Abstract
The bone marrow is an important site for assessment of the hematopoietic toxicity of new drug candidates. Here, we extended our previous work, where we developed a computer algorithm to automatically quantitate overall bone marrow cell density by analyzing digitized images of standard hematoxylin and eosin (H&E) slides of rat bone marrow and further evaluated the capability to quantify myeloid: erythroid + lymphoid (M:EL) ratio and megakaryocyte cell density. We tested the algorithm in a toxicity study, where rats were dosed with two molecules known to affect bone marrow composition, monomethyl auristatin E, and a Bcl-xL inhibitor. The image analysis method detected significant changes in M:EL and megakaryocyte number that were either not found or semiquantitatively described by manual microscopic observation of the same slides. The image analysis results were consistent with other more established but time-consuming methods that measure changes in bone marrow cell composition: smear cytology, flow cytometry, and microscopic assessment. Our work demonstrates the feasibility of a rapid and more quantitative assessment of changes in bone marrow cell lineage composition using a computer algorithm compared to microscopic examination of H&E-stained bone marrow sections.
The bone marrow is a major site of toxicity in drug safety assessment due to both on-target and off-target effects, in addition to unanticipated impact of drug targets that directly or indirectly affect hematopoietic cells (Everds and Tarrant 2013). Drugs that target the immune system may alter various molecular pathways leading to abnormal changes in specific cell populations (Reagan et al. 2011). Detection of imbalances in hematopoietic cell populations both characterizes the toxicity to specific cell populations and provides a mechanistic understanding of the toxicity of the drug candidate under investigation.
Typically, only relatively large shifts in the composition of cell populations may be detected visually by manual microscopic evaluation of standard hematoxylin and eosin (H&E) slides. Bone marrow smear cytology allows evaluation of mature, progenitor, and precursor cell populations of different cell lineages, but the data are semiquantitative, time-consuming to obtain and have high operator variability (Bolliger 2004).
If the loss of specific cell types is suspected after visual inspection or based on known activity of the drug, additional immunohistochemistry (IHC) stains on serial sections may be used to semiquantitatively evaluate cell types of interest (Rehg, Bush, and Ward 2012).
The most quantitative method currently available for measuring bone marrow subpopulations is flow cytometry, which is labor-intensive (Haley et al. 2005), and complex, as a panel of cell specific cell markers must be assembled. In addition, bone marrow must be aspirated at the time when animals are terminated, and therefore, retrospective assessment is impossible (Reagan et al. 2011).
We sought to develop a routine screening procedure that could be easily and quickly applied to standard H&E slides and could detect meaningful perturbations in the proportion of major bone marrow cell lineages. This would inform whether further specific and sensitive, but more time-consuming hematopoietic toxicity investigations are warranted. To this end, we extended our previous work where we developed an automated image analysis algorithm that could quantify bone marrow cell depletion (Kozlowski, Brumm, and Cain In Press). We added the capacity to enumerate relative numbers of differentiated hematological cell lineages including megakaryocytes, myeloid cells, and a combined group of erythroid and lymphoid cells.
In order to evaluate the image analysis method on H&E sections, we performed single-dose acute toxicity studies in rats using monomethyl auristatin E (MMAE), a microtubule inhibitor (Donaghy 2016; Stagg et al. 2016; Junutula et al. 2008) that causes bone marrow cytopenia and altered M:E ratio, or a Bcl-xL inhibitor that affects megakaryocyte differentiation and proliferation (Kile 2014). The slides were visually assessed by a pathologist, and the algorithm was applied to the scanned images. The image analysis data were compared to the manual microscopic evaluation of the sections by a pathologist and to the industry gold standards for advanced bone marrow assessments including microscopic cytology smear evaluation, manual counting on IHC of the megakaryocyte-specific marker von Willebrand’s factor (VWF; Rehg, Bush, and Ward 2012), and immunophenotyping by flow cytometry of bone marrow aspirates (Reagan et al. 2011). We further performed image analysis on the VWF IHC slides to test whether it would provide more reliable estimates of megakaryocyte relative size and numbers compared to H&E staining.
While further work is necessary to increase the sensitivity of our method, our investigation shows that image analysis could provide considerably more information about bone marrow cell composition than currently obtainable from H&E images by manual microscopic evaluation and requires little manual effort, making it a promising screening tool for preclinical safety assessment studies.
Method
Animal Handling and Treatment
The animal studies were conducted at Charles River Laboratories Inc., Reno, NV. This study complied with all applicable sections of the Final Rules of the Animal Welfare Act regulations (Code of Federal Regulations, Title 9), the Public Health Service Policy on Humane Care and Use of Laboratory Animals from the Office of Laboratory Animal Welfare, and the Guide for the Care and Use of Laboratory Animals from the National Research Council. Experimental data were obtained under Testing Facility Study No. 20066373 following approval of the procedures by the Testing Facility’s Institutional Animal Care and Use Committee.
A total of 40 Crl: CD Sprague-Dawley rats (20 males and 20 females) approximately 8 weeks of age and weighing 286 to 403 g were obtained from Charles River Laboratories, Hollister, CA. Following randomization but prior to dosing, animals were group housed (2 animals of the same sex and the same dosing group together) in solid bottom cages with appropriate bedding, with enrichment such as a hiding device and a chewing object. Environmental conditions were set to maintain 18°C to 26°C and relative humidity of 30–70%. The room had a minimum of 10 air changes/hr and had a 12-hr light/12-hr dark cycle. Water and PMI Nutrition International Certified Rodent Chow No. 5CR4 (14% protein) was provided
The animals were divided into 5 treatment groups (4 animals/sex/group). For groups 1 to 4, rats were administered MMAE (synthesized at Genentech) at 0 (0.9% Sodium Chloride), 0.03, 0.06, and 0.2 mg/kg via intravenous injection (slow bolus) to the tail vein once on day 1. For group 5, Bcl-xL inhibitor (Abcam, A-1331852, selectively disrupts Bcl-xL-Bim complexes) was dosed at 20 mg/kg once daily by oral gavage from days 1 to 3.
Dose volume for each animal was based on most recent body weight measurement. After dosing, the rats were individually housed. Animals were observed twice daily. One animal administered 20 mg/kg Bcl-xL inhibitor (group 5) was found dead on day 2. All other animals survived to scheduled euthanasia on day 4. Animals were euthanized under deep anesthesia induced with isofluorane inhalation.
IHC
At necropsy, sternum and left femur/tibial cancellous and medullary bone tissue were collected, fixed in formalin, de-calcified, paraffin-embedded, and sectioned at 5 μm. Note that because the left femur was also used to extract bone marrow for smear cytology assessment, there was consequently less tissue available for slide preparations than would be standard for a femoral preparation.
