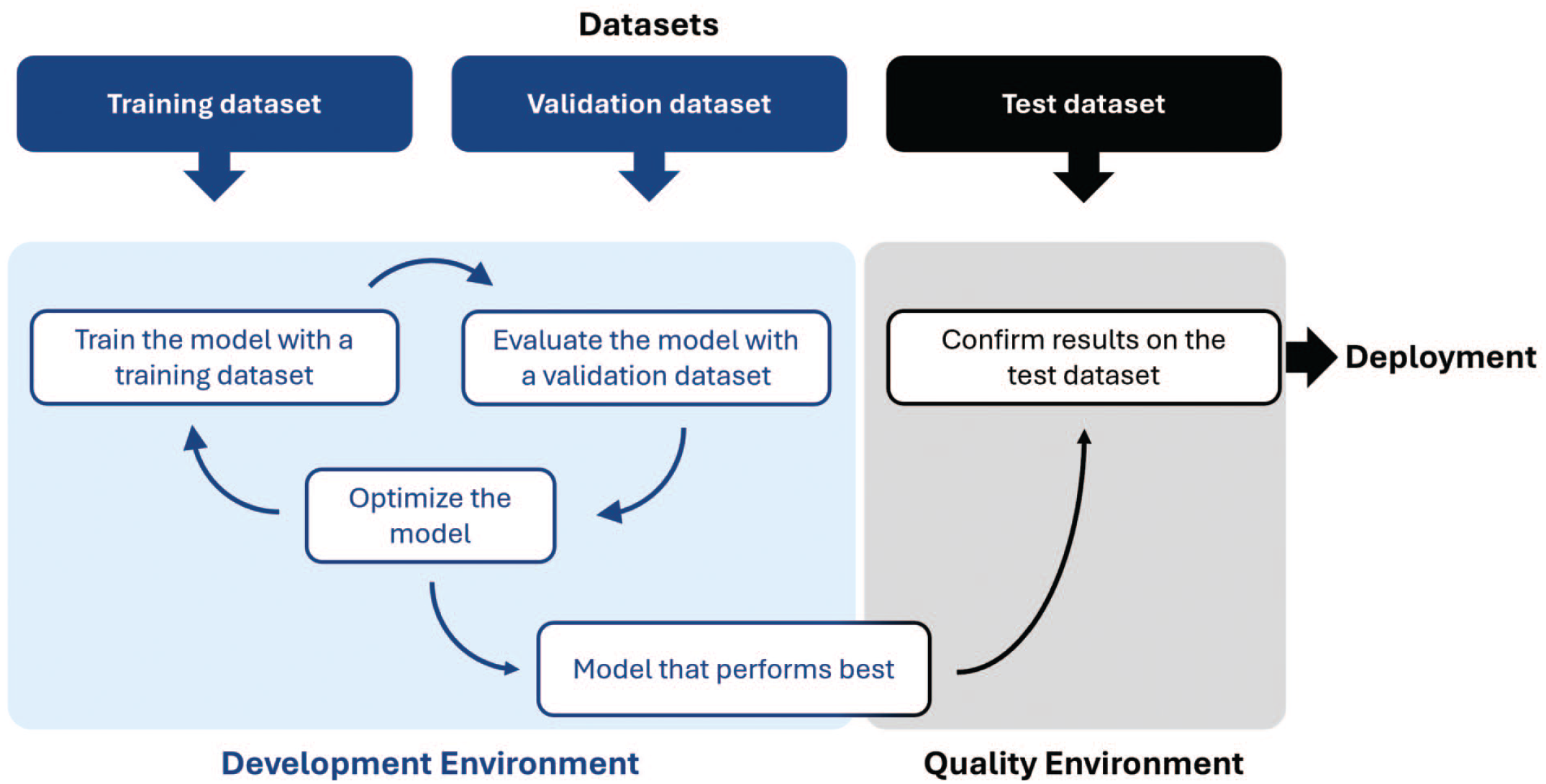
Data split during AI model development and final assessment.
Earlier in 2019, the Special Interest Group on Digital Pathology and Image Analysis of the Society of Toxicologic Pathology published an opinion piece introducing AI and ML to the toxicologic pathology community. 63 Besides defining relevant terminology, describing data generation and interpretation, and giving examples of use cases, they also suggested some basic parameters to be covered in AI model validation (i.e., AI model accuracy and precision, detection and quantitation limits, linearity, range and robustness, plausibility, relevance to target/mechanism in question, and the link to outcome of a disease or toxicity). 63 The concept of eXplainable AI (XAI),6,64 addressing the requirements of AI transparency, builds on this opinion piece. XAI aims to foster greater trust in AI, ensuring algorithms are rigorously validated and their outputs are scientifically sound, to ultimately increase the adoption and integration of AI. The U.S. FDA and European Medicines Agency (EMA) recognize this need and are working to establish guidelines that promote the responsible use of AI and ensure regulatory standards are adhered to.
U.S. FDA discussion paper and request for feedback on the use of AI/ML
In May 2023, the U.S. FDA released a discussion paper and request for feedback: “Using Artificial Intelligence & Machine Learning in the Development of Drug & Biological Products” to facilitate a discussion with stakeholders. 68 Resulting from a collaboration among the U.S. FDA’s Center for Drug Evaluation and Research, the Center for Biologics Evaluation and Research, and the Center for Devices and Radiological Health, including its Digital Health Center of Excellence, the paper addresses several key topics in the context of AI and ML. These include current and potential uses of AI/ML, considerations for the responsible use of AI/ML, and engagement with stakeholders to include, pharmaceutical companies, academia, patients and patient groups, and regulatory and other authorities. Following an overview of the current and potential future uses for AI/ML in therapeutic development, the paper then discusses the possible concerns and risks associated with these innovations and ways to address them. To this end, the U.S. FDA solicited feedback in areas such as validation and verification of AI/ML, model development, performance, and monitoring, quality and reliability of data, and governance and transparency of AI/ML model development.
