Abstract
Historically, pathologists perform manual evaluation of H&E- or immunohistochemically-stained slides, which can be subjective, inconsistent, and, at best, semiquantitative. As the complexity of staining and demand for increased precision of manual evaluation increase, the pathologist’s assessment will include automated analyses (i.e., “digital pathology”) to increase the accuracy, efficiency, and speed of diagnosis and hypothesis testing and as an important biomedical research and diagnostic tool. This commentary introduces the many roles for pathologists in designing and conducting high-throughput digital image analysis. Pathology review is central to the entire course of a digital pathology study, including experimental design, sample quality verification, specimen annotation, analytical algorithm development, and report preparation. The pathologist performs these roles by reviewing work undertaken by technicians and scientists with training and expertise in image analysis instruments and software. These roles require regular, face-to-face interactions between team members and the lead pathologist. Traditional pathology training is suitable preparation for entry-level participation on image analysis teams. The future of pathology is very exciting, with the expanding utilization of digital image analysis set to expand pathology roles in research and drug development with increasing and new career opportunities for pathologists.
Keywords
The use of morphometric tools to supplement traditional histologic analysis has been an aid to pathologists and researchers for decades (Park et al. 2013). Two-dimensional (2-D) morphometrics (cell profile counts, linear dimension measurements, etc.) enhance the qualitative diagnosis provided by a pathologist by adding both quantitative and objective support to the observations. Key advantages of employing these form-measurement approaches in biomedical research practice are that they generate quantitative data that can be easily stored in databases, the numeric data can be analyzed and compared to data collected by other tools, and that many of the morphometric methods may be generated by proficient technical staff. The 21st-century descendant of 2-D morphometrics is whole slide imaging and digital image analysis (often considered together as “digital pathology”), which greatly increase the amount of information that can be obtained from histology sections (Isse et al. 2012; Al-Janabi et al. 2012; Bauer and Slaw 2014; Rojo et al. 2006). More importantly, digital pathology allows a more consistent (i.e., objective) and quantitative evaluation than is possible by manual observation utilizing the human eye alone (Webster and Dunstan 2014; Diller and Kellar 2015; Meijer et al. 1997).
Until recently, the potential of digital pathology has been underutilized by many scientific disciplines, including pathology (Isse et al. 2012). In 2013, the College of American Pathologists’ Pathology and Laboratory Quality Center established guidelines on the validation of whole slide imaging for diagnostic purposes (Pantanowitz et al. 2013). In the same year, guidelines for validation of digital pathology systems in Good Laboratory Practice (GLP) and non-GLP studies were also published (Long et al. 2013). Just in April 2016, the Food and Drug Administration (FDA) published guidance on the use of this new technology in drug development.
Despite this growing support for increased use of digital imaging methodology, many pathologists remain reluctant to embrace the trend toward digital analysis of tissue sections for three main reasons. First, individuals believe that their livelihood will be lost either to an automated instrument and specialized analytical software or to a pathologist located in another region of the globe (i.e., “off-shoring,” as has happened in the radiology field). Second, pathologists who lack specialized computer science knowledge, and have no time (or interest) to acquire this knowledge, believe that their lack of understanding will expose their failings. Third, many pathologists still lack familiarity in using digitized slides and databases. While these assumptions are understandable, history indicates that they are not valid. Prior technical innovations (e.g., the electrocardiogram, magnetic resonance imaging) did not replace the trained biomedical scientists working in those fields (cardiologists and radiologists, respectively), although practitioners did have to adapt and take on new roles. Similarly, tissue image analysis (tIA) will not replace pathologists, as this strategy necessitates considerable oversight from an experienced morphologist to ensure that its application is efficient and effective. The current commentary describes the roles and opportunities for pathologists as invaluable members of a tissue image analysis team.
The Image Analysis Team
High-throughput digital pathology is performed most efficiently using a team approach, in which each member possesses a different expertise. The size of an image analysis team can vary greatly, as long as the following areas of expertise are represented: systems/algorithm engineering to create the analytical algorithms, biological expertise of the disease process (e.g., cancer biology) to ensure that application of the algorithm accords with current biological understanding, biostatistics expertise to analyze data sets, and pathology expertise (Grunkin, Raundahl, and Foged 2011; Webster and Dunstan 2014). Additional skilled and specially trained technicians can assist in preanalytical tasks such as scanning, image quality control, and manual image annotation. Although several kinds of expertise can be contributed by one person, it is unlikely given the complex scientific questions and increasingly sophisticated imaging hardware and software that all the needs of a tissue analysis team are met by a single person with enough expertise in all of these areas.
While all members of an image analysis team are of importance in developing successful algorithm-based solutions, a pathologist actually serves as the keystone for the entire team’s efforts. Pathology review/input should be central to all major steps of the workflow, beginning with study design and continuing through confirming sample suitability (e.g., disease diagnosis and tissue quality), assessing sample staining (e.g., sensitivity and specificity), guiding image preparation (section annotation and algorithm approval), and preparing reports (data interpretation and photomicrographic selection; Figure 1). These roles are comparable to those already performed on a regular basis by pathologists in both diagnostic laboratories and industrial research and development settings. Importantly, fulfillment of these roles requires regular, face-to-face interactions between the pathologist and other team members. As such, digital pathology offers new tools for pathologists.