For each animal, one section of femoral/tibial bone and 3 serial sections of sternum were stained with H&E, each set of sections in a single run using the Tissue-Tek Prisma system (Sakura Finetek, Tokyo, Japan). One additional serial section of sternum was stained for VWF. Sections were deparaffinized with Sakura DRS autostainer and pretreated with 0.1% pronase solution at 35°C for 30 min. Slides were washed and stained with 10.0 μg/mL primary antibody Rb a-VWF AN-G1072 (ab6994, Abcam, Cambridge, UK; 60 min) or control Rb IgG (AN-G962 Vector, Burlingame, CA), followed by Rb Envision+ (K4003, Dako, Santa Clara, CA) detection agent (30 min), and washed for 5 min in 3,3’-diaminobenzidine (DAB) solution (926504, Biolegened, San Diego, CA) in the Thermo Scientific IHC system (ThermoFisher Scientific, Waltham, MA). Slides were counterstained with hematoxylin in the Sakura DRS autostainer.
Bone Marrow Extraction and Flow Cytometry
Bone marrow was extracted from each rat’s right femur using a centrifugation method and counted using the Guava cytometry system (Millipore Sigma, St. Louis, MO). Samples were resuspended at a concentration of 10 million cells/mL in phosphate buffered solution, and 50 µL aliquots (∼5.0 × 105 viable cells/sample) were stained using Live/Dead Cell Stain kit—Far Red Dye (Thermo Fisher Scientific, Waltham, MA) according to manufacturer’s recommendations. Samples were then washed once with Stain Buffer (FBS; BD Biosciences, San Jose, CA) and Fc blocked by incubation with purified mouse anti-rat CD32 (D34-485; BD Biosciences) in stain buffer for 10 min on ice and protected from light. Subsequently, samples were stained for 45 min with combinations of BV421-conjugated anti-CD61 (2C9.G2, BD Horizon), APC-conjugated anti-rat erythroid cells (HIS49, BD Pharmingen), PE-conjugated anti-CD45 (OX-1, Thermo Fisher), PE-Cy7-conjugated anti-CD45R (B220; HIS24, Thermo Fisher), PerCP-eFluor710-conjugated anti-CD11b/c (OX-42, Thermo Fisher), or corresponding isotype controls. Following incubation, samples were washed twice with stain buffer, resuspended with 300 µL of stain buffer, and 250 µL of sample was transferred to a 96 well v-bottom plate for acquisition on a BD FACSCanto II Flow Cytometer utilizing DIVA 6.1.3 software. Fluorescence activated cell sorting (FACS) data analysis was performed using FlowJo Software v10.3 (FlowJo, Ashland, OR). Cell populations were first gated on viability and subsequently defined as B cells (lymphocytes): CD45+/CD45R (B220)+, myeloid: CD45+/CD11b/c+, erythroid: CD45dim/−/erythroid (HIS49)+, or megakaryocytes: Forward Scatterhigh/CD61+. The antibodies do not distinguish between mature and immature (progenitor and precursor) cells. Ratio values for comparison to image analysis were calculated by myeloid cell population divided by the sum of erythroid and lymphocyte populations, M:EL = (myeloid/(erythroid+B cells)). Ratio values for comparison to smear cytology were calculated by myeloid cell population divided by the erythroid populations, M:E = (myeloid/erythroid).
Bone Marrow Smear Cytology
Bone marrow cytology smears were prepared from bone marrow cells using a paint brush method, collected from the left femoral medullae of each animal. Slides were stained with a modified Wright-Giemsa stain using an automated slide stainer. Bone marrow smears were evaluated microscopically by counting 300 cells, and an M:E ratio was determined for each slide and results recorded. One animal each from the MMAE 0.03 mg/kg and MMAE 0.06 mg/kg groups and 2 animals from the Bcl-xL 20 mg/kg group had an unusable smear slide due to poor cell preservation. The bone marrows of the 0.20 mg/kg MMAE dose group were severely depleted of cells, and extracted smears were therefore all too cell-poor for evaluation. Thus, 12 of the 40 smear slides were inevaluable.
Microscopic Evaluation
Megakaryocyte numbers were counted by a pathologist in 3 random 40× magnification fields for groups 1 and 5. For IHC evaluation, DAB positive VWF+ cell counts were enumerated as the average number of cells from 3 fields at 40× magnification.
Whole-Slide Imaging and Automated Image Analysis
All slides were scanned at 20× magnification (0.46 μm/pixel) using the Nanozoomer HT 2.0 Slide scanner with an 8-bit camera (256 levels of intensity).
H&E Evaluation and Cell/Nuclei Detection
The algorithm for cell detection on H&E images is described in the companion paper (Kozlowski, Brumm, and Cain In Press). Briefly, using Definiens Developer (Definiens AG, Munich, Germany; Baatz, Zimmermann, and Blackmore 2009), images were analyzed in red, green, and blue color space. The in-built Definiens algorithm “multiresolution segmentation” (a way to divide up an image by grouping together cohesive pixels) was, with scale parameter 30, used to divide the images into smaller objects. These objects were separated into categories based on visual characteristics including area, shape features such as roundness, and pixel intensities, using the Definiens “evaluation class” (a method to manually select object features and their weights that influence how objects are classified). Areas of bone were distinguished from marrow using low (0.4×) magnification. At higher (20×) magnification, cells and nuclei within the marrow were separated as objects using multiresolution segmentation with scale parameter 85. For some marrow cells, only the nuclei were identified (lymphoid, erythroid, or myeloid cells); and in the case of mature red blood cells and megakaryocytes, the entire cell was detected (Figure 1). The 2-D cross-sectional area and average pixel brightness of all identified detected cells or nuclei were measured automatically by in-built Definiens algorithms which quantitate the pixel properties in a classified object and exported for each slide, using Definiens “Export Object Data” function, for further analysis. The entire analysis including the bone marrow cellularity measurements (Kozlowski, Brumm, and Cain In Press) as well as the analysis of subpopulations took approximately 40 min of computing time per slide on servers running Windows 7 Definiens Developer Server v2.6. However, it only required ∼30 sec per slide of manual intervention, where slides were loaded onto the scanner, and digitized images were checked for quality before automated analysis.

Image analysis tile views from different treatment groups. Representative 20× tile view of H&E (A–C) and 10× tile view of immunohistochemistry with VWF (G–I) sternal bone. Examples are shown from an animal in the control group, MMAE high dose (0.20 mg/kg) group, and Bcl-xL inhibitor (20 mg/kg) group. Image analysis overlay over the original H&E image, showing a color coding for what the identified object was classified as (D–F). Gray: bone, green: megakaryocyte, red: mature red blood cells, light blue: analyzed marrow, white: empty space/adipose, orange: myeloid cell nucleus, and blue: erythroid/lymphoid cell nucleus. Image analysis overlay for VWF (J–L). Gray: bone or empty space, red: megakaryocyte, and light blue: analyzed marrow. H&E = hematoxylin and eosin; VWF = von Willebrand’s factor; MMAE = monomethyl auristatin E.