EMA reflection paper on the use of AI
In July 2023, the EMA published a draft reflection paper on the use of AI to support the safe and effective development, regulation, and use of human and veterinary medicines, recognizing the rapid development of AI applications in the medical field and the need to prepare for the associated regulatory challenges. Open for public consultation until the end of December 2023, the final version of this reflection paper was issued in September 2024.
22
Here, it was stated that
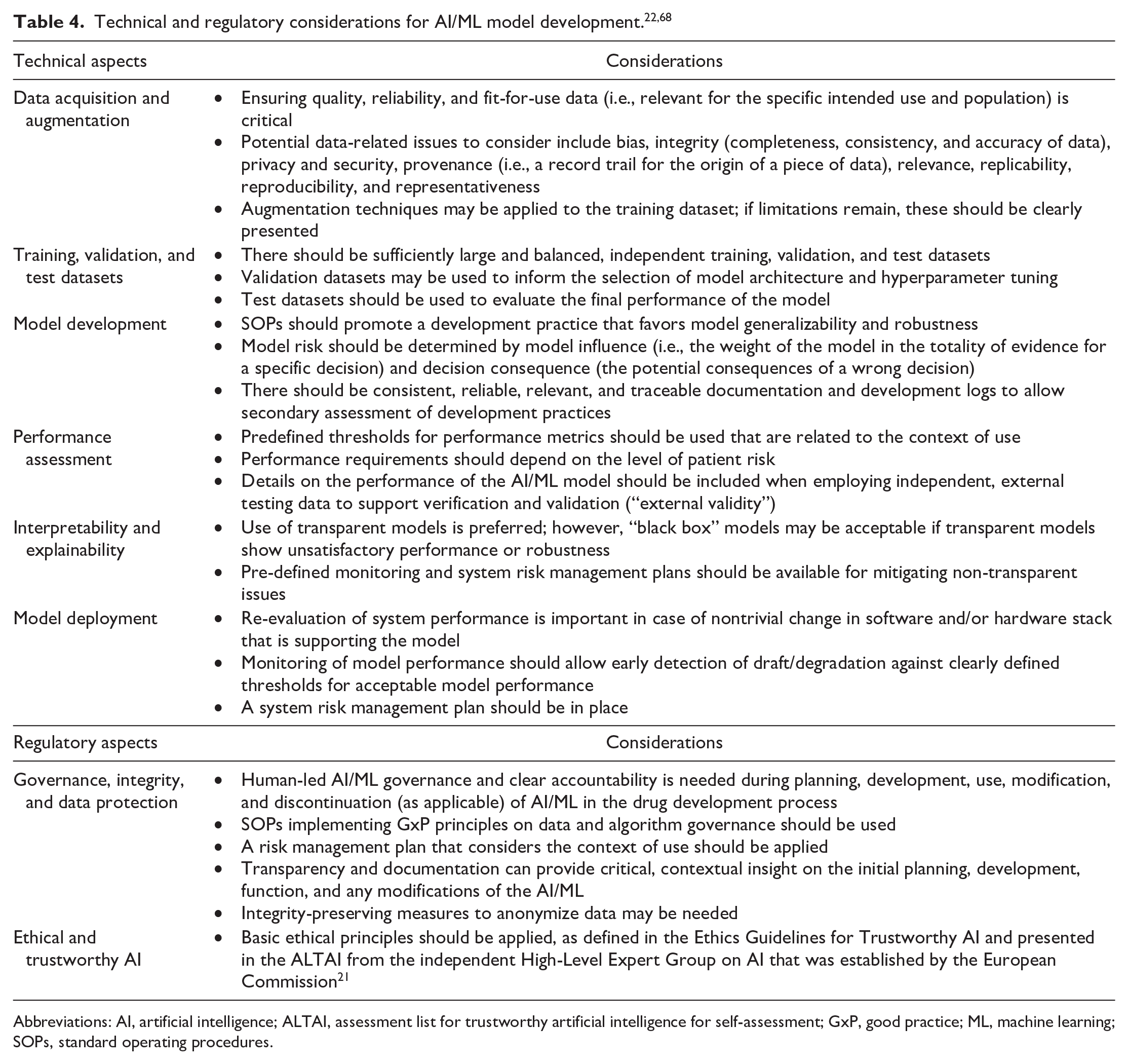
The EMA reflection paper also provides recommendations for technical aspects that should be covered in the development of AI/ML models. A combined summary of U.S. FDA and EMA considerations for the development and use of AI/ML models is outlined in Table 4.
EU AI Act
Following the U.S. FDA’s discussion paper and request for feedback on the use of AI/ML, 68 and the EMA’s reflection paper on the use of AI, 22 the EU became the first major regulatory body to have issued a legal framework for the use of AI. On August 1, 2024, the EU implemented the EU AI Act, a comprehensive legal framework with the intention to harmonize rules on the use of AI throughout the EU. 23 While fostering innovation, its main goal is to ensure that fundamental human rights are respected via safe AI systems. 23 Regulation under this Act is based on a risk-based approach regarding the potential impact of a given AI system on an individual’s health, safety, values, and fundamental rights. Only AI systems used for military or national security purposes, or for pure scientific research and development, are exempt from these regulations. 23 It is also important to note that as for the EU’s General Data Protection Regulation, the EU AI Act can apply extraterritorially to providers from outside the EU, if they have users within the EU.
Nonexempt AI applications are classified by their risk of causing harm and applications with unacceptable risks are banned (e.g., those providing “social scoring,” which may lead to discriminatory outcomes). 23 High-risk applications (e.g., medical software) must comply with security, transparency, and quality obligations, and must undergo conformity assessments. 23 Limited-risk applications, such as online chatbots, only have transparency obligations and minimal-risk applications, such as spam filters, are not regulated by the EU AI Act. 23
Ethics guidelines for trustworthy AI
Ethical guidelines for trustworthy AI have been developed by the High-Level Expert Group on Artificial Intelligence (AI HLEG), an independent expert group established by the European Commission in June 2018, and were made public on April 8, 2019. 21 Although largely centered on the human aspects of AI, requirements for trustworthy AI in toxicologic pathology can be drawn from chapter II (Seven key requirements to achieve trustworthy AI) and chapter III (AI assessment list to verify these key requirements) of the document; 46 these are listed in Table 5.
