Abstract
Digital toxicologic histopathology has been broadly adopted in preclinical compound development for informal consultation and peer review. There is now increased interest in implementing the technology for good laboratory practice–regulated study evaluations. However, the implementation is not straightforward because systems and work processes require qualification and validation, with consideration also given to security. As a result of the high-throughput, high-volume nature of safety evaluations, computer performance, ergonomics, efficiency, and integration with laboratory information management systems are further key considerations. The European Society of Toxicologic Pathology organized an international expert workshop with participation by toxicologic pathologists, quality assurance/regulatory experts, and information technology experts to discuss qualification and validation of digital histopathology systems in a good laboratory practice environment, and to share the resulting conclusions broadly in the toxicologic pathology community.
Introduction
There have been two workshops commissioned by the European Society of Toxicologic Pathology (ESTP) to discuss adoption of digital toxicologic histopathology. In the 7th ESTP International Expert Workshop (2018), a broad panel of industry, diagnostic, academic, clinical, and regulatory experts consolidated available information and experiences at the time, and aligned on a set of minimal technical and regulatory requirements for qualification and validation of digital primary and peer review evaluations in the good laboratory practice (GLP) environment. 1 In the 8th workshop, held in November 2020, experts from information technology (IT), Quality Assurance Unit (QAU), Health Authority (HA), and Toxicologic Pathology disciplines discussed the requirements in greater detail. This article is a complement to the 7th ESTP workshop paper 1 and summarizes factors that may impact qualification and validation of a digital toxicologic histopathology system and its components in a GLP environment. The workshop results shared here are the output of those expert discussions and should be taken into context with emerging guidance and literature in this rapidly evolving field.
Selected Basic Principles of the Qualification and Validation of the Pixel Pathway
The digital histopathology workflow starts when a histological tissue section is converted by a whole slide scanner (WSS) into a high-resolution digital replica, which can be displayed on a computer monitor by means of various software applications. The route taken by the “pixel data” from the scanner to the computer monitor, including any storage or archival locations is called the “pixel pathway,” and this is an important concept first described in the Digital Pathology FDA (US Food and Drug Administration) guideline in 2016. 2 The pathway will vary according to hardware and software components, the number of sites and their geographic locations, and the business need, including intended use cases. One example of a pixel pathway is illustrated in Figure 1. The key concept is that the entire pixel pathway, end-to-end, needs to be validated 2 and its components (system, equipment, and/or software) need to be qualified as part of the validation.
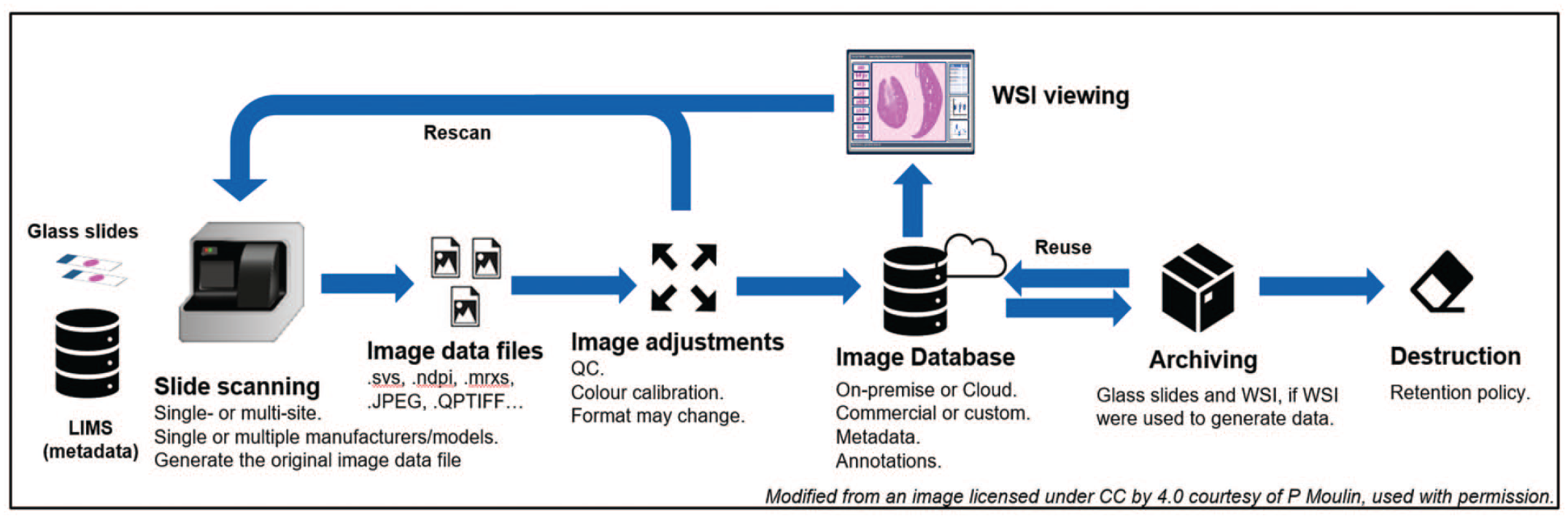
The pixel pathway. The term refers to the route taken by digital image “pixel data” from production to eventual destruction. The pathway will vary according to hardware and software components, the number of sites and their geographic locations, and the business need, including intended use cases. Note that WSI viewing might encompass primary reads, prospective or retrospective peer reviews, and pathology working group review. LIMS indicates laboratory information management systems; WSI, whole slide images.
If the entire pixel pathway occurs within a single institution (either single site or multisite), the qualification and validation are the sole responsibility of the institution. On the contrary, if the pathway is a joint venture between institutions, for example, a contract research organization (CRO) and a Sponsor, each partner is accountable for their respective phase (Figure 1). The accountability needs to be clearly stated in the study protocol and/or the respective Standard Operating Procedures (SOPs).
Although whole slide images (WSI) can be used to generate raw data, they do not strictly fulfill the definition of specimens per se since they are derived from specimens, namely glass histology slides, and not from a test system. Consequently, the term “raw data source” was coined for WSI during the 7th ESTP workshop but it is not yet a widely accepted regulatory status. 1 Despite the current lack of an appropriate term, whenever the WSI were used to generate raw data, they should be archived in addition to the glass histology slides because they are needed to reconstruct the study (21 CRF 58, 40 CRF 160, OECD 1). This is true for studies where the primary read was entirely digital but also in the event that only a subset of WSI was evaluated to take advantage of certain benefits specific to digital pathology such as side-by-side WSI evaluation in split-screen mode. In situations where WSI were not used to generate raw data, such as standard contemporaneous/prospective digital peer reviews, WSI archiving should not be required. 3 As the field matures, more complex scenarios will likely emerge that require consultation with the local QA group to determine the fate of WSI. Our recommendations regarding the archival status of WSI in the most common digital toxicologic histopathology use cases are presented in Table 1. 4
Most common digital toxicologic pathology use cases and their associated archival recommendation.
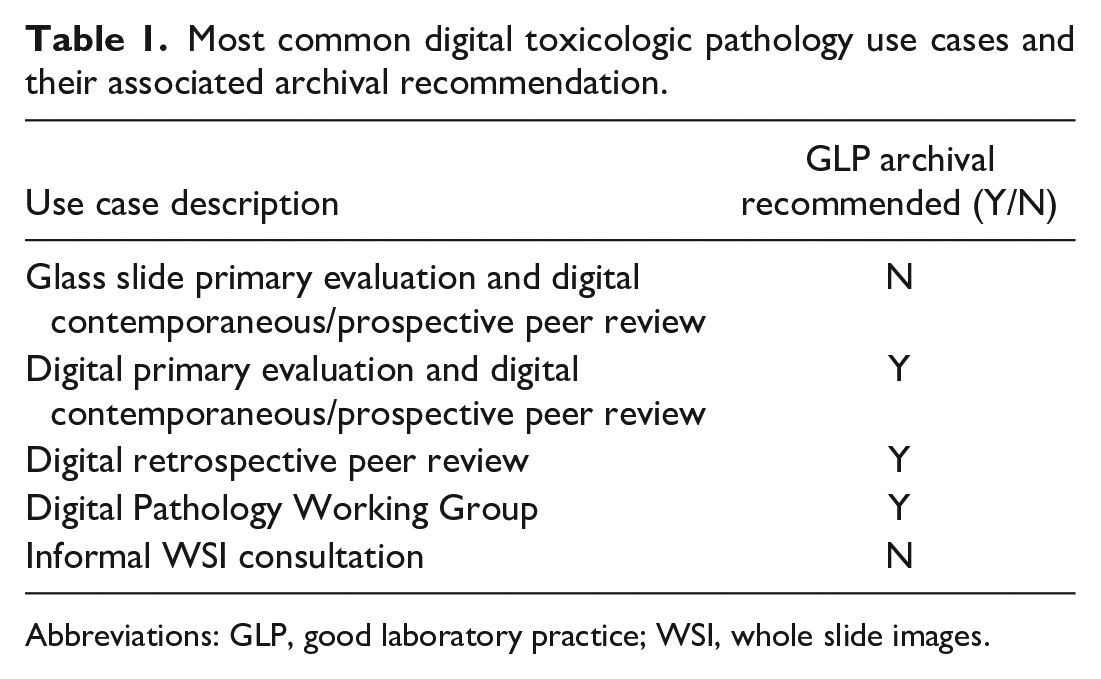
Abbreviations: GLP, good laboratory practice; WSI, whole slide images.