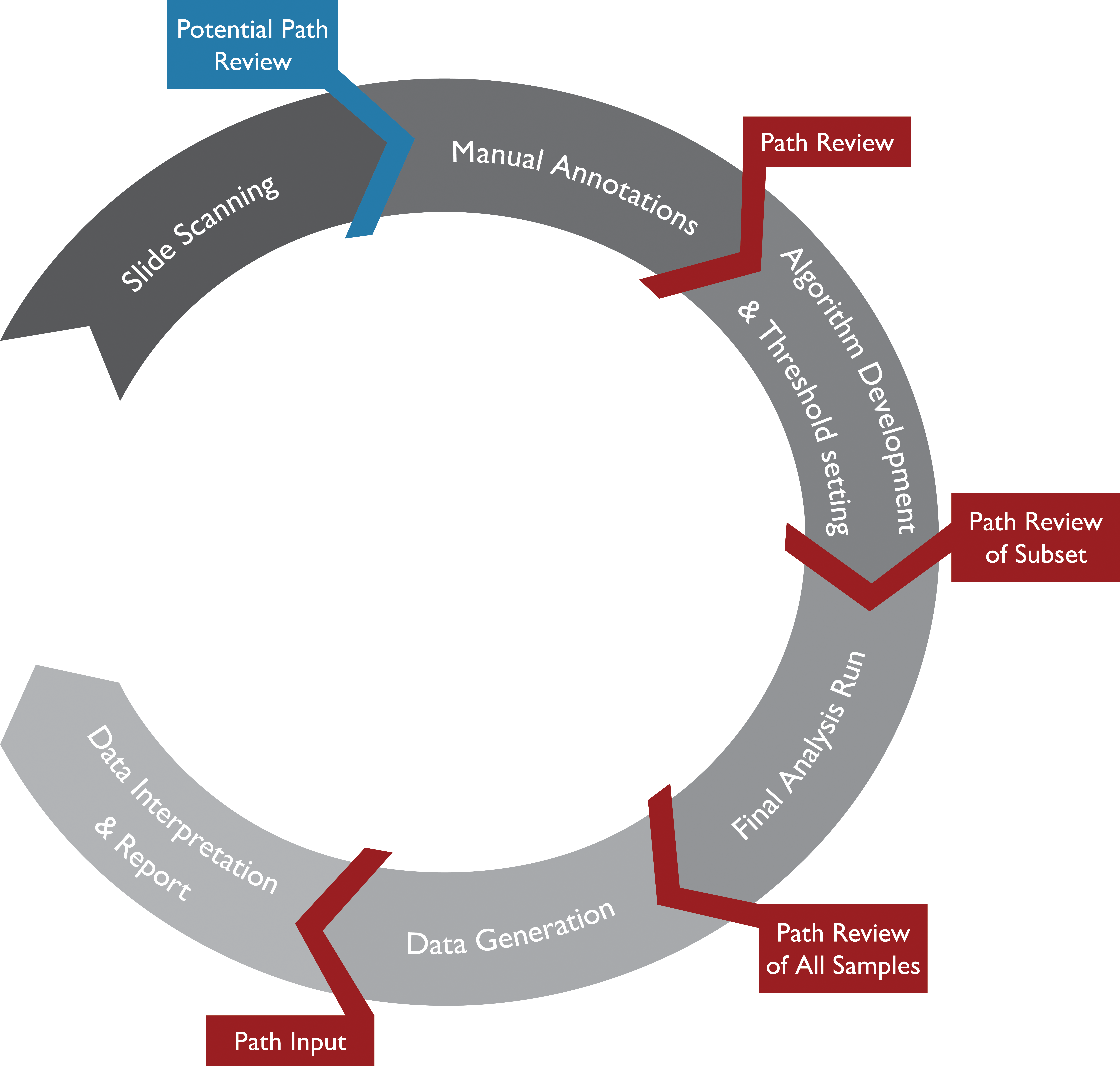
Workflow of a digital image analysis project with potential (blue) and key (red) pathology review steps as outlined in the text of this article.
Digital Image Analysis—Basic Principles
Even if application of image analysis software is delegated to other team members, the pathologist must be aware of the basic principles that underpin digital image analysis in order to interact effectively with other team members. The majority of solutions can be classified into the following categories or combinations thereof:
Area-based measurements:
Pixel-based assessment: the algorithm quantifies the color (or intensity) of staining in each pixel. Public domain turnkey solutions (e.g., ImageJ, http://rsbweb.nih.gov/ij), as well as commercially available solutions and services are accessible (Isse et al. 2012). Cell-based measurements:
Morphometry-based assessment: pixels are grouped based on similarity to defined structure profiles (e.g., cells or nuclei) that meet certain preselected criteria (e.g., size and shape). Complex algorithm-based solutions: combined (generally automated) assessment of individual pixels and tissue elements (often cells)
to identify specific structures (e.g., follicles, glomeruli); to define biologically separate tissue compartments (e.g., tumor vs. tumor microenvironment [stroma]; Figure 2); or to interrogate certain tissue regions (e.g., tumor–stroma interface). Open-source solutions (e.g., FARSIGHT, www.farsight-toolkit.org), as well as commercial algorithms and service providers are available that permit both turnkey use and deployment of customized algorithms. Object-based counting or assessment of “events”:
Specialized (individualized) algorithms: solutions to serve a particular need (often the automated identification and/or enumeration of noncell structures). Common applications range from counting chromogenic in situ hybridization (CISH) dots per cells, to assessing the size and distribution of intracellular vacuoles, to identifying and quantifying blood vessels or myofibers of various sizes. Freeware solutions (e.g., ILASTIK, www.ilastik.org) as well as commercial algorithms and service providers exist that allow both turnkey use and customized applications.
Comparing and contrasting many of the current commercially available software packages has been done elsewhere and is beyond the scope of this publication (Rojo, Bueno, and Slodkowska 2009; Mulrane et al. 2008).