Seven key requirements for trustworthy AI and selected suggestions for AI in regulated toxicologic pathology taken from AI HLEG. 21

Abbreviations: AI, artificial intelligence; AI HLEG, high-level expert group on artificial intelligence.
AI maturity model for GxP application
Once it has been established that AI models are trustworthy, their maturity concerning GxP standards should be established, aligning with the aims of the International Society for Pharmaceutical Engineering (ISPE).
20
The ISPE mission is connecting pharmaceutical knowledge to deliver manufacturing and supply chain innovation, operational excellence, and regulatory insights to enhance industry efforts to develop, manufacture, and reliably deliver quality medicines to patients. In April 2022, the ISPE issued an AI maturity model as a foundation for AI validation in a GxP environment, whereby AI system maturity is understood as “
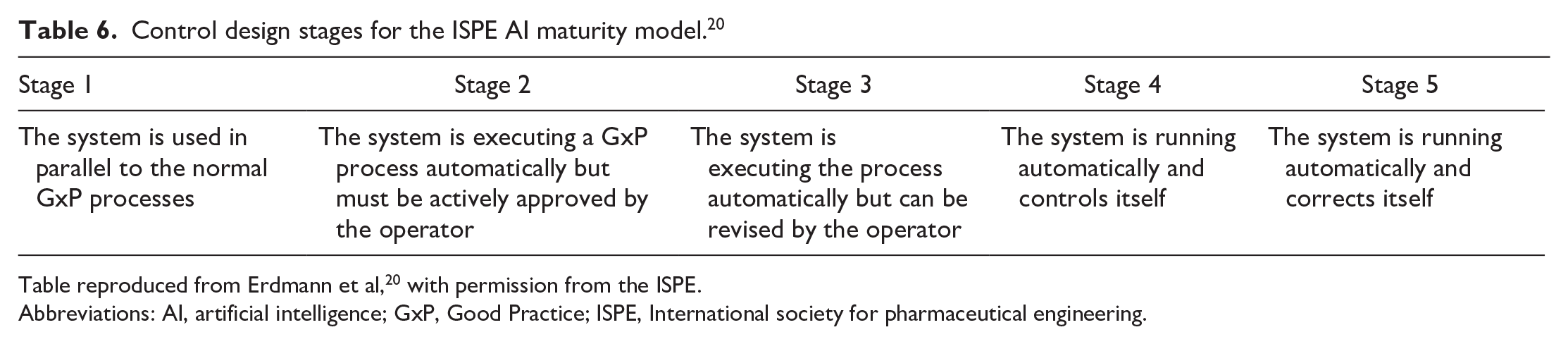
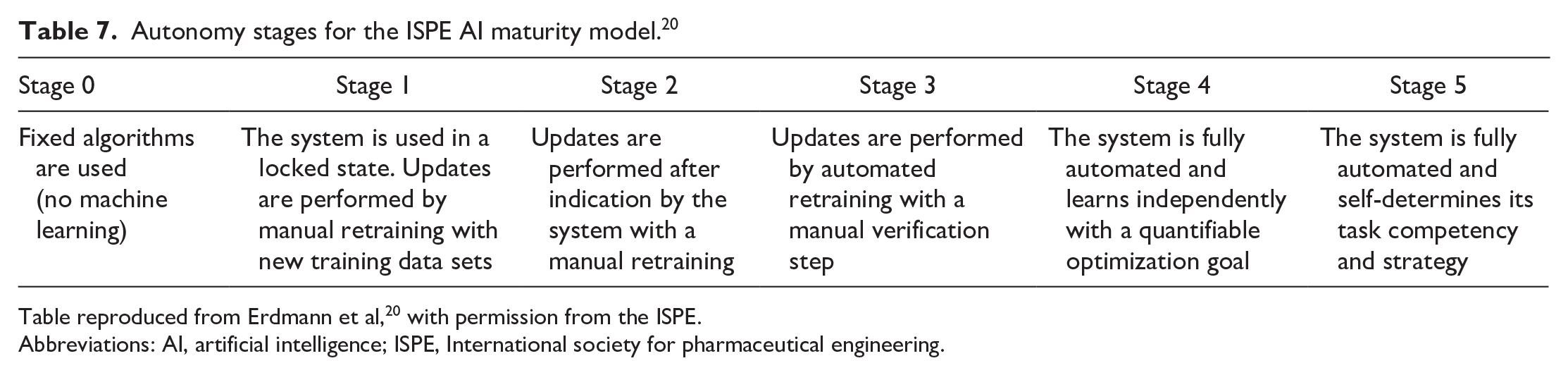
In this concept, AI model maturity is determined by control design on one hand and autonomy on the other (Tables 6 and 7). Control design refers to the system’s ability to take over controls that safeguard product quality and patient safety, while autonomy describes the system’s feasibility of automatically performing updates and improvements. 20 Depending on AI system maturity, six AI validation levels are suggested at a high level, to achieve regulatory compliance (Figure 2). However, it is understood that detailed QA requirements should be defined on a case-by-case basis.
