Abstract
Complex in vitro models (CIVMs) offer the potential to increase the clinical relevance of preclinical efficacy and toxicity assessments and reduce the reliance on animals in drug development. The European Society of Toxicologic Pathology (ESTP) and Society for Toxicologic Pathology (STP) are collaborating to highlight the role of pathologists in the development and use of CIVM. Pathologists are trained in comparative animal medicine which enhances their understanding of mechanisms of human and animal diseases, thus allowing them to bridge between animal models and humans. This skill set is important for CIVM development, validation, and data interpretation. Ideally, diverse teams of scientists, including engineers, biologists, pathologists, and others, should collaboratively develop and characterize novel CIVM, and collectively assess their precise use cases (context of use). Implementing a morphological CIVM evaluation should be essential in this process. This requires robust histological technique workflows, image analysis techniques, and needs correlation with translational biomarkers. In this review, we demonstrate how such tissue technologies and analytics support the development and use of CIVM for drug efficacy and safety evaluations. We encourage the scientific community to explore similar options for their projects and to engage with health authorities on the use of CIVM in benefit-risk assessment.
Keywords
Introduction
In 2022, the U.S. Congress passed the U.S. Food and Drug Administration (FDA) Modernization Act 2.0,
47
removing the legal requirement to use animal testing in drug development. Reducing animal use in drug development has been a long standing goal of the pharmaceutical industry, and the Modernization Act spurred further debate on the use of appropriate new approach methodologies (NAMs) for submitting new drug applications.21,65 New approach methodologies are defined as any technology, methodology, approach, or combination that can provide information on a drug and/or chemical hazard benefit-risk assessment including in silico, in vitro, in chemico, and ex vivo approaches.3,66,68 New approach methodologies are not necessarily newly developed methods, but their application to regulatory decision-making or as a replacement for a conventional testing requirement is novel.
66
Based on Center for Drug Evaluation and Research
ESTP and STP Working Group “Complex In Vitro Models & Pathology”—Enabling Collaboration Between Pathologists, Bioengineers, and Biologists
To advance the development and use of CIVM in drug development, the ESTP Pathology 2.0 Subgroup was formed which is a collaborative effort between the European Society of Toxicologic Pathology (ESTP) and the Society for Toxicologic Pathology (STP). The goal of this subgroup is to bring together diverse groups of scientists working in the field to generate a road map for the successful characterization and implementation of CIVM in drug development. This subgroup is increasing the visibility of the collaborative efforts between engineers, bioinformaticians, and computational biologists, biologists, and pathologists at forums such as the ESTP congress (Torino, 2021 and Basel, 2023) and at the two inaugural Microphysiological Systems (MPS) World Summits (New Orleans 2022 and Berlin 2023).
The aim of this review is to inform and encourage pathologists to proactively collaborate with in vitro research scientists, and vice versa, to support the advancement of CIVM and their integration into standard drug testing and development paradigms (Figure 1). In addition, this review provides an overview on the collaborative possibilities, including technical and scientific expertise pathology units can bring to such collaborations, which might not be apparent to the scientific community of MPS scientists. Industry pathologists are trained in comparative anatomy and physiology and translational animal medicine; and understand the regulatory requirements for documentation, standardization, and validation of pathology methods and data. Thus, their knowledge and expertise can greatly contribute to the characterization and validation of CIVM 77 ; and should be leveraged to pave the path toward regulatory acceptance of CIVM data. Well-defined COUs, adequate characterization and co-developed assays and readouts for CIVM will strengthen their use for comparison of nonclinical to clinical data and enhance risk-benefit assessment in the future.7,48,67
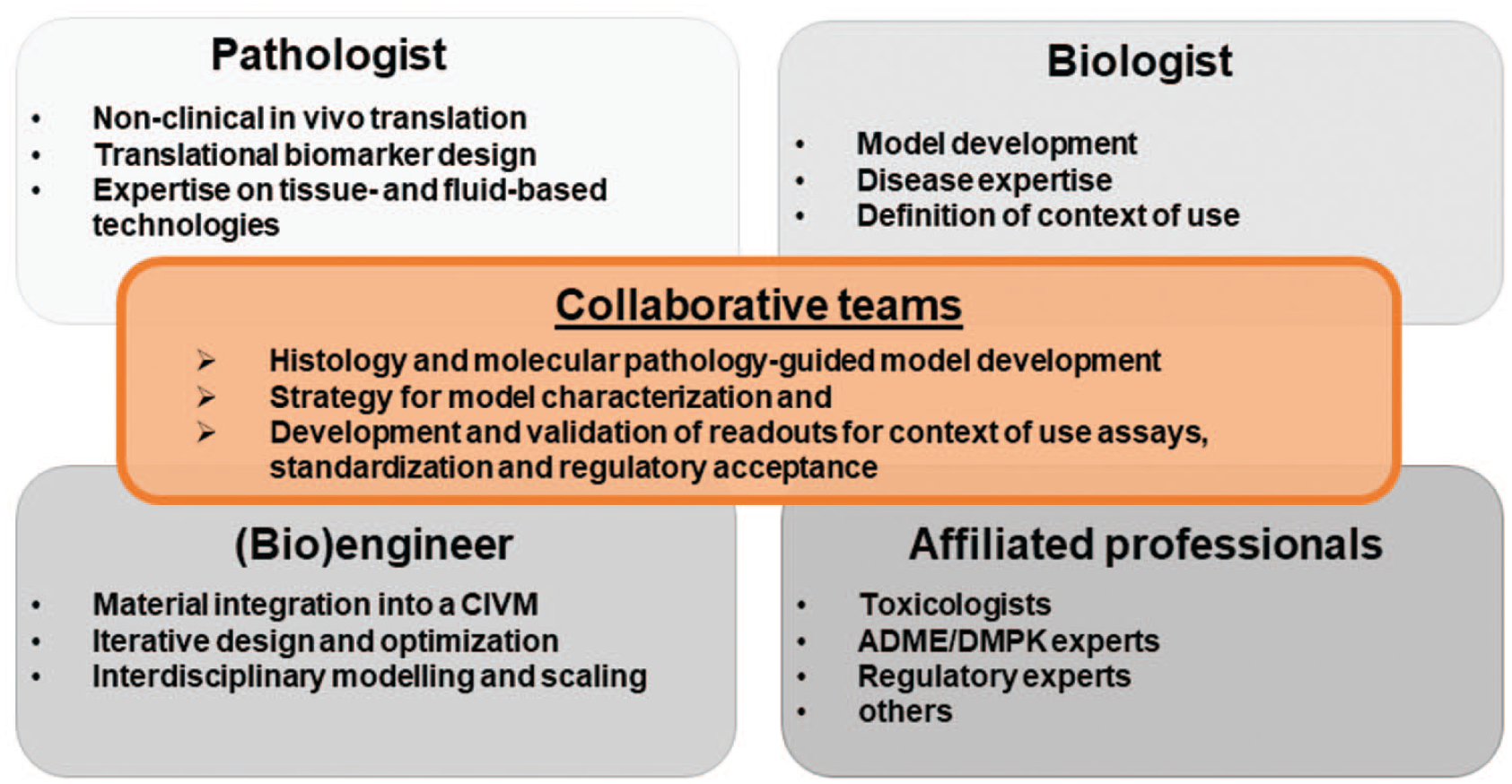
Collaborative teams of biologists, bioengineers, and pathologists define the context of use (COU) in drug development. We recommend that biologists and bioengineers work closely together with pathologists to develop complex in vitro models (CIVM) where using the relevant cellular input in the suitable microenvironment is one of the key drivers that determines successful development and implementation of the model. Furthermore, these collaborative teams can work on the strategy for CIVM, apply assays for characterization and drive standardization for regulatory acceptance.
Pathologists’ Expertise and Contribution to Advance CIVM Use in Drug Development
Pathologists are trained to identify and characterize the myriad ways normal biological systems respond to injury and disease. In addition, they are able to recognize and acknowledge the normal physiological processes in the steady state, as well as understand the mechanisms by which injuries occur and the host’s associated response to such injuries. Traditionally, this training is applied to animal models, underlying the belief that animal biology is relevant to humans. Anatomic pathologists assess macroscopic, light microscopic, and ultrastructural morphologic changes in cells, tissues, and organs to provide insights into the cause and pathogenesis of injury and disease. On the contrary, clinical pathologists assess cytologic, immunologic, and biochemical changes for similar purposes. In the pharmaceutical and chemical industry, anatomic and clinical pathologists integrate these complex data sets to understand efficacy models, identify hazards, and assess tissue and fluid-based biomarkers for new drugs under the umbrella of toxicologic pathology. 61 In addition, pathologists contribute to interdisciplinary teams by applying novel technologies such as spatial transcriptomics, single cell RNA sequencing61,64 (scRNA-seq), and digital pathology69,78 on tissues from preclinical and clinical studies to increase the translatability and human prediction in drug development. 56
Since pathologists are not specifically trained to apply their expertise to CIVM, they need to be consultatively informed on and exposed to the particular CIVM and their COU by the model engineers to be able to contribute to the morphologic evaluation and application of standard tissue technologies to CIVM experiments (Figure 2). Accomplishing this may require modification of methods and readouts ranging from in device analysis (Figure 2A), development of a novel nomenclature to even measurement of different biomarkers and endpoints (Figure 2B). In addition, pathologists may contribute toward defining relevant questions for CIVM experimentation, its limitation and predictive value, given their knowledge of the inadequacies of animal models in predicting certain aspects of human physiology and disease.