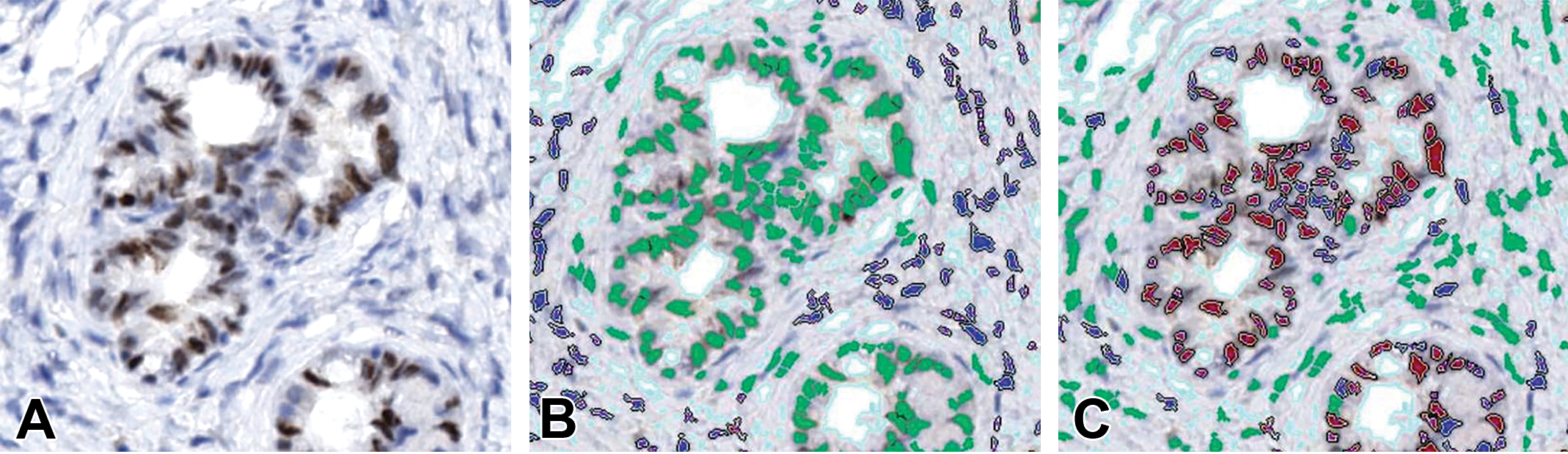
Pancreas with pancreatic intraepithelial neoplasm (PanIN), Ki-67 immunohistochemical staining, demonstrating effective use of tissue image analysis to discriminate between a target cell population and other tissue elements. (A) Histology image of the lesion prior to image analysis, showing labeled tumor nuclei (brown) on a hematoxylin counterstained field. While it is apparent that there are structures representing PanIN foci, many of the nuclei in both the tumor epithelium and adjacent stroma are morphologically similar. It is the relationship of nuclei to each other that aids the algorithm to separate stroma (B, shown in blue markup) from the epithelial compartment (C, shown in red and blue markup, where red = Ki-67-positive tumor cell nucleus and blue = Ki-67-negative tumor cell nucleus).
Independent of the complexity of an algorithm, at the very beginning of each solution, human beings must create digital analysis solutions and oversee their application, including partially or completely subjective decisions at a subset of steps throughout the process of algorithm design (Tadrous 2010). The pathologist’s expertise is instrumental during this process because our subjective or semiquantitative assessments often are considered the “gold standard” upon which automated digital pathology measurements will be based (Diller and Kellar 2015). One example is the scoring of tumors based on immunohistochemical (IHC) labeling for a particular biomarker, in which algorithms are developed and validated by comparing their performance to manual pathology scores. Failed algorithms are modified and retested until they recapitulate the manually acquired data. Once concordance has been achieved between an automated algorithm and a manual scoring scheme, quantitative computer-based assessments are expected to be more accurate, consistent, and less biased (e.g., when counting structures or applying the same thresholds across a large sample cohort) than semiquantitative scores made by a human being, no matter how well trained or experienced (Webster and Dunstan 2014; Diller and Kellar 2015; Wang et al. 2013).
The Pathologist’s Roles Within an Image Analysis Team
Even without detailed knowledge of algorithm development and how to refine software so that failed algorithms will meet expectations, a pathologist is crucial to the success of digital image analysis projects because meaningful analysis algorithms cannot be devised without their specialty expertise. Necessary technical knowledge includes, but is not limited to, understanding the impact of tissue handling, fixation methods, tissue processing, and histological staining procedures (e.g., CISH, IHC) on sample quality and molecular integrity. Of even greater importance, pathology training imparts broad conceptual knowledge of biological systems, including biomarker expression as well as comparative anatomy, histology, and pathology; these are prerequisites for rational evaluation of diseases and their pathogeneses. This deep comprehension of fundamental pathophysiological principles makes the pathologist uniquely suited to undertake the following critical roles within the digital image analysis workflow (Figure 1).
Slide, Staining, and Scanning Quality Review
Before any image analysis should be attempted, a pathologist must evaluate whether or not the specimens are of sufficient quality (e.g., not necrotic, well preserved) and the target tissue is present in suitable quantity. Image analysis can be hampered by low slide quality and a lack of consistency in slide preparation, and results can be greatly affected by preanalytical variables (Webster and Dunstan 2014). For example, in almost all human clinical trials with tissue analysis as an end point, the interval between tissue harvest and fixation, the fixation time, and staining quality are poorly standardized across institutions (Dunstan et al. 2011; Potts, Young, and Voelker 2010). In addition, competency testing of histology technicians and pathologists can vary greatly among different laboratories and hospitals that participate in any given clinical trial. To demonstrate quality, institutions need to provide regular training opportunities for laboratory staff and document that individuals have completed the institutional learning objectives needed to produce and analyze tissue specimens.