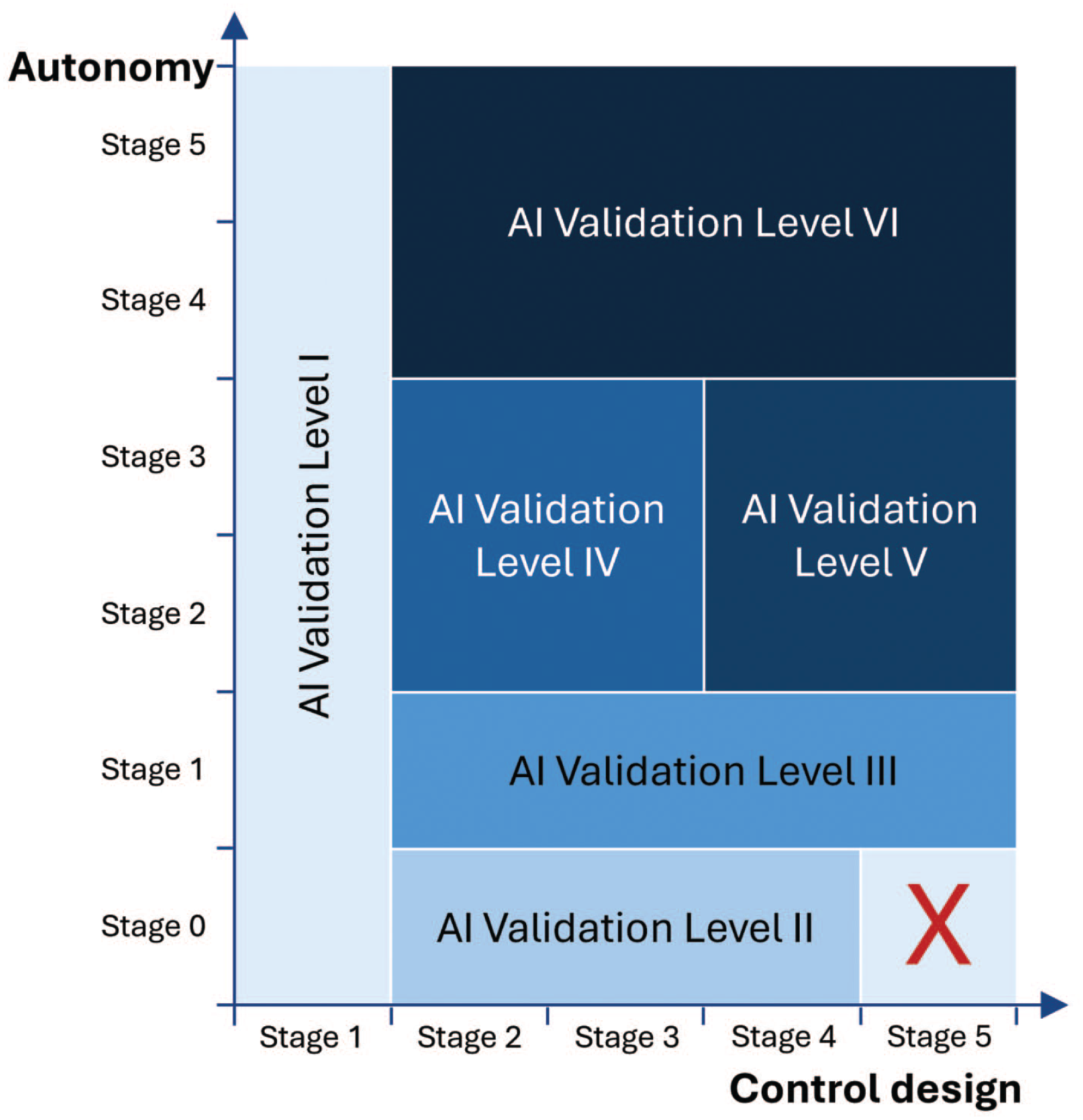
ISPE AI maturity model. 20
The six ISPE validation levels from the model are described in Table 8. 20 Briefly, systems in validation level I have no influence on product quality and patient safety; therefore, validation is not mandatory. However, safeguards are recommended to ensure that the operator is handling results based on critical thinking and does not use these results to justify decisions. Systems in validation level II are AI applications that are not based on ML and, therefore, do not require training. Instead, validation can be performed with a conventional computerized system validation approach. Validation level III systems are AI applications that are based on training with data for generating their outputs and that operate in a locked state until a retraining process is performed. Additional validation aspects regarding data integrity and model quality should also be performed. Validation level IV systems include AI applications with greater autonomy (i.e., with partially automated update processes). For these, validation should focus on controlling model quality during operation. Systems in validation level V are AI applications with a high level of autonomy and self-control, meaning that stronger controls should be in place and should be reflected in the validation strategy. The systems in validation level VI are self-learning AI systems and currently, a validation concept for this system level does not exist but is under development.