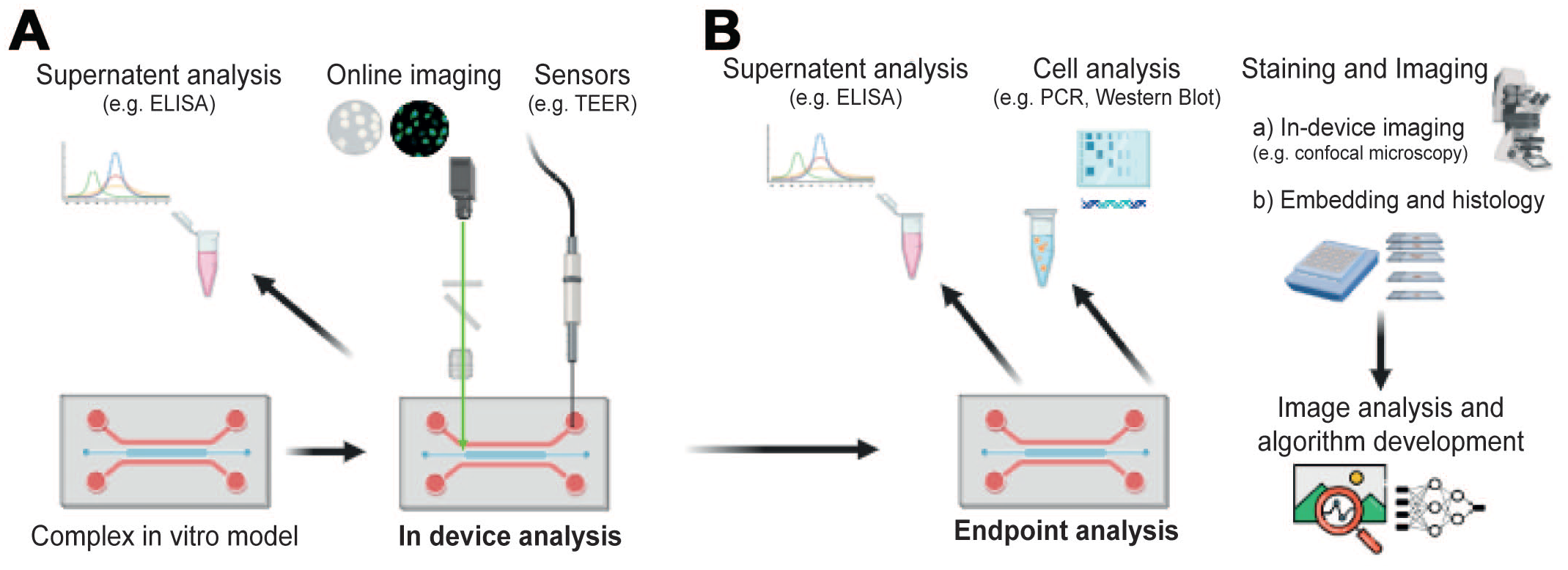
Overview of different readouts from complex in vitro models (CIVM). (A) CIVM can be assessed in the device while in culture by analyzing the supernatant, eg, enzyme-linked immunosorbent assay (ELISA), on-line imaging or different sensors such as the transepithelial–transendothelial electrical resistance (TEER) measurements. (B) Endpoint analysis can be carried out by various methods. Supernatants can be analyzed by ELISA, fluorescence-activated cell sorting (FACS) or other clinical pathology assays. Cells can be collected and analyzed by polymerase chain reaction (PCR), fluorescence-activated cell sorting (FACS) or Western Blot (WB). After fixation of the cells, staining and imaging can be performed by directly imaging in the device (eg, confocal imaging), or by embedding of CIVM and applying various tissue technologies (eg, light microscopy and electron microscopy). Once images are acquired, image analysis, and algorithms can be developed and applied to recognize and classify specific patterns. Figure created with biorender.com.
Recapitulating the complexity of in vivo biology by CIVM are challenging at the whole organism level. Thus, pathologists have a significant collaborative role to play in supporting the development and application of these novel systems, which often use reductionism principles to model biology in a scalable format11,59,67 (Figure 1).
Considerations From Biologist’s and/or Bioengineer’s View on Collaboration With Pathologists on CIVM Experimental Set Up
With the goal of replacing and reducing animal models, bioengineers are working on increasingly complex cellular models to closely mimic the in vivo physiology of target organisms. Such advancements are particularly crucial in the agrochemical sector. The Environmental Protection Agency (EPA) recently announced their goal to significantly reduce the use of animals in the development of new active substances by 2035. 20 Complex in vitro models are expected to play a key role in testing hypotheses derived from an NAMs assay battery. This battery is based on the Integrated Approaches to Testing and Assessment (IATA) 76 which combines several sources of information such as in silico prediction tools, high-throughput cellular assays, and “omics” data, aiming to achieve an organ-relevant, functional level of testing.
Developing CIVM require a deep understanding of in vitro cell culture systems combined with novel approaches such as three-dimensional (3D) cell printing as well as incorporation of cells into microfluidic devices. Model development requires developmentally correct combination of different relevant cell types to mimic differentiated 3D structures, the implementation of mechano-stimulation for specific tissues such as lung and heart to increase maturation, and the use of growth factors and specific biomaterials, scaffolds, and extracellular matrix (ECM) to facilitate cell and tissue growth, maturation, and function. 26 By far, the most important for tissue reconstruction is the selection of an optimal representative cell source, ideally sourced from a single donor. Nowadays, induced pluripotent stem cells (iPSCs) represent a significant breakthrough due to their potential to recreate complex tissue structures of uniform origin with high physiological relevance. 44 Furthermore, patient-specific iPSC-derived cells enable the replication of disease phenotypes in vitro, thereby providing valuable insights into disease mechanisms and potential therapeutic investigations. 38 During the development of CIVM, continuous collaboration with pathologists is envisioned to enhance realistic recapitulation of the in vivo 3D tissue cytoarchitecture, required for relevance of these models, as well as to ensure that tools for morphological and functional characterization are available.27,77 Morphological markers may be used to verify cell differentiation status, assess cellular polarization by staining basolateral and apical marker proteins, and monitor the cellular organization and stability of the CIVM over time. However, this represents only the initial step toward a toxicologically relevant CIVM.
The ultimate goal of CIVM experiments relies on predicting in vivo-relevant human and animal toxicity, which requires close collaboration with pathologists (Figure 1). Validating CIVM to successfully predict specific toxicological outcomes requires comparison with relevant data from a multitude of in vivo studies. 13 Confidence can be built in the CIVM only if the in vitro data aligns with the pathological outcomes observed in vivo. For a successful prediction of drug and environmental safety, several allometric scaling factors have to be taken into account to extrapolate from in vitro data to the clinical outcome. This may include differences in the nominal exposure, metabolic rates and differences in cell and fluid ratios. While comparisons between human CIVM and clinical data are ideal, the use of animal cell-derived CIVM may also be useful in this respect as animal toxicology studies provide a rich data set for comparison to CIVM outcomes. Evaluation of a CIVM response to toxicological challenges often relies on functional assessments rather than morphological changes, which may include assays for secreted biomarkers, barrier function, or other organ-specific functions. Biomarkers such as the concentrations of soluble “synthetic” proteins (eg, albumin), cell membrane leakage enzymes activity (eg, ALT), and levels of inflammation (eg, various cytokines) in the medium at different time points can be indicative of toxicological effects. Also, oxygen and glucose/lactose concentrations can be measured in the system by enzyme-based sensors and provide a status of the CIVM metabolic state in real-time. 5 Monitoring of these biomarkers is analogous to what is done in vivo (eg, hematology and clinical chemistry) and provides vital information about the CIVM and potentially insight as to how a toxicity may be monitored in vivo (Figure 2).