Perhaps the most important deficiency in this regard is staining quality, as the majority of image analysis studies focus on assessing a specific biomarker and its distribution within a target tissue. Therefore, a well-optimized staining protocol is essential not only to address the mere presence or absence of a biomarker but also to answer more detailed and sophisticated questions such as the intracellular compartmentalization, degree of expression, and sometimes colocalization with another biomarker (Dunstan et al. 2011). Keystones for creating and sustaining well-optimized staining protocols that produce consistent staining results are the use of automated histostainers and the regular use of within-run controls, especially inclusion with each staining run of a positive control slide cut from the same tissue block to compare for consistency in labeling between staining runs (Dunstan et al. 2011). A pathologist is best suited to assess that staining occurs in the expected cellular compartment of the target tissue and is absent in the nontarget tissues also present on the slide. Furthermore, it is of the utmost importance that pathologists verify that the staining intensity covers a dynamic range (i.e., that different intensity levels of staining are present within each slide and across all slides of a study cohort), which is required to permit differential assessment of staining intensity between target and nontarget cells. Image analysis is meaningless if slides are so overstained that all stained cells fall within the highest intensity group. Among other reasons, this is because overpowering chromogenic localization that obscures nuclear hematoxylin staining prevents the detection of nuclei (and thus cells), which often is a crucial first step in generating analytical algorithms.
Therefore, pathologist review of all slides prior to algorithm development is a critical first quality control step to ensure that the material meets a quality standard that will allow collection of meaningful and reproducible data (Shakeri et al. 2015; Ameisen et al. 2014). This slide review is ideally done after digitization of the study material so that scanning quality can be assessed within the same review. Scanning artifacts that can affect image analysis results include improper cleaning of slides prior to scanning (i.e., the presence of dust particles that can be counted as cells or obscure cells), poorly focused scans (i.e., improper selection of focal points), and compensation lines (i.e., artifacts at the edges where scanned lines are stitched together to form a whole slide image). It goes without saying that the presence of the target tissue in suitable quantity needs to be confirmed as well.
Annotation Strategy and Review
While manual image annotation (also known as “masking”) is not always crucial for image analysis, it should be employed whenever possible. Manual annotations entail drawing digital inclusion lines around tissue areas intended for 2-D analysis (i.e., defining the region of interest [ROI]; Figure 3). For each slide, an algorithm will only analyze areas within the “inclusion” annotation. Within these inclusions, smaller areas can be omitted by “exclusion” annotations. Excluded areas usually entail histological artifacts such as tissue folds, staining artifacts, scanning artifacts such as small focal blurs, artifacts present within the tissue that can hamper meaningful analysis (e.g., anthracosis, surgical dye), or areas unsuitable for analysis (e.g., large vessels, necrosis). Annotations need to be performed by well-trained technicians to ensure that minimal corrections need to be implemented by the reviewing pathologist.
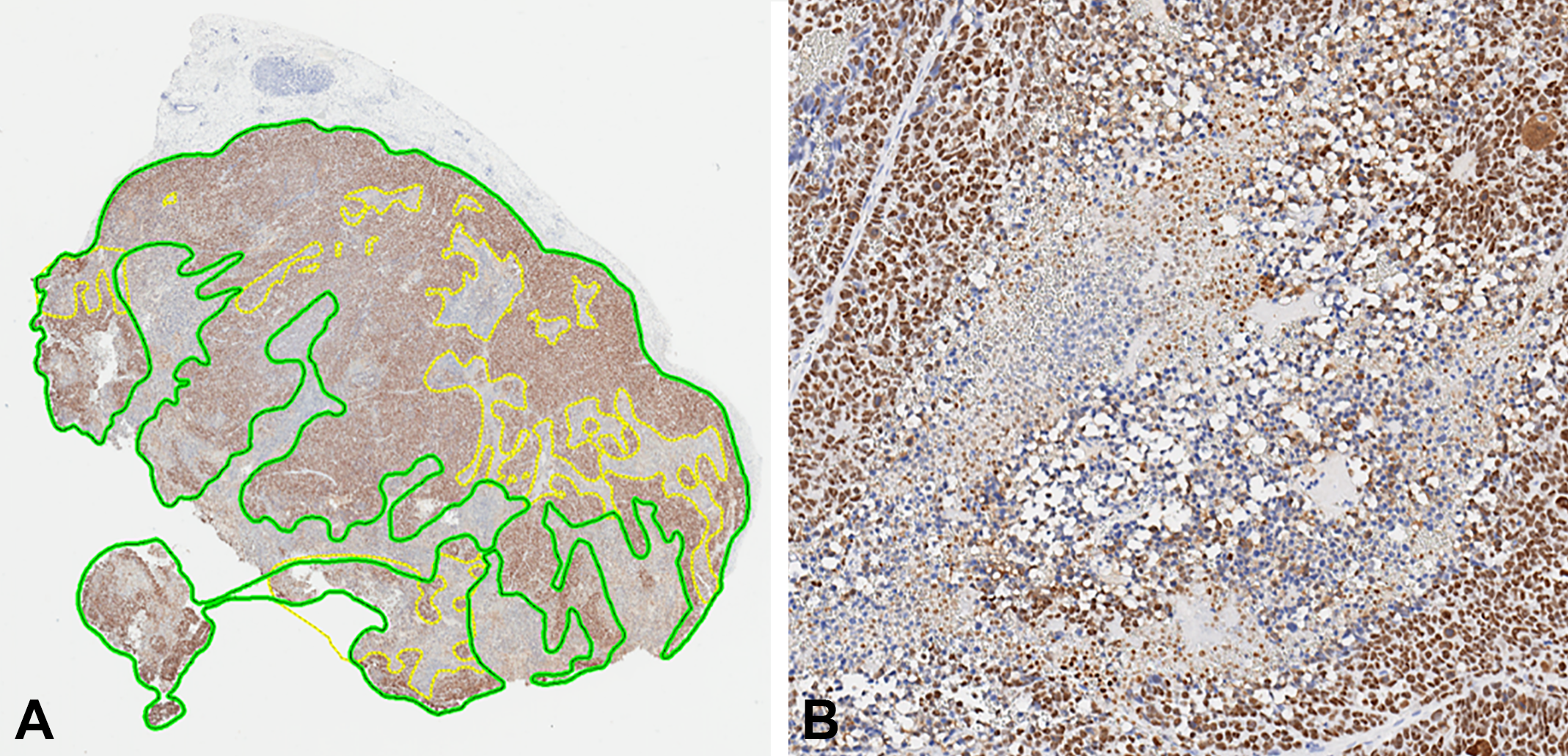
Manual image annotation of a representative tumor xenograft. Panel A shows a single large “inclusion” annotation (green outline) with multiple “exclusion” annotations (yellow) to exclude larger areas of necrosis from analysis. Panel B shows that smaller areas of necrosis may exist within the inclusion annotation, which will be excluded most efficiently using a software solution (i.e., during algorithm development) rather than a manual exclusion annotation.