Ultimately, it is the responsibility of the study pathologist to employ appropriate procedures and safeguards to assure the validity of any histopathology data generated from images. 5
The Basic Validation Principles Applied to Digital Toxicologic Pathology
The terms validation and qualification are often confused. The equipment and software that are part of a digital pathology system are qualified and the qualification establishes that they work as intended by the manufacturer(s). The validation establishes that for a specific purpose, for example, primary digital histopathological evaluation of toxicity studies, the processes in place as defined in SOPs and protocols performed by trained personnel to use the qualified digital pathology system are fit for purpose, that is, allow the pathologist to reliably perform their histopathological assessment.
Validation of the pixel pathway is subject to the same requirements as any GLP system. The best approach is considered to be risk-based, because it ensures that the phases with highest business and compliance risk receive foremost priority. The adopted approach will vary among institutions. Recommended validation steps will be reviewed in this section but the reader is also encouraged to consult the work by Jacobsen et al. 4 which also describes the validation of WSI and the associated workflow for the digital primary evaluation and peer review of a GLP-compliant toxicity study.
The ultimate aim of the validation process is to provide documented evidence that the digital evaluation is equivalent to reading slides using a light microscope (see Figure 1 depicting the WSI life cycle and Figure 2). 6
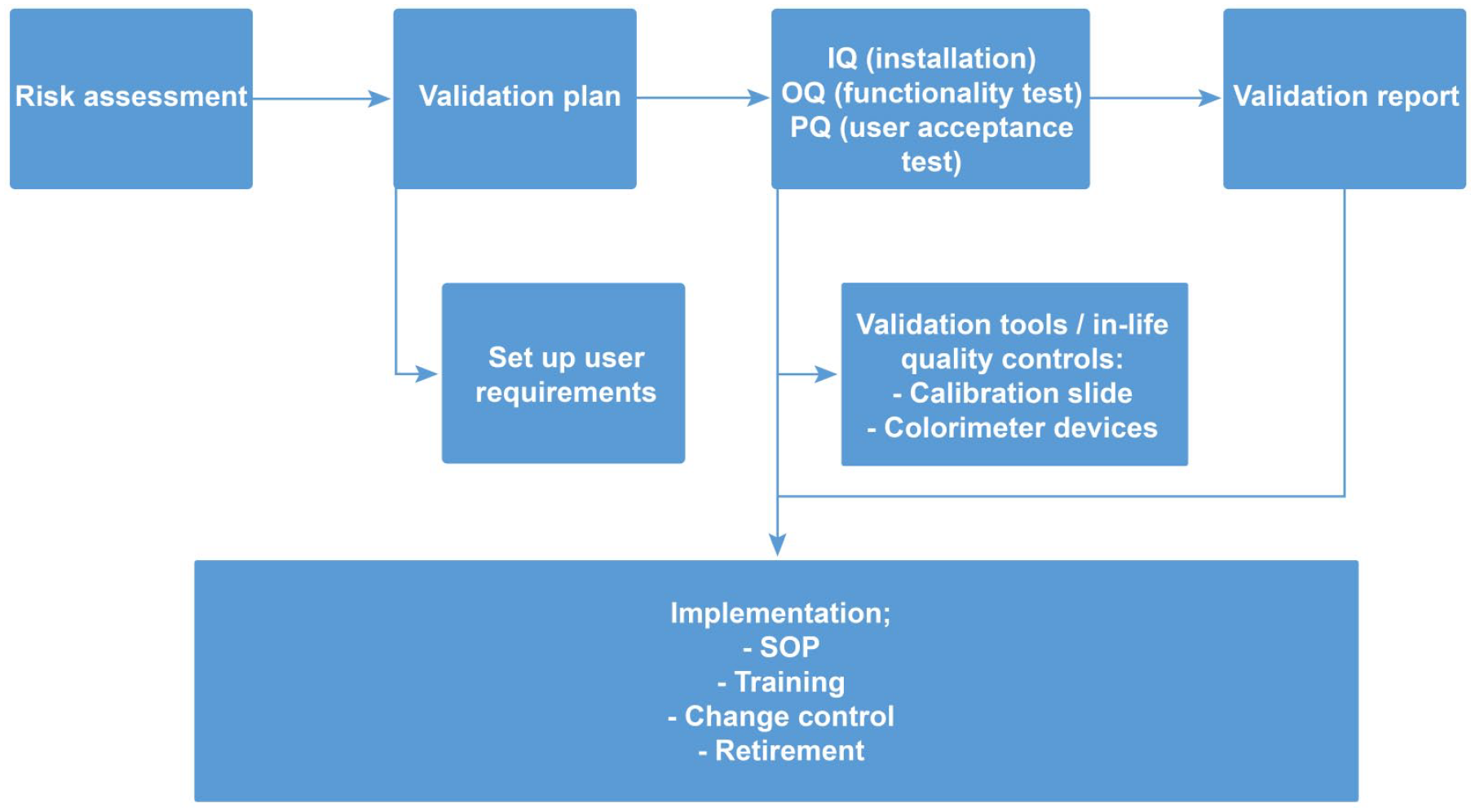
Validation process for digital pathology. The validation of a digital pathology workflow comprises the key steps encountered in computer system validation, from the definition of use cases to retirement of the system. The specificity lies in ensuring integrity of the pixel pathway, in particular when several institutions are involved (refer to Figure 1 for more details). IQ indicates installation qualifications; OQ, operational qualifications; PQ, performance qualification; SOP, Standard Operating Procedure.
An important first step is to draft a validation plan, which should define the intended use cases, including end user requirements and associated functional and design/system specifications, and the testing strategy. The user requirements specification (URS) will include the requirements of pathologists, regulatory authorities, and cybersecurity. Examples of URS have been previously published.4,7
The validation plan should disclose and define all components and steps in the pixel pathway, unless this information is already supplied by the vendor in the case of a commercial end-to-end solution (see Table 2). Once the validation plan is in place, testing can commence.
Design specifications for a digital pathology system.
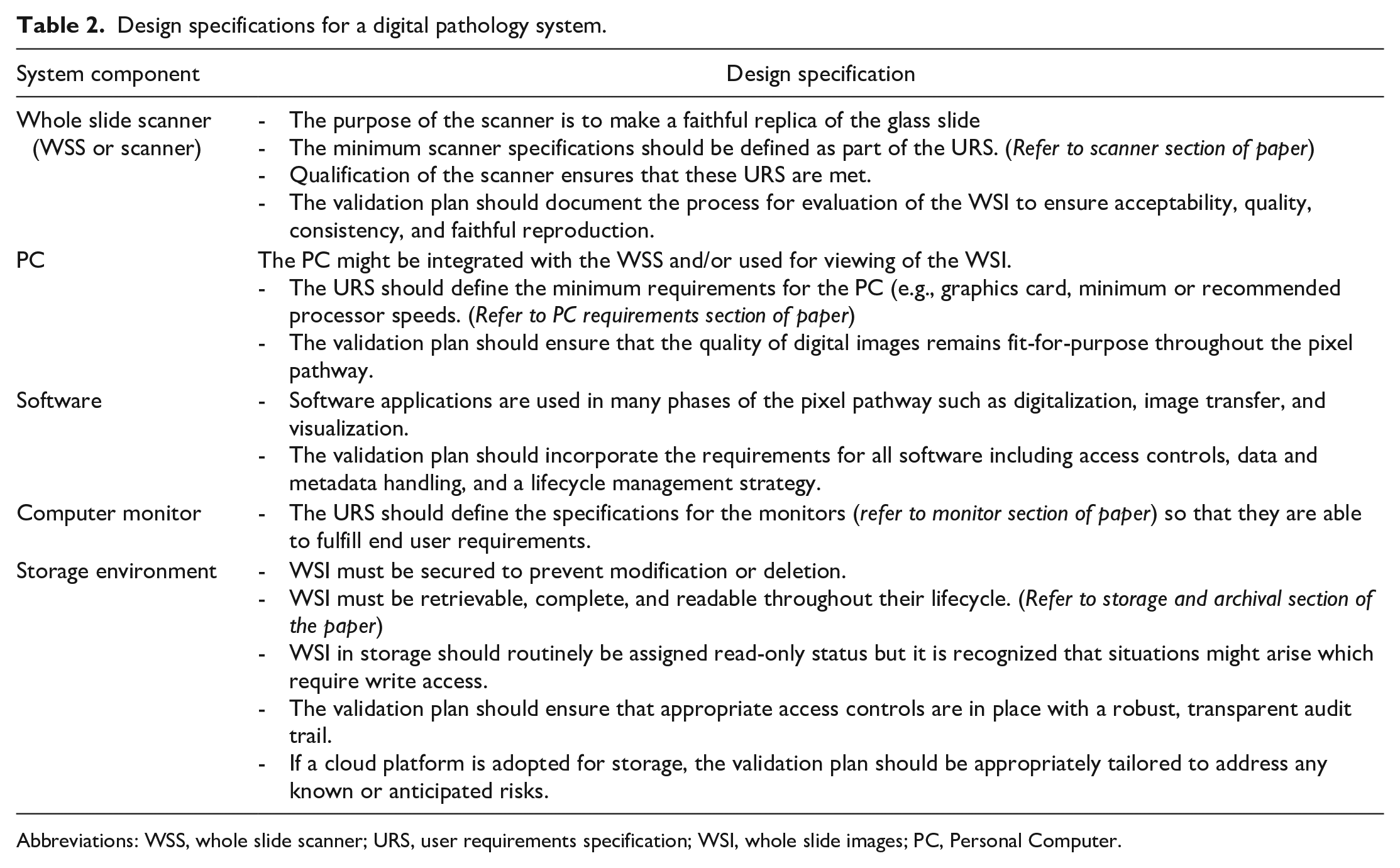
Abbreviations: WSS, whole slide scanner; URS, user requirements specification; WSI, whole slide images; PC, Personal Computer.