ISPE AI maturity validation levels. 20
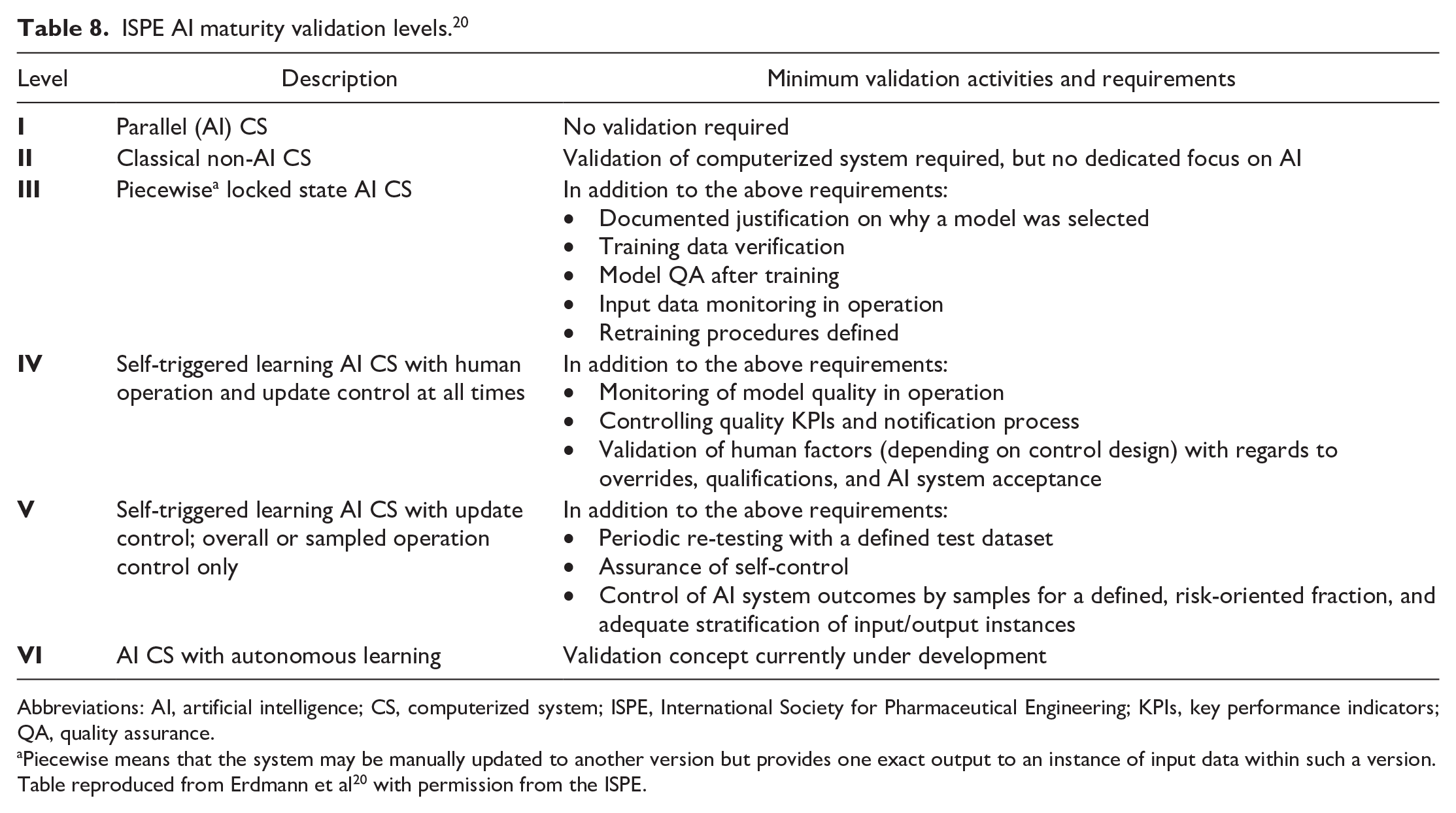
Abbreviations: AI, artificial intelligence; CS, computerized system; ISPE, International Society for Pharmaceutical Engineering; KPIs, key performance indicators; QA, quality assurance.
Piecewise means that the system may be manually updated to another version but provides one exact output to an instance of input data within such a version.
Table reproduced from Erdmann et al 20 with permission from the ISPE.
It is important to note that a dynamic development path for AI applications is suggested in this ISPE model for AI maturity. 20 For instance, initially a model may be developed with less autonomy and more control, resulting in lower validation requirements. During the system’s lifecycle, based on continuous evaluation of model success, there may be changes in use cases and newly identified risks, resulting in the adaptation of model design, either to allow more human control or to expand the system’s autonomy. While these systems and processes discussed above were not designed with toxicologic pathology specifically in mind, their principles are, nevertheless, applicable to the field of toxicologic pathology.
Use of AI to Support Regulatory Decision-Making in Drug Development
In January 2025, the U.S. FDA issued a draft guidance on the role of AI in the regulatory decision-making process for determining a drug’s safety, efficacy, and quality. 65 The draft guidance provides a risk-based credibility assessment approach to determining the reliability of an AI model based on risk and specific to the model’s context-of-use. It outlines a framework that sponsors may use to demonstrate risk and credibility of an AI model deployed in a decision-making environment during drug development. 65
Considerations for AI model development
Although the literature and regulatory guidelines published so far are generally meant for clinical applications, most of the principles developed in these articles may be applied to toxicologic pathology. AI algorithms for toxicologic pathology applications usually involve some type of pattern recognition task, such as detecting specific features, structures, or lesions in a histological image. Pattern recognition typically occurs at the patch level, object level, or pixel level. In some instances, the task could involve label prediction at the WSI level, for example, to determine whether an image is normal or abnormal.
How to determine ground truth
A fundamental requirement for developing successful AI algorithms is the availability of high quality, ground truth datasets. Creating high-quality ground truth annotations requires access to WSIs from multiple sources, such as different institutions, scanners, and staining protocols, and so on. Assembling such a dataset is not straightforward. However, the IMI Bigpicture project, a European public-private initiative, aims to address this issue by creating a repository of more than 3 million WSIs with associated metadata from multiple clinical and nonclinical sources. 44 Together, it is hoped that more than 2 million WSIs will be sourced from animal models and nonclinical toxicity studies from different organizations using a diversity of slide scanners and staining protocols.
Human-generated ground truth annotations are invariably susceptible to intraobserver and interobserver variability, but also to inconsistencies in image magnification, annotation tools (e.g., polygon vs free-hand drawing tools), and protocols (overlapping vs nonoverlapping annotations), to name a few. To address these challenges, recent efforts have proposed a consensus-driven approach, particularly the use of reference datasets. These reference datasets comprise small, curated sets of ground truth annotations, accompanied by detailed guidelines for annotation practices. Such datasets can reduce variability and enhance the robustness of ground truth data.42,70
The curated reference datasets serve two purposes; they enable annotators to benchmark their annotations against a standard, improving consistency, and the detailed guidelines help minimize inaccuracies and variability during the annotation process. Nevertheless, this consensus approach still relies on human annotations, which are often a bottleneck due to the limited availability of subject matter experts (i.e., pathologists). The recruitment of multiple pathologists for ground truth generation is typically restricted to large consortia and well-funded programs, making it impractical for many applications. To address this issue, crowdsourcing-based approaches have gained traction. These methods leverage nonexperts to generate ground truth data, often by embedding the annotation process in a virtual or gaming environment that incentivizes active participation. Typically, the same dataset is annotated by multiple participants and interrater agreement is used as a measure of annotation quality. However, in highly specialized fields such as histopathology, subject matter expertise is crucial for generating reliable ground truth due to the inherent contextual complexity of histological data. Currently, there are numerous efforts to improve the quality of ground truth labels for these specialized applications.