Examples of monitoring toxicity in vivo can be found in specific complex in vitro kidney models.24,72 For example, potential toxicity to renal transporter function can be investigated in experiments using transporter-specific substrates and inhibitors. 72 Megalin-cubulin-directed endocytosis of albumin by renal proximal tubule epithelial cells (RPTECs) can be measured in CIVM and was demonstrated to be increased in a 3D versus a 2D system. 25 Cisplatin-induced toxicity to RPTECS was abolished in a 3D kidney model when the cells were exposed to an inhibitor of the OCT2 transporter. 46
Biomarkers (i.e. secreted albumin), when combined with spatial data (i.e. histology), can be powerful tools for evaluating toxicity in vitro, and although not commonly, could be used as readouts in clinical pathology. For example, changes in albumin secretion are a common indicator of hepatotoxicity in vitro. 10 Nevertheless, this is not a reliable biomarker for early liver injury in vivo, thus the direct comparison between in vitro and in vivo studies is difficult. This should be considered when comparing in vivo versus in vitro data. Thus, the implementation of pathological and nonpathological readouts in each CIVM should remain in constant discussion between bioengineers and pathologists to find the most powerful parameters to predict more accurate in vivo outcomes and together define the correct frame of interpretation.
Adaptation of Tissue Technologies to CIVM to Support CIVM Engineering, Characterization, and Enable Pathology and Toxicity Evaluation
Complex in vitro model can be used for higher throughput drug screening and potentially more translational biomarker-driven mechanistic toxicity research.
Generation of CIVM to investigate tissue pathology, drug-induced or spontaneous, will benefit from a pathologist’s input on the salient histopathological, ultrastructural, or molecular pathology features as well as confirmation that the aspects to be investigated are maintained. In addition, pathologists can also evaluate and comment on the inter-CIVM heterogeneity to improve the robustness of system when used as a tool to investigate toxicity or molecular mechanisms. 77 In general, CIVM cannot fully recapitulate all features of in vivo tissues or a disease process. For toxicity assessments, biologists, toxicologists, and pathologists will benefit from working together to identify and delineate the scientific questions that can be addressed and define the specific COU for each individual model (Figure 1) being developed.
As an example, tumor organoids are promising tools for cancer biology investigations and preclinical drug screenings because they are often representative of the histology and drug responses of patients.23,45,58 To be able to compare and assess their translatability, assays need to be harmonized and the CIVM need to be co-developed together with the endpoint. 45 High throughput histology is a valuable method to guide the engineering of novel CIVM, perform morphologic characterization, and evaluate the best treatment time point when utilized for efficacy and toxicity assessments.23,54 Histology of CIVM can also be used to study molecular pathways and compare treated versus nontreated CIVM, as this has been described for a microfluidic micro-histological platform that enables ex vivo culture of a large array of prostate and ovarian cancer microdissected tissue 63 (MDT) (Figure 3). Our recommended strategy for model characterization is first to compare the morphology and cell composition of the CIVM with the corresponding tissue of origin and/or functional unit, respectively, with routine histology of Hematoxylin and Eosin (HE)-stained samples (Figure 3A). Second, the expression of key markers and targets needs to be confirmed and again benchmarked to the tissue of origin (Figure 3B). Thirdly, a deeper characterization by molecular pathology and/or ultrastructural imaging can be applied, to understand similarities and differences between the CIVM and their tissue of origin and/or functional unit (Figure 3A, C), as well as any inter-CIVM variation. Depending on the COU, the characterization needs to include the CIVM cell compositions, cell maturation states, functionality, morphology, and spatial distributions.
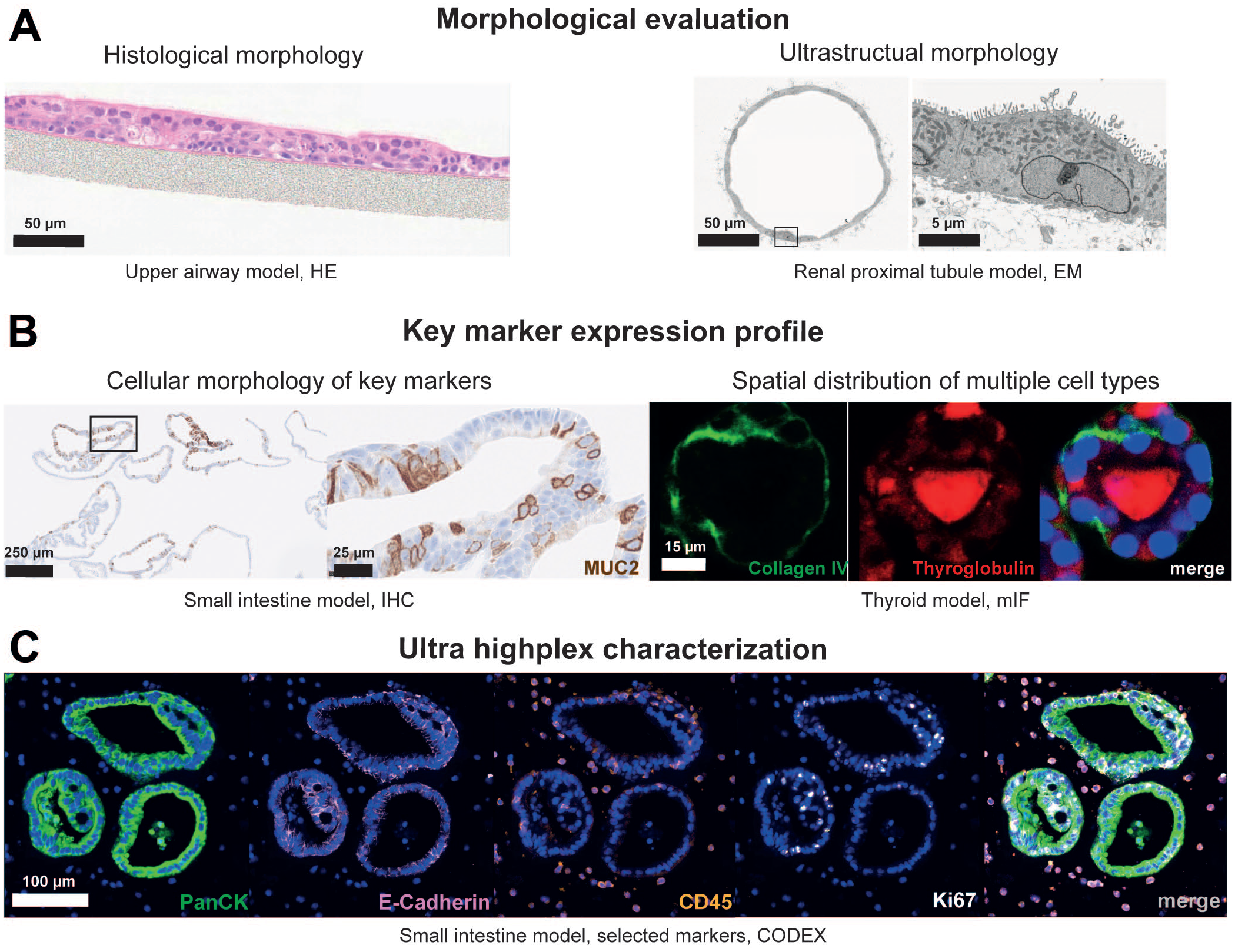
Molecular pathology methods and readouts for characterization and model validation. (A) Morphologic evaluation of complex in vitro models (CIVM) can be achieved on a histological or ultrastructural level by Hematoxylin and Eosin (HE) stain (example: upper airway model) or transmission electron microscopy (TEM)/scanning electron microscopy (SEM) (kidney on a chip), respectively. Here, TEM demonstrates appropriate polarity of the cells with presence of tight junctions, apical brush border-like ultrastructure, the endocytic system, and basal cellular interdigitations in a complex in vitro model for renal proximal tubules. (B) Expression profile of key markers for a given CIVM can be achieved by immunohistochemistry (IHC) or multiplex immunofluorescence (IF). Cellular morphology of key markers can be assessed by IHC, eg, Mucin 2 (MUC-2) expression was detected in intestinal organoids. Multiplex IF can be used to monitor cellular polarization (apical and basal) and visualize tissue architecture, eg, basally expressed collagen IV (green) and apically secreted thyroglobulin (red) in a human 3D thyroid model, image modified from Kühnlenz et al. 34 (C) Complex characterization of CIVM can be achieved by high-plex proteomics as exemplified by a 25-plex CODEX® staining on a section of formalin-fixed and paraffin-embedded (FFPE) organoids of the small intestine co-cultured with immune cells.