During the testing phase, the pixel pathway is qualified which refers to the action of proving that all hardware and software components operate correctly and are fit for purpose. Qualification includes the following 6 :
Installation and operational qualifications (IQ/OQ)—IQ confirms that the components are installed per design specification and the OQ confirms they meet all the functionalities defined in the user requirements. IQ/OQ might be done by a commercial vendor.
User acceptance testing, also called performance qualification (PQ)—A set of WSIs that represent typical preparation types is evaluated by qualified pathologists to confirm acceptability. The PQ should include confirmation of noninferiority relative to glass slides by means of digital to optical equivalence testing at different phases of the pixel pathway. For example, when viewed in the digital pathology system, both the WSI file format used for primary evaluation and the one retrieved from archive, if different, are faithful replicas and allow pathologists to do a diagnosis just like the light microscopic evaluation of the originator glass slide.
Hardware or software components that have previously been validated for different purposes should be revalidated for the new intended purpose. In addition, if components are commercially procured, the vendor-documented validation strategy and findings should be thoroughly assessed. Any relevant documents must be retained by the vendor. The OECD Advisory Document Number 17 6 includes detailed guidance on considerations relating to assessment of vendors for computerized systems and related services.
Qualification has been successfully completed when the entire pixel pathway meets the predetermined requirements outlined in the validation plan.
Finally, a validation report should be drafted by the institution(s), which summarizes the components in the pixel pathway and the qualification results. Prior to implementation, the system owner must ensure that relevant SOPs and appropriate personnel training for operating/using the digital pathology system and access controls are established in the test facility/site. Once completed, the digital pathology system can be declared to be validated and it can be implemented in a GLP environment. The implementation phase is detailed in the last section.
WSS
WSS Qualification
The concepts of scanner qualification will not be detailed here as they were described in the 7th ESTP Expert Workshop Paper. 1 The FDA and the European Union (EU) have published differing guidance on the qualification of WSS used for clinical purposes. 2 However, after May 26, 2024, EU guidance on WSS will change and place requirements as rigorous as those established by the FDA. Current EU regulation only requires a self-declaration of conformity by the manufacturers. After May 26, 2024, the EU will consider WSS to be class C medical devices, which is the penultimate risk classification, 8 and requires WSS to meet standards approved under the new EU regulation 2017/746. 9 These new standards will require manufacturers to conduct a performance evaluation similar to the testing scheme requested by the FDA, which will include a scientific validity report, an analytical performance report, and a clinical performance report. Whole slide scanner manufactured with the new EU CE label for clinical use will be supported with documentation that a toxicologic pathology test facility could use to support the validation of a digital histopathology system using such a WSS. In published accounts of digital histopathology system validation, the WSS used has not been classified as clinical devices and other validation strategies have been employed to satisfy GLP requirements. 3
Recommended Procedures for Best-Quality WSI
Requirements for creating of high-quality WSI depend on the intended use case; however, there are some commonalities:
Proper maintenance and calibration of the WSS including color performance per vendor specification and periodic scans of well-characterized slides.
Cleaning and quality inspection of slides before and after scanning.
Labeling suitable for creating and linking metadata. This step can be based on label barcodes or validated optical character recognition of labels.
Quality control of the WSI (color, tissue completeness, and presence of scanning artifacts).
WSI File Format
Also critical is a system design that supports the lifecycle management plan by incorporating use of an appropriate file format. Scanner-specific proprietary file formats including their lack of interoperability were discussed in the 7th ESTP workshop paper 1 and some progress on this issue has been made in the intervening years. For example, libraries like Fiji’s Bio-Formats plugin render most proprietary formats viewable 8 and there is increased interest in standardized formats such as DICOM or OME-Tiff, which can containerize metadata with the image data. Widespread adoption of one of these image formats will improve interoperability between various digital histopathology systems and lead to standardization in metadata. However, the consensus on a nonproprietary format has not yet been reached by the scientific community.9,10
Metadata and Associated Data
Metadata is any information that identifies, describes, or relates digital images and their elements. 1 It gives identity, meaning, provides context, defines structure and enables retrieval across systems and usability, authenticity, and auditability across time. From a regulatory standpoint, the absolute minimum metadata set that should accompany a WSI is all the information currently captured on the glass slide label, because it links a digital image with the study data. Metadata can be expanded according to the local data ecosystem capability and end user requirements to encompass additional attributes of the study (e.g., study identifier, design, duration, and route of administration), animal (e.g., species, strain, gender), sample (e.g., tissue name, necropsy date, type of fixative), and digital image (e.g., pixel count, pixel size, light source). Workshop participants recommend that industry explore the creation of a standard for metadata terminology akin to the successful SEND initiative (www.cdisc.org/standards).
Associated data may include any ancillary data, which might be helpful during review of digital slides, including, but not limited to, in-life animal data, organ weight data, macroscopic data and clinical pathology/biomarker data, microscopic findings in the case of a peer review, but also image analysis (IA)/artificial intelligence (AI) results. The format of associated data could vary from one organization to another but if external sharing is the intended use, then associated data should be format agnostic.
To support the benefits of digital pathology, metadata and its associated data should be linked to WSI in a searchable and retrievable database or image management system (IMS). 2
Laboratory Information Management Systems
The antemortem and postmortem phases of nonclinical toxicology studies generate large amounts of data that are potentially relevant to the histopathology interpretation by the study pathologist. The ability of the pathologist to access these metadata and associated data easily during the digital histopathology evaluation depends on how well and how fully the IMS of the digital histopathology platform is integrated with the various existing laboratory information management systems (LIMS; e.g., in-life, histology, and pathology LIMS systems). As most test facilities currently use different vendors for digital histopathology and LIMS, integration can be a technical challenge. Workarounds to incomplete technical integration, such as manual free-text entry, may compromise data integrity or involve costly quality check processes. Therefore, use of drop-down lists, automating data flows among integrated LIMS, and implementing a centralized repository of controlled terminology lists may ultimately improve quality and manage the long-term costs of using a digital histopathology platform.
In-life LIMS typically collect biometric data such as supplier ID, animal ID, and body weight, and various other procedural data, for example, dosing and blood collection regimens, and clinical and dosing observations. In addition, in-life LIMS often also capture necropsy data such as organ weights, macroscopic findings, and tissues collected. Histology LIMS typically capture procedural data from receipt of tissues in fixative pots to stained histological sections on glass slides, including trimming, decalcification, processing, embedding, staining, histology review notes, audit trials, and slide barcodes. In laboratories that do not have a histology LIMS, the data may instead be captured on paper. Pathology LIMS collect microscopic findings, pathologist comments, and macroscopic to microscopic correlations, and also typically allow easy access to in-life and necropsy data.
Image management systems have developed from simply storing, searching, and displaying WSI to the point where a large amount of metadata and associated data can be linked at either a study or WSI level. This opens the potential for the pathologist to sort and re-sort WSI quickly and conveniently into subcollections based on the linked data to facilitate the microscopic examination. Such developments in the functionality of IMS are allowing pathologists to examine studies, tissues, and cases from a new perspective which previously would have been prohibitively time consuming to achieve.
LIMS Integration
Harvesting metadata and associated data from the institutional LIMS and integrating the information with WSI might seem daunting. Close collaboration between technical and business owners of the various LIMS and IMS, as well as IT professionals, is best established early in the effort. The linkage of information to the WSI is dependent on the workflow. A single unidirectional (i.e., moving data only from the study data LIMS to the IMS) linkage might suffice for digital peer reviews because it will provide the pathologist enough information to carry out the task. The extraction and linkage of information might be facilitated by a SEND-compliant LIMS output. On the contrary, primary digital pathology microscopic examinations will benefit from real-time bidirectional linkage between the LIMS and IMS. This interface is much more complicated and requires cooperative product development efforts from the manufacturers of LIMS and IMS. Some of these interoperability efforts between vendors are being piloted by a group of users.
An early and key step in the accurate linkage of information to WSI is determining the identity of every WSI. In the conventional glass slide workflow, the identity is established via the human-readable information on the slide label which usually includes the study number, animal number, dose group, and block number. In the digital workflow, the identity can be captured in various ways. For example, the key metadata can be containerized with the image in the DICOM or OME-tiff file formats. Alternatively, it can be manually added to the image file names, but a thorough quality control will need to be put in place to minimize human error and thereby the risk of misidentification which will help ensure data integrity. The application of a barcode to the slide label by a histology LIMS is a modern and practical solution which enables automation. Minimally, a 2-dimensional barcode should contain the human-readable information available on the slide label. Additional information can be included, as deemed necessary by the institution. Following digitization, the barcode is read by the LIMS, which the triggers retrieval and linkage of the correct key metadata required for identification. Histology LIMS can also associate the barcodes with all upstream histology procedures, including all entered metadata and associated data and it might even be possible to harvest the relevant pathology and associated data. Once the WSI has been correctly identified and linked to the various LIMS, multiple metadata fields can be automatically attributed to the slide, animal, dose group, and/or study, thus greatly improving the digital primary read or peer review.