In Lopez-Perez et al., the authors trained an AI algorithm to detect tumor, stroma, and immune infiltrates from tumor biopsy hematoxylin and eosin (H&E)-stained images using two different training datasets. 37 One of the training datasets, which the authors considered as the gold standard, was generated by board certified pathologists, while the other training dataset was generated by medical students. Importantly, the authors used statistical ML methods to enrich the training dataset generated by the medical students, which resulted in comparable performance of the AI algorithm when the gold standard was used as training data.
Quality requirements for ground truth datasets
The quality and the size of ground truth datasets depend on the type of labels being predicted and the complexity of the AI algorithm. A good ground truth dataset must exhibit accuracy, class balance, completeness, and diversity. Accuracy is essential, as errors in ground truth can lead to poor model performance. Class balance ensures that the dataset provides appropriate proportions of labels, minimizing any imbalances that may affect training outcomes. The specific approach to achieving class balance depends on the application. For image-level classification tasks (e.g., normal vs abnormal slides), balancing the number of WSIs per class is crucial. For region-level detection (e.g., glomeruli in the kidney), the proportion of annotated pixel areas matters. For instance-level tasks (e.g., cell phenotyping), a balanced count of labeled instances for each category (e.g., tumor cells, lymphocytes, and endothelial cells, etc.) is essential.
Ground truth data should be representative of a wide range of real-world scenarios that the AI algorithm could encounter, which will enable it to perform well with new, unseen data. The ground truth should also cover a wide range of data scenarios that account for pre-analytic and analytic variables, such as variations in tissue source (e.g., different strains/species), staining (e.g., different H&E staining kits), sample format (e.g., tissue microarrays and longitudinal vs transverse sections), and slide digitization (e.g., slide scanner and magnification, etc.).
The roles of training and test datasets in algorithm development and validation introduce subtle but important differences in their quality requirements. While error-free labels are generally preferred for training, techniques exist to handle label noise during this phase, making minor inaccuracies manageable. 34 However, for regulatory approval and performance metrics calculations, the test data must be error-free. Similarly, while real-world datasets often exhibit intrinsic imbalances due to the natural prevalence of certain labels, care should be taken during test dataset preparation to avoid over- or under- sampling specific classes, which could bias performance metrics.
How to detect and manage bias
Bias in ground truth data is a major concern for AI algorithms, particularly in pathology applications. Bias can arise from visual and cognitive traps that affect pathologists’ annotations. 4 Visual traps occur when perception deviates from objective reality, such as optical illusions impacting the size, intensity, or color of features. Cognitive traps include diagnostic drift, where diagnoses or scores systematically shift across a study. Confirmation bias can also be an issue, where knowledge of treatment groups influences diagnosis.
Bias can also stem from nonhuman factors that inadvertently influence AI algorithms. For example, recently, an AI algorithm trained to classify tumor types using H&E images from The Cancer Genome Atlas learned to identify tumors based on their hospital of origin rather than morphologic tissue features. 18 This underscores the importance of appropriate data preparation such as removing metadata from WSIs and sourcing data from diverse origins, to prevent algorithms from learning non-histologic, source-specific patterns.
Bias reduction strategies include minimizing class imbalance, using data augmentation, human-interpretable features, and model cross-validation. Cross-validation evaluates model performance by training on different subsets of the ground truth data and testing on the remaining data. This approach helps assess generalization to unseen data and identify potential overfitting, provided biases in the ground truth are controlled. In toxicologic pathology, cross-validation poses additional challenges, as patches extracted from WSIs may lead to contamination if similar lesions from the same experiment appear in both training and testing sets. This could lead to some spillover/contamination and the resulting validation results do not adequately generalize the algorithm performance to unseen data. Thus, a mixing of image patches from the same animal in test and training datasets should be controlled for. In addition, rare classes may lack sufficient ground truth data, leaving them potentially unrepresented in either the training or test datasets during cross-validation.
How to use data augmentation
Training deep neural networks on histopathology image datasets poses unique challenges due to high data dimensionality, limited labeled samples, and variability in staining protocols and imaging equipment. Data augmentation is a key strategy to enhance dataset diversity, mitigating overfitting and improving model generalization to unseen data. Augmentation methods for histopathology must provide domain-specific needs, such as handling color variations from staining and preserving the spatial structure of biological features. Current research spans stain-specific color transformations, geometric augmentations, generative synthetic data, and fully automatic augmentation pipelines, but no consensus yet exists on the optimal data augmentation strategies for toxicologic pathology.
Color augmentations tailored to H&E staining are well-studied for addressing variability in slide preparation and imaging.52,60,62 Techniques like H&E color transformations introduce realistic stain variations, improving robustness in tasks like mitosis detection and tumor classification. 61 Automated frameworks, such as bilevel optimization and RandAugment, further refine augmentation by eliminating manual selection and outperform traditional methods. 24
Generative models, especially generative adversarial networks (GANs), offer another approach by creating synthetic histopathology images to address data scarcity and class imbalance.19,29,39 Methods like Auxiliary Classifier GAN and diffusion models have shown promise in augmenting underrepresented classes and boosting model performance. However, concerns remain regarding the realism and biological fidelity of synthetic images. Geometric and patch-wise augmentations, including patch stitching and region-level transformations have also been explored.59,71 These methods enable biologically meaningful augmentations while maintaining spatial consistency, making them particularly useful for tasks like multiple instance learning, where structural information at the subregion level is critical.