Tissue embedding techniques to enable morphologic evaluation of CIVM
Novel technologies around CIVM biology are evolving, but there is sparse information on simple and efficient methods to evaluate microtissue (i.e. organoid) morphology. Structural and ultrastructural assessment of CIVM can provide key information during the validation of a model including the identification of cell-types present and features of cellular viability and maturity compared to normal tissues. Models that are more complex can involve more than one cell type, for example, co-culture of intestinal or liver cells together with the respective epithelial/endothelial cells in microtissues. In this case, structural assessment can aid in ensuring that all cell types are present in the correct ratios to form proper 3D orientation and that they are stable over time.29,37 The application of conventional histopathology techniques allows the observation of multiple microtissues in a single cross-section, maintains good morphology, and enables further morphological readouts and quantitative image analysis in a high throughput manner.17,51 Ultrastructural techniques allow investigation of cell-cell and organelle interactions, which can provide important additional information in the characterization of a CIVM or as a readout in mechanistic studies.
The adoption of specific embedding methods is necessary for each model based on their prior cultivation, ease of handling and subsequent application, both in terms of staining and further image evaluation. Depending on the in vitro device and type of cells, various workflows for histology embedding techniques can be applied in a high-throughput manner (Figure 4). Advantages and disadvantages for each technique are summarized in Table 1. In general, the generation of optimal cutting temperature compound (OCT) blocks (Figure 4A) is faster (only 2 hours) compared to formalin-fixed paraffin-embedded (FFPE) blocks (Figure 4B–E), but results in inferior morphology of the sections. In contrast, formalin fixation better preserves morphology and can be either initiated from organoid pellets (Figure 4B) 17 or directly on the device (Figure 4C) which also reduces the manual handling. 10 Slides from both OCT and FFPE blocks can be used for several applications, eg, HE staining, immunohistochemistry (IHC), or multiplex immunofluorescence (IF), followed by whole slide imaging up to 40×. Depending on the hypothesis to be tested, any one or more of these techniques are indicated. For example, for organoids and spheroids, the pellet method—either OCT and/or FFPE—represents a reliable high-throughput embedding approach (Figure 4F, G). If there is an additional need to retain the spatial information between co-culture systems, eg, peripheral blood mononuclear cells (PBMCs) co-cultured with intestinal organoids23,54 (Figure 4H), the droplet method or direct in well fixation (by HistoGelTM) 10 is indicated (Figure 3C, D). Owing to their special construction, MPS models often require an adaptation by direct fixation and embedding in the device in a lithographic manner 63 to preserve the morphology of the cellular network in the chamber (Figure 4E). As exemplified by the human inner blood-retinal barrier-on-a-chip 42 (Figure 4J), the fixed and embedded cell networks and layers can usually only be extracted when removing the chip material surrounding them.
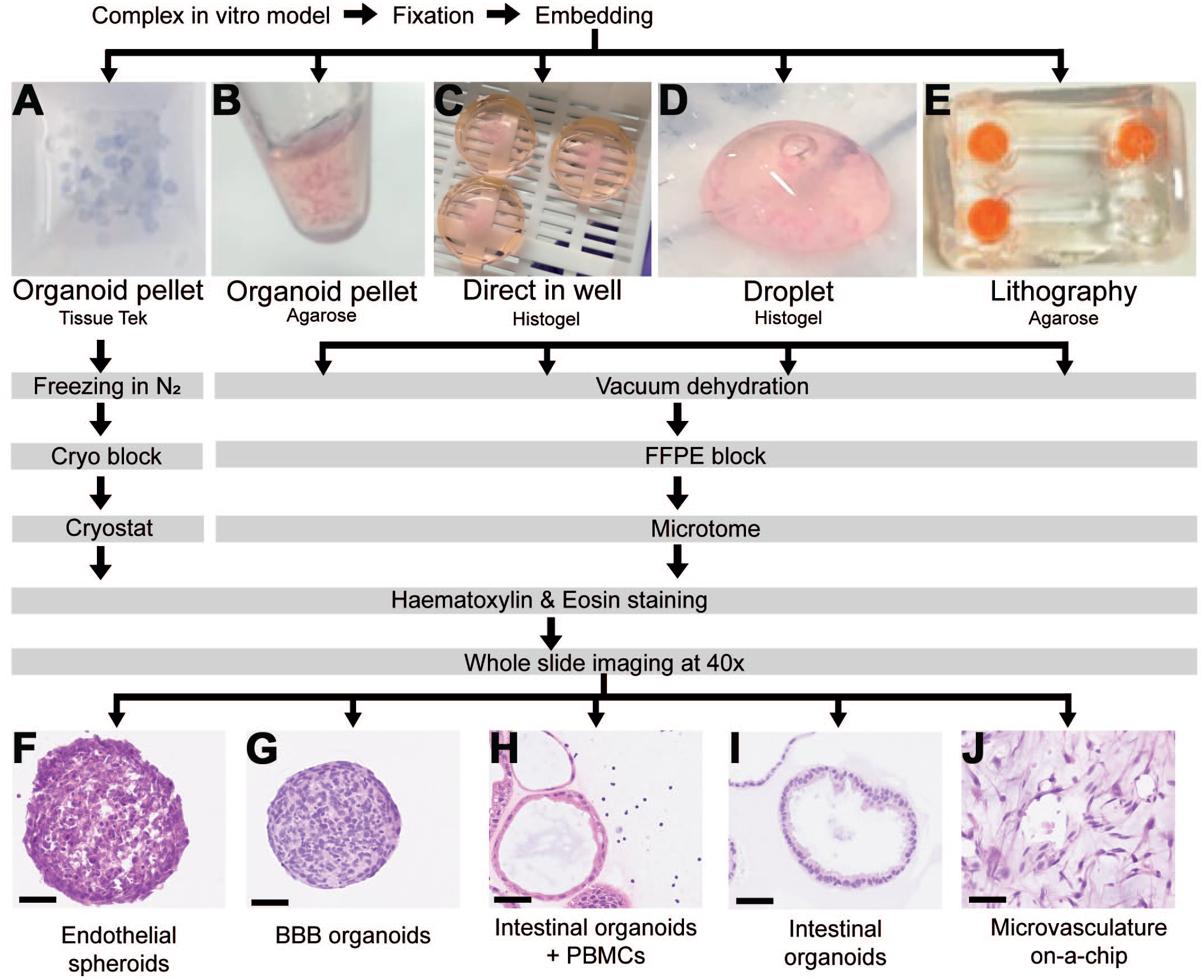
Adaptation of tissue embedding techniques to complex in vitro models (CIVM) to enable morphologic and molecular pathology evaluation. (A)-(E) Optimization of high-throughput histo-techniques for CIVM depending on their individual needs. Fixed CIVM can be embedded as organoid pellet to produce (A) cryo blocks or (B)-(E) FFPE blocks. Embedding (C) direct in well or as (D) droplet can be performed using HistoGel™. (E) Perfusable microphysiological systems (MPSs), such as microvascular on-a-chip can be embedded in agarose in a lithography-like manner. (F)-(J) Imaging of embedded CIVM as whole slide images to enable histopathological evaluation. (F) Endothelial spheroids were collected as an organoid pellet and embedded as a cryo block in OCT. (G) BBB organoids were also collected as an organoid pellet but processed as an FFPE block. (H) Intestinal organoids co-cultured with PBMCs required direct in-well embedding to retain the spatial distribution of the PBMCs around the organoids. (I) Intestinal organoids grown on Matrigel™ were subjected to droplet embedding to decrease the manual handling. (J) The cellular channel of a microvascular-on-a-chip 42 was embedded in agarose from inside and outside to preserve the morphology (lithography). Scale bar = 50 µm.
Advantages, disadvantages, and examples of different histology embedding techniques for CIVM.