Obtaining Megakaryocyte Number and M:EL Ratio from H&E Images
The area and pixel brightness of all detected objects were plotted, producing a graph resembling flow cytometry data (Figure 2). Megakaryocytes separated from other cells due to their relatively large area. Other objects were more difficult to classify. Consistent with known morphologic characteristics of different cell types, dark and compact nuclei were attributed to lymphocytes and erythroid cells, and larger, brighter nuclei to myeloid cells (Travlos 2006). The myeloid and erythroid or lymphoid lineage cells did not clearly separate on the image analysis export data (Figure 2); therefore, a threshold for area and brightness was set to categorize them into the two cell populations, and the same threshold values were used to analyze all the images from slides in that experiment. Since the threshold was applied in the same way for all samples, it was possible to estimate the change in ratios of M:EL cell populations relative to the control animals. The algorithm does not distinguish between mature and immature cells. Overall, approximately 150,000 to 300,000 myeloid or erythroid cells and 1,000 to 2,000 megakaryocytes were analyzed per slide.
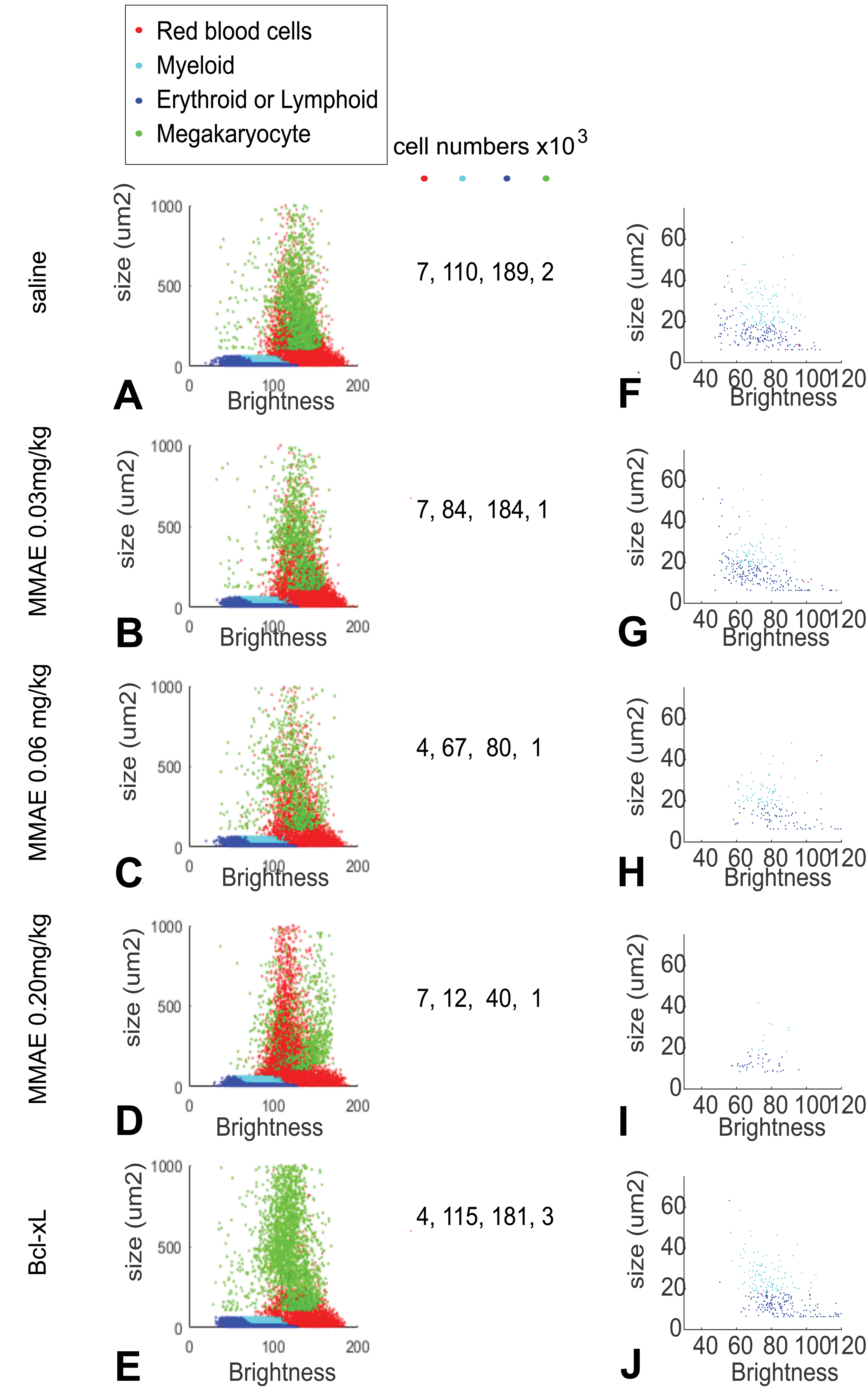
Image analysis method for cell subpopulation classification and strategy for M:EL estimation. Left (A–E) panel: each data point in the graph represents a detected cell or nucleus “object” found in the marrow of a single sternal bone marrow H&E section from a representative animal in each treatment group. The average brightness (the averaged value of the red, green, and blue color layers in the image, see Kozlowski, Brumm, and Cain In Press) of the object is plotted on the X-axis, and the size of the object in μm2 is plotted on the Y-axis. On all graphs in the left panel, every cell detected on the slide is represented. Because the megakaryocytes had a larger variation in sizes, they appear as a large smear compared to the myeloid or erythroid cells, but the megakaryocytes are much fewer in number. The actual number of cells detected is indicated on the right of this panel. For example, for the control animal A, there were ∼7,000 red blood cells, ∼110,000 myeloid nuclei, ∼189,000 erythroid or lymphoid nuclei, and ∼2000 megakarcyocytes detected. Note that the myeloid nuclei and erythroid or lymphoid nuclei numbers are not exact and are only meant to be interpreted in treatment groups relative to control groups. Accordingly, we can deduce that for animal B, where there are ∼84,000 myeloid nuclei, but ∼184,000 erythroid or lymphoid nuclei detected, there was a relatively myeloid-specific cell depletion compared to control. But we cannot be certain that there are ∼84,000 myeloid nuclei in this bone marrow sample. The right panel (F–J) displays zoomed in views of the graphs in the left panel and just show 1/1,000 cells of those shown in the left panel. It is provided to emphasize how difficult it is to resolve the myeloid and erythroid or lymphoid cell populations; the cyan and dark blue dots form one large cloud. But the brightness and size thresholds are set the same way in all the dose groups, and thus relative cyan: dark blue ratio is used to estimate the relative change in M:EL ratio. M:EL = myeloid: erythroid + lymphoid.