Assessing accuracy and precision of AI algorithms
Performance metrics for AI algorithms in pathology can be broadly categorized by task: classification or segmentation. Segmentation algorithms commonly use metrics like the Dice similarity coefficient (F1 score) or Intersection over Union, which measure the overlap between the true and predicted areas of a class on a scale from 0 to 1. A value of 0 indicates no overlap (poor performance), while a value of 1 indicates perfect overlap (ideal performance). Classification algorithms often rely on the area under the receiver operating characteristic (ROC) curve, which also ranges between 0 and 1. The ROC curve plots the true positive rate against the false positive rate across varying thresholds, and for multiclass problems, partial ROC curves are generated for each class by treating one as positive and the rest negative. However, these metrics have limitations, particularly due to challenges like class imbalance and WSI-level aggregation. In addition, their interpretability in downstream decision-making can be problematic. For example, while an F1 score near to 1 is desirable, practical considerations (data quality and diversity, task complexity, and algorithm selection) often result in less-than-ideal scores (e.g., 0.8). While higher scores generally indicate better performance, they should not be the sole acceptance criteria for the algorithm. Instead, calculating confidence intervals for metrics can provide a clearer picture of their reliability by showing the range of possible values.
Using independent datasets with orthogonal endpoints, such as pathologist scores, can further validate algorithm performance on real-world data. For example, if an AI algorithm detects a specific lesion, comparing its output to pathologists’ score on a study with control and treated groups provides a more realistic assessment of its utility. More broadly, it is important to evaluate whether the AI algorithm delivers tangible benefits, such as improved speed, accuracy, or scalability in processing images and generating meaningful information. Finally, interpreting and reporting these metrics requires careful consideration of the specific domain and task, and incorporating expert feedback (“pathologist in the loop”) is critical for evaluating the algorithm’s practical relevance.
Explainability of AI algorithms
Beyond AI performance metrics lies explainability. Toxicologic pathology AI models should be interpretable so their decision-making processes can be understood by pathologists and other stakeholders. XAI addresses the “black box” nature of AI by offering insights into how conclusions are reached, fostering trust and credibility. Regulatory agencies such as the U.S. FDA and the EMA emphasize the importance of human interpretable features (HIFs) for AI algorithms, although guidelines specific to toxicologic pathology remain limited.22,68
Heat maps and saliency maps are commonly used HIFs to visualize how AI models make decisions.43,55,58 Saliency maps highlight the contribution of individual pixels to a classification decision. For example, incorrect classification can arise if an algorithm prioritizes background pixels over those of interest. In contrast, heat maps show how broader regions in an image can influence decisions and provide a complimentary, higher-level view. While saliency maps represent bottom-up perceptual features, heat maps represent top-down semantic or contextual information.
Despite their widespread use, these qualitative tools have limitations. Their interpretation is often subjective and requires expertise from pathologists. Moreover, heat and saliency maps are model-specific, meaning the same task performed by different AI architectures could produce inconsistent visualizations. This variability can reduce their utility across models. Recent advancements aim to address these issues by introducing quantitative endpoints for HIFs. For example, Boggust et al, 7 have proposed a saliency score that is a feature-wise importance score, and describes the feature’s influence on the model’s output for a given label.
Life cycle management of AI algorithms.
The successful development of an AI algorithm is typically marked by its routine use for the intended application. In the GLP space, the sustained use of validated AI algorithms imposes the need for life cycle management to maintain its credibility. Life cycle management refers to the monitoring and management of changes to AI algorithm inputs or outputs, whether accidentally or deliberately, to ensure that the algorithm remains fit for use. Draft guidance from the U.S. FDA recommends the implementation of a life cycle maintenance plan for AI algorithms that support/generate regulatory decision-making data. 65 Specifically, the guidance recommends that the life cycle plan include checks for data drifts and periodic monitoring of model performance to ensure that the algorithm remains fit for purpose for the intended use. A risk-based approach for life cycle management is recommended to assess the impact of changes to the input data for the AI algorithm. In toxicologic pathology, changes in the input data could occur when the histology images are generated at a different source (e.g., a new CRO), or if there are systematic changes to the staining protocol that could affect the color profile of the stained slide. Depending on the outcome of the assessment, some retraining and/or retesting may be required.
Challenges and Outlook for AI
Overcoming the specific challenges to the use of AI in toxicologic pathology
The comprehensive pathology assessments routinely conducted by pathologists in nonclinical toxicology studies involve complex tasks that are challenging to replicate computationally. In addition, the field of toxicologic pathology poses significant challenges for AI due to its extensive diversity, which includes numerous species, up to 60 organs, each with substructures, and a wide range of potential artifactual, background, procedural, or test article-related findings per organ or even substructure. In established AI applications for pathology, models are typically trained and tested on the same set of predefined classes, allowing them to quantify expected treatment-related findings for a specific organ and species. However, this “closed-set” approach has limitations in the diverse and unpredictable scenarios that are encountered in toxicologic pathology. To address the variability and ensure real-world applicability, an “open-set” of recognition techniques should be aimed for. These methods allow AI models to create more robust models, identify and classify novel or unexpected features not included in their training, making them better suited for routine assessments in toxicologic pathology.
Histology-based foundation models, and generative and multimodal AI applied to pathology are, at the time of this publication, promising emerging approaches that, if deployed alone or in combination in routine pathology workflows, could significantly impact the field of toxicologic pathology. The integration of AI tools into the toxicologic pathology workflow offers numerous benefits for the future. These include streamlining histological processing, reducing animal usage, enhancing the efficiency, consistency, and accuracy of pathology assessments, minimizing bias and variability between observers, speeding up reporting, and uncovering new insights in pathological evaluations.
Foundation models
To address the unique toxicologic pathology challenges, AI models must go beyond clinical foundation models, which are primarily focused on oncology and cancer-specific tasks.38,69 Toxicologic pathology demands a broader and, therefore, a more versatile framework capable of capturing the full diversity of tissue structures and histopathological patterns across multiple organ systems and species. This inclusivity is essential to manage the wide variability of lesions encountered in toxicologic pathology. In this regard, HistoNet, developed by Novartis, represents a pioneering effort. 31 It is the first published foundation model specifically trained on toxicologic pathology data and was created even before the term “foundation model” was formally established. 31 HistoNet demonstrated the potential of such models to generalize across tasks and datasets, enabling applications like artifact detection, lesion grading, and CBIR. Looking ahead, the IMI Bigpicture project, with its ambitious collection of over 2 million nonclinical slides, provides a critical resource for advancing toxicologic pathology AI. By leveraging this dataset, future models will be able to achieve a new level of diversity and specificity, supporting developments tailored to toxicologic pathology’s unique challenges.