Viewing Experience
The importance of color calibration along the pixel pathway was one of the highly debated topics during the workshop. When compared with a high-performing digital camera integrated in a slide scanner, the human visual cognitive system is variable, biased and limited in terms of color and hue perception.11 -13 However, thorough evaluation of histopathology slides entails more than just color assessment as evidenced by the fact that pathologists can recognize lesions in black and white images. 13 Therefore, information other than color, such as contrast, depth, shape, densities, structure, architecture, and texture, are additional cues integrated by pathologists’ visual cognitive systems to recognize lesions and arrive at a diagnosis. 14 While all attendees agreed that the standardization of WSI color is important for quantitative image analytic approaches such as IA, AI, and machine learning, several were of the opinion that it is less important for digital primary read or peer review due to the adaptability of toxicologic pathologists to different stain profiles. However, workshop participants recommend that routine calibration of WSS as specified by the vendor should be regularly performed.
All the attendees were convinced of the benefit of minimum standards for monitor quality and calibration (luminance, contrast, color). 7 Like the adjustments available on analogue microscopes, pathologists should understand the effects on the viewing experience of environmental factors such as ambient light and the different adjustable monitor specifications.13,15 A point-of-use tool has been developed to assess the suitability of the display for digital pathology assessment, which also takes environmental factors into account.16,17 Pathologists should be aware of the environmental factors, which contribute to computer vision syndrome, a condition characterized by symptoms ranging from eyestrain to blurred and double vision which is brought about by long-term use.1,18 Monitor height, distance-to-display, and back lighting should ideally be optimized and the 20-20-20 rule practiced as far as possible, that is, for every 20 minutes spent looking at a computer screen, the person should look at an object 20 feet away for 20 seconds, to reduce eye strain and visual fatigue.
Hardware Requirements: Pathologist Computer
The computer is an integral component of the digital pathology workstation and the search for the “best” computer is complex. This section will provide important points to consider without being overly prescriptive. Early engagement of the institution’s IT and purchasing departments to understand policy and limitations will be advantageous given that the final choice is often a balancing act between personal and corporate considerations. For example, some institutions do not permit consumer-grade purchases due to their relatively high turnover and low lifespan. The first consideration is laptop versus desktop, and the mobility required by the pathologist will be a key determinant. Next, one needs to consider the processing power. The processing power (central processing units [CPU] and graphics processing units [GPUs]) of computers continues to rapidly improve. Thus, the minimal standards recommended by the viewing software vendor today might be out of date by the time of publication. As a general rule, a high-quality CPU is important but might not be as pivotal as the GPU for image viewing. Random Access Memory (RAM) works with the on-board CPU to enable function. The newest and strongest GPUs are typically available in consumer-grade computers earlier than business-grade computers. Selecting a GPU well above the minimal standards recommended by the viewing software vendor will improve the shelf life of a computer and improve the monitor options and available configurations.
The on-board storage capacity of the computer is another important consideration. The selected image-viewing application will determine to a large extent, the storage requirements. Generally, a 2 Tb SSD drive might be sufficient; however, additional storage might be needed if a local cache is required for rapid access of images in remote settings, that is, home working. The storage capacity can also be increased via an external, high-volume, SSD drive.
Vendors of image-viewing software typically define the minimum requirements needed for good performance.
Hardware Requirements: Monitor
Toxicologic pathologists will potentially need to spend many hours examining digital images. Therefore, selecting a computer monitor is an important step in the configuration of a digital pathology workstation. The monitor should be of sufficient quality, include all relevant functionality, and offer an ergonomic and comfortable viewing experience. This section recommends minimum standards without being overly prescriptive.
Modern monitors can have up to 30 different specifications. The most important specifications for digital slide display are typically available in consumer gaming monitors. A 16:9 format, large display (30″ or larger) with high resolution (4K: 3840 × 2160), high pixel density, and low pixel pitch, as well as adequate luminance and contrast ratios are recommended by workshop participants. While yet higher resolution will result in a sharper image with more details (for a similar screen size) and enable display of a correspondingly larger field of view, it can also result in objects appearing smaller than normal when using the maximal magnification. In addition, a higher-resolution monitor might not achieve the targeted frame rate when combined with the display format and the image viewer settings, resulting in image tearing. 15
Workshop participants further recommend that for a smooth and seamless viewing experience with fast navigation of the digitized slide, responsiveness in case of touch operation, and free from stutter, flickering, or ghosting artifacts, a ≥120 Hz refresh rate together with a ≤1 ms response time is recommended. NVIDIA G-SYNC or a similar technology is another gaming-focused feature that should be taken into consideration, but which necessitates a compatible graphics processing hardware. It synchronizes the display’s refresh rate to the GPU computer, helping to eliminate screen tearing and minimize input lag. A DisplayPort (1.2 minimum) connectivity supporting G-SYNC technology and 4K resolution is another feature to consider.
A built-in calibration system or the use of an external sensor to periodically calibrate the display is considered essential. The color range of the monitor should also be matched to the scanner color range.
A monitor which is adjustable for height, swivel, pivot, and tilt, as well as a 170° viewing angle and an anti-glare display surface will help set up an eye-friendly, ergonomically designed workspace.
Hardware Requirements: Input Device/Peripherals
Although the stage and focus control mechanisms and objective selection design of light microscopes have been refined to improve ergonomics over many years to the point where there are few, relatively minor, differences between the varying commercial brands and models, such refinement is yet to take place in the digital slide navigation realm. Several groups have highlighted the options open to the pathologist as well as attempted to compare them in terms of comfort, speed, and accuracy.19 -21 An extensive range of novel peripherals have been evaluated including vertical/ergonomic mice, trackballs, trackpads, a 6 degrees-of-freedom navigator, games console controllers, a hanging gimbal device replicating light microscope stage control, and head and eye tracking software. However, according to a 2019 STP survey undertaken prior to the Covid19 pandemic, there had been very little uptake in the toxicologic pathology field with all respondents reporting that the conventional mouse and keyboard are still used for digital slide navigation in their institution. 22 This is likely a result of conventional mice and keyboards being ubiquitously available, and many companies and pathologists being relatively early in their adoption of digital pathology platforms are only now examining sufficient numbers of digital scans to warrant the investment in, and see the benefit from, dedicated peripheral input devices.
Many of the previously mentioned devices have inbuilt functionality to program the various available buttons and controls to complete different actions. This includes being able to alter how the pan and zoom functions are controlled, navigation to the next slide, annotation creation (e.g., marker or text comment) or fine tune the input to output rate to the pathologist’s personal preference. Other peripheral devices are also available to supplement data or annotation entry, for example, shortcut keyboards. However, having an increasing number of peripheral devices concurrently in use introduces its own complexities both in terms of design and ergonomics of the pathologist’s workspace and how these devices interact and to which active program they function alongside. Ability for devices to concurrently provide input to different specified open programs would be ideal, to prevent the pathologist from repeatedly having to swap active programs to allow input.
Most of the references discussing digital slide navigation have, understandably, compared different input devices using a small number of slides or cases and have compared the time taken to reach a diagnosis as a comparative measurement. The conclusions drawn by these papers, therefore, need to be taken in context of the typically high number of slides examined per day by many toxicologic pathologists, and also due to the purpose of the examination, and many tissues being unremarkable, less time is typically spent examining each section. This will result in comparatively more time spent interacting with the input device to pan, zoom, and switch slides, which will likely exacerbate any perception of increased workload and result in fatigue. 19
Storage and GLP Archival
In the context of the cases mentioned in Table 1 and workflow depicted in Figure 1, a WSI file may change storage location and/or format throughout its lifecycle. In general, storage can be separated into three tiers: initial, intermediate, and long term. Initial storage is the first location from the scanner and is usually associated with an image database provided by the scanner vendor. It may be accessed by multiple groups, lack retention policies, be associated with metadata or not, and might be unstructured. This storage might contain a mix of images for GLP and non-GLP projects. For studies where GLP compliance is claimed, a dedicated storage must be created, requiring the use of validated servers and the database must be subject to minimum requirements (including, but not limited to, defined roles, user account management, audit trail, etc.). This storage might not be optimized for study review and require the use of a dedicated WSI viewer to allow primary or peer review.
Whole slide images might remain in the initial location, be copied, or be moved to an intermediate storage for review and analysis. The intermediate location might be associated with a specialized IA system, be structured by study, associated with scanner- and study-specific metadata or associated data, and have tight controls. The user should be aware of any encryption or compression occurring during transfer to evaluate their impact. Images remain in intermediate storage for the duration of a study. The intermediate storage should be optimized for evaluation and analysis.
Once a study is complete, it would be moved to long-term storage for archival or future analysis. This storage could be either online network storage or removable media for off-line storage.
Types of Storage
Various types of storage support the use cases, as shown in Table 3.
Types of storage.