IHC Evaluation
VWF-stained slides were analyzed at low magnification (0.4×) to identify marrow using multiresolution segmentation at scale parameter 50, followed by a Definiens evaluation class to classify marrow, based on brightness and standard deviation of the red layer. At higher magnification (10×), marrow was distinguished from bone by multiresolution segmentation with scale parameter 100 and by applying the same evaluation class for marrow as in the low-resolution classification (Figure 1). A “brown” image layer was defined as: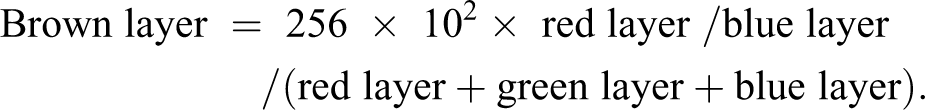
DAB+ cells were identified by a pixel intensity threshold on this brown image layer. Analysis took approximately 15 min per slide of computing time on servers running Windows 7 Definiens Developer Server v2.6.
Statistical Analysis
MATLAB (version R2016a; Mathworks, Natick, MA) software was used to perform all statistical analyses. Statistical differences between treatment groups and the control group were performed by Student’s t-tests. Pearson’s cross-correlation coefficient was used to compare methods.
Results
Routine Histopathologic Examination Detects Overall Bone Marrow Cell Depletion, but Not Changes in M:E or M:EL Ratios
To perturb total bone marrow cell populations, rats were treated with varying doses of a known cytotoxicant, MMAE (0.03, 0.06, and 0.20 mg/kg), or 20 mg/kg Bcl-xL inhibitor.
Microscopic pathology evaluation and automated cell density evaluation of H&E sections of both sternal and femoral/tibial bone revealed a dose-dependent minimal to severe depletion of total bone marrow cells in rats dosed with ≥0.03 mg/kg MMAE, and minimal depletion in rats given 20 mg/kg Bcl-xL inhibitor (Figure 2B in Kozlowski, Brumm, and Cain In Press). Routine histopathologic examination of sternal and femoral/tibial bone H&E slides did not reveal any changes in the M:E or M:EL ratios for rats given MMAE or Bcl-xL inhibitor, but minimally to moderately increased megakaryocyte numbers were noted in the Bcl-xL inhibitor group (data not shown).
Automated Quantification of M:EL Ratio Changes Correlate with Flow Cytometry Assessment
Automated assessment of M:EL ratios on the images of sternal and left femoral/tibial bone marrow H&E slides was compared to M:EL ratios computed from flow cytometry immunophenotyping data in the right femoral bone. Both methods detected an increase in M:EL ratio for the MMAE 0.03 and 0.06 kg/mg groups and a steep drop in M:EL ratio in the MMAE 0.20 mg/kg dose group (Figure 3).