Generative and multimodal AI
Generative AI refers to AI systems trained on vast datasets that can create new content, such as text, images, or videos, based on the patterns and structures learned from existing data. Large language models can provide draft lesion descriptions using standardized terminology or generate draft pathology reports by synthesizing data from observation tables. They can also compare tabulated data with draft reports to flag discrepancies and suggest refinements. For image models, generative AI will enhance data diversity by generating synthetic examples of rare lesions or patterns for model training. Multimodal AI combines data from various sources or modalities, such as histopathological images, clinical pathology, and molecular profiles, to provide a more comprehensive or holistic assessment. This approach leverages the strengths of different data types and provides a holistic view of an animal’s condition. These approaches will help AI algorithms to model the adversity of lesions, identify the no-observed-adverse-effect level, and streamline the integration of relevant pathology descriptions into toxicology reports.
Applying AI tools in routine toxicologic pathology
The implementation of AI-assisted artifact detection, such as identifying technical or digital artifacts on WSIs, could automate decisions regarding the need for recutting, resampling, or rescanning, and function as a QC step in the workflow. Integrating such a model during the histology processing phase can save time and resources, while segmenting artifacts for downstream AI models. Similarly, AI-driven outlier or lesion detection algorithms could identify tissue areas from specific organs that deviate from the norm, such as when compared to concurrent or virtual controls. Eventually, WSIs from a study could be uploaded to an AI-powered platform, which would analyze the images, determine potential target organs at a group level, flag outliers at the slide level, and segment the relevant regions. By identifying possible target organs early in the process, AI tools could trigger the histology processing of intermediate and recovery groups before pathologist evaluation, thus saving time. The pathologists would then review the regions flagged by the AI platform, allowing them to focus their expertise on the most critical areas, improving overall efficiency and accuracy. In addition, AI models could propose preliminary diagnoses and grading for the flagged lesions, which the pathologist would then confirm, thereby reducing interobserver and intraobserver variability. Ultimately, further productivity gains could be achieved with generative AI models that automate report generation based upon the outputs of pathologist-confirmed lesion detection models, streamlining the entire workflow.
CBIR systems retrieve visually similar images from large databases based on the content of a query image. 14 This approach is beneficial for background changes by providing access to similar cases and their corresponding diagnoses in an extensive, centralized virtual control database. Pathologists could then use CBIR systems to search for similar cases by uploading a query image. The AI would retrieve and display images with similar features, along with their diagnostic information and animal metadata, helping the pathologist assess the spontaneous nature of a finding.
The concept of synthetic controls relies on the creation of artificial control organs or animals using generative AI models that have been trained on extensive control datasets. These datasets can be either monomodal (e.g., WSIs only) or multimodal (e.g., including both WSIs and a selection, or all in vivo and ex vivo parameters, measured in animals from toxicological studies). These synthetic controls could augment or ultimately replace existing control datasets by providing standards for comparison in toxicologic studies, thereby reducing animal use.
Conclusion
The future of toxicologic pathology is closely tied to the integration of digital pathology and the full transition to AI-assisted digital pathology represents a game-changing opportunity for the field. Pathologists must now actively shape how AI is adopted and applied to mitigate the influence of other stakeholders, potentially diminishing the pathologist’s central role in toxicity assessments. This moment calls for growth and adaptation; pathologists need to upgrade their skills to work effectively with digital pathology and AI, not only to use them but to guide their development and applications in ways that best serve the field. Training programs are critical to ensure that pathologists are equipped to understand and collaborate with these tools, enhancing their expertise while maintaining their vital role in decision-making. At the same time, it is important for pathologists to engage with health authorities and regulatory bodies to communicate the shared benefits of digital pathology and AI adoption—streamline workflows, improve consistency and accuracy, and ultimately, reduce the use of animals in preclinical testing. Despite the expected challenges of implementing such novel approaches into the highly regulated world of drug development, AI-assisted digital pathology will not only enhance efficiency but will also strengthen the overall quality and impact of pathologists’ contributions. By embracing these advancements and leading their implementation, pathologists can ensure that AI becomes a tool to enhance their expertise, not replace it. In essence, this is a pivotal era for toxicologic pathology, which requires an industry-wide commitment to growth and innovation, thus paving the way for more accurate data interpretation, more efficient workflows, and ultimately, the advancement of scientific discovery in nonclinical toxicology.
Footnotes
Acknowledgements
This publication is a deliverable of the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement No. 945358 (Bigpicture, https://www,bigpicture.eu/). This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation program and EFPIA (![]() ). The authors would like to thank Thomas Lucotte, Holger Höfling, and Manuel Hermann for their critical and impactful review comments on this manuscript. They also thank Ben McDermott of the Bioscript Group (Macclesfield, UK) for medical writing support in accordance with Good Publication Practice guidelines, which was funded by Boehringer Ingelheim Pharma GmbH & Co. KG.
). The authors would like to thank Thomas Lucotte, Holger Höfling, and Manuel Hermann for their critical and impactful review comments on this manuscript. They also thank Ben McDermott of the Bioscript Group (Macclesfield, UK) for medical writing support in accordance with Good Publication Practice guidelines, which was funded by Boehringer Ingelheim Pharma GmbH & Co. KG.
Authors’ Contributions
GPE conceived and drafted the initial manuscript. CH, JB, and SR provided substantial writing, reviewing, and editing of the initial draft. All authors provided scientific expertise associated with the topic, critically revised the manuscript and made additions and edits to reach the final submitted and approved version.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Publication costs associated with this article were funded by Boehringer Ingelheim Pharma GmbH & Co. KG.