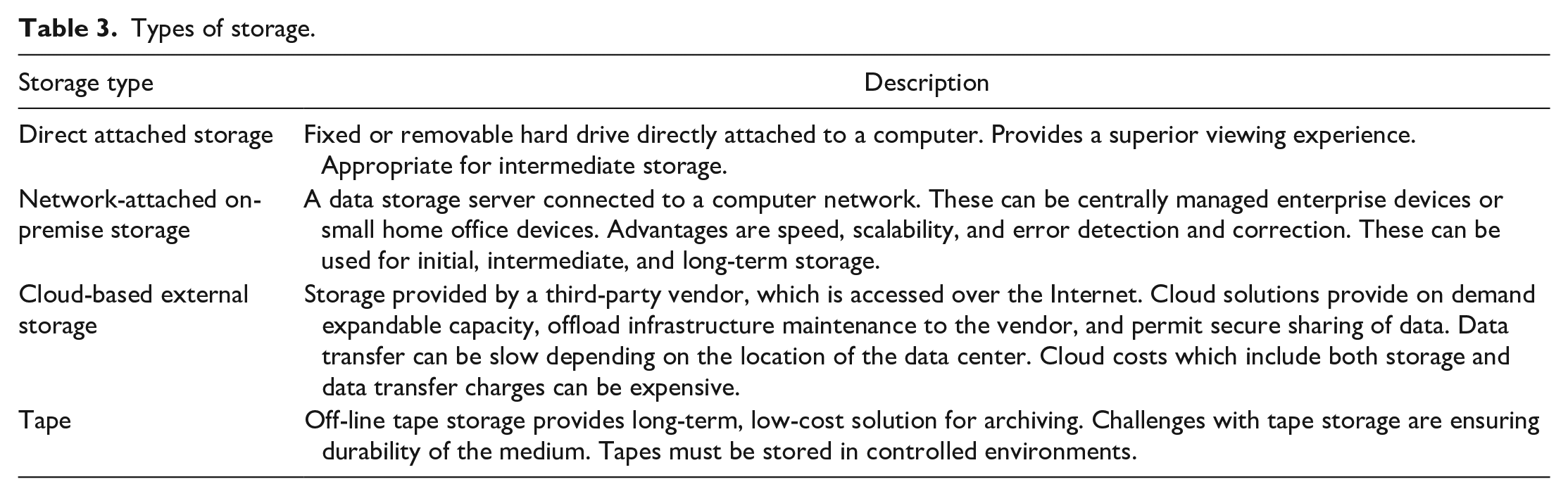

Whichever type of storage is used, the general considerations summarized in Table 4 should be borne in mind.
Data storage considerations.
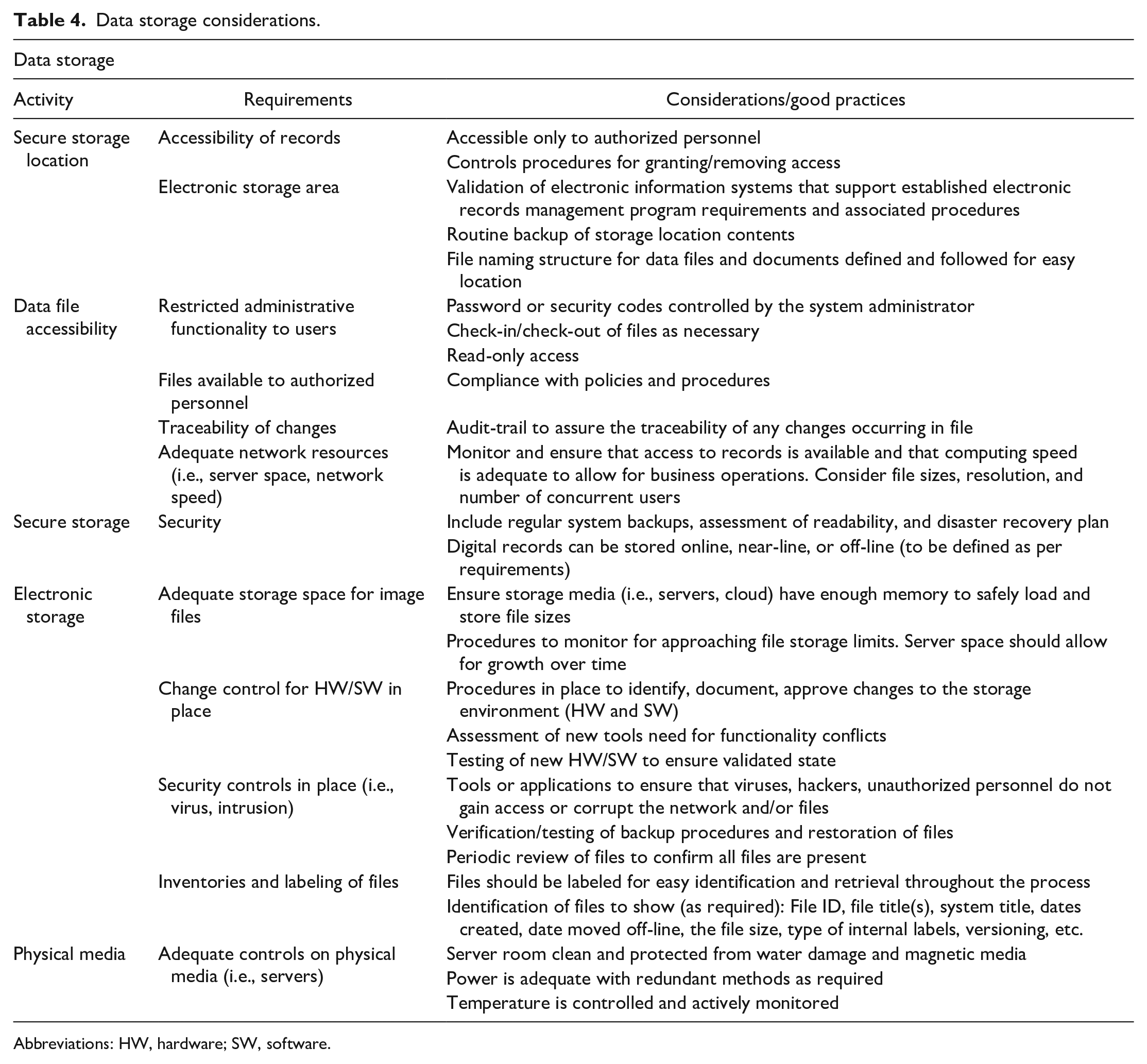
Abbreviations: HW, hardware; SW, software.
Technical Considerations
Regardless of the types of storage, technical considerations for each tier of storage include data transfer rates, capacity, integrity, and cost. It is beyond the scope of this white paper to cover all aspects of these topics. A general overview is presented here, and the reader is advised to discuss these considerations with their IT organization.
Capacity
The storage space required for a single toxicology study can be variable. Individual WSI range from a few hundred megabytes (MB) to single-digit gigabyte (GB) in size, depending on tissue size, scanning resolution, and image format. The number of images scanned for a study varies from study design. Tables 5 and 6 show some typical storage requirements based on study type. 23
Examples of some use cases in toxicology studies commonly conducted in the pharmaceutical industry.
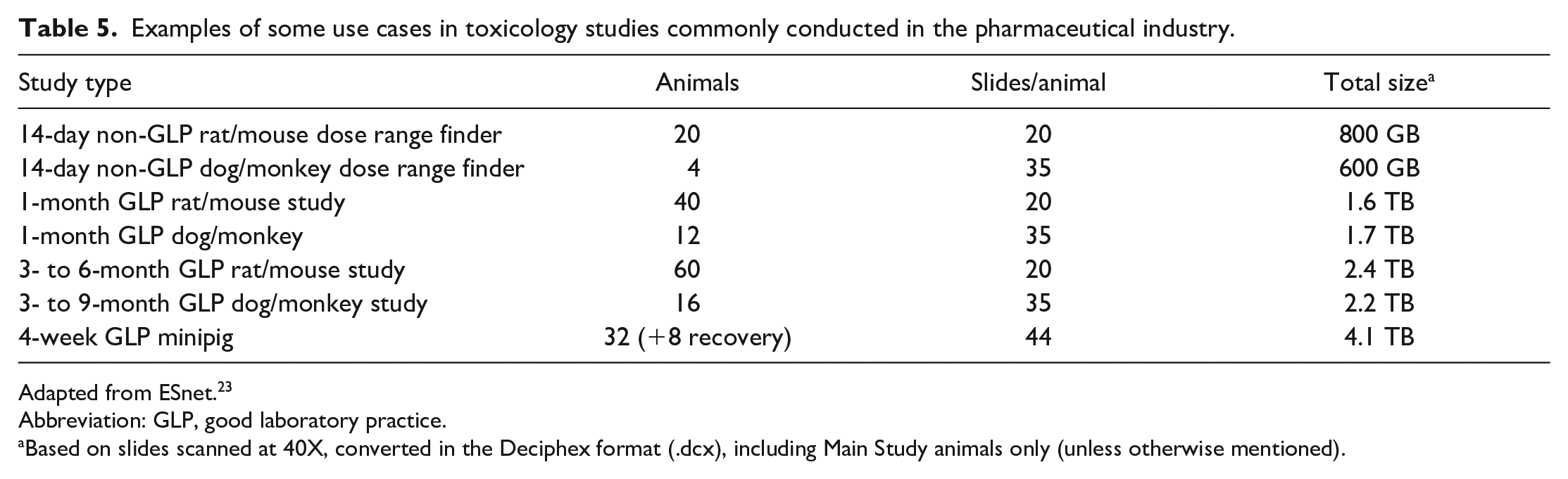
Adapted from ESnet. 23
Abbreviation: GLP, good laboratory practice.