For small studies, annotations can be done by a pathologist simultaneously with the slide quality review described previously. In larger studies, however, pathologists typically will devise an annotation strategy, train technicians to perform the manual annotations, and then review the accuracy of the technicians’ annotations. While annotations of nodular masses such as tumor xenografts can be uncomplicated and require little time per slide, other indications or tissue locations may require more complex and time-intensive annotations. Therefore, this division of labor within the image analysis team ensures efficient completion of annotations, while limiting the pathologist’s direct involvement in the task. To fully take advantage of the increased efficiency to be drawn from this team approach, the number and complexity of annotations should be limited. The most straightforward means of simplifying the annotation process for a sample is to ensure both high tissue quality (e.g., appropriate fixation and processing, minimal or no necrosis) and high section quality (e.g., tissues with relatively consistent staining and few or no folds or tears).
Analytical Algorithm Development and Review
During the initial reviews outlined above, the pathologist can also gather valuable information to provide to the team members who will be developing 2-D tIA solutions. Such attributes may entail assessment of variations in staining quality and tissue morphology. For example, in a study of renal cell carcinomas, the presence of carcinomas of the “clear cell” type may require a unique algorithm solution to recognize tumor cells since baseline algorithms suitable for assessing other histological types will tend to misclassify tumor cell nuclei as entire cells while ignoring the clear cytoplasm. This information suggesting that the sample set has multiple phenotypes that require distinct algorithms may result in the pathologist’s involvement in selecting a subset of study slides to establish a “training set” to aid in algorithm development. In contrast, studies in which the sample set exhibits greater homogeneity in staining presentation and tissue morphology permit the use of a statistical or random approach to select slides for the algorithm training set. In any case, detailed knowledge of the study’s goal and main end points permit the pathologist and study analyst to collaborate closely on a study-by-study basis to determine the most appropriate path forward when devising suitable algorithms for 2-D digital analysis of tissue sections.