Bone marrow smears prepared from the left femoral bone were examined microscopically, and a change in M:E ratio was observed at both 0.03 and 0.06 mg/kg doses. Image analysis on sternal bone detected an M:EL ratio increase in the MMAE 0.06 mg/kg dose group (Figure 3). Flow cytometry data of the MMAE 0.20 mg/kg dose were indicative of a large reduction in M:EL ratio. This was observed to a lesser extent by the automated method (sternum) but was inevaluable by smear cytology. Flow cytometry (right femur) and image analysis (sternum), but not smear cytology (left femur) or image analysis (left femur/ tibia), detected a statistically significant change in M:EL or M:E ratios in the Bcl-xL group compared to control.
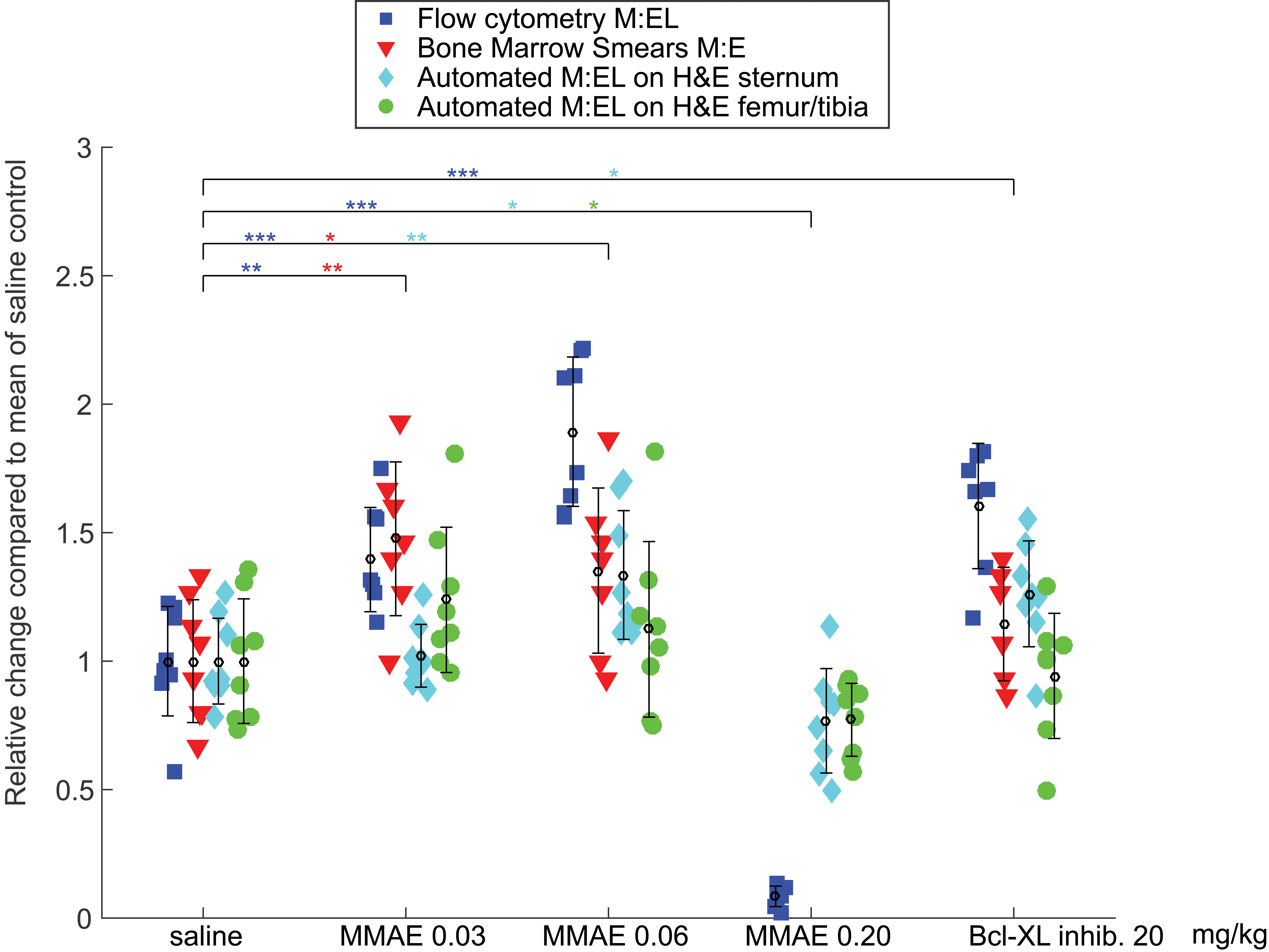
M:EL ratios for all measurement methods. All data plotted and normalized to saline control group average. Blue is M:E ratio estimated from cytology smears, red is M:EL ratio from flow cytometry, cyan is M:EL ratio from image analysis on H&E slides from sternal bone, and green is image analysis on H&E slides from femoral/tibial bone. Asterisks (*) are shown if there was significant difference in the particular treatment group, compared to the control group, and are color-coded according to which method the data originated from. For all tests between groups, no * in a particular color between groups means the different was insignificant (
At the individual animal level, automated image analysis of sternal bone M:EL and flow cytometry M:EL ratios (Figure 4) correlated (
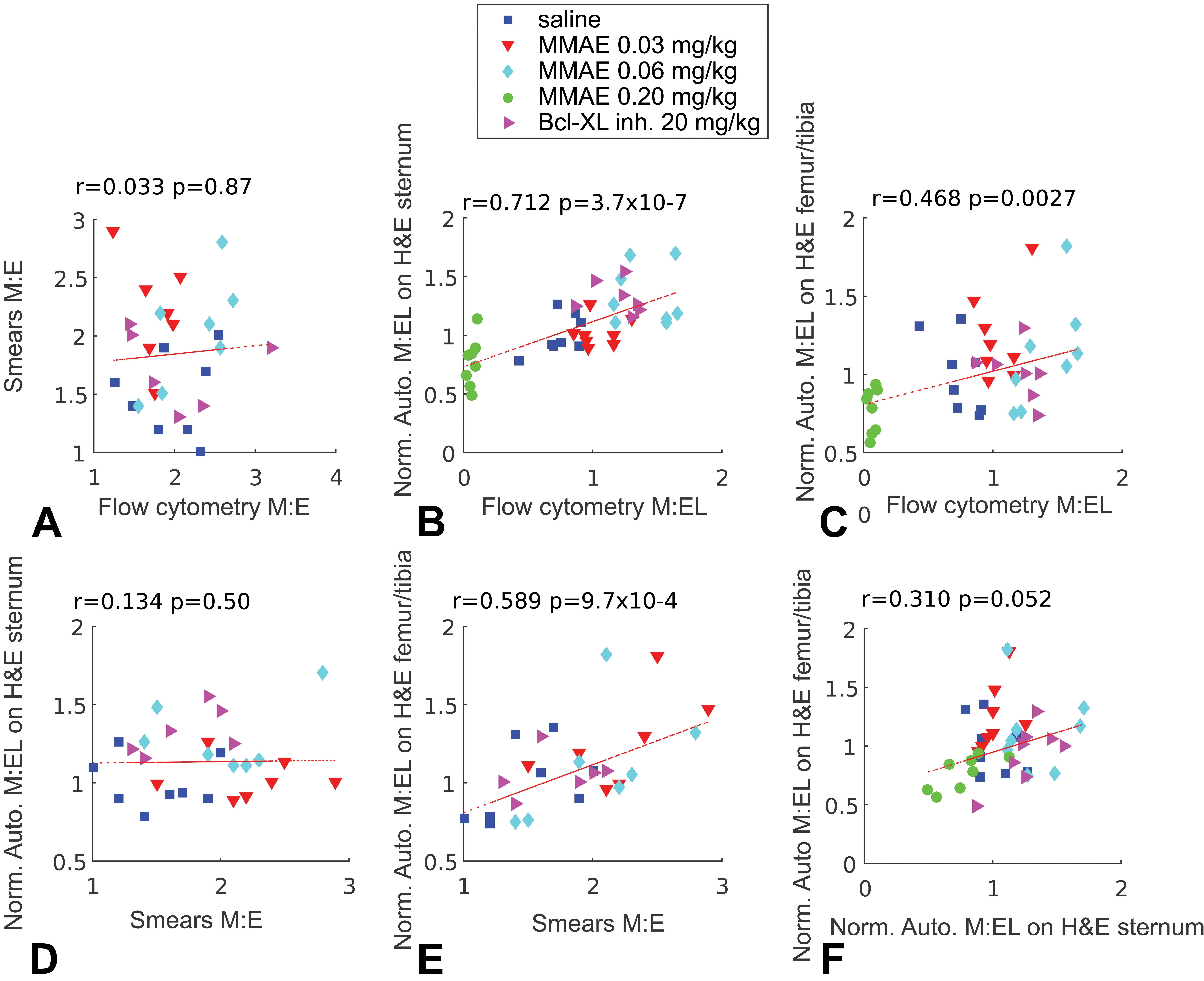
M:EL ratio estimate from flow cytometry correlates better to image analysis on H&E sternum than with any other method. Scatter plots to compare measurement methods for the M:E or M:EL ratios for all animals in the study. Colors indicate different groups shown in the legend. The linear correlations coefficients and their associated
Image Analysis on H&E and IHC Slides Detected Changes in Megakaryocyte Density and Relative Size in Bcl-xL Inhibitor–Treated Group
Using image analysis, it was possible to obtain a measure for density of the number of cells/unit area for megakaryocytes because this population has sufficient morphologic distinction to count individually (as opposed to myeloid, lymphoid, or erythroid cells).
We hypothesized that megakaryocytes may be challenging to enumerate in H&E marrow sections due to morphologic changes associated with cellular maturation (Ru et al. 2015) and their color (balance of basophilic and eosinophilic features) that can resemble bone and osteoclasts. Therefore, we incorporated analysis of a slide stained with the megakaryocyte specific marker, VWF.