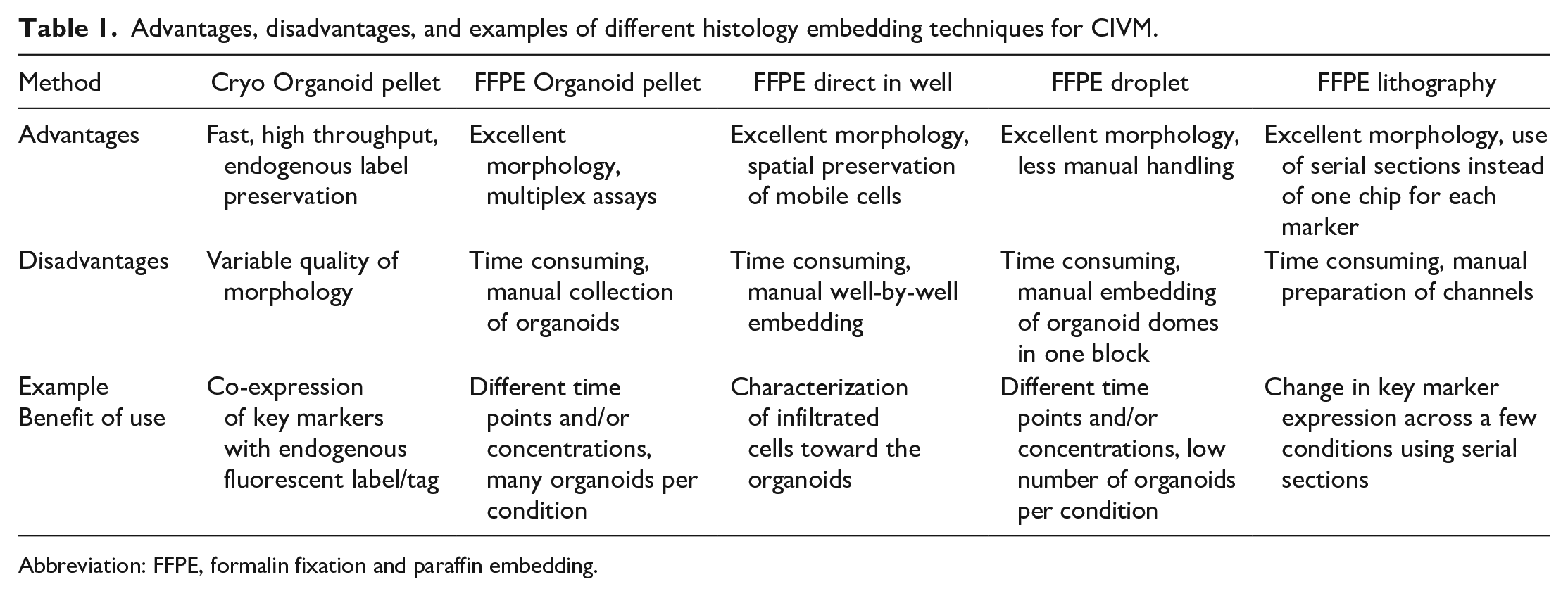

Abbreviation: FFPE, formalin fixation and paraffin embedding.
Ultrastructural assessment and embedding techniques for CIVM
The initial fixation of CIVM for ultrastructural assessments are similar to that described for FFPE above. For example, a microfluidic chip model may be perfused with Karnovsky’s EM-grade fixative before the tissues are carefully dissected for further, standard processing. Ease of access to the cellular tubules, and ECM in MPS models varies significantly and may require cutting through materials such as polydimethylsiloxane (PDMS), glass or plastic coverslips/encasements with currently available models. Recently, one author within The ESTP and STP working group has adopted the use of backscattered electron scanning electron microscopy (SEM) for the ultrastructural assessment of tissues, organoids, and organ-on-a-chip. 55 This technique has the advantage of allowing multiscale imaging that provides transmission electron microscopy (TEM)-like ultrastructure image quality without the loss of histological context.24,25,75
For example, ultrastructural assessment of RPTECs in a kidney CIVM was able to demonstrate appropriate polarity of the cells including development of a robust apical (lumen adjacent) brush border, tight junctions, and the endocytic system in addition to appropriate basal cellular interdigitations.24,25,75 Appropriate distribution of renal transporters on the apical or basolateral aspects of the RPTECs can be confirmed by IF. Functional transporters can be further investigated in experiments using transporter-specific substrates and inhibitors.25,46
An example of an ultrastructural assessment of human 3D renal proximal tubules cultivated in a microfluidic chip (Nortis Bio Inc) is provided in Figure 3A.
Multiplex and highplex IHC and in situ hybridization provide valuable readouts for CIVM
Histology embedding and processing of CIVM enables their subsequent assessment as performed for tissue-derived histology sections. Immunohistochemistry/immunofluorescence, and in situ hybridization (ISH) enables the visualization of multiple structural or functional markers/ribonucleic acid (RNA) in CIVM and can thus support basic model characterization (Figure 3B). Moreover, high-plex proteomics platforms can be used to simultaneously stain more than 100 protein markers on a tissue slide and provide complex structural and functional readouts for CIVIM. Using a similar approach, Lee et al used the CODEX® platform to characterize ducts, lobules, and associated stromal cells of a human breast tissue mimetic organoid model by spatially localizing more than 25 antigens associated with different cell types. 36 Likewise, Whale et al applied iterative indirect IF imaging (4i) on histological sections of retinal organoids to assess the time course of human retinal organoid development using 63 antibodies covering major retinal cell types, subcellular compartments, morphological structures, and signaling pathways.73,33 An example of an intestinal organoid characterization using a 25-plex CODEX® panel18,30 is shown in Figure 3C. Considering the small size of some CIVM and that a limited number of sections can be produced from one paraffin block, multiplex and high-plex proteomics and corresponding image analysis pipelines provide valuable tools to comprehensively characterize cellular marker expression and their spatial association in CIVM. Furthermore, these data are useful when comparing biomarker profiles with the tissue of origin.
Spatial transcriptomics: A novel toolbox to characterize CIVM
Single-cell and single-nucleus RNA sequencing enable the comparative assessment of the transcriptome of defined cell types as well as their developmental or disease state in CIVM and the corresponding tissues.9,12,15,53,73 However, these technologies lack spatial resolution and do not enable the association of transcriptomic signatures with morphologic features. In this regard, embedding of CIVM, especially spheroids, organoids, assembloids, and co-cultures as cryo or FFPE material allows their analysis on novel spatial transcriptomics platforms. These technologies provide whole transcriptome or panel-based readouts of up to 1000 RNA targets and localization of the transcripts within the tissue slide. Spatial whole transcriptome RNA sequencing has been applied to generate a spatial atlas of human cortical organoid development 71 and to characterize the influence of loss-of-function mutations in disease-associated genes such as phosphatase and tensin homolog (PTEN) on cell-type specific developmental events in brain organoid models. 50 Moreover, spatial transcriptomics data from consecutive sections have been integrated to reconstruct a 3D molecular cartography of human brain organoids. 41 These studies and recent reviews underline the potential of combining single cell and spatial transcriptomics data to locate different cell types in organoids and to assess their molecular states.35,62 Pathologists can support tissue preparation and guide the association of transcriptomic and morphological features, which will support validation and efficient application of these complex methods for CIVM characterization and toxicology readouts.
Case Examples of Pathology-Biologist-Bioengineer Collaborations
Imaging and artificial intelligence of an upper airway model
Artificial intelligence (AI)-based methods can be applied to two-dimensional (2D) histology slides of CIVM to facilitate the analysis of complicated data sets. In addition, AI-based machine-learning methods provide opportunities for applying quantitative and automated approaches to computer vision (i.e. the field AI-enabling computers to derive information from images and videos). 70 Recent and rapid advancements in machine-learning methodology include deep learning AI which uses convolutional neural networks (CNNs) to build classifiers that can detect and quantify tissue and cell endpoints (For a review, see Li et al 39 ). Most computer vision deep learning classifiers are built under the supervision of pathologists who participate in the training of the computer and the verification and qualification of the computer output. 78
Once a qualified model is developed, the pathologist has a tool that improves their speed and consistency by limiting observational biases and helping to quantify and integrate diverse and large data sets.39,70 These methods can be used by pathologists to replace qualitative scoring methods using ordinal scales (i.e. 1 [minimal] to 5 (severe)) or semiquantitative feature-based scoring systems, the latter of which are complicated and at risk for inconsistency in their application between studies, laboratories, and pathologists. 31 By using deep learning CNN classifiers more quantitative and automated measurement systems are possible that improve consistency between studies by eliminating bias and diagnostic drift. These tools will also support the scaling of pathology measurements for CIVM by eliminating laborious and repetitive manual functions that are performed by pathologists and researchers.39,70 Finally, AI can be used to integrate the computer vision data with other CIVM endpoints to produce multimodal methods for CIVM analysis.43,52
Examples of the application of deep-learning-based computer vision solutions in CIVM evaluation may include everything from the assessment of time-lapse images and video to quantification of circulating biomarkers or cells to morphological changes from histological preparations. 39 The pathologist can provide recommendations for adapting their in vivo tool kit for use with deep-learning-based AI solutions for CIVM. A recent application of deep learning AI was used as a proof-of-principle to simplify a feature-based, manual histologic scoring system which measured several key alterations in a Transwell® upper airway 3D model system (personal communication). 74 Included in the feature set for manual evaluation of the CIVM histological sections by the pathologist were degeneration/necrosis, vacuolation, epithelial detachment, and erosion/ulceration (Figure 5). The HE-stained sections were digitized, and a deep learning CNN classifier was built to differentiate these features automatically for the pathologist. After qualification of the CNN classifier, pixel areas of each lesion class could be quantified to produce an automated “area score” for the upper airway CIVM (versus the manual ordinal score provided by the pathologist). Other algorithms could be layered on top of this type of analysis to enable readouts such as number of necrotic cells, depth of erosion/ulcer, or other endpoints deemed important for the intended use of the model.