Based on slides scanned at 40X, converted in the Deciphex format (.dcx), including Main Study animals only (unless otherwise mentioned).
Examples for some use cases for image analysis in multigeneration studies according to OECD 443 and mechanistic cell proliferation studies in the chemical industry.
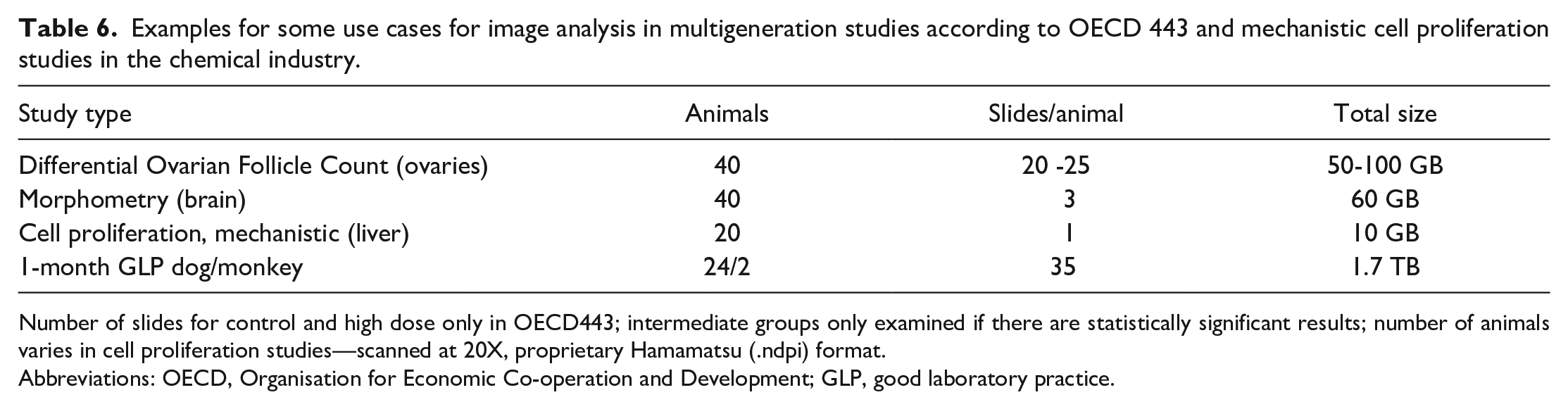
Number of slides for control and high dose only in OECD443; intermediate groups only examined if there are statistically significant results; number of animals varies in cell proliferation studies—scanned at 20X, proprietary Hamamatsu (.ndpi) format.
Abbreviations: OECD, Organisation for Economic Co-operation and Development; GLP, good laboratory practice.
Transfer Rates
Data transfer rates impact workflow and viewing experience in a digital pathology system. Transferring whole studies between two locations can take from a few hours to a few days (Table 7).1,2,23 Rates are dependent on network speed and storage read/write rates.
Examples of data transfer duration/network speed. 2
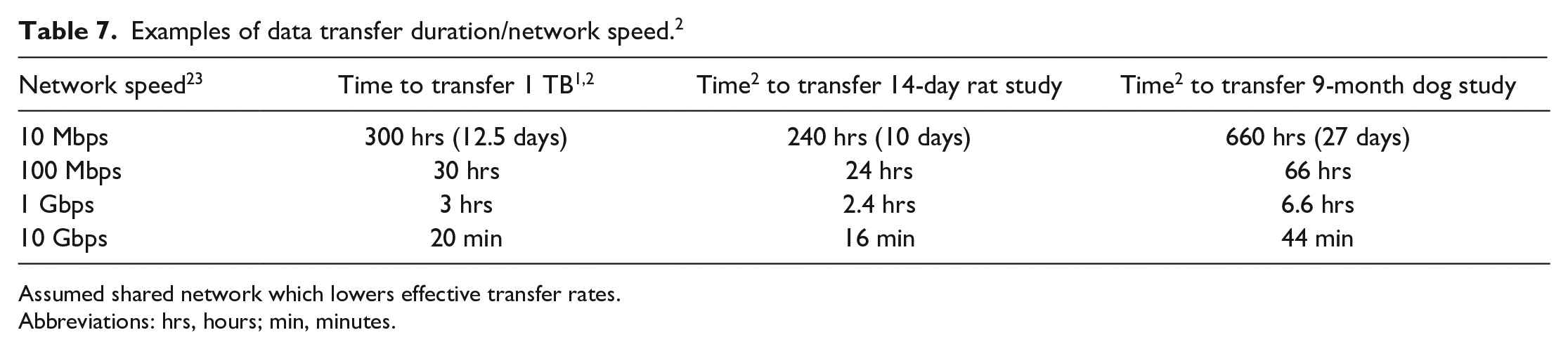

Assumed shared network which lowers effective transfer rates.Abbreviations: hrs, hours; min, minutes.
When computing transfer rates, it is important to use the same units of measure. File sizes are referenced in bytes (B), while network speeds are often quoted in bits(b) per second (bps). A byte is 8 bits resulting in 1 bps being 8 times slower than 1 bps. A second consideration is network throughput. Networks are shared among many users and computer systems. Throughput is the actual rate data transferred on the network. This could range from 10% to 90% of the network speed depending on network activity. In addition, the slowest device in the transfer chain will ultimately determine the transfer rates. For example, though USB3 offers a speed of 5 Gbit/s (625 Mbps), a disk with 250 Mbps write speed limits the transfer.
Data transfer rates are the critical factors for best viewing experience required for primary read and peer review. A pathologist needs to navigate and switch between images rapidly. Rapid selection and movement are best achieved when images are on fast, local disk or a networked storage device located at the pathologist work site. Images stored in a remote data center or in the cloud load noticeably slower than those locally stored. A downside of direct attached storage is their relatively small capacity and can only handle a few studies. Local network storage should be considered when more than a few studies need to be accessed.
Data rates have little impact on initial and long-term storage. Transfer rates only add a modest amount of time to the scanning process.
Additional data transfer considerations are listed in Table 8.
Steps involved in electronic data transfer and related requirements.
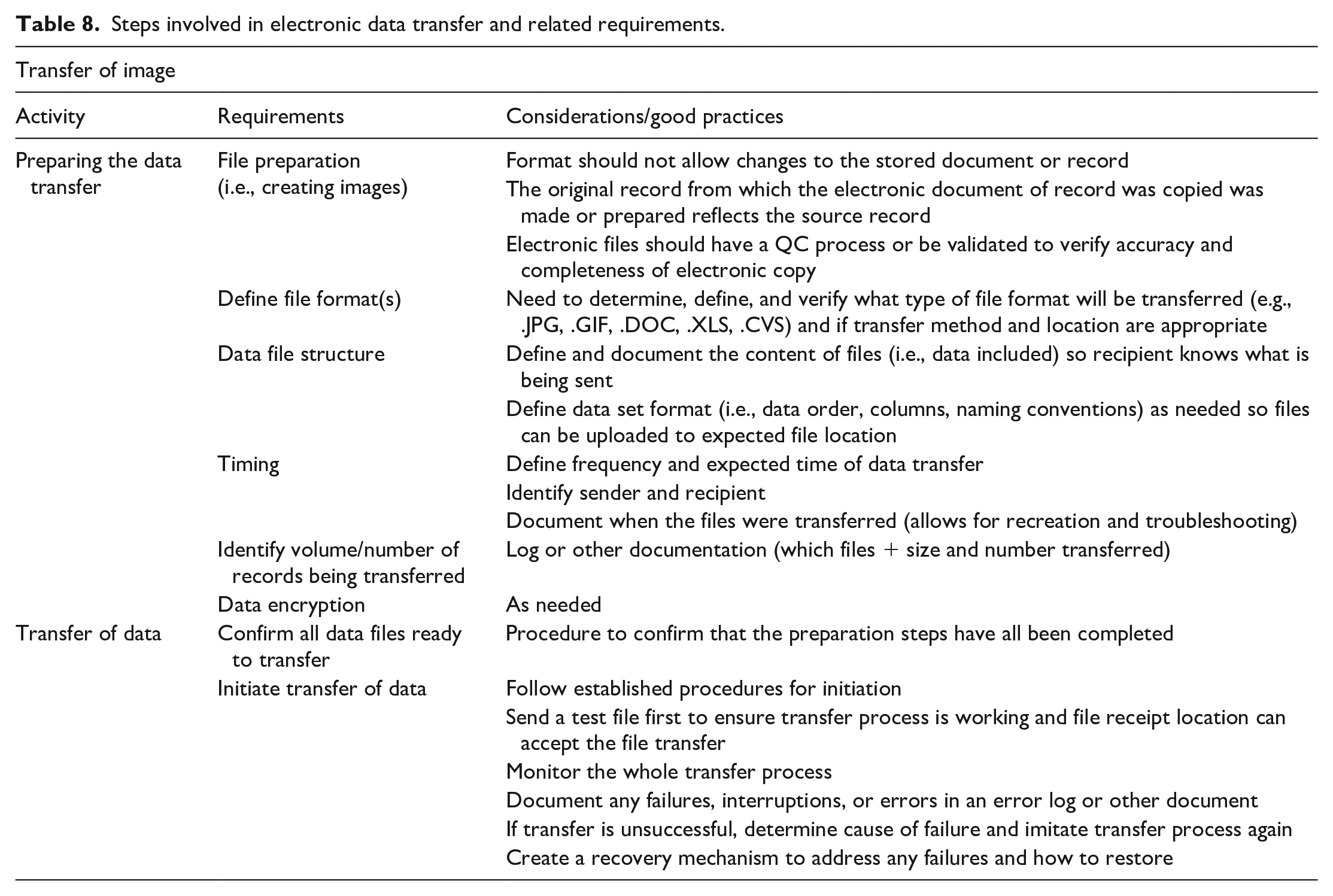
Integrity
When transferring data between storage locations, steps must be taken to ensure data integrity and completeness. When transferring files, there are two failure modes to consider: (1) the data are corrupted in transit or (2) data are corrupted when it is read or written. Encrypted transfer protocols such as sFTP and HTTPS verify the data integrity of the transmission but do not verify that the data were read or written correctly. Windows operating system has capabilities to verify the data are written correctly but does not check for corruption during transmission. Protocols such as FTP and HTTP have no integrity checks.