Flow cytometry and manual semiquantitative counts of IHC sections, as well as image analysis of IHC (Figure 5A), detected a significant decrease in megakaryocyte number in the MMAE 0.20 mg/kg group compared to the control group. However, automated analysis of H&E sections, both on sternum and femoral/tibial bone, did not identify the decrease in the MMAE 0.20 mg/kg group. Only manual counts and image analysis of the H&E and IHC slides detected a significant increase in megakaryocyte number in the Bcl-xL inhibitor–treated group compared to the control group. Although flow cytometry detected a trend of increased megakaryocyte number in the Bcl-xL inhibitor treated group, it did not reach statistical significance. Overall, there was an unexpectedly large intergroup variation in the megakaryocyte density measurements by flow cytometry. However, the fact that image analysis on H&E failed to identify megakaryocyte depletion in the MMAE 0.20 mg/kg dose group, while the image analysis on IHC identified this change, suggests that the analysis on IHC is likely more sensitive than on H&E sections.
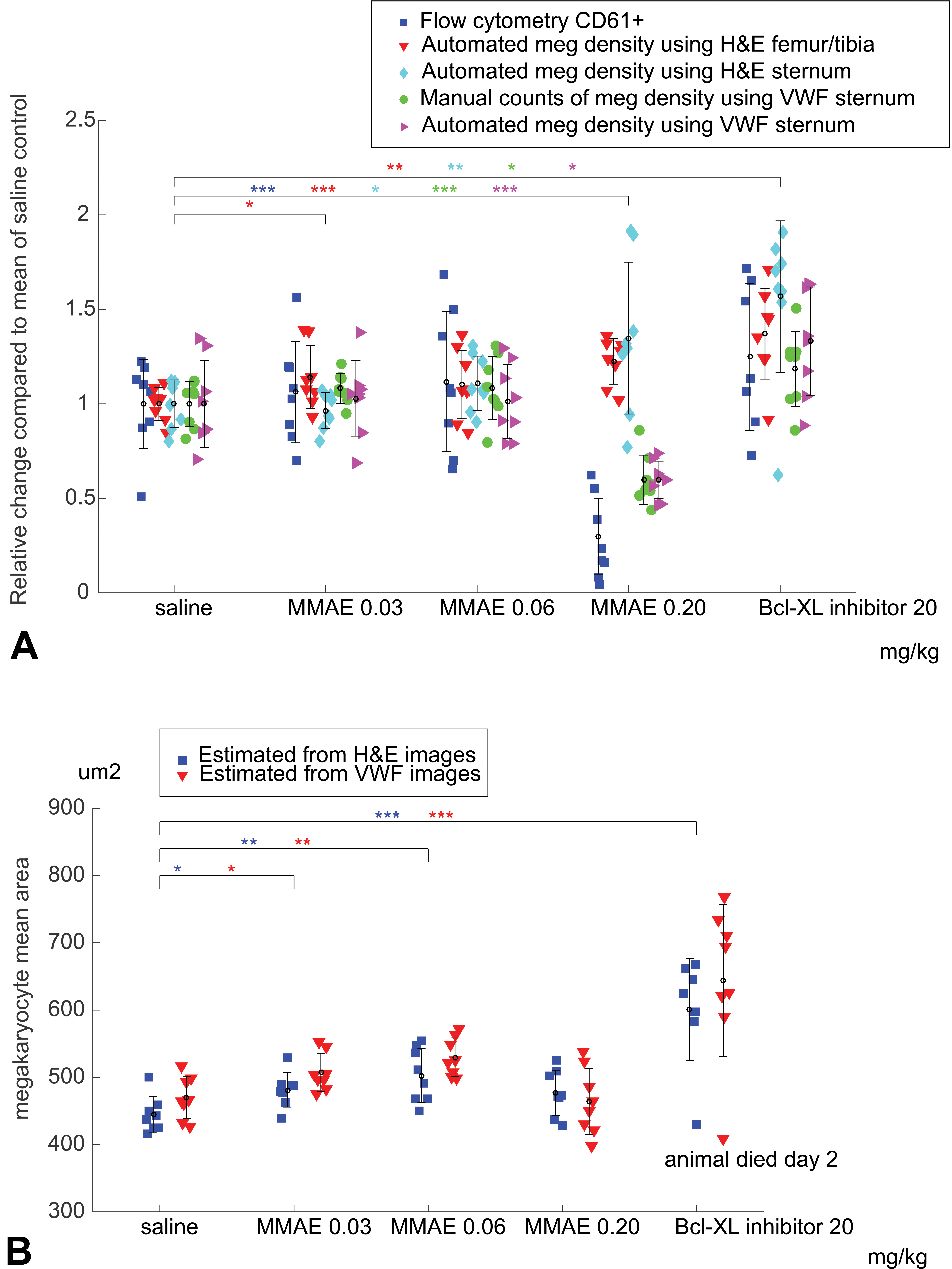
Megakaryocyte measurement for all methods. (A) Megakaryocyte (meg) density measurements. All data plotted as relative to saline control group average. (B) Image analysis on VWF slides quantification of average megakaryocyte relative size in each slide. Note that the animal that died on day 2 did not have elevated megakaryocyte relative size. For both A and B, the significance detected between the control group and treatment group is shown by a presence of an asterisk * in the color corresponding to the type of measurement, shown in each legend. For all tests between groups, no * between groups means the different was insignificant (
In order to further investigate the changes in megakaryocytes in this group, we examined the relative area of these cells in the 2-D sections. We did not attempt to measure absolute 3-D volumes of these cells but aimed to measure whether there was a relative increase in areas of 2-D sections of megakaryocytes in treatment groups compared to controls. Image analysis on both H&E and the VWF IHC revealed that there was a significant increase in average megakaryocyte size in the Bcl-xL inhibitor–treated group (Figure 5B) compared to the control group, except in the animal that died on day 2. There was no notable increase in sensitivity in measuring megakaryocyte relative size in using image analysis on VWF IHC slides compared to that on H&E slides.
The Reproducibility of the Automated Measures of M:EL Change and Megakaryocyte Density
The reproducibility of the automatic method on sternal bone was assessed for M:EL ratio change and megakaryocyte density. Three serial sections of the sternal bone marrow were automatically analyzed for each measurement (Figure 6). Coefficient of variation (CV) was computed for each animal. The average CV of the M:EL ratio was 14.3 ± 10.9%, and for megakaryocyte density, 14.6 ± 10.6%. For megakaryocyte density, we found a particularly high degree of variation concentrated in the MMAE 0.20 mg/kg dose group (Figure 6B). Visual inspection revealed that in a few instances, the large areas of serum left in the bone marrow after severe depletion of cells were misclassified by the algorithm as megakaryocyte cells. For M:EL ratio, the animals with large interslide CVs were not concentrated in specific treatment groups, and visual confirmation revealed that larger errors were associated with smaller amounts of marrow tissue available for evaluation in specific sections.