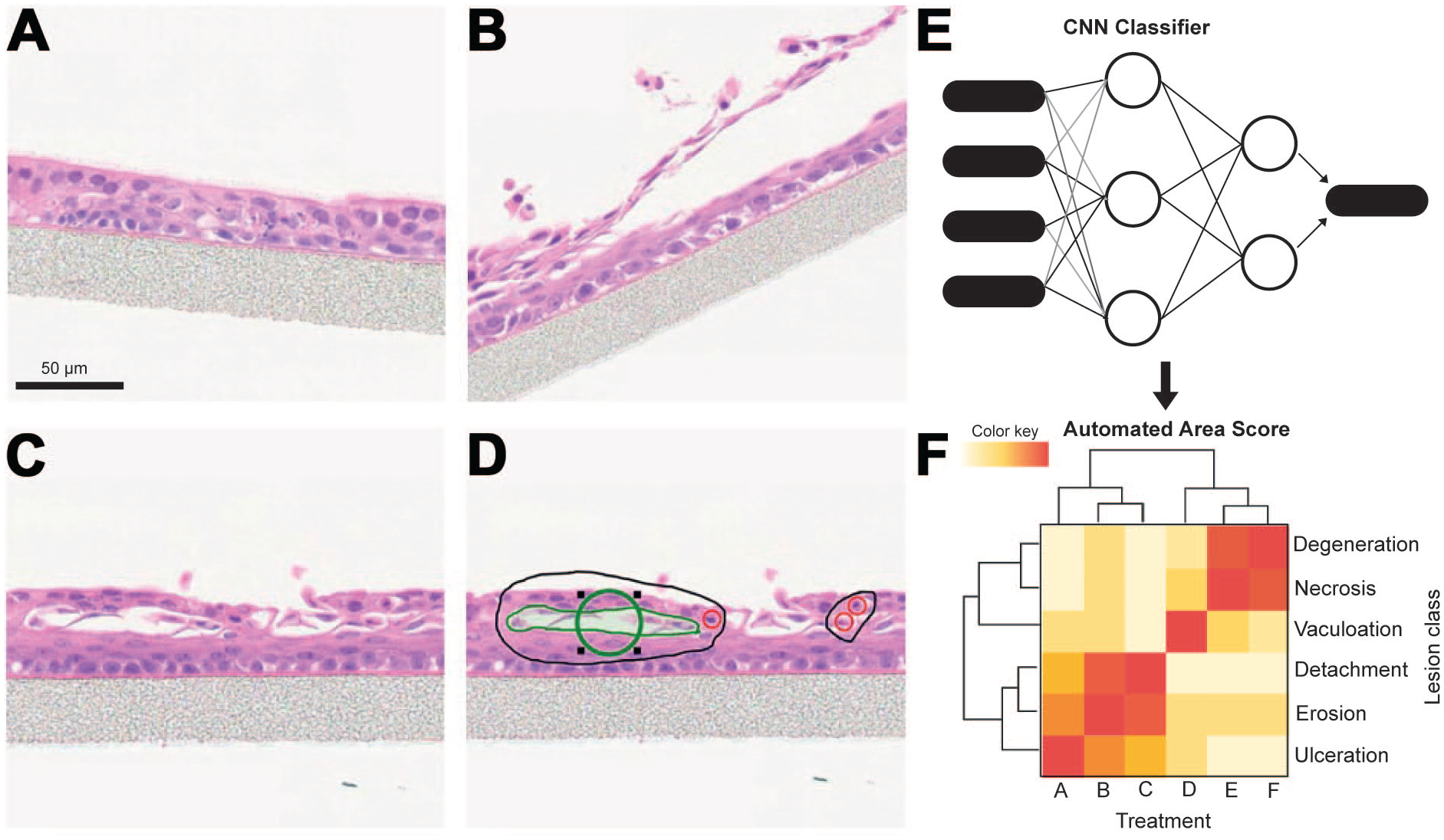
Algorithm-assisted histologic scoring measuring key pathologies in a Transwell® upper airway 3D model system. (A)–(D) manual annotation and classification of representative images. Included in the feature set for manual evaluation of the complex in vitro models (CIVM) histological sections by the pathologist were degeneration/apoptosis, pseudocystic degeneration, valuolation, squamous differentiation, and epithelial thinning. (E) The digitized HE images and a deep learning CNN classifier was built to differentiate these features automatically for the pathologist. (F) After qualification of the convolutional neuronal network (CNN) classifier, pixel areas of each lesion class could be quantified to produce an automated area score for the upper airway CIVM. This figure has been partially created with biorender.com. Scale bar = 50 µm.
Morphology-based characterization of a Liver-Thyroid model
During the development of new 3D models, morphological assessment is key for the confirmation of structures consistent with the in vivo orientation that are often critical in determining the metabolic functionality of a tissue. Provided here is a case example of the development of a microfluidic thyroid-liver system to assess thyroid perturbations relevant to humans (Figure 6). While rodents are a standard species used in the nonclinical evaluation of the toxicity of chemicals, they are also very sensitive to perturbations of thyroid homeostasis. Long-term perturbations in rodents can induce thyroid neoplasia, which may overestimate the human safety risk. 16 Both rodent and human 3D thyroid and liver models have been created to understand species translation of thyroid toxicities.28,34
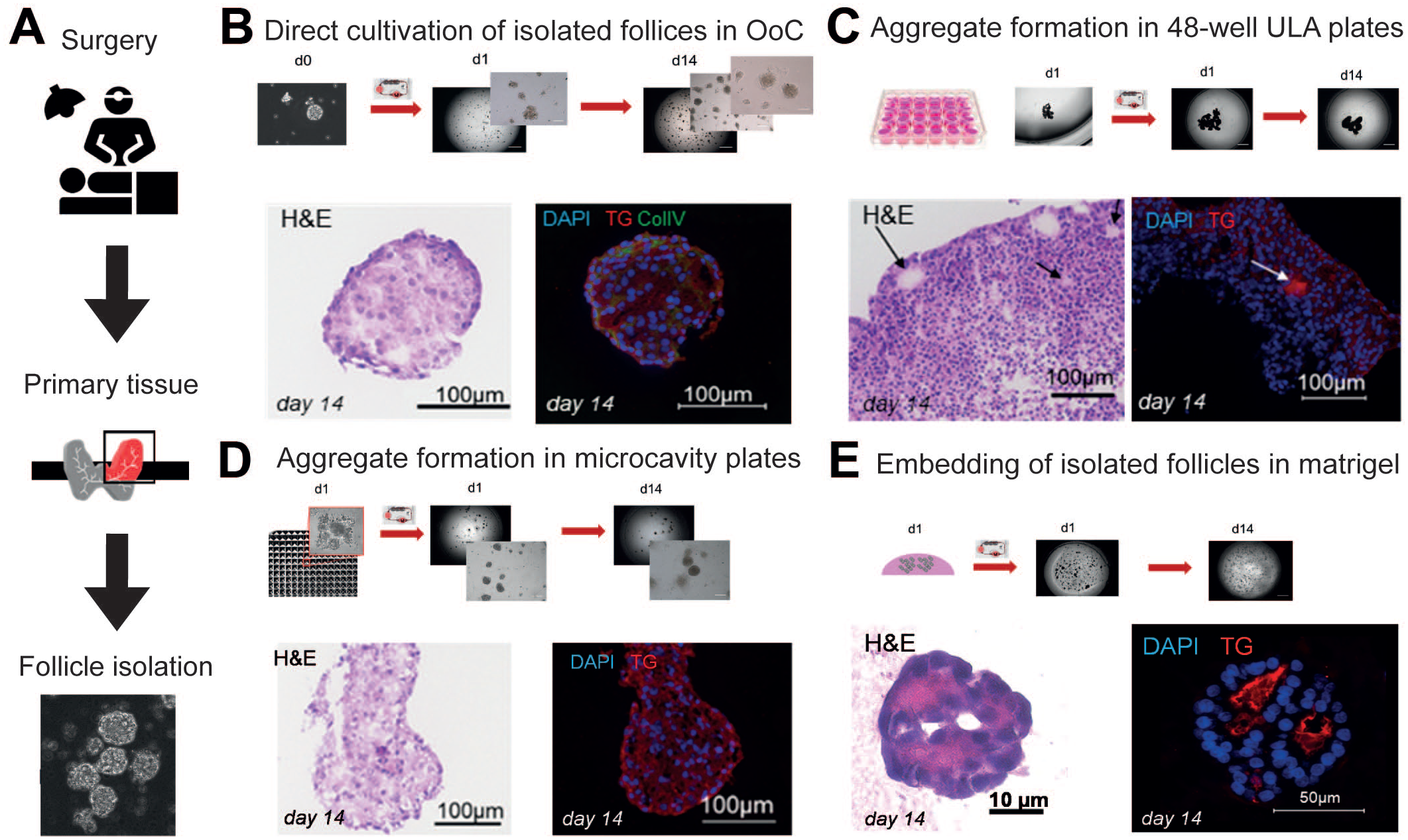
Morphological characterization of four human 3D in vitro thyroid models after a 14-day cultivation in the HUMIMIC Chip2 (TissUse GmbH) to identify conditions that preserve the follicular in vivo tissue structure. (A) Human thyroid tissue leftovers from surgeries were used as a source for enzymatically isolating thyroid follicles. Extracted follicles served as a starting material for the subsequent model evaluations. (B) Thyroid follicles were directly cultivated on the glass bottom in the HUMIMIC Chip2. (C) Thyroid-follicle aggregates were formed in 48-well ultra-low attachment (ULA) plates for 24 hours. (D) Extracted follicles were pre-aggregated for 24 hours in KUGELMEIERS Sphericalplate 5D containing 750 microcavities per well. (E) Thyroid follicles were embedded in growth factor-reduced matrigel. Brightfield images of the models prior to chip transfer and within the culture compartment of the HUMIMIC Chip2 at culture day 1 and 14 can be seen. For each model, end-point morphology was assessed by HE staining and IF staining of 8-µm cryosections. IF merges of nuclei (DAPI, blue) and thyroglobulin (TG, red) are depicted. Collagen IV (green) additionally was stained in condition (B).