The checksums are used to ensure data integrity across the entire transfer process. A checksum is a unique number which is computed based on the data within a file. When files are transferred, the checksum of the destination file is compared with that of the source file. If the checksums differ, then the destination file was corrupted. Common checksums are MD5 and SHA.
Archival
Archival of WSI used for non-data-generating contemporaneous peer review should not be required per GLP guidelines; each individual institution should reach a decision about the trustworthiness of these WSI. All materials needed to reconstruct the study must be archived. It has been agreed among the scientific community that the following elements must be retrieved 1 :
The glass slides and signed peer review statement, for a digital contemporaneous peer review where primary review was done on glass slides.
The WSI, metadata, and associated data (if applicable), when a retrospective peer review is conducted.
The glass slides, WSI, metadata, and associated data when primary assessment is done on digitized slides.
Annotations or overlays generated for morphometric or IA purposes. Any marking done by the study pathologist or peer reviewer with a view to facilitating discussions and consensus (e.g., pin marking of focal lesions) is considered “working notes” and is not critical to study reconstruction.
The overall recommendations for electronic data archival are summarized in Table 9.
Recommendations for data archival and retrieval.
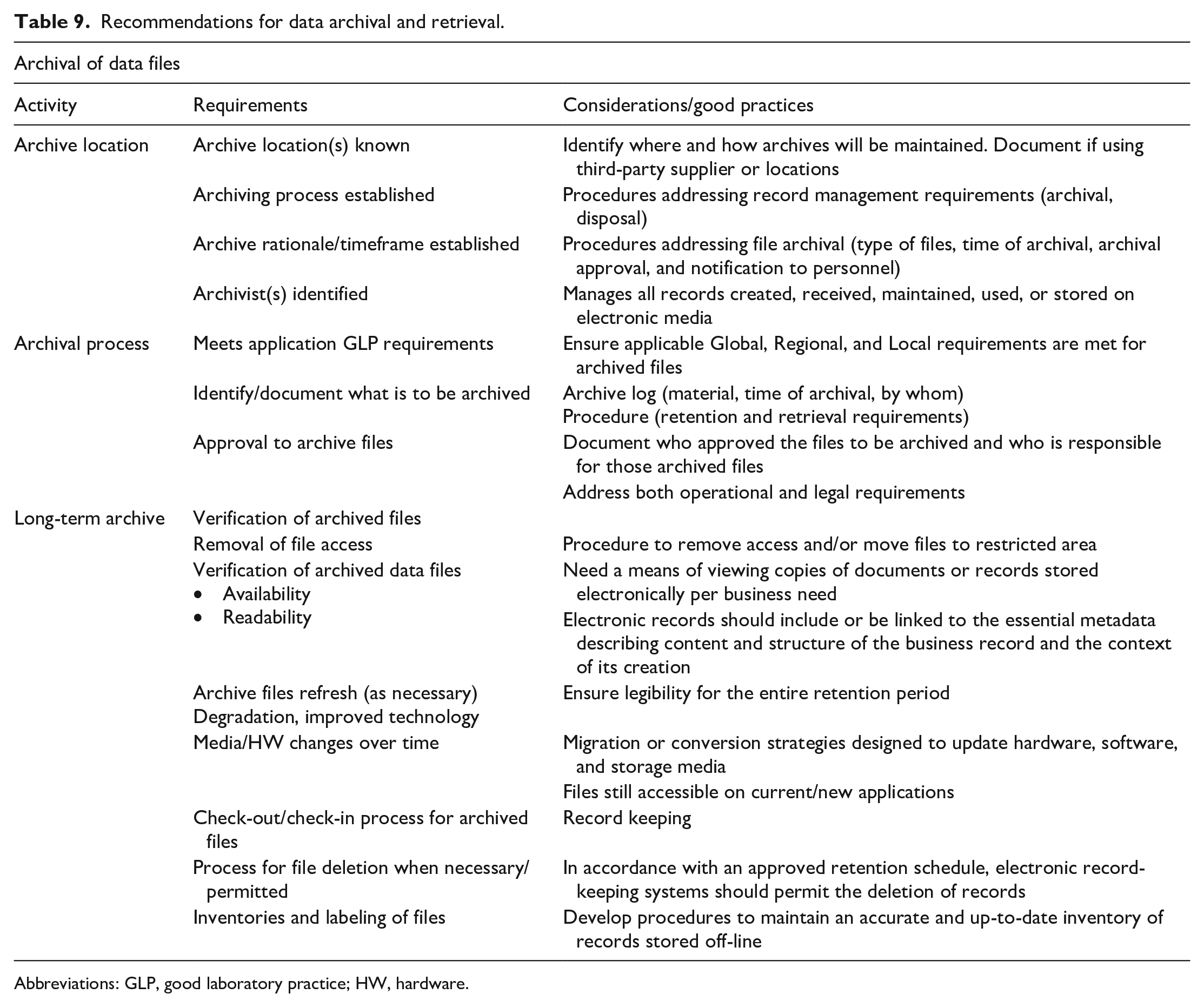
Abbreviations: GLP, good laboratory practice; HW, hardware.
As mentioned above, central to GLP-compliant archival is the concept of faithful replication of the glass slide, as defined in the 7th ESTP Workshop paper, 1 and the ability to recreate a study. It is important to bear in mind that some digital slide viewers may provide functionalities facilitating the pathologist’s viewing and navigating experiences; these sometimes imply conversion of the native file to a proprietary format.
To ensure legibility of the files over the retention period, WSI format should ideally be vendor-independent. The compatibility with image viewers should be assessed.
The whole pixel chain should be considered during validation, including cases where WSI need to be de-archived for re-evaluation. In case of format conversion across the data transfer chain, it is key that the validation process clarifies which file(s) will be archived. Whole slide images should be archived in a format that (1) enables the system lifecycle management plan 6 and (2) is a faithful representation of the glass slide. As the validation of the pixel chain has already proven that the WSI file format used for assessment by the pathologist is a faithful replica of the glass slide in the validated viewer, archiving this format is the easiest. However, other formats may be archived as long as the validation process showed that these were faithful replicas to each other and to the glass slide when visualized in the validated viewer. The computerized system that allowed the creation and visualization of the WSI will need to be maintained in a validated state at the institution. 6
For GLP archival, tape and local servers (using their own disk storage) are acceptable storage forms. For archiving on removable media, multiple copies of the data should be considered. Regardless of the media used for archiving, processes should be in place to verify readability over time. In addition, for long retention periods considerations should be made for migrating archives off legacy technology over time.
The IT infrastructure supporting long-term storage of images files should guarantee that the integrity of data is maintained over time and allow access to data in an established timeframe for retrieval, for example, in the event of regulatory inspection. The physical address of data centers hosting digital slides and their associated metadata may be required by regulatory agencies, which is being considered as an issue for broader discussion at OECD level by the Belgian GLP Compliance Monitoring Authority (Martijn Baeten, GLP Coordinator at Sciensano [Compliance Monitoring Authority for Belgium], personal correspondence with Anja Gilis, October 5, 2021).
Secured access and GLP compliance, including disaster recovery plan, should be specified in the service-level agreement when WSI storage and archival is taken care of by a third party, for example, software-as-a-service and cloud storage provider.
At every step of storage during active use, security should be maintained by ensuring password-protected access to authorized personnel (administrator, user, auditor) with the adequate profile, this list being reviewed and adapted regularly. After archival, requirements as described in OECD 17 6 should be met.
All the details related to data storage and archival should be specified in the study protocol and/or relevant SOPs, giving elements as to the location of storage, retention and deletion policy, and possible transfer of ownership. These recommendations are summarized in Table 9.
Implementation
In essence, upon implementation, it must be possible to demonstrate through a validation report that within the existing infrastructure, the new digital histopathology workflow is fit for purpose, that is, validated and controlled for the intended use. By “controlled” it is meant that risks have been identified and mitigations have been put into place as needed to ensure data integrity and trustworthy study outcomes. The best way to achieve this is to document in SOPs and/or protocol the complete end-to-end process flow and for each step to identify, assess (impact and likelihood), and mitigate (or accept) any risks. Such a risk-based approach with focus on data integrity fits the emphasis of OECD17 6 and the new OECD advisory document on GLP data integrity, 24 where the key message is for users to demonstrate that a computerized system is suitable throughout its life cycle and for its intended purpose. The documented risk-based life cycle can be used both for internal purposes (e.g., as basis for procedure and training, upon change control, as a basis for QA audit scoping) as well as to provide evidence of oversight and compliance with external parties (clients of CROs or monitoring authorities).