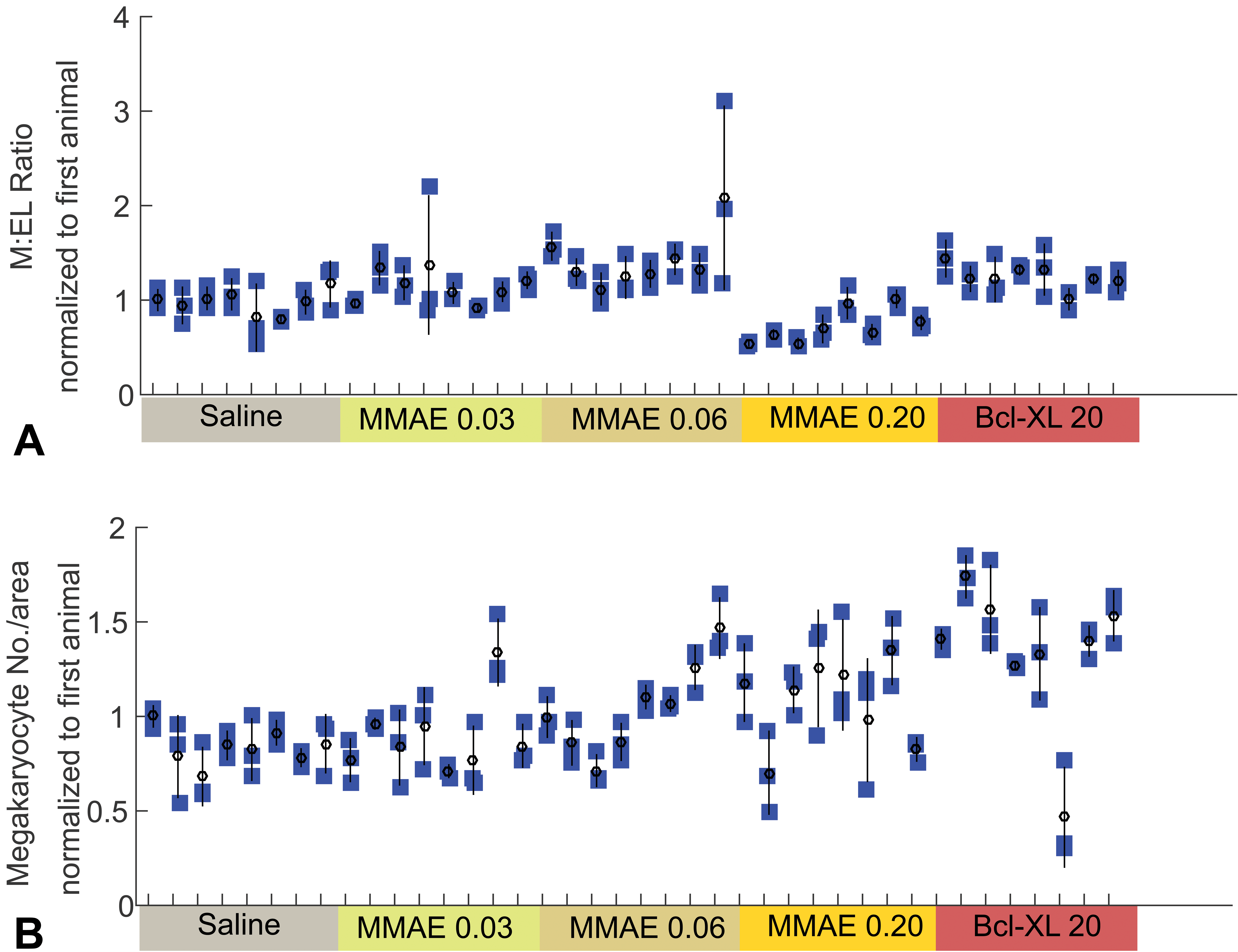
Image analysis reproducibility analysis. Three sets of H&E sternal serial sections were analyzed using the image analysis algorithm, and the results plotted normalized to the first animal. The average CV of the M:EL ratio was 14.3 ± 10.9%, and for megakaryocyte density, 14.6 ± 10.6%. H&E = hematoxylin and eosin; M:EL = myeloid: erythroid + lymphoid; CV = coefficient of variation.
Discussions
We developed a method to screen for drug-induced changes in the hematological composition of rat bone marrow, using images of H&E slides scanned at 20×. To this end, we extended an automated method that was previously designed to quantify the total cell density in bone marrow (Kozlowski , Brumm, and Cain In Press), without the requirement of significant additional computing time. In order to identify different cell types, we employed a strategy analogous to a technique used in flow cytometry (Virgo and Gibbs 2012), where cells may be categorized by their relative size and optical properties, without the use of cell-type specific markers. Because of the difficulty in distinguishing between myeloid, erythroid, and lymphoid cells by optical properties, we could not count these cells directly but developed a method to measure their ratios (M:EL) relative to control animals that are comparable provided that all study animals are stained in the same H&E run. Megakaryocytes, however, could be counted directly due to their large area that distinguishes them clearly from other cell types.
We measured the sensitivity of our method to detect changes in M:EL by dosing rats treated with varying doses of MMAE, a molecule known to affect hematological composition, and compared the automated methods with other established methods that can be used to measure shifts in bone marrow cells of different lineages. Because our algorithm can only measure M:EL ratios relative to control, we computed the analogous values using flow immunophenotyping data. On a single animal level, M:EL ratios derived from flow cytometry correlated well with image analysis (
On a group level, the automated algorithm on sternal bone detected M:EL changes at the middose (0.06 mg/kg) but not the lowest dose (0.03 mg/kg). Since changes at the lowest dose were detected by flow cytometry data and by bone marrow smear cytology, we conclude that the automated algorithm is less sensitive than these methods. It was notable that 12/40 animals in the study had inevaluable cytology smears. Therefore, image analysis could provide a backup methodology to evaluate M:EL ratios if bone marrow smears for an animal are compromised in quality. Importantly, the pathologist did not detect an M:E or M:EL change at 0.06 mg/kg, demonstrating that our method has utility in flagging a moderate change in M:EL that may be otherwise missed by manual examination of H&E slides alone. The relative decrease in M:EL ratio of the MMAE 0.20 mg/kg group compared to control was greater when measured by flow cytometry compared to the automated algorithm, although both methods detected a statistical difference in that treatment group. This suggests that the sensitivity of the algorithm to detect change in M:EL in cases of severe pancytopenia may be suboptimal compared to higher cellularity sections.