If scoring of biomarker expression is part of the analysis strategy, it is crucial that all threshold settings are approved by a pathologist before the final analysis is performed (Tadrous 2010; Webster et al. 2011). Determining threshold values for scoring can be done employing one of the following basic strategies:
Comparison to manual scoring: pathologists can provide scores (e.g., simple intensity scores or formal H scores) for entire slides on a subset of study samples. The algorithm is then engineered to closely match those scores. Estimation of staining: the pathologist trains an algorithm to recognize staining thresholds by trial and error until an acceptable solution is achieved.
In both cases, the pathologist works closely with a technician well versed in the image analysis software because this division of labor is more efficient and cost effective.
Test algorithms should be reviewed by a pathologist prior to final analysis runs. Review criteria vary between studies and may include, but are not limited to, multiple features:
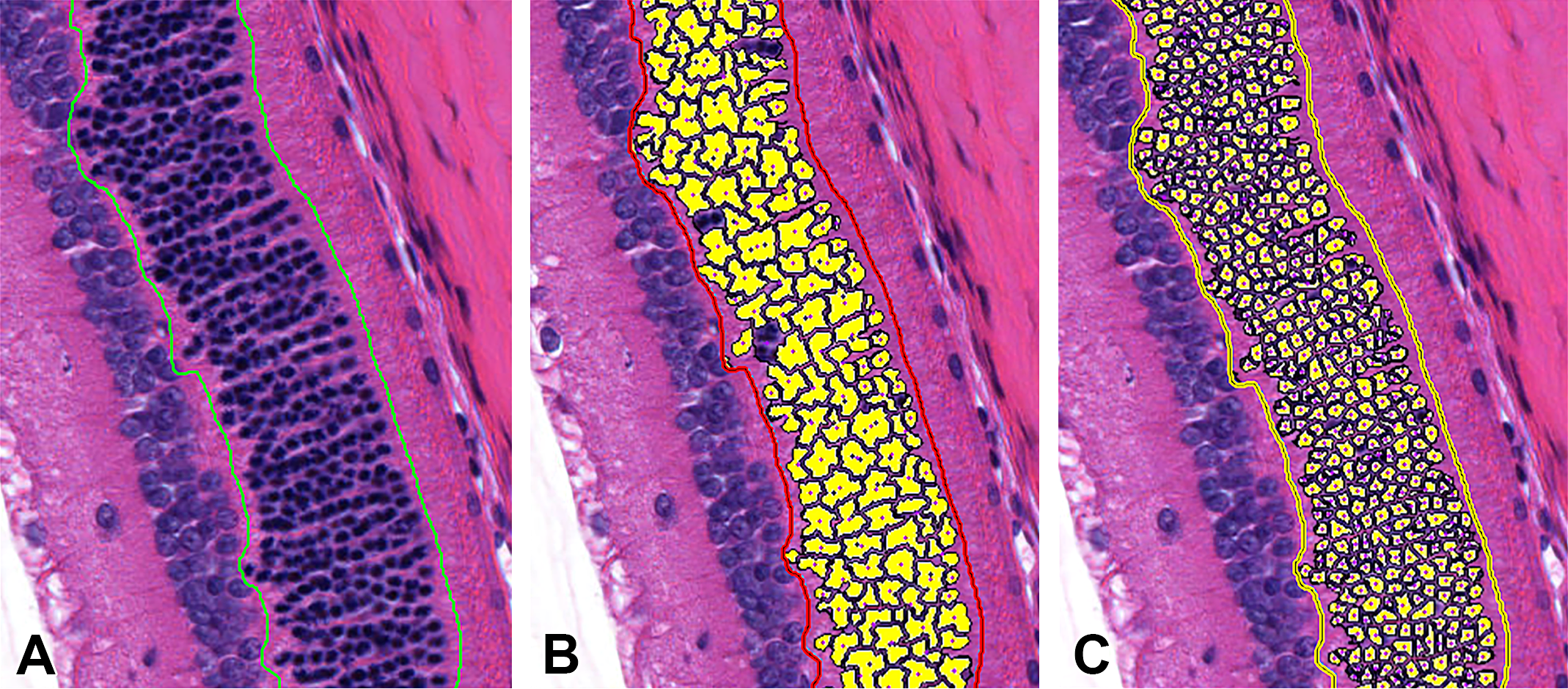
Analytical algorithm development showing possible solutions when seeking to adequately enumerate nuclei in the outer nuclear layer (ONL) of the retina. (A) H&E-stained image delineating the ONL. (B) Algorithm markup that should fail review as it does not detect all nuclei and—more importantly—is overmerging (combining) nuclei, thus creating a significant and false decrease in the cell profile count. (C) Algorithm markup that passes review as nuclei are adequately identified for enumeration.
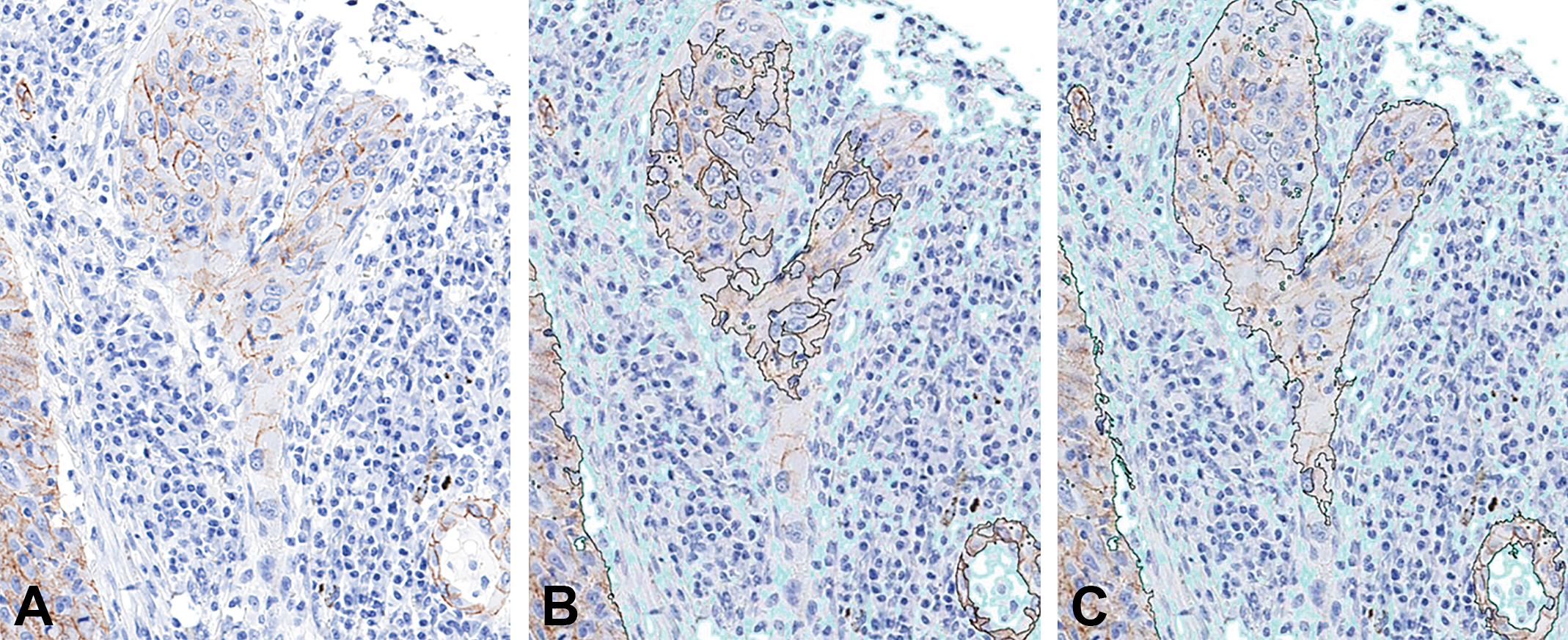
Analytical algorithm development showing adequate separation of tissue compartments (tumor cells vs. stroma) for a typical epithelial neoplasm. (A) Baseline image demonstrating the distribution of a membrane marker expressed in some neoplastic cells. (B) Algorithm markup that should fail review as it does not identify enough neoplastic cells within the region of interest (ROI, found inside the black line); areas excluded from ROI/analysis are marked in light teal. (C) Algorithm markup that passes review because it adequately distinguishes neoplastic epithelial cells from surrounding stroma. This separation is completely independently of immunohistochemical (IHC) staining characteristics and solely relies upon differences in cell morphology between elements in the two compartments. IHC staining: brown color using diaminobenzidine as the chromogen with a hematoxylin counterstain.