The thyroid can be directly affected by chemicals inhibiting the production of thyroid hormones. Alternatively, xenobiotic induction of phase 2 enzymes in the liver can result in increased clearance of thyroid hormones via the formation of inactive glucuronidated and sulfated forms, which can be significantly different in rats and humans. 2 The resulting decrease in circulating thyroid hormones reduces the feedback to the pituitary gland, which results in increased production of thyroid-stimulating hormone (TSH). Elevated TSH drives overstimulation of thyroid activity that can lead to tumor formation.2,49,57
Successful in vitro thyroid hormone metabolism requires functional thyroid and liver organ models with specific morphological characteristics understood by studying in vivo physiology and anatomy. The thyroid’s smallest functional subunits are thyroid follicles, which synthesize thyroid hormone within a central lumen. 19 By reproducing this architecture, thyroid hormone synthesis can be expected in vitro. Human thyroid follicles were enzymatically isolated from fresh primary tissue (Figure 6A) and their maintenance tested using different approaches (Figure 6B–E). Neither a direct cultivation of the isolated follicles in the HUMIMIC Chip2 (2-organ chip device, manufactured by TissUse) (Figure 6B) nor the chip-based culture of follicle aggregates (Figure 6C, D) maintained the follicular cell organization as shown by sections stained for HE and fluorescently labeled thyroglobulin (TG), the precursor molecule of the thyroid hormones. These models did not fulfill the goal to produce thyroid hormones. Embedding the isolated follicles into Matrigel allowed the follicular structure to be maintained, and this was validated by sectioning and staining the 14-day chip-cultured thyroid microtissues (Figure 6E). Confocal microscopy was used to visualize the 3D structures in a more representative manner as more than one z-lane can be simultaneously analyzed by this technique and generally can be recommended for a fast morphological structure analysis of 3D organ models. 34 In addition, morphological differences in the structures were demonstrated between human thyroid follicles treated with and without TSH by confocal analysis indicating different states of follicular activity. 34 In accordance with the morphological findings, the established rat and human thyroid follicle models produced thyroid hormones in a TSH-dependent manner.28,34 Thus, successful thyroid hormone secretion could ultimately be used as a functional metabolic marker for the detection of direct thyroid hormone perturbation. This was tested with 4-day methimazole treatment which decreased the T3 and T4 secretion in the human and rat models, respectively.28,34
Reproduction of thyroid hormone metabolism in vitro requires characteristic morphological features in the liver model. Liver spheroids were derived either from fresh primary hepatocytes from Ham Wistar rats or the HepaRG cell line for the human model. The latter is currently replaced by cryopreserved human hepatocytes due to increased relevance (unpublished data by Bayer). The protein expression of the thyroid hormone transporter MCT8 as well as a bile canaliculi network, visualized by immunolabeling the MRP2 transporter and ZO1 required for the thyroid hormone catabolite efflux, was shown in the liver spheroid models. 34 In line with these morphological characteristics, active thyroid hormone metabolism was demonstrated in both species.28,34 The human liver model was able to produce the sulfated and glucuronidated thyroid hormone metabolites from the addition of T4 to the system, whereas the rat liver model mainly demonstrated the formation of the glucuronidated form. These thyroid hormone metabolites were used as functional parameters to evaluate indirect hepatocyte-mediated thyroid toxicities. The rodent-derived liver spheroids responded with increased glucuronidated metabolite formation when treated with pregnenolone-alpha-carbonitrile (10 uM), whereas the human model was not responsive to this rat-specific nuclear receptor activator.28,34
Once the models were characterized functionally and for toxicity, the co-culture of the thyroid and liver organ models was evaluated over 21 days in the HUMIMIC Chip2 to demonstrate the maintenance of morphology and function. In summary, morphology and function must be confirmed to recapitulate key in vivo endpoints to interrogate the model for the questions being asked. This work demonstrates the importance of joint efforts by pathologists and bioengineers.
Assessing developmental neurotoxicity with an AI-enabled morphology-based readout of neural organoid morphogenesis
While there are myriad methods for assaying CIVM, imaging combined with AI-based analysis using CNNs is perhaps the most scalable, cost-effective, and versatile pipeline for collecting information on 3D microscale tissue function. Either in real-time or post IHC/IF treatments (as reviewed above), CIVM can be imaged using scanning optical coherence tomography, brightfield, confocal, 2-photon, or light-sheet microscopy to assess pathological endpoints of cell morphology, viability, metabolism, proliferation, and migration as well as the microscale tissue’s physiology and structure. Moreover, when CIVM are based on human pluripotent stem cell (hPSC)-derived organoid cultures, such analysis can also provide information on tissue morphogenesis or organ development and function. As stated previously, the challenge with this approach is designing the CIVM and identifying the correct pathological endpoints for extraction with AI-based image analysis to correlate the model’s readouts with clinical human pathology. Thus, it is critical to develop the CIVM and their analysis pipeline via a collaboration between clinicians, pathologists, bioengineers, and computer vision experts.
Such an approach was used to develop the RosetteArray® platform (Neurosetta LLC) for detecting developmental neurotoxicity (DNT) and modeling neural tube defect and autism spectrum disorder risk (Figure 7). As published by the Ashton lab and others, micropatterned culture substrates can be used to regulate the morphology of hPSC aggregates and induce reproducible neural rosette formation, the inceptive stage of neural organoid morphogenesis, during neural differentiation 32 (Figure 7A). The rosette structure models a slice of the developing human neural tube, which is the anlage of all brain and spinal cord tissues. Disruption to rosette formation has been shown to be a potent indicator of DNT and useful for detecting a chemical or genetic mutation’s risk for causing congenital neural tube defects 60 (eg, Spina bifida), Autism Spectrum Disorder, 6 and even Huntington’s Disease. 22 In a proof-of-concept publication, the Ashton group demonstrated how micropatterned hPSC-derived neural rosette morphogenesis could be standardized and combined with computer vision algorithms to detect various facets of rosette morphogenesis from confocal Z-stack images. Now at Neurosetta (www.neurosetta.com), this technology has been used to develop the RosetteArray® screening platform (Figure 7B). It combines the micropatterned neural rosette formation assay with rapid scanning confocal imaging and AI-based image analysis to rapidly produce dose-response curves of neuroepithelial cell number, neural differentiation, and AI-detected rosette emergence. These endpoints are sensitive to perturbations in cell viability and proliferation, epigenetic pathways that regulate neural differentiation, and planar cell polarity, migration, and other cytoskeletal-relevant pathways that govern rosette morphogenesis, respectively. Thus, the RosetteArray® platform’s combination of bioengineered, microarrayed neural organoid culture, rapid scanning confocal imaging, and AI-based image analysis using cloud computing enables scalable, cost-effective use of the neural rosette formation assay for DNT screening as well as neurodevelopmental and neurodegenerative disease modeling.