Typically, bone marrow cytology smears are prepared from femoral bone, and histopathology evaluation is performed on sternal bone. In this study, we also performed flow cytometry on the right femur bone. Since we were comparing bone marrow composition using multiple different methods, we were concerned that the site of bone collection, rather than analysis technique, may affect the results. It is known that bone marrow cellularity content is similar but not identical between tibial, femoral, and sternal rat bone marrow (Cline and Maronpot 1985). Therefore, we also prepared H&E histopathology sections from the left long bone after extracting enough material for bone marrow smears, which consisted of the stifle joint that contained femoral bone and portions of the tibia. Unfortunately, this provided less material for analysis than would be standard for femoral preparations. This may explain why the algorithm did not detect a change in M:EL in the left femoral/tibial bone at the 0.06 mg/kg dose of MMAE, while this was detected on the sternal bone. However, image analysis of the left femur/tibia, rather than the sternal bone, correlated better with cytology smears sampled from the left femoral bone. This would suggest that there may be some discrepancy between M:EL ratios in the femoral/tibial and sternal bones, though there are two few samples in this study to provide conclusive evidence of this. In the future, the image analysis algorithm may be used to further probe differences in bone marrow subpopulations in different anatomic sites of bone collection.
We also tested the ability of the algorithm to detect changes in megakaryocyte density using the Bcl-xL inhibitor molecule, which is known to disrupt the pathway in megakaryocytopoiesis and platelet survival (Kile 2014). Genetic deletion of Bcl-xL causes a depletion of mature megakaryocytes and platelet production; yet due to a compensatory mechanism, there is an increase in the number of megakaryocytes (Kodama et al. 2012). Consistent with this, an increase in megakaryocyte density was detected by image analysis of H&E slides in Bcl-xL-treated animals, compared to controls. Flow cytometry did not detect a statistically significant difference from control in this group, which may have been due to poor performance of our antibody (CD61), since there was generally a high variance in megakaryocyte numbers detected in all groups by this method. Although the flow cytometry data for megakaryocytes were variable, we obtained additional quantitative data on megakaryocytes density using an alternative approach: serial sections to the analyzed H&E slides were stained with VWF, which labels megakaryocytes as well as platelets and endothelial cells (Rehg, Bush, and Ward 2012). In general, for the human eye, it is easier to enumerate cells with a cell-type-specific stain rather than H&E, though such stains are not routinely prepared during toxicologic studies without cause. For these studies, 3 fields of view and 8 rats per group were adequate to detect differences manually, though the number of fields of view that are needed will depend on the biological variance of the data. We found that manual counting of megakaryocytes on these IHC slides tracked with our image analysis on the entire IHC slide images, in that they both showed a significant increase in the Bcl-xL inhibitor–treated group. The manual counts and algorithm (both H&E and IHC) detected that this increase was absent in the one animal in the Bcl-xL group that had died on day 2, whereas other animals were euthanized on day 4. However, the manual counts detected a significant decrease in the MMAE 0.20 mg/kg dose group, whereas the algorithm on H&E was unable to detect it. In fact, it produced particularly variable results in the latter group when serial sections were analyzed. This was due to the algorithm mistaking features of the cell-depleted marrow as megakaryocytes in H&E, highlighting the challenge of identifying cells by appearance alone when they are not specifically labeled. Because the algorithm was built to detect megakaryocytes in control animals, it was not optimized to distinguish the cells in conditions of extreme cell depletion. In the future, the solution may be to test the algorithm in multiple toxicity studies and optimize it to utilize invariant features of the specific cell type to identify them. Alternatively, IHC images can be used for more accurate quantitation in cases of marked cell depletion, as supported by the closer agreement with both flow cytometry and manual counts in the MMAE 0.20 mg/kg group.
Interestingly, our algorithm applied to H&E sections was robust enough to identify the increase in both number and relative size of megakaryocytes in the Bcl-xL-treated group, demonstrating that in situations of small perturbations, it is still able to identify cells correctly. The increase in relative size of these cells is consistent with previous observation in Bcl-xL mice, reflecting an increased population of large megakaryocytes with high nuclear ploidy (Kodama et al. 2012) due to disruption in cell maturation.
Our analyses show that the H&E algorithm may be best applied to situations where relatively subtle changes in bone marrow cell composition may be suspected. Our original algorithm is able to detect substantial overall decrease in cellularity (see Kozlowski, Brumm, and Cain In Press), but when there is a profound drop in cell number, as in the MMAE 0.20 mg/kg dose, the changes in megakaryocytes and M:EL ratios become somewhat “masked.” Typically toxicity studies have low, mid, and high doses, so that the mechanism of toxicity may be studied at lower doses. Our algorithm is optimal for detecting relatively small changes that may be missed by a pathologist examining H&E slides. In these cases, the algorithm can detect changes in cell density as well as appearance, which may be used to shed light on pathways of drug-induced toxicity. Based on our current data, we recommend using the M:EL and megakaryocyte measurements in cases where the cellularity portion of our algorithm reports that the overall cell depletion of any animal does not exceed 40% (i.e., 60% of the cells remain), which in this experiment corresponds to our mid-dose group (MMAE 0.06 mg/kg).
Overall, a remarkably large amount of information was extracted from H&E slides imaged at the 20× magnification, in contrast to the typical magnification used to analyze cytology smears, for example, which is performed at 60× to 100×. With advancing scanning technology, we may soon be able to analyze cells at higher magnifications, which would allow more accurate cell subtyping and more feasible application of this approach with the trade-off between time, computing power, and improved method performance. Furthermore, in the future, we may take advantage of recent progress in machine vision to more finely categorize cell phenotypes, separate mature and immature cells, and measure response to stress by their appearance on H&E slides.
In conclusion, we have extended our image analysis method to enumerate and characterize bone marrow cell subpopulations that can be performed at the same time as automated cell density computations. Further optimization is needed to increase the sensitivity and decrease the variability of the method. However, this is a proof-of-concept work that leverages routine availability of bone marrow H&E sections prepared in preclinical rodent toxicity studies and has the advantage of rapid processing steps and data output in comparison to manual histopathology evaluation. This image analysis approach could provide an ancillary quantitative test to complement standard hematotoxicity assessments and inform the need for follow-up testing.
Footnotes
Acknowledgments
The authors would like to thank Jochen Brumm for statistical advice and Charles River Laboratories for conducting the
Author Contribution
Authors contributed to conception or design (CK, PK, JT); data acquisition, analysis, or interpretation (CK, AF, GC, PK, JB, JT); drafting the manuscript (CK, AF, PK, JB); and critically revising the manuscript (CK, AF, GC, PK, JB, JT). All authors gave final approval and agreed to be accountable for all aspects of work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of Conflicting Interests
All authors were employees and stockholders of Genentech Inc.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was fully funded and supported by Genentech Inc. No support from external sources was received.