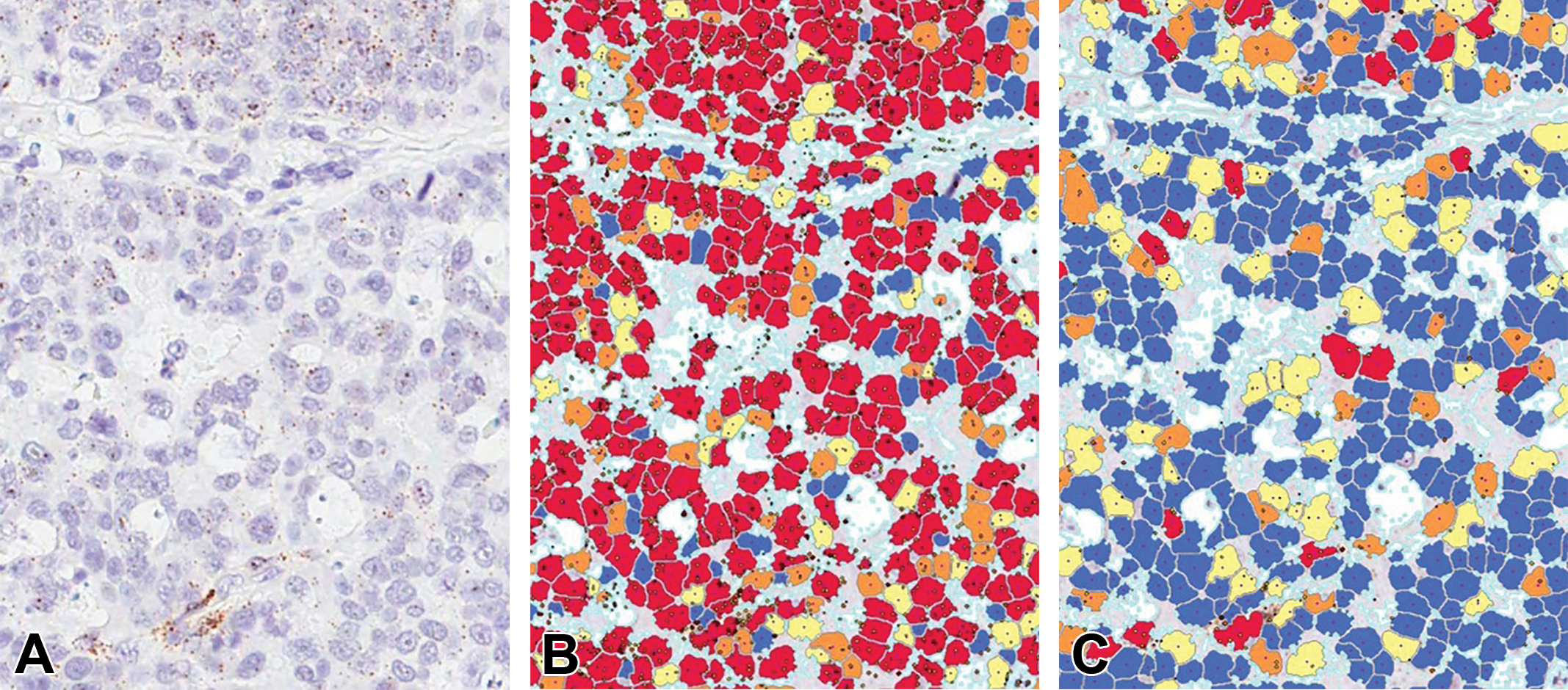
Analytical algorithm development to detect cells expressing genes that have been labeled by chromogenic in situ hybridization (CISH) in a carcinoma. (A) Baseline image showing small CISH-labeled dots superimposed on hematoxylin-stained tumor cells. (B) Algorithm markup that should fail review as the solution identifies dots in many areas where there are none (via visual assessment) and assigns a falsely high count of average dots per nucleus. (C) Algorithm markup that should pass review as it adequately identifies dots in areas where CISH dots are visually present. Color code: blue = no dots (negative), yellow = low dots count, orange = moderate dot count, and red = high dot count.
The algorithm must execute the analysis with acceptable accuracy and precision of all slides in a study, even though it cannot be expected that the solution will perform with absolute precision and accuracy on each individual slide within a diverse set. Thus, it is important to communicate the review criteria to the entire team (e.g., what is the minimum percentage of correctly classified cells for the algorithm to be accepted?). Should an algorithm fail to meet the established criteria, feedback from the pathologist to the analyst regarding why the algorithm failed is essential to improve the solution. As outlined above, it is not crucial that the pathologist be cognizant of software applications well enough to personally troubleshoot any issues. These refinements are usually performed by other members of the image analysis team. However, it is helpful if the pathologist knows the fundamental limitations of the image analysis strategy and software, so that team expectations are reasonable and communications are effective.
Some commercially available software solutions for image analysis leave very little ability to change parameters during the process of algorithm development. Even parameters that can be altered are often part of a “black box” solution, in which the end user cannot always predict how changing a certain setting will affect the algorithm performance or the final data extracted (Webster and Dunstan 2014). In contrast, other commercially available solutions are very complex and provide such a plethora of options for algorithm modifications that they require image analysis experts to be part of any analysis team in order to use the software to the extent of its abilities (Webster and Dunstan 2014; Dunstan et al. 2011). In our experience, when used in a high-throughput setting, image analysis software requires the ability to manipulate multiple parameters to define the most suitable algorithm and necessitates the cooperation of a pathologist and experienced analytical scientist to achieve the best solution in the shortest time frame.