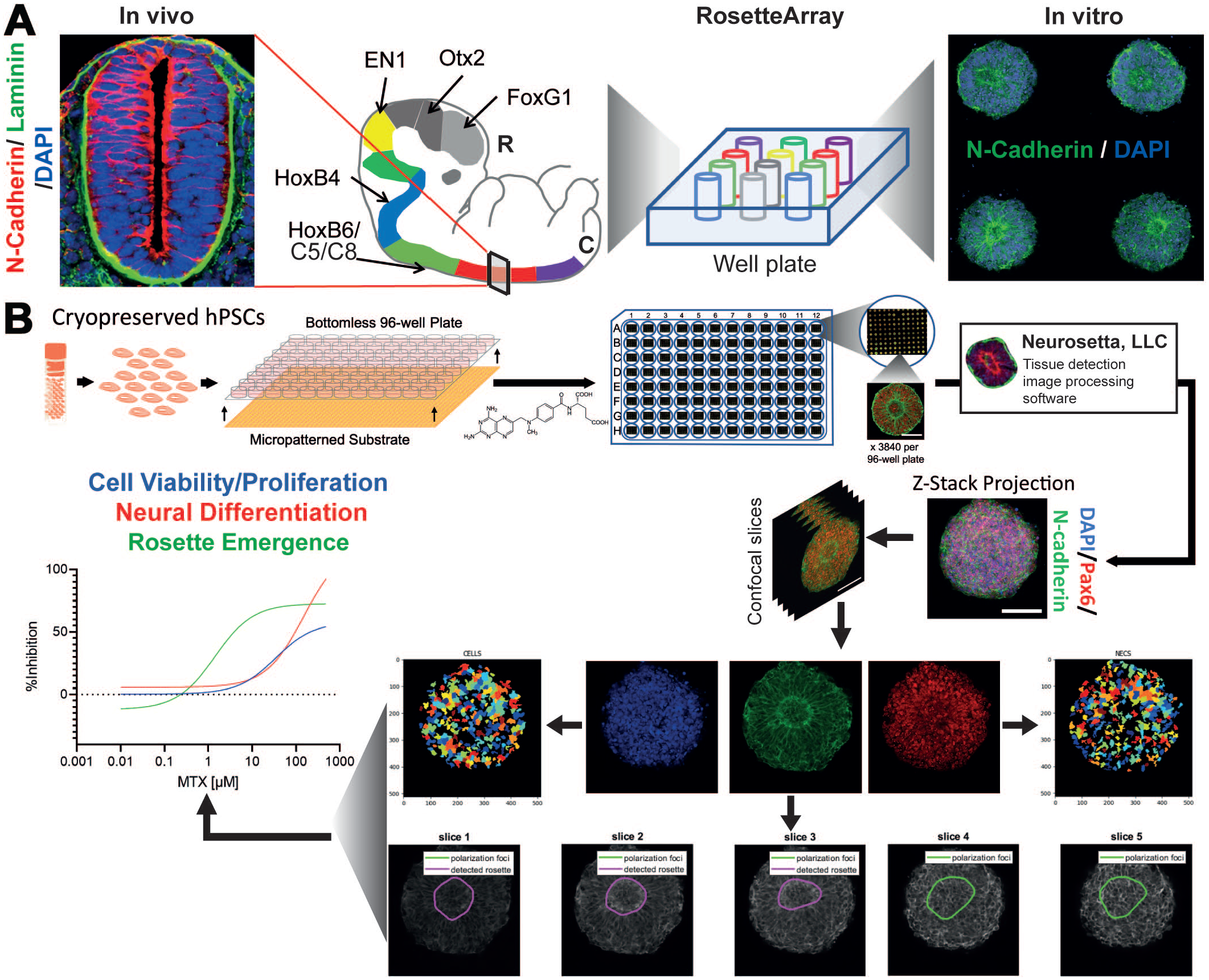
RosetteArray® Platform. (A) Comparison between in vivo slice of the developing neural tube and micropatterned neural rosette structure. (B) Schematic of screening pipeline that entails seeding of cryopreserved human pluripotent stem cells (hPSCs) into custom micropatterned well plates to generate RosetteArrays after 5-8 days of culture in the presence or absence of chemicals. Within each well hundreds of 3D micropatterned rosette tissues are generated and imaged using laser scanning confocal microscopy. Z-stack images are processed slice-by-slice to quantify the number of DAPI+ cells and Pax6+ neuroepithelial cells, and a custom AI algorithm is used to identify N-cadherin+ rosette structures (magenta traces) within imaged tissues. Then, these data are used to create dose-response curves to detect chemical (eg, methotrexate, MTX) developmental neurotoxicity or risk associated with genetic mutations in gene edited or patient-derived hPSC lines.
Conclusion and Outlook
Complex in vitro model offer the opportunity to evaluate biological responses, both mechanistically and quantitatively, in situations where the complexity and variability of the response in animal models precludes such precision. Such responses can be problematic in an evaluative science like safety assessment which leans heavily on historical experiences to determine the significance of a morphological or biochemical change. Dissecting through complexity and variability is necessary to translate those responses to potential health consequences for human patients. As demonstrated with several case reports and in-depth discussion of techniques, CIVM are becoming more mature in terms of their capabilities and potential as alternative methods to animal-based drug development tools and assays. 40 The limitations of these models are becoming better understood and addressed on a broader scale. The efforts to focus on platforms and methodologies that can be reproduced consistently, with minimal heterogeneity, will further build confidence in the use of these new platforms. This will enable collaborations between vendors, researchers and pharmaceutical companies that are required to implement new testing strategies in the industry. The collaboration between ESTP and STP through the working group and newly formed Special Interest Group (SIG) “Complex In Vitro Models & Pathology” are working to increase awareness of pathologists, investigative scientists and device engineers on the integrated role that they all play to optimize the characterization/qualification of new CIVM. Ideally, together, they drive the application and use cases of those CIVM in future benefit-risk assessments in the pharmaceutical and chemical industry.
Benchmarking through these efforts that pathologists can assist in (1) defining the specific COU, (2) defining biological content, architecture, and function (characterization), (3) benchmarking to tissues of origin and defining limitations for CIVM and their specific COU, (4) defining and establishing biologically relevant endpoints, and (5) extrapolating outcomes to the human in vivo context. The willingness of scientists and engineers to engage with pathologists is already high and needs to be met with interest and confidence among pathologists to contribute to these new platforms and ultimately to reduce animal use and follow the 3R concept. This is already evolving through open dialogue, and it is essential that this gap continue to be filled to ultimately shift the traditional evaluation of the safety of pharmaceuticals and chemicals into a new era with more streamlined approaches.
As CIVM continue to evolve, pathologists will be able to help bridge the gap between the traditional approaches that require higher animal usage and new approaches that can decrease animal use while maintaining confidence in the overall risk: benefit of the pharmaceutical or chemical being evaluated. A combination of animal-derived and human-derived models will provide important additions to our modeling platform portfolio as we further resolve questions around translation between nonclinical species and people.
The future of our ability to incorporate CIVM into standard testing strategies is bright, and there are new promising COUs being developed. The three case examples provided in this manuscript exemplify the ability of CIVM to represent pertinent aspects of human physiology and development when the system focuses on the clinically relevant question that can be asked with the specific model that has been developed. Understandably, the goal will never be to recapitulate human anatomy and physiology in its entirety, thus it is crucial to fully understand and acknowledge the limitations of each CIVM individually. In addition, the techniques summarized in this review continue to evolve, and these capabilities are only enhanced with the use of imaging and AI-assisted methods of evaluation. As long as these advancements are made with detailed understanding of the necessary cell-cell interactions and physiology, the field will continue to progress.
Footnotes
Acknowledgements
The authors would like to thank Christelle Zundel (Roche), Rachel Neff (Roche), Thomas Maurissen (Roche), Marius Harter (Roche), Timothy Recaldin (Roche), Michael Bscheider (Roche), Nikolche Gjorevski (Roche), and the SciLifeLab for their contribution to Figures 3 and 4; James Baily (CRL) and Mary McElroy (CRL) for contribution to Figures 3 and ![]() . They also thank the ESTP (Pathology 2.0) and STP (SIG) for their support and providing the frame of the working group.
. They also thank the ESTP (Pathology 2.0) and STP (SIG) for their support and providing the frame of the working group.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: RSA is the CEO of Neurosetta LLC and holds equity in the company.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