Throughout implementation, compliance with international GLP OECD1 25 must be considered mandatory and applicable guidance documents such as OECD15 26 and OECD16 27 should be taken into account and built into the process flow. If several parties are involved in the workflow (i.e., CRO and Sponsor), all parts of the chain should guarantee the overall GLP footprint. Implementation of such a GLP-compliant workflow for primary assessment and contemporaneous peer review was described recently. 4
If a significant alteration is introduced into a validated system, for example, the inclusion of an entirely new hardware or software component or a software update, the impact and potential additional validation requirement is to be assessed via a change control process in consultation with the QAU. Consideration of the need for change control is also applicable for internal control/periodic review, which evaluates and records for correct functioning of the complete system at predetermined intervals to keep it in a validated state. Like for the change control, the decision to perform periodic review is risk-based. For example, as it constitutes a higher business and GLP risk, one can decide to perform periodic review for digital primary evaluation but not for digital peer review. During a digital peer review, the reviewing pathologist constantly compares his own histopathological assessment on WSI with the study pathologist’s assessment, which in itself constitute a kind of “internal control/periodic review.”
Certain specific challenges (e.g., use of cloud storage for archiving, use of AI) related to digital pathology have not yet been covered by any GLP regulatory guidance document. Risk-based approaches should be followed, keeping trustworthiness of study conclusions and reconstructability as central drivers and keeping in mind that even in case of subcontracting some part of the digital pathology process (hosting), the study director remains responsible for the GLP compliance of the entire study. Confidence in the quality of the WSI submitted for assessment should be the main driver and any needed action (e.g., rescanning or going back to the glass slide) should be taken to ensure the quality of the pathology interpretation.
Worth mentioning are the potential challenges with archiving of GLP data on cloud servers. In short, GLP regulations require that the physical location of the storage location is identified such that the actual protection measures can be inspected. Some countries (e.g., Germany) only allow archiving of GLP data in GLP-certified facilities, which is often not the case for data centers. 1 Until OECD GLP guidance on this topic is available, this challenge should be carefully considered and may be worthwhile to discuss with local GLP Monitoring authorities (Martijn Baeten, GLP Coordinator at Sciensano [Compliance Monitoring Authority for Belgium], personal correspondence with Anja Gilis, October 5, 2021).
Table 10 can be used as a basis for a checklist on what to take into account before and after implementation of GLP digital pathology.
Key items to consider before and after implementation of GLP digital pathology.
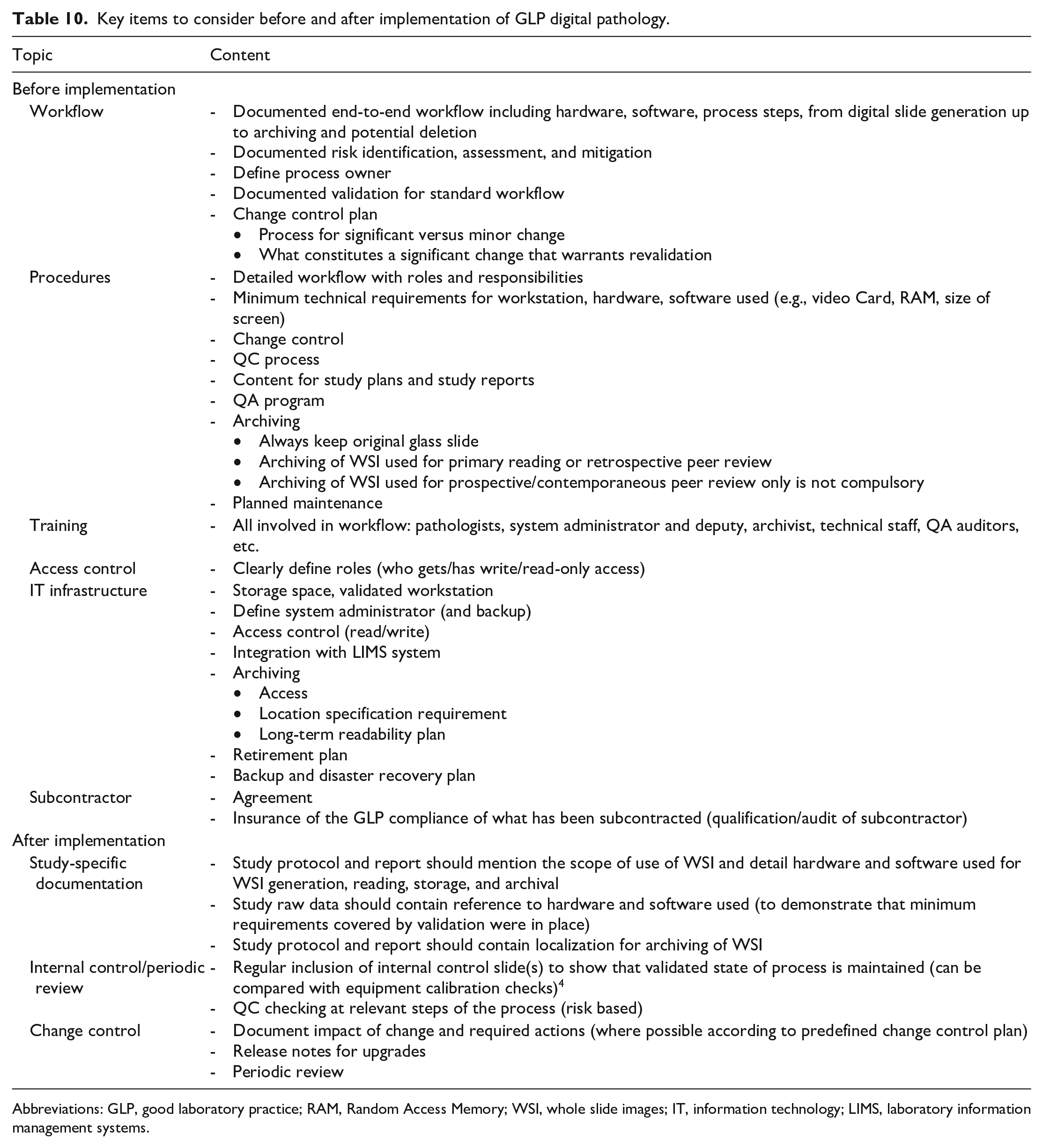
Abbreviations: GLP, good laboratory practice; RAM, Random Access Memory; WSI, whole slide images; IT, information technology; LIMS, laboratory information management systems.
Conclusion
Despite some specific complexities, which certainly do not outweigh the benefits of implementing GLP digital toxicologic pathology, the general validation, qualification, and implementation path is highly similar to any other GLP process. The pixel pathway needs to be validated from end-to-end and is highly dependent on the use cases applying to digital pathology within and between institutions. The IT requirements specific to each part of the pixel pathway need to follow vendor’s and institution’s IT requirements to optimize the end user experience. The close collaboration of members from a multidisciplinary team composed of pathologists, IT, and regulatory (QA) experts will assure the success of digital pathology in GLP preclinical toxicology.
Footnotes
Acknowledgements
The following colleagues are gratefully acknowledged for their great contributions during the preparation and participation in the workshop: Joel Aguettant, Sanofi; Andreas Popp, AbbVie; Patric Burkhardt, Roche; Pierre Maliver, Roche; Charlotte Dalsgaard, Novo Nordisk; Celine Contet, Sanofi; Christian Rapp, Merck; Gundi Mueller, Merck; Jan Baumann, Merck; Kevin Keenan, GSK; Christoph Kramer, Novartis; Linzi Gillbanks, Covance; Ronald Menner, Pfizer; Markus Haak, Roche; Martin Wolf, AbbVie; Matt Jacobsen, AstraZeneca; Roberta Loughins, GSK; Thamas Kallal, AbbVie; Timo Daniel, AbbVie; Oliver Turner, Novartis; Maarten Vergauen, Janssen; Yaniv Cohen, Roche; Ellen Kneer, Boehringer Ingelheim; Fra Lugarini, Janssen; Nicolas Rudinger, Bayer; Kathleen Stango, Novartis; Wolfgang Kalsch, Boehringer Ingelheim.
Authors’ Note
This article is a product of a European Society of Toxicologic Pathology (ESTP) Working Group commissioned by the Executive Committee (EC) of the ESTP. It has been reviewed and approved by the Committee of Scientific Standards and EC of the ESTP. The opinions in the article are those of the authors and do not necessarily represent the policies, positions, or opinions of their respective agencies and organizations. Mention of trade names or commercial products does not constitute endorsement or recommendation for use. This article has been reviewed and endorsed by major toxicologic pathology organizations, including the Society of Toxicologic Pathology, the British Society of Toxicological Pathology, the Dutch Society of Toxicologic Pathology, the French Society of Toxicologic Pathology, the Society of Hungarian Toxicologists, International Academy of Toxicologic Pathology, and the Japanese Society of Toxicologic Pathology.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