Final Review and Data Interpretation
After the final analysis run, each slide and its analytical solution should undergo a final review by a pathologist, who confirms that results for a given sample are suitable for inclusion in the final study data. Criteria for approving or failing a sample should be objective (e.g., the analytical algorithm correctly identifies a certain percentage of tumor cells within a tissue section, a section will only be analyzed if it contains a given amount of diseased tissue). Only results from approved slides should be included in the final data set. Failed specimens for which the original algorithm provided a nearly acceptable solution may receive a cycle of algorithm refinement and reanalysis (as described in the section above) to define whether or not slight adjustments might permit useful data to be acquired from the sample. Even when pushed to their limits, in some cases, tIA algorithms may fail to meet a priori criteria for including data from a specimen in the study. In such instances, the sample should be excluded from the data collection phase of the study; for samples in which serial sections have been processed to evaluate different markers, the exclusion may apply to only some of the sections for that specimen. Excluded specimens should be categorized as “unsuitable for analysis” in final study reports, and a rationale for their exclusion should be communicated.
If the disease expertise within the image analysis team is solely provided by the pathologist, data interpretation should be performed by or with the assistance of the pathologist as well. Larger or very specialized teams should include scientists with specialty expertise, for example, a cancer biologist as part of the team analyzing a cancer study. In this setting, data interpretation and report preparation can be performed primarily by the scientist with input from a biostatistician, but the final report should be reviewed by the pathologist as well. If the pathologist has played a significant role through multiple stages of the study design, specimen quality assurance, and annotation and analytical reviews, the pathologist typically should be included as a signatory on the report.
Selecting the Team Pathologist
The type of pathologist incorporated into the team will depend on the nature of the study. For drug discovery studies focused on basic research questions, individuals with advanced comparative pathology training (e.g., PhD or DSc) but no medical background may be suitable team members. In general, studies intended for submission to regulatory agencies, which are conducted in accordance with GLP or Clinical Laboratory Improvement Amendments guidelines, should include pathologists with suitable medical (MD or equivalent) or veterinary medical (DVM or equivalent) training and, ideally, board certification in pathology. Small projects may be served adequately by a single pathologist, but larger studies may require assignment of several pathologists in order to complete the work in a timely manner. In the latter case, one pathologist (often the most experienced in digital pathology) will be chosen to lead the pathology subgroup of the image analysis team.
Summary
Digital image analysis is positioned as a transformative technology for biomedical science in general and the pathologist in particular. Pathology data sets can now be analyzed in new and exciting ways, enriching the impact of data on basic science and medicine. While oftentimes the apprehension is expressed that image analysis may replace pathologists, the truth is that no part of the image analysis process can be performed successfully without input from pathologists. The wide-ranging pathophysiological expertise that pathologists bring to an image analysis team makes them invaluable, and indeed vital team members, and their input and feedback are essential for creating fit-for-purpose tissue analysis solutions and high-quality data. Pathologists contribute to digital pathology research teams by (a) ensuring that image analysis data are meaningful and reproducible and (b) that image analysis tools are defined with pathologists’ input to ensure maximum utility and applicability.
As with any scientific methodology, certain limitations apply to the utilization of tIA, and other data-generating approaches may prove more appropriate for addressing specific research questions than will 2-D morphometric analysis. Examples of other techniques include stereology (3-dimensional [3-D] analysis of a structure using 2-D evaluation of structures in step-sectioned specimens), other 3-D noninvasive imaging approaches (e.g., magnetic resonance microscopy), or tissue homogenization for flow cytometric analysis of biomarker expression on dispersed cells. For instance, a recent position paper advocates that accurate assessment of common morphological end points in lung (e.g., alveolar number or size) requires the application of 3-D stereology (Hsia et al. 2010). However, while mandated use of 3-D measurements may be justified in some settings, several recent studies across various disease indications in humans have shown that 2-D tIA of digital images yields data sets that are (1) concordant with scores acquired by conventional manual evaluation and diagnostic scoring and (2) at times provide an even more accurate measurement for the severity and stage of a disease (Daunoravicius et al. 2014; Zhong et al. 2016; Laurinaviciene et al. 2011; Turashvili et al. 2009; Vennalaganti et al. 2015; Lee et al. 2013; Farris et al. 2014; Snead et al. 2016; Ohlschlegel et al. 2013). The utility of 2-D tIA gains additional importance in tissue specimens harvested in the course of human clinical trials because 2-D digital imaging approaches still provide useful information when tissue sample sizes are too small (e.g., needle core biopsies) to permit 3-D evaluation or when ethical considerations preclude optimal processing of tissue (i.e., euthanasia with rapid removal and fixation of tumor-ridden organs as practiced in animal experiments is not possible in human patients).
Another possible limitation of digital pathology involves data misinterpretation due to an incorrect understanding of the basic principles by which 2-D tIA should be conducted. A classic example of misguided data interpretation occurs when scientists assume that a net reduction in absolute numbers of a target cell population has occurred solely because the number of cell profiles is decreased as a percentage of the total number of cells that are present. This misinterpretation discounts the possibility that the target cell population might have been stable (or even increasing) in absolute terms, which is undesirable behavior in a tumor cell population even if tumor burden as a percentage of a tissue is low. This limitation is best surmounted by entrusting data interpretation to individuals with suitable training and experience in both 2-D tIA methods but also the fundamental biological principles that underpin a given disease—in short, pathologists with expertise in modern digital pathology.
Footnotes
Acknowledgments
The authors would like to thank the entire staff of Flagship Biosciences, Inc., who contributed to the many learning experiences that are incorporated in this article. Special thanks to Matthew Steaffens, Justin Major, Stefan Pieterse, Benjamin Landis, and Shane Cogar for their aid in figure preparation.
Author Contributions
Authors contributed to conception or design (FA, KW, BB, KS, CM, DR, EC, GY), drafting the manuscript (FA), and critically revising the manuscript (KW, BB, KS, CM, DR, EC, GY). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors received financial support in the form of a salary as employees of Flagship Biosciences Inc., a company specialized in the development of digital image analysis solutions.
